Calcium handling precedes cardiac differentiation to initiate the first heartbeat
Figures
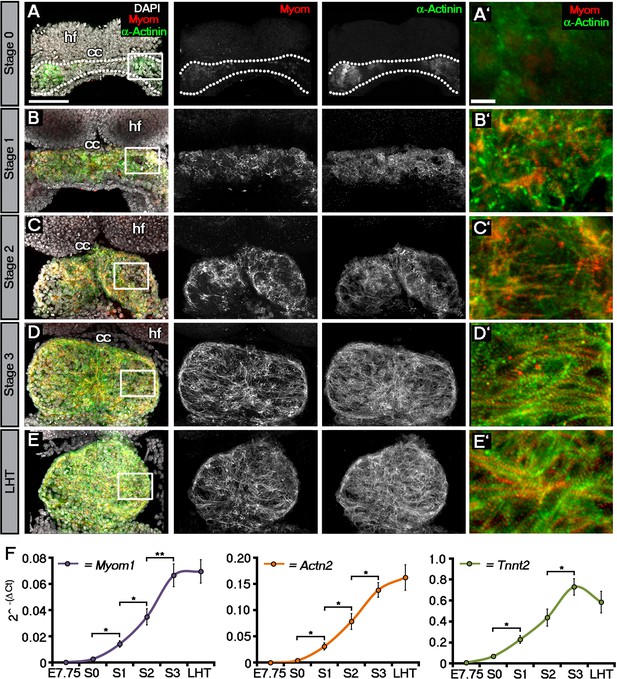
Sarcomeric assembly occurs in the forming cardiac crescent during heart development.
Maximum intensity projections of alternating myomesin (Myom) and sarcomeric alpha-actinin (α-Actinin) immunostaining from cardiac crescent formation to the linear heart tube stage (LHT; A–E; A, 11 stacks; B, 36 stacks; C, 35 stacks; D, 31 stacks; E, 36 stacks). Analysis by qRT-PCR revealed a significant increase in the expression of Myom1, Actn2, Tnnt2 (encoding Myomesin, sarcomeric alpha-actinin and cardiac troponin t), in isolated cardiac crescents between stage 0 and stage 1 (F). cc, cardiac crescent (lateral plate mesoderm); hf, head folds (neural ectoderm). Scale bars: A–E, 100 μm, A’–E’, 10 μm. Statistics: ANOVA and Tukey test for multiple comparisons (*p<0.05; **p<0.01; ***p<0.001).
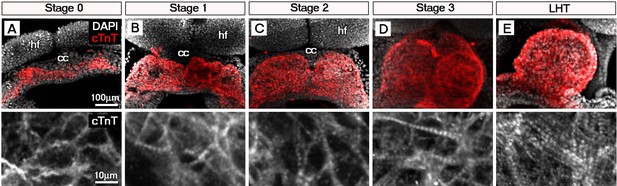
Sarcomeric assembly occurs in the forming cardiac crescent during heart development.
Maximum intensity projections of cardiac troponin (cTnT) immunostaining revealed progressive differentiation and sarcomeric assembly during stages of cardiac crescent through to linear heart formation (LHT; A–E; A, 30 stacks; B, 31 stacks; C, 40 stacks; D, 26 stacks; E, 21 stacks). qRT-PCR of Myom1, Actn2 and Tnnt2 expression during stages of cardiac crescent through to linear tube formation (F, n = 5 per stage). cc, cardiac crescent; hf, head folds. Scale bars: A, top panel 100 μm, bottom panel 10 μm.
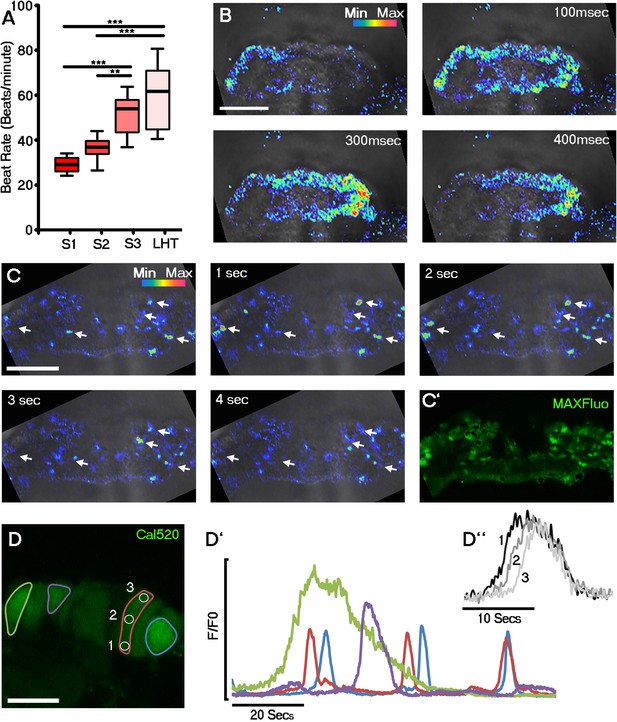
Initiation of contraction begins within the forming cardiac crescent and is preceded by spontaneous asynchronous Ca2+ oscillations during heart development.
