Myofiber-specific TEAD1 overexpression drives satellite cell hyperplasia and counters pathological effects of dystrophin deficiency
Figures
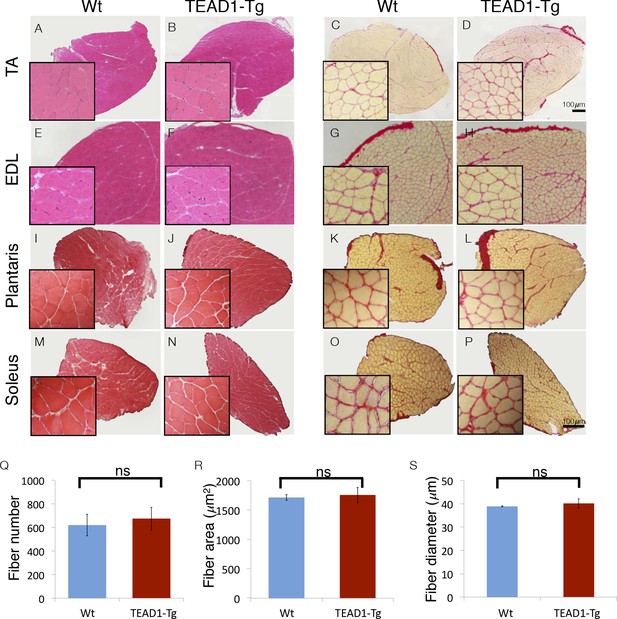
Skeletal muscle is histologically indistinguishable between Wt and TEAD1-Tg mice.
(A–P) H and E (A–B, E–F, I–J, M–N) and Sirius Red (C–D, G–H, K–L, O–P) stains of TA (A–D), EDL (E–H), Plantaris (I–L), and Soleus (M–P) from Wt (A, C, E, G, I, K, M, O) and TEAD1-Tg mice (B, D, F, H, J, L, N, P). Insets are magnified images of the representative areas within the tissue. (Q–S) EDL fiber number (Q), fiber area (R), and fiber diameter (S) are quantified for both genotypes (n > 3 adult mice for all measurements).
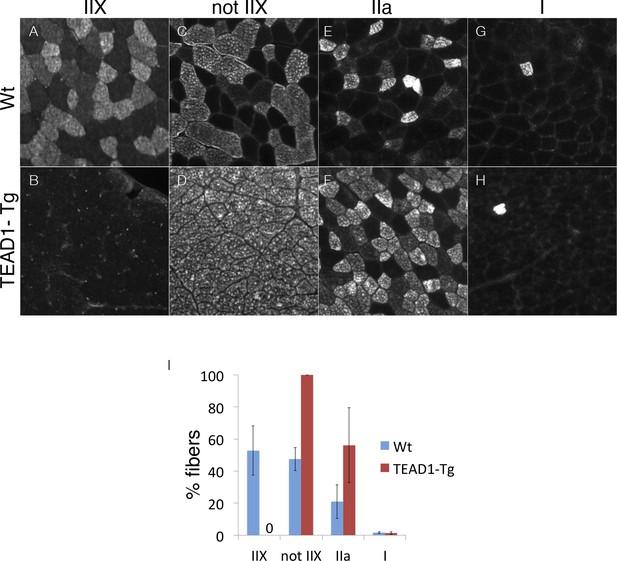
Ratios of type II fibers are changed in TEAD-Tg muscle.
Wt (A, C, E, G) and TEAD1-Tg (B, D, F, H) TA muscles were queried for percentages of type IIX fibers (A, B), fibers that were not IIX (C, D), type IIa fibers (E, F), and type I fibers (G, H). The TA muscle was chosen for analysis because it is comprised of all muscle fiber types. The results were then quantified (I). n > 3 adult mice for all measurements, p<0.05 represented by (*), p<0.005 represented by (**), p<0.0005 represented by (***).
