Rational design of aptazyme riboswitches for efficient control of gene expression in mammalian cells
Figures
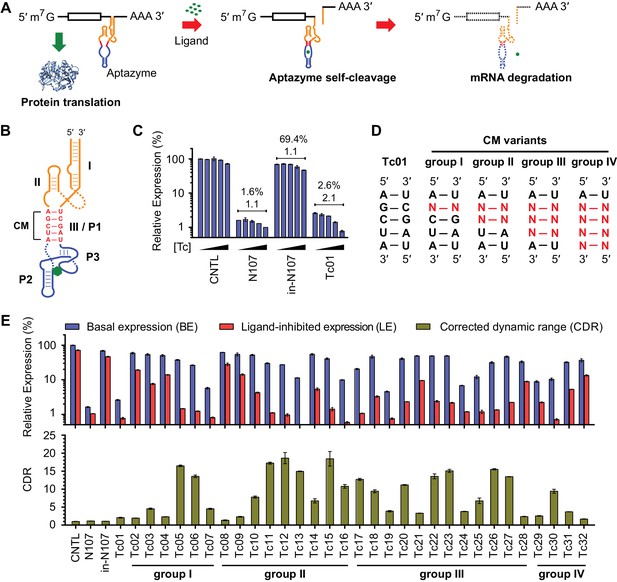
A test panel of 32 tetracycline (tc) aptazymes.
(A) A diagram representing aptazyme-mediated control of gene expression. An aptazyme riboswitch, composed of a ribozyme (orange), a communication module (red) and an aptamer (blue), is inserted at the 3' untranslated region of an mRNA transcript. Ligand binding to the aptamer activates the ribozyme, which cleaves itself in cis, leading to degradation of the RNA message. (B) Secondary structure of the Tc01 aptazyme, composed of a hammerhead ribozyme (orange), a communication module (CM, red), and a Tc-binding aptamer (blue). Tc is represented as a green hexagon. The original stem III of the hammerhead ribozyme and P1 stem of the Tc aptamer are replaced with the communication module, which connects the ribozyme and aptamer and signals the binding state of the aptamer to the ribozyme. (C) Enzymatically active (N107) and inactive (in-N107) forms of the N107 hammerhead ribozyme, or the aptazyme Tc01 were placed 3´ to a Gaussia luciferase (GLuc) gene. HeLa cells transiently transfected with plasmids expressing GLuc regulated by N107 or Tc01 were cultured with 0, 3, 10, 30, or 100 µM Tc for 2 days. Secreted GLuc in the culture supernatant was measured by a luminescence assay. Luciferase expression levels are normalized to that of an expression construct ('CNTL') lacking any regulatory elements. The percentages above the figure indicate 'basal expression' (BE), GLuc expression from each construct in the absence of Tc relative to that from the CNTL control construct. The lower number above each figure indicates the 'corrected dynamic range' (CDR) at 100 µM Tc. CDR is the Tc-induced fold-change in aptazyme-regulated GLuc expression divided by the Tc-induced fold-change in GLuc expression from the CNTL control construct. (D) To gain insight into the relationship between communication-module sequence and CDR, 32 aptazyme variants, differing only in their communication modules and represented here in four groups, were generated. Full communication-module sequences are shown in Figure 1—figure supplement 1B. (E) HeLa cells transiently transfected with plasmids expressing GLuc regulated by these 32 aptazyme variants were cultured in the presence or absence of 100 µM Tc for 2 days. Secreted GLuc in the culture supernatant was analyzed as in (C). The upper panel shows the BE (blue) and 'ligand-inhibited expression' (LE; red) of each variant. LE is the relative luciferase expression of each plasmid in the presence of Tc compared to the luciferase expression level of the CNTL control plasmid in the absence of Tc. The lower panel shows the CDR of each variant. All data shown are representative of two or three independent experiments. All data points represent mean ± S.D. of three biological replicates.