Quantitative analysis from the onset of cardiac contraction at stage 1 of crescent formation to formation of the LHT (see Figure 1 and Supplementary file 1a for morphological staging. Stage 1, n = 12; stage 2, n = 8; stage 3, n = 10; LHT, n = 7), revealed a significant increase in heart rate from stages 2 to 3 (A). Ca2+ signal following Cal520 loading of stage 1 embryos revealed lateral propagations of transients across the crescent that correlated with the onset of beating at stages significantly earlier than previously described (B). Ca2+ signal following Cal-520 loading of stage 0 embryos revealed spontaneous asynchronous Ca2+ oscillations in individual cells prior to beating, highlighted by white arrows (C), temporal maximum intensity projection of Cal-520 fluorescence (MaxFluo) over a period of 30 s. Higher resolution imaging of stage 0 single cell Ca2+ oscillations represented as a temporal maximum intensity projection over a period of 100 s (D) revealed variation in SACO transient size and frequency (D’) and could be observed slowly propagating throughout cells (D’’). Scale bars: B, C, 100 μm, D, 20 μm. Statistics: ANOVA and Tukey test for multiple comparisons (*p<0.05; **p<0.01; ***p<0.001).
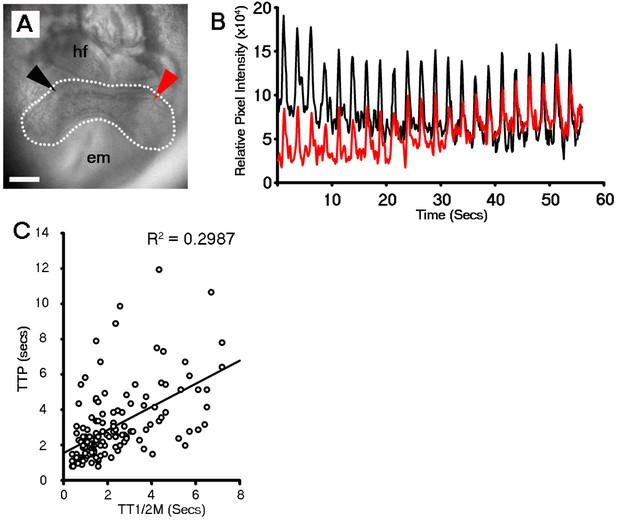
Discrete foci either side of the embryonic midline beat at the same rate within the stage 1 cardiac crescent.
DIC image of a stage 1 embryo with foci of beating highlighted on the left (red arrowhead) and right (black arrowhead) of the embryonic midline and cardiac crescent outline highlighted by white dashed line (A). Discrete foci on the left and right of lateral region of the crescent beat at the equivalent frequency (B), indicative of synchronization across the midline or intrinsic pacing. hf, head folds; em, embryonic midline. (C) Single cell Ca2+ oscillations were observed in all stage 0 cardiac crescents (n = 35) and varied in time to peak (TTP) and time to ½ maximal (TT1/2M) fluorescent intensity (n = 145 cells), revealing variable Ca2+ wave duration as well as differences in speed of Ca2+ influx and efflux. All scale bars 100 μm.
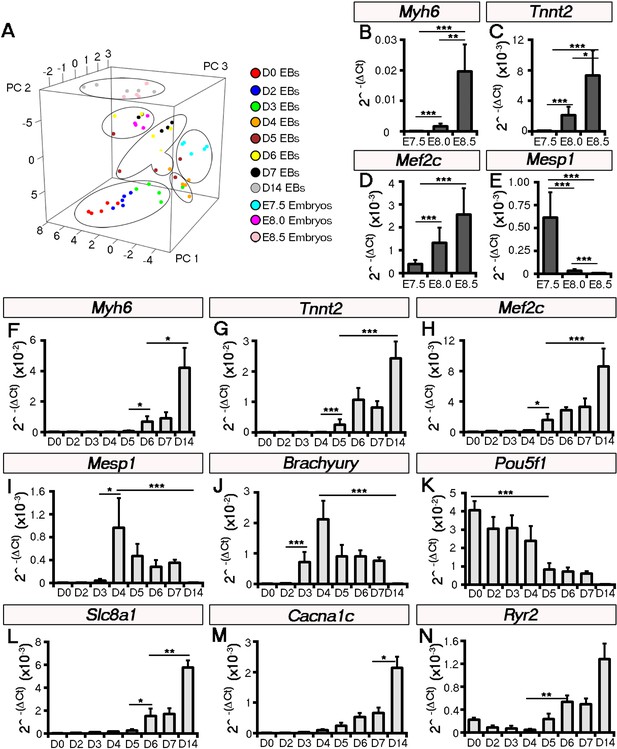
Principal component analysis of temporal gene expression profiles to cluster embryonic stage with stage of ESC-derived cardiomyocyte differentiation.
Principal component analysis (PCA) comparing embryonic stages E7.5-E8.5 (n = 5 per stage) and days 0–14 (n = 5 per stage) of ESC induced cardiomyocyte differentiation (embryoid bodies; EBs; A). Groups were calculated by hierarchical clustering: E7.5 embryos clustered weakly with day 4/5 EBs, E8.0 embryos clustered with day 6/7 EBs and E8.5 embryos clustered with day 14 EBs as an outlier (B). Temporal gene expression profiles were assessed on whole embryos by qRT-PCR: significantly increased expression of Myh6 (B), Tnnt2 (C) and Mef2c (D) was evident from E7-5-E8.5 (n = 5 per stage), coincident with increased cardiomyocyte differentiation. In contrast Mesp1 was significantly down-regulated from E7.5 following early cardiac specification (E). Comparative gene expression across the time course of ESC differentiation (n = 5 per stage), revealed equivalent trends of increased Myh6 (F), Tnnt2 (G) and Mef2c (H) from days 4 through 14, consistent with cardiomyocyte differentiation and embryonic stages E7.5-E8.5. Mesp1 was similarly downregulated from day 4 of differentiation (I; as mapped against E7.5 from the PCA A) and this followed Brachyury expression indicating (lateral plate) mesoderm formation (J) and loss of pluripotency, as demarcated by a down-regulation of Pou5f1 (encoding Oct-4) from day 4 (K). Ca2+ handling genes were expressed during the time course of ESC differentiation, with Slc8a1 encoding NCX1 (L) expressed from day 4 and significantly increased from day six, Cacna1c encoding LTCC (M) and Ryr2 (encoding the Ryanodine Receptor; N) significantly increased from day 7. All error bars are mean ± S.E.M. Statistics: one-way ANOVA and Tukey test for multiple comparisons (*p<0.05; **p<0.01; ***p<0.001).