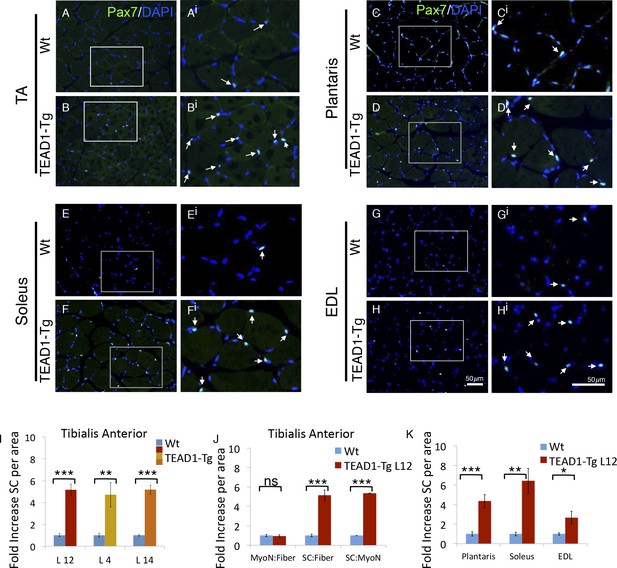
TEAD1-Tg hind limb features SC hyperplasia.
(A–Hi) Pax7 IF in hind limb muscle groups: TA (A–Ai, B–Bi), Plantaris (C–Ci, D–Di), Soleus (E–Ei, F–Fi), and EDL (G–Gi, H–Hi). Wt (A–Ai, C–Ci, E–Ei, G–Gi) and TEAD1-Tg muscles (B–Bi, D–Di, F–Fi, H–Hi) are represented. Ai–Hi are zoomed images of areas represented by the white boxes in A–H, arrows indicate Pax7 (green) positive nuclei (blue). (I–K) Quantification of SC hyperplasia (fold increase above Wt) in TA sections from three independent TEAD1 transgenic lines with different transgene insertion sites (I). Quantification of TA sections from Wt and TEAD1-Tg line L12 for fold increase of myonuclei per fiber, SC per fiber, and SC per myonuclei (J). Quantification of plantaris, soleus, and EDL muscle group sections for fold increase in SC number in TEAD1-Tg line L12 compared to WT (K). n > 3 adult mice for all measurements, p<0.05 represented by (*), p<0.005 represented by (**), p<0.0005 represented by (***).
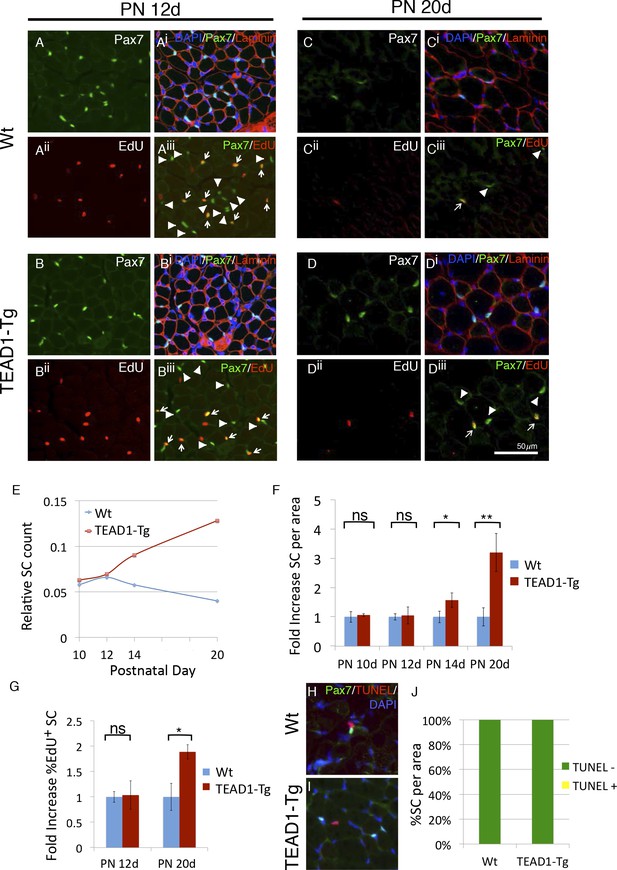
SC hyperplasia arises during perinatal stages in TEAD1-Tg mice.