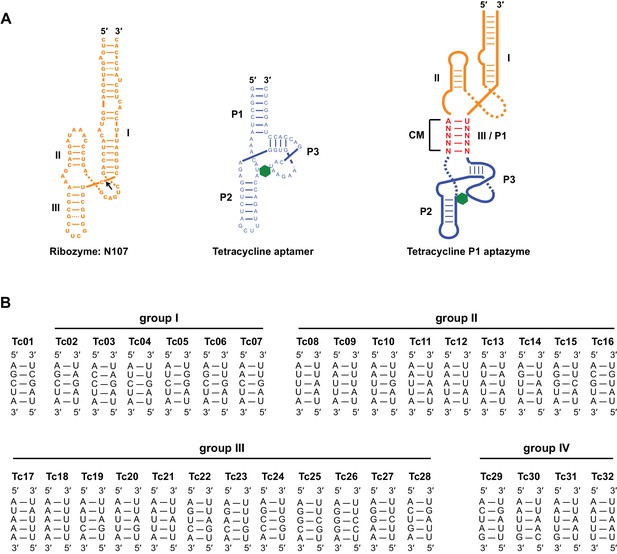
Sequence and secondary structure of test-panel Tc aptazymes.
(A) The secondary structures of the hammerhead ribozyme N107 (left), a Tc-binding RNA aptamer (center), and Tc aptazymes characterized in Figure 1e (right). The N107 hammerhead ribozyme consists of three stem structures (I, II, III) that stabilize a conserved enzymatic core. A black arrow indicates the self-cleavage site. The Tc aptamer also consists of three stems (P1, P2, and P3). Tc is represented as a green hexagon. Tetracycline P1 (Tc-P1) aptazymes were generated by fusing stem III of the ribozyme N107 (orange) with the P1 stem of the Tc aptamer (blue). The five base pairs connecting the ribozyme to the aptamer form a communication module (CM; red) that transmits the ligand-binding state of the aptamer the ribozyme. (B) Communication-module sequences of the 32 test-panel aptazymes are grouped as in Figure 1D and E.
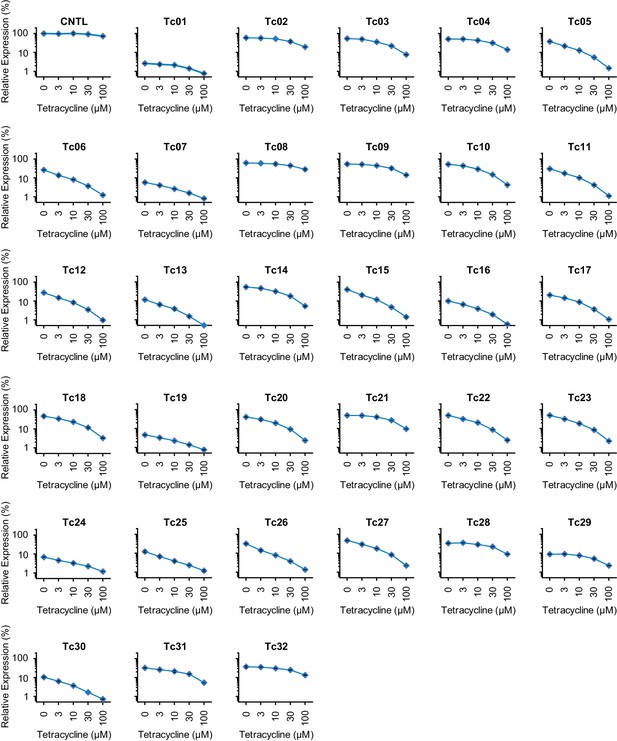
Characterization of test-panel Tc aptazymes.