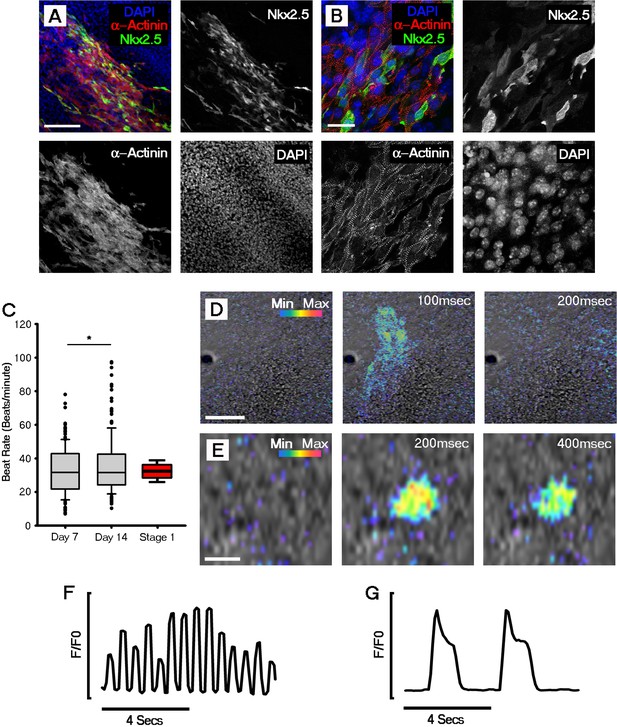
Cardiomyocyte formation, onset of beating and Ca2+ transients are evident by day 7 of ESC-derived cardiomyocyte differentiation.
Immunocytochemistry revealed the expression of sarcomeric α-actinin in Nkx2.5 positive regions, indicating formation of ESC-derived cardiomyocytes (A); higher magnification images of α-actinin filaments revealed the formation of sarcomeres (B). Beating was first observed in day 7 (D7) ESC-derived cardiomyocytes. Rate of beating was assessed using bright field movies and calculated at day 7 (n = 176 EBs) and day 14 (n = 180 EBs), revealing a significant increase between the two time points (C). Comparative beat rates were observed in ESC derived cardiomyocytes and stage 1 embryos (n = 12). Cal520 Ca2+ imaging of day 7 ESC-derived cardiomyocytes revealed large fast propagating Ca2+ waves (D) as well as slow Ca2+ oscillations in isolated single cells (E). Representative Cal-520 traces for the time series shown in D (F) and E (G). Scale bars: A, D, 100 μm; B, 25 μm; E, 10 μm. All error bars are mean ± standard deviation. Statistics: ANOVA and Tukey test for multiple comparisons (*p<0.05).
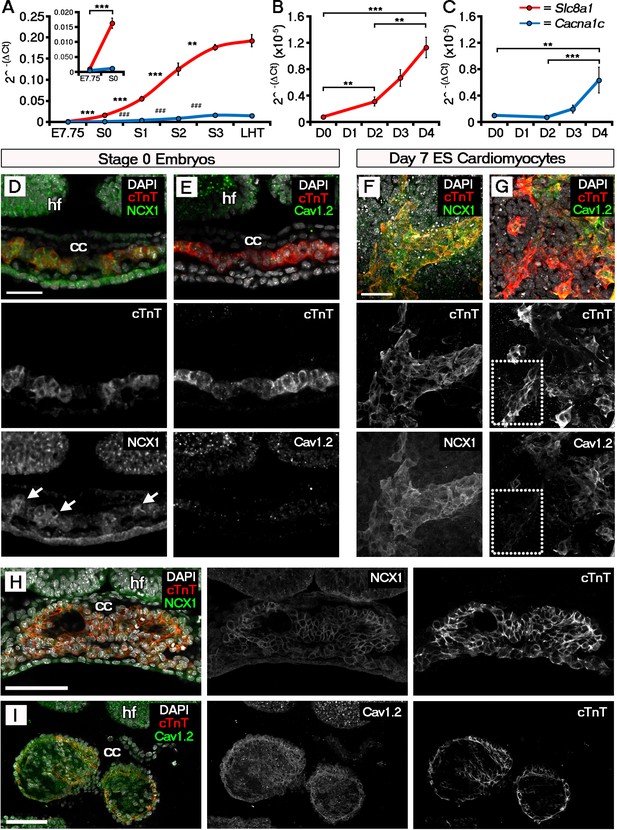
The ECC components NCX1 and LTCC are expressed within the early embryonic heart and ESC-derived cardiomyocytes.