(A–Diii) Wt (A–Aiii, C–Ciii) and TEAD1-Tg (B–Biii, D–Diii) SC numbers and proliferation were assessed. Postnatal day 12 (PN 12d, A–Biii) and day 20 (PN 20d, C–Diii) are represented in IF images. Pax7 (green) alone is shown in A, B, C, and D, and combined with DAPI (blue) and laminin (red) in the merged images Ai, Bi, Ci, Di. EdU (red) alone is represented in Aii, Bii, Cii, Dii and combined with Pax7 (green) in merged images Aiii, Biii, Ciii, Diii, arrows indicate nuclei labeled with both EdU and Pax7 while arrowheads are Pax7 positive nuclei that show no EdU label. (E–G) Numbers of SCs normalized to total myofibers per image is quantified for PN 10d, 12d, 14d, and 20d Wt or TEAD1-Tg TA sections (E). Fold increase in SC numbers in TEAD1-Tg TA sections compared to Wt was quantified for PN 10d, 12d, 14d, and 20d (F). Fold increase in percent EdU positive SCs in TEAD1-Tg TA sections compared to Wt was quantified for PN 12d and 20d (G). (H–J) Apoptosis rates of SCs were assessed for PN 14d. Representative images for Wt (H) and TEAD1-Tg muscle (I) show Pax7 (green) and TUNEL (red) stained nuclei (blue). Percent of SCs that were TUNEL positive or negative was quantified (J). n > 3 mice for all measurements, p<0.05 represented by (*), p<0.005 represented by (**), p<0.0005 represented by (***).
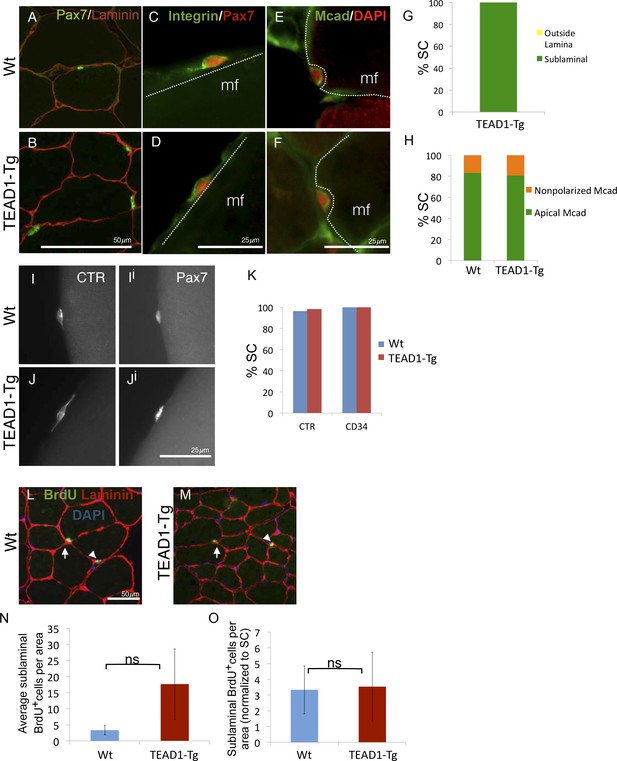
SC localization, marker expression, and long-term proliferation are no different in TEAD1-Tg compared to Wt mice.
(A–H) Localization of SC (Pax7, green) within the muscle fiber basal lamina (laminin, red) shown by IF of adult Wt (A) and TEAD1-Tg (B) TA sections and quantified for TEAD1-Tg TA sections (G). Polarized Integrin-β1 expression (Integrin, green) in SCs (Pax7, red) was assessed on TEAD1-Tg and Wt isolated EDL fibers (myofiber, mf) and found to be basally localized in all instances where the position of the SC on the fiber allowed for assessment (C,D). M-Cadherin (Mcad, green) polarization in SCs (Pax7, nuclei shown in red) was assessed in TA sections for apical localization (E,F), which is quantified in H. Non-polarized Mcad in Wt and TEAD1-Tg samples is likely due to imperfect SC orientation within the section. I–K) Assessment of quiescent marker (calcitonin receptor, CTR) expression in SCs (Pax7, Ii and Ji) on Wt (I) and TEAD1-Tg (J) isolated EDL fibers are represented by IF and quantified in K along with the general stem cell marker, CD34. L–N) Following a month-long BrdU treatment, Wt (L) and TEAD1-Tg (M) adult TA sections were assessed for long-term proliferation. Since myonuclei are non-proliferative and SC nuclei present the only other sublaminal nuclear species, this assay cumulatively captures any SC proliferation over the one-month long period. Very low numbers of BrdU nuclei were detected in both sublaminal and interstitial compartments of TA muscles from TEAD1-Tg and Wt mice. BrdU (green) labeled nuclei (blue) within the basal lamina (red) of the myofiber were quantified (N) and normalized to SC number (O). For G, H, and K n > 50 cells, while for N, 3 mice were quantified for each genotype.