HeLa cells transiently transfected with plasmids expressing Gaussia luciferase (GLuc) regulated by the indicated aptazymes were cultured for 2 days with the indicated concentrations of Tc. GLuc secreted in the culture supernatant was measured by a luminescence assay. Luciferase expression were normalized to that of an expression construct ('CNTL') lacking any regulatory elements. All data points represent mean ± S.D. Data shown are representative of two or three independent experiments.
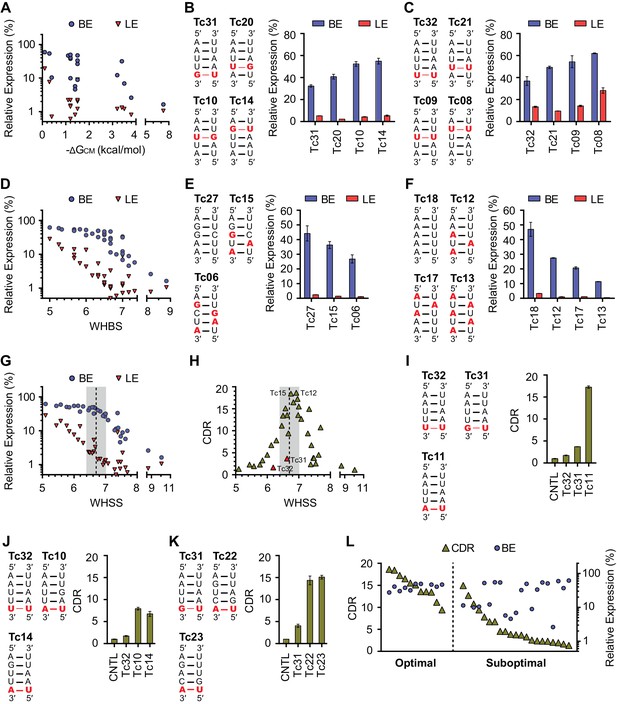
Development of rational-design principles.
(A) The predicted hybrid free energies of the communication modules (ΔGCM) of test-panel aptazymes were calculated with RNAstructure Web Server (Bellaousov et al., 2013). BE (blue circles) and LE (red triangles) were then plotted against the negative ΔGCM value for all aptazymes whose ΔGCM could be determined. Note the poor correlation between ΔGCM and BE or LE. (B–C) BE and LE of the indicated aptazyme variants are shown. Expression levels are normalized to that of the CNTL control construct. The communication-module sequences of these variants are displayed to the left of the figures. Red indicates base pairs different from the consensus. Note that all constructs in each set have the same base pairs, and that order of these pairs impacts BE. (D) Accordingly, every CM base pair was assigned a weight based on its proximity to the ribozyme, with distal bases assigned lower weights. A Weighted Hydrogen Bond Score (WHBS) was then calculated as the weighted sum of hydrogen bonds in the communication module. BE (blue circles) and LE (red triangles) of each variant were plotted against WHBS. Note that WHBS better correlates with the rank order of BE than does ΔGCM or the unweighted sum of hydrogen bond numbers in the communication module (Figure 2—figure supplement 1B). (E–F) The BE and LE of the indicated aptazymes are shown, with their communication-module sequences indicated to the left. Nucleotides with the potential to form inter-strand base-stacking interactions are highlighted as red. (G) Potential inter-strand purine base-stacking interactions were weighted by proximity to the ribozyme, and added to the WHBS to form a modified score (WHSS). BE (blue circles) and LE (red triangles) were plotted against WHSS of the corresponding aptazyme variant. Note that the WHSS better correlates with the rank order of BE than does the WHBS or ΔGCM. An inflection point of the BE at WHSS 6.7 is indicated with a dashed line. Shading indicates a region with WHSS values 6.7 ± 0.3. (H) The