Analyses by qRT-PCR revealed a significant increase in the expression of Slc8a1 (encoding NCX1) in the heart from E7.75 to stage 0 (A, n = 5 per stage) and from day 2 of differentiation of ESC-derived cardiomyocytes (B, n = 5 per stage). In contrast expression of Cacna1c (encoding the LTCC subunit CaV1.2), increased at a later stages from stage 0 to stage 1 and from day 4 of ESC-derived cardiomyocyte differentiation (C, n = 5 per stage). Confocal imaging section of a stage 0 embryo following immunostaining for cTnT (red) and NCX1 (green), indicated membrane localization of NCX1 within the forming crescent (D; white arrows in lower panel), whereas CaV1.2 (green) was absent from cTnT+ (red) regions at the same stage (E). A maximum intensity projection of day 7 ESC-derived cardiomyocytes revealed complete overlap of staining for cTnT (red) and NCX1 (green; F, 33 stacks), whereas CaV1.2 (green) overlapped in part with cTnT (red) but there were also extensive cTnT+/CaV1.2- regions (dotted box) emphasizing the later requirement for LTCC (G, 22 stacks). Confocal imaging section of a stage 2 embryo following immunostaining for both NCX (H) and CaV1.2 (I) revealed the expression of both proteins at later stages of heart development. cc, cardiac crescent; hf, head folds. Scale bars: D, F 50 μm, H, I 100 μm. All error bars are mean ± S.E.M; Statistics: one-way ANOVA and Tukey test for multiple comparisons (*p<0.05; **p<0.01; ***p<0.001).
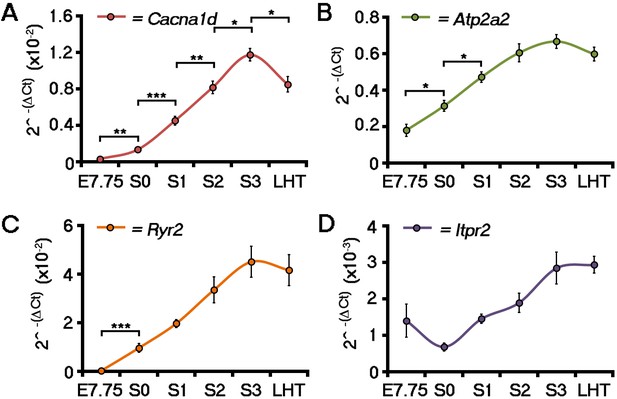
ECC component expression increases during cardiac crescent formation.
Analyses by qRT-PCR revealed a significant increase in the expression of the Cacna1d (encoding the LTCC subunit Cav1.3) from E7.75 to the stage 0 cardiac crescent (A), suggesting both L-type Ca2+ channel subunits Cav1.2 and Cav1.3 maybe required for cardiac function from stage 1 onwards. Expression of Atp2a2 (encoding the SR Ca2+ pump SERCA2a; B) and Ryr2 (C) significantly increased between E7.75 and stage 0, whilst Itpr2 (encoding the IP3 type 2 receptor) was unchanged over the complete timecourse from E7.75 to the LHT (D). All error bars are mean ± S.E.M. Statistics: A–D, one-way ANOVA and Tukey test for multiple comparisons (*p<0.05; **p<0.01; ***p<0.001).
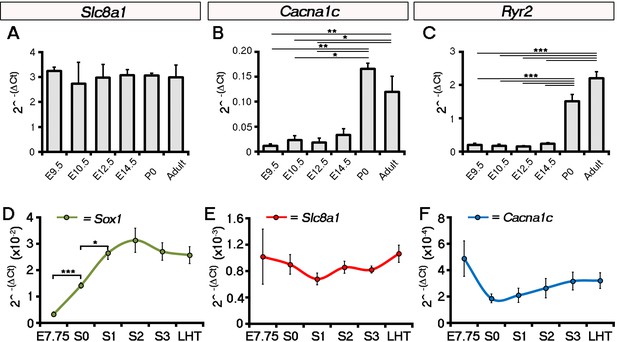
Expression of Slc8a1 and Cacna1c does not increase in the head folds between E7.75 and the LHT stages of development.
Analysis by qRT-PCR showed that expression of Slc8a1 (encoding NCX1 was maintained throughout later stages of embryonic development with no significant change at birth (newborn pups, P0) or into adulthood (A), in contrast to Cacna1c (encoding the LTCC subunit Cav1.2; B) and Ryr2 (encoding the Ryanodine receptor; C) which increased significantly at P0. In the head folds, qRT-PCR revealed a significant increase in the expression of the neural ectoderm marker Sox1 from E7.75 to stage 0 (A), however, in contrast expression of both Slc8a1 (B) and Cacna1c (C) were unchanged over the entire time course. All error bars are mean ± S.E.M, n = 3, A–C; n = 5, D–F. Statistics: one-way ANOVA and Tukey test for multiple comparisons (*p<0.05; **p<0.01; ***p<0.001).