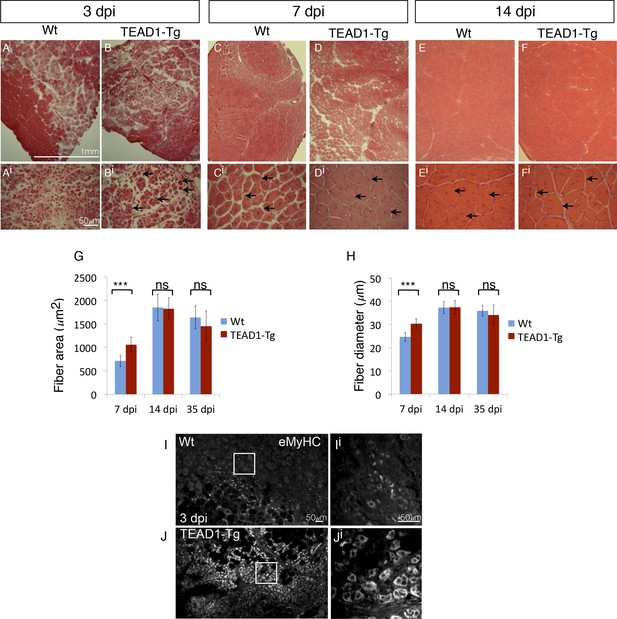
TEAD1-Tg muscle exhibits faster regeneration than Wt muscle upon injury with cardiotoxin.
(A–Fi) H and E stains of Wt (A–Ai, C–Ci, E–Ei) and TEAD1-Tg (B–Bi, D–Di, F–Fi) TA muscle 3 days (3 dpi, A–Bi), 7 days (7 dpi, C–Di), or 14 days (14 dpi, E–Fi) after injury with cardiotoxin (CTX). Panels Ai–Fi show a higher magnification of the regenerating muscle (indicated by arrows and central nuclei). A lack of arrows in Ai is due to a lack of identifiable young fibers at that stage in Wt muscle. G–H) Quantification of fiber area (G) and fiber diameter (H) for 7 dpi, 14 dpi, and 35 dpi. I–Ji) IF of embryonic myosin heavy chain (eMyHC, I–Ji) localized to regenerating fibers in Wt (I–Ii) and TEAD1-Tg (J–Ji) TA muscles 3 days after injury by CTX. Images Ii and Ji show a zoomed-in region indicated by the white boxes in images I and J. n > 3 mice for all samples quantified. p<0.05 represented by (*), p<0.005 represented by (**), p<0.0005 represented by (***).
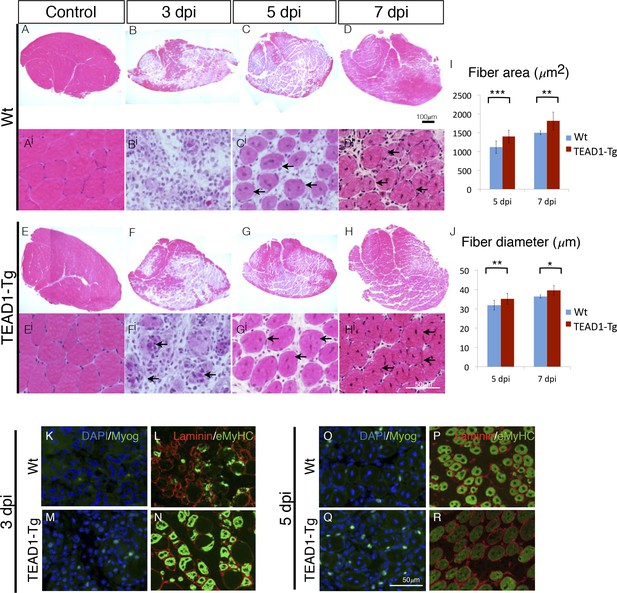
TEAD1-Tg muscle exhibits faster regeneration than Wt muscle upon injury with BaCl2.