CDR values of all 32 test-panel aptazymes were plotted against their WHSS. Note the CDR optimum near the WHSS value of 6.7 (shaded). Aptazymes that exhibited the highest (Tc12, Tc15) or lower-than-expected (Tc31 and Tc32) CDRs are indicated. (I–K) The CDRs of the indicated aptazymes are shown, with their communication-module sequences displayed to the left. Note that aptazymes in each panel share identical sequences (I) or similar WHSS (J,K), but differ in the stability of the communication-module base pair immediately proximal to the Tc aptamer (red). Aptazyme WHSS values range from 6.13 to 6.29 in (J), and from 6.58 to 6.63 in (K). Tc31 and Tc32, with 1 and 0 hydrogen bonds at this position, are outliers in Figure 2H, likely because these weak bonds destabilize the aptamer, and lower its affinity for Tc. (L) Efficient aptazymes met two criteria: a communication-module WHSS value within the range of 6.7 ± 0.3, and an aptamer-proximal communication-module base pair with stability similar to or higher than that of the original aptamer. Aptazymes that meet both criteria have high BE and CDR ('optimal' aptazymes), whereas those that fail to meet one have low BE or narrow CDR ('suboptimal' aptazymes). The CDR and BE of the both optimal and suboptimal aptazymes are ordered by CDR and plotted. Data points in panels A, D, G, H and L represent mean of three biological replicates. Data points in panels B, C, E, F, I, J, and K represent mean ± S.D. of three biological replicates. Numerical data for all the figures are available in Figure 2—source data 1.
-
Figure 2—source data 1
Communication-module scores and expression values of test-panel Tc aptazymes.
Predicated ΔG of the communication module (ΔGCM), total number of communication-module hydrogen bonds (HBN), a Weighted Hydrogen Bond Score (WHBS), and Weighted Hydrogen-bond and Stacking Score (WHSS) of each test-panel Tc aptazyme are listed along with its observed basal expression (BE) at 0 µM Tc, ligand-inhibited expression (LE) at 100 µM Tc and corrected dynamic range (CDR). n.d. indicates that ΔGCM could not be determined. BE and LE are shown as the percentage of GLuc expression normalized to expression of the CNTL control construct lacking any aptazyme. Aptazymes classified into the optimal group in Figure 2l are shaded. BE, LE and CDR are shown as mean ± S.D. Data are representative of two or three independent experiments.
- https://doi.org/10.7554/eLife.18858.006
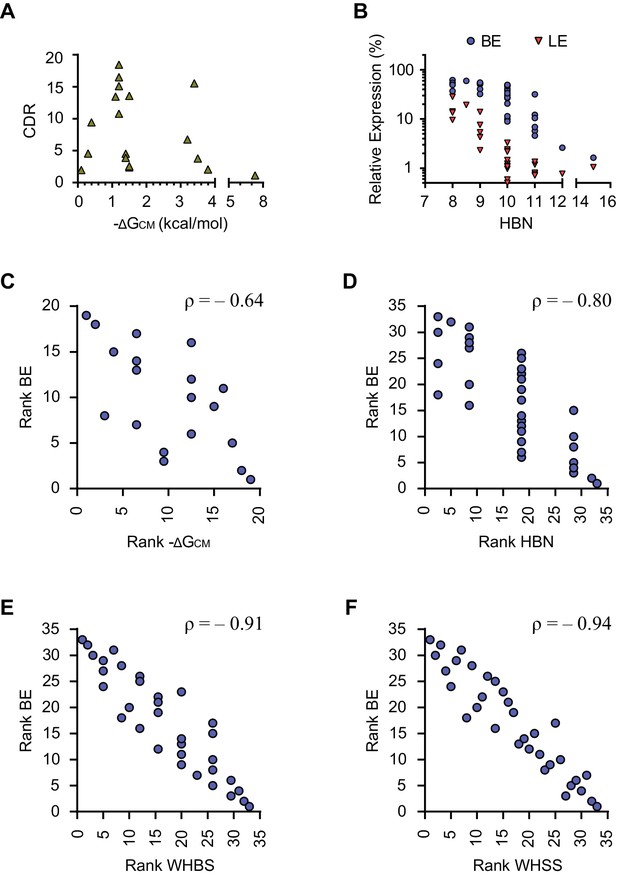
Correlation analysis of energy-like functions describing the communication modules.