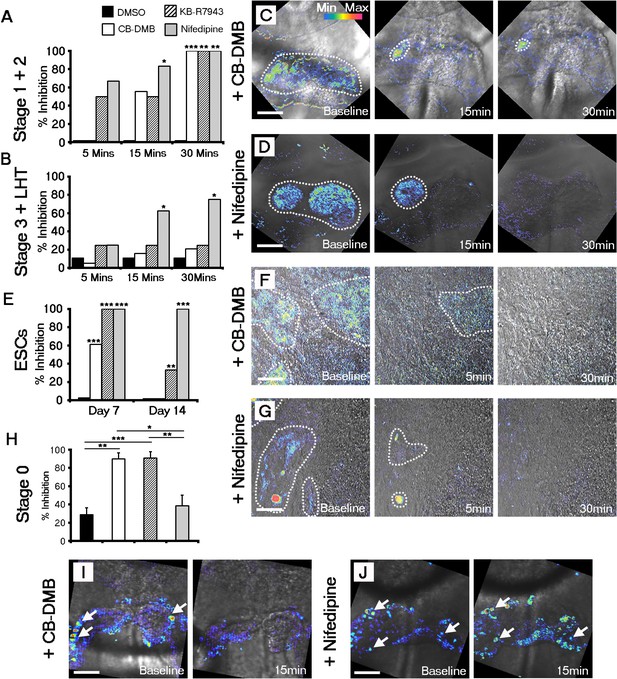
Both NCX1 and LTCC are required for Ca2+ transients associated with beating cardiomyocytes during cardiac crescent development.
Inhibition of Ca2+ transients upon treatment of stage 1-LHT embryos with either NCX1 inhibitors CB-DMB, KB-R7943 or the LTCC inhibitor nifedipine, relative to DMSO control after 5, 15 and 30 min of drug application (A, B). Inhibition of NCX1 with either CB-DMB (20 μM) or KB-R7943 (30 μM) affected only stage 1 and 2 embryos (A), whereas inhibition of LTCC with nifedipine (10 μM) effected both stages 1 and 2 and the later stage 3/LHT (B; stage1/2: DMSO, n = 5; CB-DMB, n = 9; KB-R7943, n = 6; nifedipine, n = 6; stage3/LHT: DMSO, n = 9; CB-DMB, n = 19; KB-R7943, n = 8; nifedipine, n = 8). Time series of Ca2+ transients on stage1-2 embryos at different time points of either CB-DMB or nifedipine treatment, revealed a confinement to the right side of the embryo prior to complete block (C, D). ESC-derived cardiomyocytes at different days of differentiation were treated with the same channel blockers: inhibition of NCX1 with CB-DMB (10 μM) significantly reduced contractions only in day 7 cardiomyocytes, whereas KB-R7943 (30 μM) affected cardiomyocytes at both day 7 and 14 (E). Inhibition of LTCC with nifedipine (10 μM) significantly reduced contractions in both day 7 and 14 cardiomyocytes and to a much greater extent than KB-R7943 at the later stage (F; day 7: DMSO, n = 38; CB-DMB, n = 36; KB-R7943, n = 7; nifedipine, n = 10; day 14: DMSO, n = 25; CB-DMB, n = 36; KB-R7943, n = 15; nifedipine, n = 15). Time series of Ca2+ transients on day 7 ESC-derived cardiomyocytes at different time points of either CB-DMB or nifedipine treatment, revealed a confinement prior to complete block (F, G), equivalent to that observed in the treated embryos (C, D). Treatment of stage 0 embryos prior to the onset of beating with CB-DMB (20 μM) and KB-R7943 (30 μM) resulted in inhibition of slow asynchronous Ca2+ transients after 15 min application relative to baseline (H, I; DMSO, n = 10; CB-DMB, n = 8; KB-R7943, n = 9) whereas treatment with nifedipine (10 μM) had no discernible effect on the slow transients (H, J; nifedipine n = 8), supporting the earlier role for NCX1 in initiating Ca2+ handling and beating. All scale bars 100 μm. Statistics: Freeman-Halton extension of Fisher exact probability test for embryos for embryos; Chi-square test with Bonferroni correction for ESCs (*p<0.05; **p<0.01; ***p<0.001).
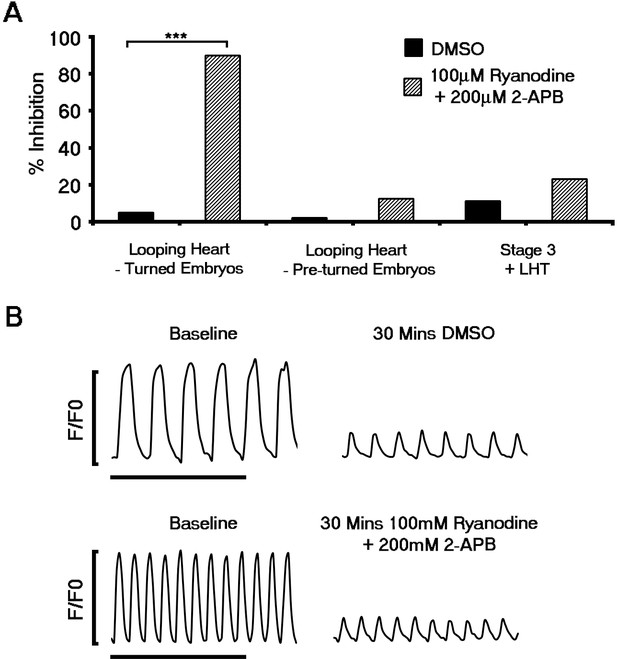
Contribution of the sarcoplasmic reticulum (SR) to Ca2+ transients does not occur until looping stages of heart development.