(A–Hi) H and E stains of Wt (A–Di) and TEAD1-Tg (E–Hi) TA muscle 3 days (B–Bi, F–Fi), 5 days (C–Ci, G–Gi), or 7 days (D–Di, H–Hi) after injury with BaCl2 or uninjured (A–Ai, E–Ei). Images Ai–Hi are higher magnification views of regenerating areas of muscle (regenerating fibers indicated by black arrows). I–J) Quantification of fiber area (I) and fiber diameter (J) for 5 days and 7 days after BaCl2 injury. K–R) IF images show myogenin (green) and DAPI (blue; K,M,O,Q) or embryonic myosin heavy chain (green) with laminin (red; L,N,P,R) localized to regenerating fibers in Wt (K–L, O–P) and TEAD1-Tg (M–N, Q–R) TAs 3 days and 5 days after BaCl2 injury. For quantification of fiber number and diameter n=3 mice were used. p<0.05 represented by (*), p<0.005 represented by (**), p<0.0005 represented by (***).
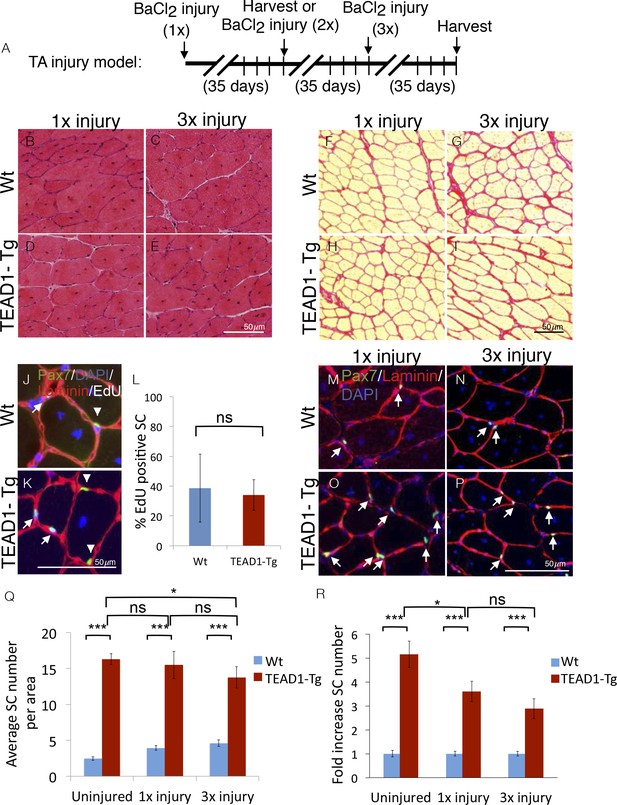
SC hyperplasia persists through multiple injuries.
Adult mouse TA muscles were injured with BaCl2 multiple times with 35-day regeneration periods between injuries as indicated (A). This injury paradigm applies to panels B–I, M–P) H and E stains show muscle after 35 days of regeneration following one injury (B,D) or 3 injuries (C,E) for Wt (B,C) or TEAD1-Tg muscle (D,E). (F–I) Sirius Red stains show the connective tissue after 35 days of regeneration following one injury (F,H) or 3 injuries (G,I) for Wt (F,G) or TEAD1-Tg muscle (H,I). (J–L) Wt (J) and TEAD1-Tg (K) TA muscles were injured with CTX and regenerated for 2 weeks. EdU was given on days 2–5 of regeneration. IF of Pax7 (green), laminin (red), EdU (white), and counterstained with DAPI (blue) allowed for quantification of sublaminal Pax7 and EdU positive cells (L). Myonuclei in images are EdU positive but reduced in intensity due to larger nuclear volume. M–P) Numbers of Pax7 expressing cells were assessed by IF of Pax7 (green) and laminin (red), counterstained with DAPI (blue) after 35 days of regeneration following one injury (M,O) or 3 injuries (N,P) for Wt (M,N) or TEAD1-Tg muscle (O,P) and is quantified as SC averages per area (Q) and fold increase relative to Wt (R). Uninjured data from Figure 2 displayed for comparison. n > 3 adult mice for all measurements, p<0.05 represented by (*), p<0.005 represented by (**), p<0.0005 represented by (***).