(A) The predicted hybrid free energies of the communication modules (ΔGCM) of the test-panel aptazymes were calculated with RNAstructure Web Server. Corrected dynamic ranges (CDR) of the test panel aptazymes were then plotted against the negative ΔGCM values for communication modules whose ΔGCM could be determined. (B) Basal expression (BE) and ligand-inhibited expression (LE) were plotted against total hydrogen bond numbers (HBN) of the communication modules of the test-panel aptazymes. (C) BE and negative ΔGCM of the test-panel aptazymes were ranked separately for Spearman’s rank correlation analysis. Ranked BE were plotted against ranked negative ΔGCM. The Spearman’s rank correlation coefficient (ρ) is shown above the figure. (D–F) Spearman’s rank correlation between BE and HBN (D), Weighted Hydrogen Bond Score (WHBS; E), or Weighted Hydrogen-bond and Stacking Score (WHSS; F) were analyzed as in (C). Note that WHSS correlates better with BE than does ΔGCM, HBN, or WHBS.
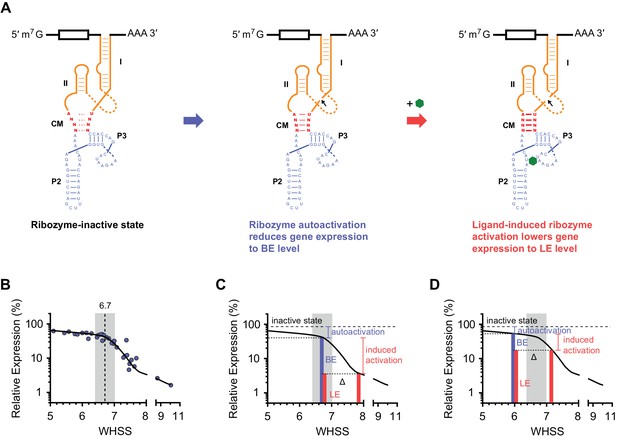
The relationship between aptazyme activation and dynamic range.
(A) When messenger RNAs (mRNA) containing an aptazyme are transcribed, a fraction of the mRNA population will undergo ribozyme autoactivation, resulting in mRNA degradation. Ribozyme autoactivation reduces gene expression to BE and is primarily determined by the communication module sequence (CM; red). Ligand-aptamer binding triggers a conformational change in the aptamer. The communication module signals this change to the ribozyme. Subsequent activation of the ribozyme lowers gene expression to LE. (B) A plot of BE versus WHSS as shown in Figure 2G. The curve shows how gene expression responds to differences in an inferred energy-like function of the communication-module sequence (WHSS), and contains an inflection point at a WHSS value of 6.7. A region near this inflection point (± 0.3) is shaded. (C–D) Diagrams explaining why high levels of ligand-induced ribozyme activation (BE – LE) were observed at WHSS 6.7, the observed inflection point in (B). Energy contributed by the communication module triggers ribozyme autoactivation (blue brackets), lowering gene expression to BE (blue bars) from expression observed with an inactive ribozyme (dashed lines). When the ligand binds the aptamer, it stabilizes the communication module by adding additional energy (Δ), increasing the frequency of ribozyme activation (induced activation, red brackets), and further decreasing gene expression to LE (red bars). Note that identical Δ values have greater impact on the dynamic range of aptazymes with WHSS values near the inflection point (C) than on those with WHSS values far from this point (D)