The functional role of SR derived Ca2+ was assessed using pharmacological inhibition of RyR using Ryanodine and IP3Rs with 2-APB. Inhibition of RyR and IP3 receptors prevented beating in the looping hearts of turned embryos relative to DMSO (Ryanodine + 2-APB, n = 20; DMSO, n = 20; A) and significantly after formation of the LHT. In contrast, application of ryanodine and 2-APB did not inhibit contraction of looping hearts in non-turned embryos (Ryanodine + 2-APB, n = 24; DMSO, n = 13) or block Ca2+ transients in stage 3 cardiac crescents or LHT (Ryanodine + 2-APB, n = 13; DMSO, n = 9), as shown in representative Ca2+ traces (B, scale bar 5 s). All error bars are mean ± S.E.M. Statistics: A, Freeman-Halton extension of Fisher exact probability test (*p<0.05; **p<0.01; ***p<0.001).
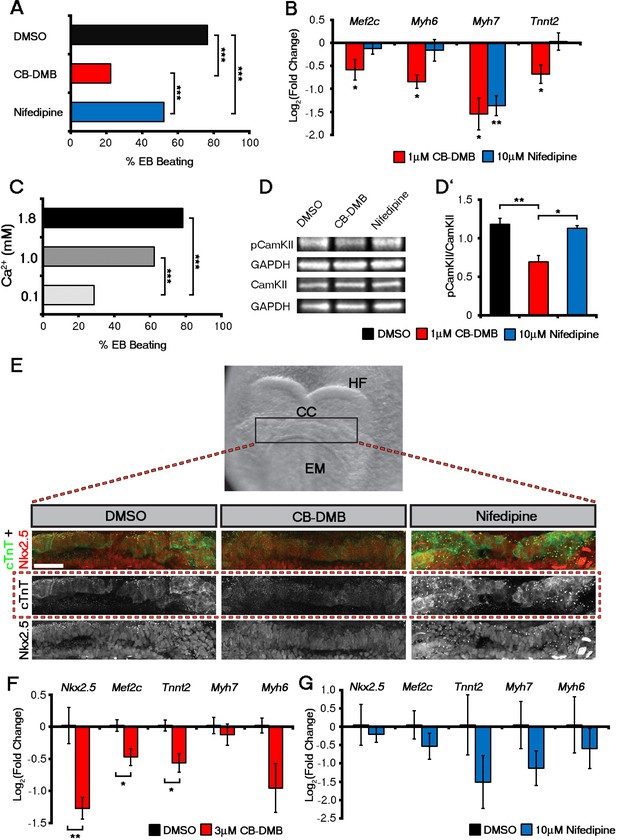
Influx of Ca2+ and CaMKII signalling are required for early and late cardiac gene expression and crescent formation.
Following chronic exposure of embryoid bodies for 14 days to CB-DMB (1 μM) there was a significant decrease in the incidence of beating from 80% to 22% as compared to a reduction to 52% following nifedipine treatment (10 μM) (A; DMSO, n = 196; CB-DMB, n = 152; nifedipine, n = 71). Prolonged exposure resulted in a significant decreases in mature cardiomyocyte genes, Mef2c, Myh6, Myh7 and Tnnt2 (B) gene expression following treatment with CB-DMB (n = 6) but not with nifedipine (n = 6). EBs cultured for 14 days in different concentrations of extracellular Ca2+ (1.8 mM is the normal culture medium concentration) revealed significantly decreased incidence of beating following culture with reduced Ca2+ when assessed in media containing 1.8 mM Ca2+ (C; 1.8 mM, n = 74; 1.0 mM, n = 61; 0.1 mM n = 80). Cultured ESC derived-cardiomyocytes exposed to NCX1 inhibitors effected downstream Ca2+ signalling via alterations in the levels of phosphorylated CaMKII (pCaMKII; D). pCamKII to total CamKII ratio was decreased in the presence of 1 μM CB-DMB as compared to 10 μM nifedipine and DMSO (D’; n = 3). E7.5 embryos were dissected and cultured for 12 hr in media containing either DMSO, nifedipine (10 μM) or CB-DMB (3 μM) and stained for cTnT and Nkx2.5 (E; maximum intensity projections, 30 stacks each). Embryos developed normally in culture, as indicated by head fold formation, coalescence of the cardiac crescent and addition of somites (not shown). Embryos cultured in CB-DMB were delayed in terms of cardiac crescent formation and show a weaker cTnT signal compared to either DMSO alone or nifedipine-treated (E; number of affected embryos: DMSO – 1/7; CB-DMB – 7/8; Nifedipine – 1/6). Cultured E7.5 embryos in the presence of either CB-DMB or nifedipine for 12 hr, revealed that CB-DMB significantly down-regulated the expression of both early Nkx2.5 and Mef2c and late Tnnt2 (F, G) cardiac genes, coincident with impaired cardiac crescent formation, whereas nifedipine-treatment did not appear to have any effect on cardiac gene expression (H). All error bars are mean ± S.E.M. Statistics: B, D, G and H: one-way ANOVA and Tukey test for multiple comparisons; A, C: Chi-square test with a Bonferroni correction for multiple comparisons (*p<0.05; **p<0.01; ***p<0.001). CC, Cardiac crescent; HF, Head folds; EM, Embryonic midline. All scale bars 50 μm.