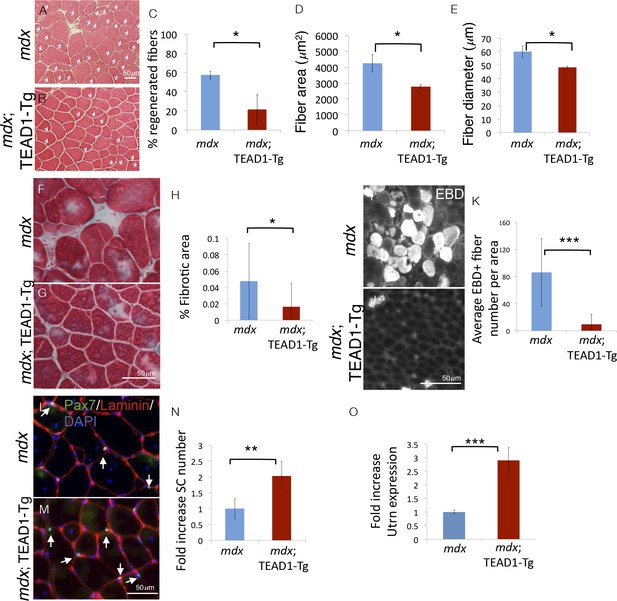
Pathologies of muscular dystrophy are ameliorated by TEAD1-Tg expression.
Histology, IF analyses, and qRTPCR on dystrophic muscle from mdx mice modeling chronic injury. A–C) Asterisks in H and E stained muscle sections indicate regenerated fibers in mdx (A) and mdx; TEAD1-Tg mice (B). The percent of fibers that are regenerative is quantified for each genotype in C. D–E) The fiber area (D) and fiber diameter (E) for mdx or mdx; TEAD1-Tg TA muscle is also quantified. F–H) Trichrome staining was employed to label fibrotic areas in mdx (F) or mdx; TEAD1-Tg (G) TA muscle. Percent fibrotic area was quantified (H). I–K) Membrane leakage was assessed by Evan’s Blue Dye incorporation into the myofibers of mdx (I) or mdx; TEAD1-Tg (J) TA muscle. Positive fibers per area were quantified (K). (L–N) Numbers of Pax7 expressing cells were assessed by IF of Pax7 (green), and laminin (red), counterstained with DAPI (blue) in mdx (L) or mdx; TEAD1-Tg (M). TA muscles are quantified as fold increase relative to mdx in N. Utrophin (Utrn) expression was quantified in mdx or mdx; TEAD1-Tg TA muscle via qRTPCR (O). n > 3 adult mice for all measurements, p<0.05 represented by (*), p<0.005 represented by (**), p<0.0005 represented by (***).
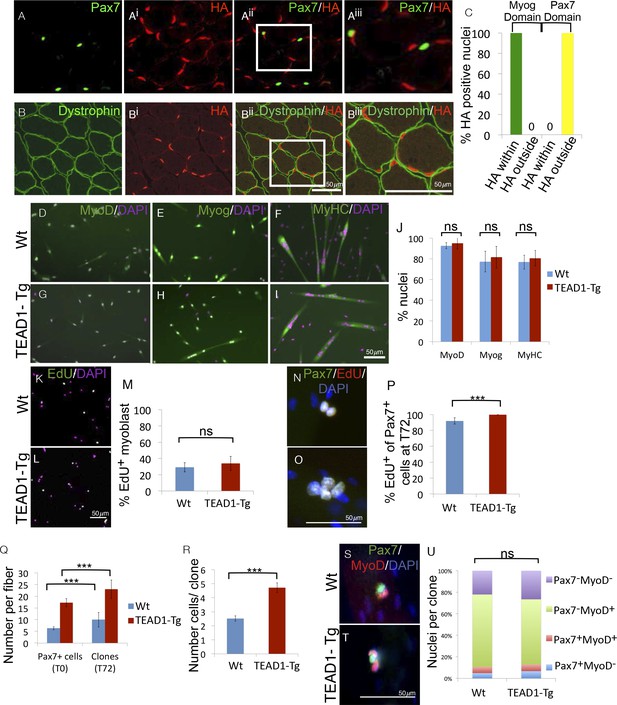
SC hyperplasia derives from cell-cell signaling between the myofiber and SCs.