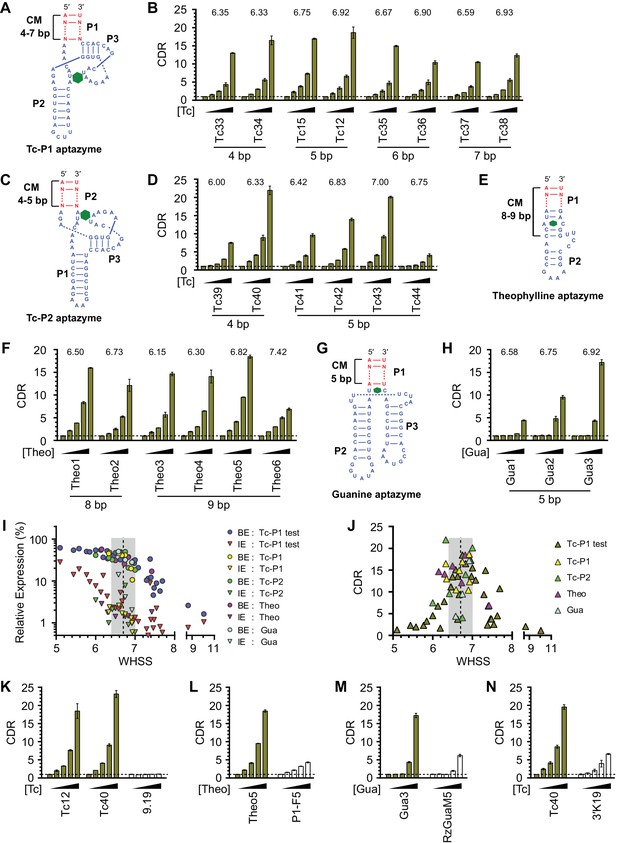
Validation of the design principles.
(A–D) Secondary structures of Tc-P1 (A) or Tc-P2 (C) aptazymes, varying in communication-module lengths and the stem to which the ribozyme is attached, are shown without their ribozyme domains. HeLa cells transiently transfected with plasmids expressing GLuc regulated by Tc-P1 (B) or Tc-P2 (D) aptazymes were cultured with 0, 3, 10, 30, or 100 µM of Tc for 2 days. GLuc secreted into the culture supernatant was measured by a luminescence assay, and CDRs were calculated as in Figure 1E. WHSS values for each aptazyme are shown above figures. () Secondary structure of a theophylline (Theo) aptazyme, with its ribozyme domain omitted. (F) Experiments similar to (B) except that HeLa cells transiently transfected with plasmids expressing GLuc regulated by Theo aptazymes were cultured with 0, 0.1, 0.3, 1.0, 3.0 mM theophylline for 2 days. WHSS values are shown above figure. (G) Secondary structure of a guanine (Gua) aptazyme, with ribozyme omitted. (H) Experiments similar to (B) except that 293T cells transiently transfected with plasmids expressing GLuc regulated by Gua aptazymes were cultured with 0, 7, 20, 60, or 180 µM of guanine for 1 day. WHSS values are shown above figure. BE, LE, and CDR values of all validation-panel aptazymes are available in Figure 3—source data 1. (I) BE and LE of all aptazyme variants from the test panel and the four validation panels were plotted against WHSS of the corresponding variant. (J) CDRs of all aptazymes were similarly plotted against their WHSS values. (K–M) Aptazymes with the highest CDRs from each class (olive) are compared to previously described aptazyme off-switches (white) with the highest reported dynamic ranges. Tc-P1 aptazyme Tc12 and Tc-P2 aptazyme Tc40 are compared to 9.19 (Wittmann and Suess, 2011) (K); Theo5 is compared to P1-F5 (Auslander et al., 2010) (L); and Gua3 aptazyme is compared to RzGuaM5 (Nomura et al., 2012) (M). (N) Tc40 is compared to a previously described aptazyme on-switch, 3’K19 (Beilstein et al., 2015). Data shown are representative of two or three independent experiments. Data points in panels I and J represent mean of three biological replicates, and data points in remaining figures represent mean ± S.D. of three biological replicates.
-
Figure 3—source data 1
Communication-module scores and expression values of validation-panel aptazymes.