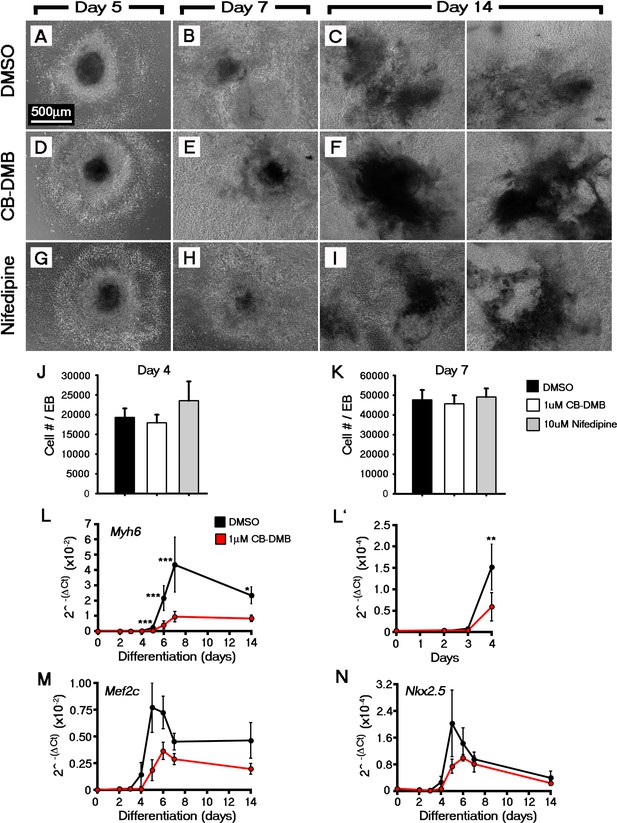
Inhibition of NCX1 or Cav1.2 did not overtly effect embryoid body formation or cell outgrowth during Eomes-GFP ESC differentiation.
Analysis of Eomes-GFP ESC differentiation using qRT-PCR at multiple timepoints revealed the CB-DMB inhibition blocks the expression of key cardiac genes during the formation of cardiomyocytes (Myh6 (A), Mef2c (B), Nkx2.5 (C); n = 3). Treatment of Eomes-GFP ESCs with 1 μm CB-DMB or 10 μm nifedipine did not prevent EB formation or cell outgrowth. Bright field images confirmed that cultures were grossly unaffected by CB-DMB or nifedipine at all stages of differentiation (D–L). The number of cells per EB were also unaffected at both day 4 (DMSO, 20744 cells/EB (n = 9); CB-DMB, 19,184 cells/EB (n = 7); nifedipine, 26,359 cells/EB (n = 3); L) and day 7 (n, DMSO, 46,800 cells/EB (n = 10); CB-DMB, 44,922 cells/EB (n = 8); nifedipine, 50761 cells/EB (n = 3); M). Scale bars: 500 μm. All error bars are mean ± S.E.M. Statistics: A–C, two-way repeated measures ANOVA and Tukey test for multiple comparisons; M–N, one-way ANOVA and Tukey test for multiple comparisons (*p<0.05; **p<0.01; ***p<0.001).
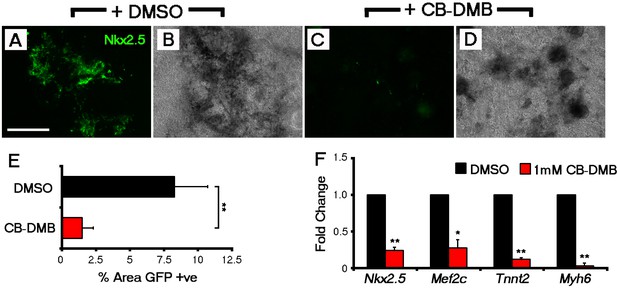
NCX1 blockade from the outset of ESC differentiation reduces the incidence of Nkx2.5+ cardiac progenitors.
Control Nkx2.5-EGFP EBs (DMSO-treated) reveal extensive GFP+ cardiac differentiation after 14 days in culture (A, B) as compared to a loss of GFP+ cells when treated with CB-DMB (day 0–14) (C, D). Fluorescence (A, C) and bright field (B, D) images shown for each treatment group; bright field confirmed the cultures were grossly unaffected by CB-DMB treatment. To quantify levels of GFP expression, the percentage area of Nkx2.5-EGFP regions was calculated from epifluorescent images using Image J and revealed a significant decrease in the percentage area of GFP+ cardiac differentiation (n = 9; E). This was also observed in gene expression as assessed by qRT-PCR, with significant decreases in cardiomyocyte genes, Nkx2.5, Mef2c, Tnnt2 and Myh6 (F), following treatment with CB-DMB (n = 5). All error bars are mean ± S.E.M. Statistics: one-way ANOVA and Tukey test for multiple comparisons (*p<0.05; **p<0.01; ***p<0.001). Scale bar: A, 500 µm.
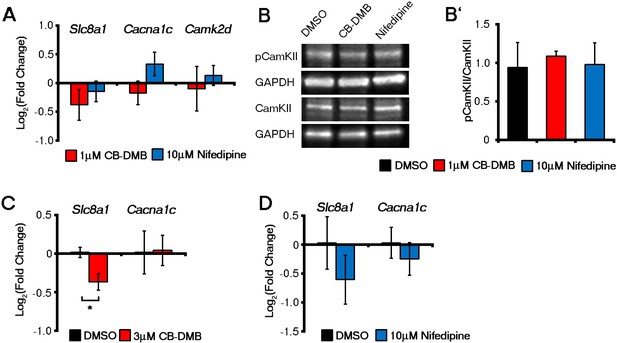
Late administration of NCX1 and LTCC inhibitors did not affect gene expression in ESC-derived cardiomyocytes whereas NCX1 inhibition reduced Slc8a1 in E7.5 embryos ex vivo.