(A–Aiii) IF images of TEAD1-Tg TA adult muscle show Pax7 expressing SCs (green; A) and HA tagged TEAD1 (red; Ai), and are merged in image (Aii) and a zoomed-in merged image (Aiii) to show that these genes are expressed in distinct compartments. B–Biii) IF of HA-TEAD1 (red, Bi) and dystrophin (green, B) show that the TEAD1 transgene is expressed in nuclei of the myofiber in TA adult muscle. Transgene expression was also queried by HA colocalization with Myogenin (Myog) and Pax7-expressing reserve cells in differentiated cell culture experiments after 3 and 5 days of differentiation, respectively (C). (D–J) In vitro differentiation of myoblasts derived from Wt (D–F) or TEAD1-Tg (G–I) hind limb muscle shows equivalent expression of MyoD (D,G; green), Myogenin (E,H; green), and myosin heavy chain (F,I; green) and is quantified in J (n> 480 cells for all). (K–M) Cultures of Wt (K) or TEAD1-Tg (L) myoblasts show equivalent EdU label (green), which is quantified in M (n>375 cell for each). N–P) Muscle fibers and associated SC were isolated and grown in culture for 72 hr with EdU in the growth media after 24 hr in culture. A minimum number of 50 fibers were quantified. Cell clones labeled with Pax7 (green), DAPI (blue), and/or EdU (red) can be observed on isolated muscle fibers from Wt (N) or TEAD1-Tg (O) mice and are quantified in P. The average number of Pax7-positive cells (T0) and the average number of clones on muscle fibers after 72 hr (T72) in growth media are quantified in Q. Clone sizes for TEAD1-Tg and Wt fibers at 72 hr post myofiber isolation in R (n=3 mice each). After 72 hr in culture, the ratio of self-renewing SCs to differentiating cells within these clones is quantified in U (n>150 clone) for both Wt (S) and TEAD1-Tg (T) isolated fibers cultured in 10% FBS/ DMEM. p<0.05 represented by (*), p<0.005 represented by (**), p<0.0005 represented by (***).
Tables
PCR genotyping and RT primers, and conditions.
| Gene | Forward primer | Reverse primer | Size |
|---|---|---|---|
| TEAD-HA | 5’-ATCCATGCTTGTTACCTTCAG-3’ | 5’-ACTACAAGGACGATGACAAG-3’ | 460 bp |
| Internal Control | 5’-CAGCTCTACATCACCTGCCA-3’ | 5’-CACTGGGAAGAGACACTCAG-3’ | 520 bp |
| PCR conditions | Denature: 94°C for 30 s Anneal: 56°C for 30 s Extend: 72°C for 60 s (34 cycles) | ||
| Dystrophin (WT) | 5’-GCGCGAAACTCATCAAATATGCGTGTTAGTGT-3’ | 5’-GATACGCTGCTTTAATGCCTTTAGTCACTCAGATAGTTGAAGCCATTTTG-3’ | 134 bp |
| Dystrophin (mdx) | 5’-GCGCGAAACTCATCAAATATGCGTGTTAGTGT-3’ | 5’-CGGCCTGTCACTCAGATAGTTGAAGCCATTTTA-3’ | 117 bp |
| PCR conditions | Denature: 94°C for 20 s Anneal: 60°C for 20 s Extend: 72°C for 20 s (5 cycles) Denature: 94°C for 20 s Anneal: 64°C for 20 s Extend: 72°C for 20 s (23 cycles) | ||
| utrophin | 5’-AGTATGGGGACCTTGAAGCC-3’ | 5’-CGAGCGTTTATCCATTTGGT-3’ | |
| cycloA | 5’-ATTTCTTTTGACTTGCGGGC-3’ | 5’-AGACTTGAAGGGGAATG-3’ | |
| actin | 5’-CCCTAAGGCCAACCGTGAA-3’ | 5’-CAGCCTGGATGGCTACGTACA-3’ | |
Primary antibodies used for immunofluorescence. The monoclonal antibodies for Pax7, Myog, Myosin, IIx type myosin, IIa type myosin, non-IIx type myosin, I type myosin, and embryonic myosin, developed by Stefano Schiaffino, C Lucas, A Kawakami, WE Wright, DA Fischman, and HM Blau were obtained from the Developmental Studies Hybridoma Bank, created by the NICHD of the NIH and maintained at The University of Iowa, Department of Biology, Iowa City, IA 52242.