BE and LE (100 µM Tc for Tc aptazymes, 3 mM theophylline for Theo aptazymes, or 180 µM guanine for Gua aptazymes) are shown as the percentage of GLuc expression relative to the GLuc expression of the CNTL control construct. BE, LE and CDR are shown as mean ± S.D. Data are representative of two or three independent experiments.
- https://doi.org/10.7554/eLife.18858.010
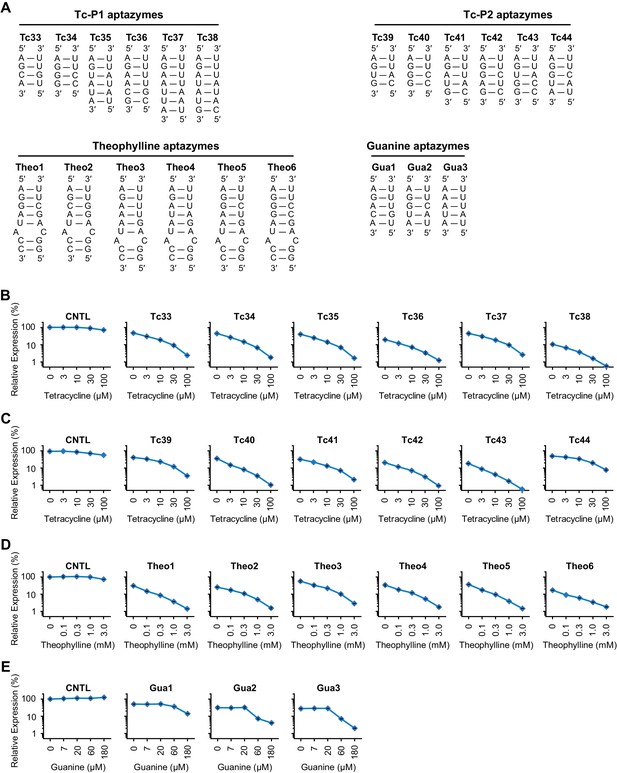
Characterization of four validation panels of aptazymes.
(A) Communication-module sequences of the validation-panel aptazymes tested in Figure 3. (B–D) HeLa cells transiently transfected with plasmids expressing GLuc regulated by Tc-P1 (B), Tc-P2 (C), or Theo (D) validation-panel aptazymes were cultured with the indicated concentrations of tetracycline or theophylline for 2 days. GLuc secreted in the culture supernatant was measured by a luminescence assay. Luciferase expression levels were normalized to that of the CNTL control construct. (E) Experiments similar to (B) except that 293T cells transiently transfected with plasmids expressing GLuc regulated by Gua aptazymes were cultured with the indicated concentrations of guanine for 1 day. All data points represent mean ± S.D. Data shown are representative of two or three independent experiments.
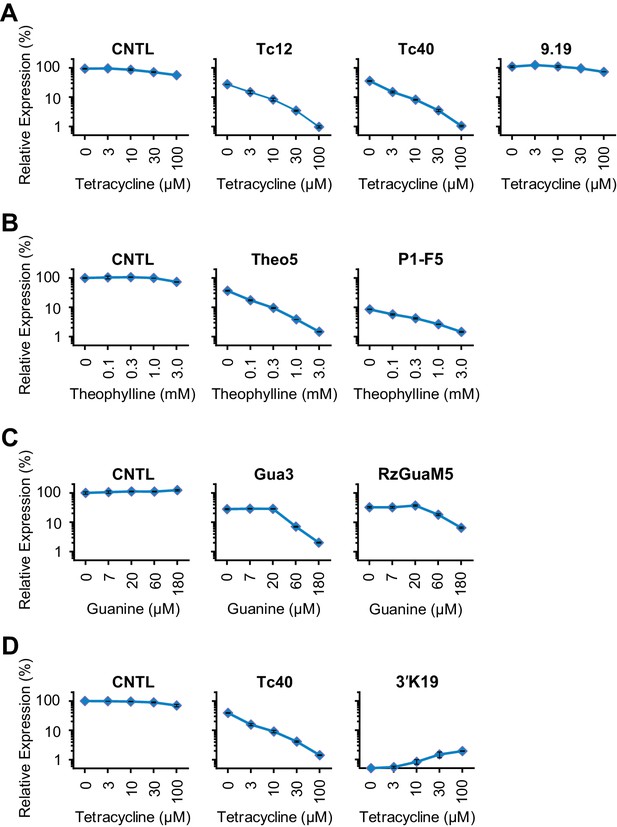
Comparison of our best Tc, Theo, and Gua aptazymes with previously described aptazyme off- or on-switches.