Expression of Slc8a1, Cacna1c or Camk2d (A), as determined by qRT-PCR on day 14 ESC-derived cardiomyocytes were unaffected by culture in the presence of either DMSO, CB-DMB or nifedipine (n = 6). Activation of CaMKII as determined by the ratio of pCaMKII/total CaMKII levels was unaltered in ESC-derived cardiomyocytes after four days in culture in the presence of either DMSO, CB-DMB (1 μM) or nifedipine (10 μM) (B, n = 3). In E7.25 embryos cultured for 12 hr in the presence of either DMSO, CB-DMB or nifedipine, there was a significant decrease in Slc8a1 in embryos cultured in CB-DMB (C, n = 3) consistent with loss of crescent cells (see Figure 5F), but no change with treatment of nifedipine (D, n = 4) consistent with the lack of effect on crescent formation (see Figure 5F). Cacna1c remained unchanged in the presence of either inhibitor (C, D), suggesting non-cardiac (neural fold) expression in CB-DMB treated embryos (see Figure 3). All error bars are mean ± S.E.M. Statistics: one-way ANOVA and Tukey test for multiple comparisons (*p<0.05; **p<0.01; ***p<0.001).
Videos
Representative movie of beating regions in the cardiac crescent at Stage 1.
DIC imaging of a stage 1 embryo highlighting beating regions in the lateral regions (dotted circle) of the developing cardiac crescent. Acquisition was performed at 10 frames per second (fps) with a 20x objective and movie played at 8 fps. Scale bar: 100 µm.
Representative movie of a Ca2+ transient at Stage 1.
Confocal time-lapse of a stage 1 embryo loaded with Cal-520. Cal-520 emission (rainbow) was captured simultaneously with DIC imaging (gray). Embryo the same as that shown in Figure 2D. Acquisition was performed at 10 fps with a 40x water immersion objective. Background fluorescence was removed by subtracting the signal at a resting phase. Movie played at 8 fps to better show the propagation of the Ca2+ transient. Scale bar: 100 µm.
Representative movie of SACOs at Stage 0.
Confocal time-lapse of a stage 0 embryo loaded with Cal-520. Cal-520 emission (rainbow) was captured simultaneously with DIC imaging (gray). Acquisition was performed at 10 fps with a 20x objective. Background fluorescence was removed by subtracting the signal at a resting phase. Scale bar: 100 µm.
Representative high-resolution movie of SACOs at Stage 0.
Confocal time-lapse of a stage 0 embryo loaded with Cal-520. Cal-520 emission (rainbow) was captured simultaneously with DIC imaging (grey). Acquisition was performed at 10 fps with a 20x objective. Movie playback is at 5x original speed. Background fluorescence was removed by subtracting the signal at a resting phase. Scale bar: 10 µm.
Representative movie of a propagating Ca2+ transient at day eight of ESC cardiomyocyte differentiation.
Confocal time-lapse of a day eight EB loaded with Cal-520. Cal-520 emission (rainbow) was captured simultaneously with DIC imaging (grey). Acquisition was performed at 10 fps with a 20x Objective. Background fluorescence was removed by subtracting the signal at a resting phase. Scale bar: 100 µm.
Representative movie of SACOs at day six of ESC cardiomyocyte differentiation.
Confocal time-lapse of a day six EB loaded with Cal-520. Cal-520 emission (rainbow) was captured simultaneously with DIC imaging (grey). Acquisition was performed at 10 fps with a 20x objective. Background fluorescence was removed by subtracting the signal at a resting phase. Scale bar: 10 µm.
Additional files
-
Supplementary file 1
Embryo staging specific to the early developing heart from cardiac crescent to linear heart tube.
(a) Morphological criteria of stages of cardiac crescent development. Different stages of cardiac crescent development (stage 0 to stage 3) were defined based on the ratio between the weight and maximum width of the cardiac crescent. As development progresses the width decreases and the maximum height increases. These stages are a more accurate representation of developmental stage than more widely used staging criteria such as somite number and embryonic day. (b) Statistical comparison of qRT-PCR results of isolated cardiac crescents. Table of p-values obtained from ANOVA and a post-hoc Tukey test on the qRT-PCR results obtained from isolated cardiac crescents, comparing whole embryos pre-cardiac crescent formation (E7.75), at all stages of cardiac crescent development (stage 0 to stage 3) and linear heart tube (LHT) stage. (c) Statistical comparison of qRT-PCR results of isolated head folds. Table of p-values obtained from ANOVA and a post-hoc Tukey test on the qRT-PCR results obtained from isolated head folds, comparing whole embryos pre-cardiac crescent formation (E7.75), at all stages of cardiac crescent development (stage 0 to stage 3) and linear heart tube (LHT) stage. (d) Analysis from SACO inhibition experiments at Stage 0. Table of results detailing individual inhibitor experiments carried out on SACOs at stage 0. Information includes embryo ID, inhibitor used, absolute number of SACOs observed, before and after treatment, ratio of SACOs maintained after treatment, percentage inhibition, area containing SACOs and length of imaging. (e) List of antibodies. List of primary antibodies used for immunostaining and western blot analyses and secondary antibodies used for all the experiments with source and dilution used. (f) List of primers for qRT-PCR. List of primer pairs used for qRT-PCR analysis and respective amplicon size.
- https://doi.org/10.7554/eLife.17113.024