| Antibodies | Host | Dilution | Source |
|---|---|---|---|
| Anti-Pax7 | mouse (IgG1) | 1:4-5 | DSHB Pax7 (RRID:AB_528428) |
| Anti-Laminin | rabbit | 1:3000 | Sigma L9393 (RRID:AB_477163) |
| Anti-HA | rabbit | 1:1000 | Invitrogen PA1-985 (RRID:AB_559366) |
| Anti-HA | rabbit | 1:100 | Sigma H6908 (RRID:AB_260070) |
| Anti-HA | mouse (IgG1) | 1:100 | Covance mms-101P (RRID:AB_2314672) |
| Anti-B1 Integrin | rat | 1:200 | Abcam AB95623 (RRID:AB_10676803) |
| Anti-Mcadherin | mouse | 1:200 | Santa Cruz SC81471 (RRID:AB_2077111) |
| Anti-Calcitonin Receptor | rabbit | 1:250 | AbD Serotec AHP635 (RRID:AB_2068967) |
| Anti-CD34 | rat | 1:25 | BD Biosciences 553731 (RRID:AB_395015) |
| Anti-Dystrophin | mouse | 1:1000 | Genetex GTX27164 (RRID:AB_386029) |
| Anti-Dystrophin | rabbit | 1:100 | Abcam AB15277 (RRID:AB_301813) |
| Anti-MyoD | rabbit | 1:50 | Santa Cruz SC760 (RRID:AB_2148870) |
| Anti-MyoD | mouse (IgG1) | 1:200 | Santa Cruz SC32758 (RRID:AB_627978) |
| Anti-Myogenin | mouse | 1:20 | DSHB F5D (RRID:AB_528355) |
| Anti-Myogenin | mouse (IgG1) | 1:200 | Santa Cruz SC12732 (RRID:AB_627980) |
| Anti-MyHC | mouse | 1:20 | DSHB MF20 (RRID:AB_2147781) |
| Anti-eMyHC | mouse (IgG1) | 1:400 | DSHB F1.652 (RRID:AB_528358) |
| Anti-myosin (IIX) | mouse (IgM) | 1:2 | DSHB 6H1 (RRID:AB_2314830) |
| Anti-myosin (non-IIX) | mouse (IgG1) | 1:2 | DSHB BF-35 (RRID:AB_2274680) |
| Anti-myosin (IIA) | mouse (IgG1) | 1:2 | DSHB SC-71 (RRID:AB_2147165) |
| Anti-myosin (I) | mouse | 1:2 | DSHB BA-D5 (RRID:AB_2235587) |
| Anti-cleaved caspase 3 | rabbit | 1:100 | Cell Signaling 9664S (RRID:AB_2070042) |
Secondary antibodies used for immunofluorescence.
| Host | Antigen | Fluorophore | Dilution | Source |
|---|---|---|---|---|
| Goat | Rabbit IgG | Alexa 488 | 1:1000 | Invitrogen A11034 (RRID:AB_2576217) |
| Goat | Mouse IgG | Alexa 488 | 1:1000 | Invitrogen A11001 (RRID:AB_2534069) |
| Goat | Mouse IgG1 | Alexa 488 | 1:100 - 1:1000 | Invitrogen A21121 (RRID:AB_2535764) |
| Goat | Rabbit IgG | Alexa 568 | 1:1000 | Invitrogen A11011 (RRID:AB_2534078) |
| Goat | Rabbit IgG | Alexa 546 | 1:100 | Invitrogen A11035 (RRID:AB_2534093) |
| Goat | Mouse IgG | Alexa 568 | 1:1000 | Invitrogen A11031 (RRID:AB_2534090) |
| Goat | Rat IgG | Alexa 546 | 1:200 | Invitrogen A11081 (RRID:AB_2534125) |
| Goat | Mouse IgM | Alexa 594 | 1:1000 | Invitrogen A21044 (RRID:AB_2535713) |
Weights (mgSD) of respective muscle groups in Wt vs. TEAD1-Tg mice. TEAD1-Tg hind limb muscles are equivalent in weight to Wt. Weight measurements (mg) and standard deviation for TA, EDL, Soleus, and Plantaris muscles from TEAD1-Tg or Wt mice. By t-test no significant difference was found between genotypes (n > 5 muscles for all measurements).
| TA | EDL | Soleus | Plantaris | |
|---|---|---|---|---|
| Wt | 41.0 ± 5.5 | 9.3 ± 1.9 | 8.3 ± 1.4 | 16.5 ± 2.5 |
| TEAD1-Tg | 38.6 ± 3.4 | 9.5 ± 1.5 | 9.7 ± 0.9 | 14.3 ± 2.7 |