Experiments similar to those shown in Figure 3—figure supplement 1 were performed to measure the CDR of the indicated aptazymes. Tc-P1 aptazyme Tc12 and Tc-P2 aptazyme Tc40 are compared to 9.19 (Wittmann and Suess, 2011) (A); Theo5 is compared to P1-F5 (Auslander et al., 2010) (B); Gua3 aptazyme is compared to RzGuaM5 (Nomura et al., 2012) (C); and Tc40 is compared to the on-switch 3’K19 (Beilstein et al., 2015) (D). All data points represent mean ± S.D. Data shown are representative of two or three independent experiments.
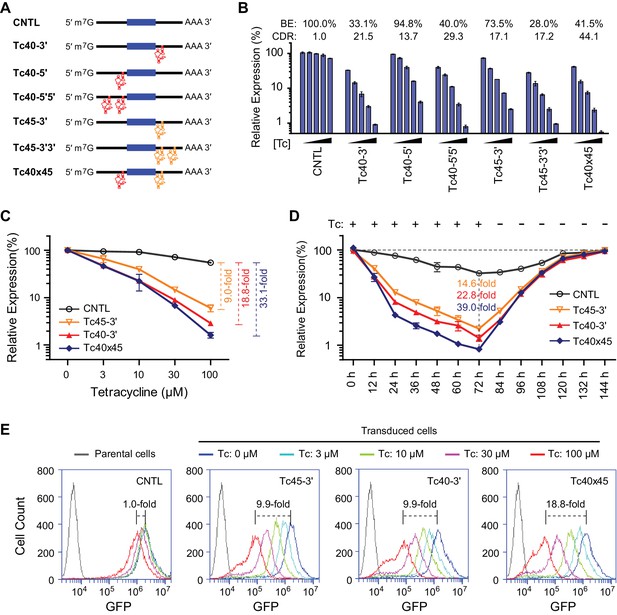
Control of retroviral vectored transgene expression using rationally designed Tc aptazymes.
(A) A representation of the reporter constructs tested. Tc40 (red) or Tc45 (orange) were inserted either upstream (5′) or downstream (3′) of the GLuc gene (filled blue box), either as a single copy (Tc40-5′, Tc40-3′, or Tc45-3'), in tandem repeats (Tc40-5′5′ or Tc45-3′3′), or across the Gluc gene (Tc40×45). (B) Plasmids containing the Tc variants represented in (A) were tested by transient transfection. Transfected HeLa cells were cultured with 0, 3, 10, 30, or 100 µM Tc for 2 days. Secreted GLuc in the culture supernatant was measured by a luminescence assay. Luciferase expression levels are normalized to that of the CNTL control construct. BE and CDR at 100 µM Tc are shown above the figure. (C) HeLa cells were transduced with murine-leukemia virus (MLV) vectors expressing GLuc genes regulated by the indicated Tc aptazyme variants and selected by puromycin. Stably transduced cells were cultured with the indicated concentrations of Tc for 48 hr. Secreted GLuc in the supernatants was measured by luminescence assay. Luciferase expression level was normalized to that observed without Tc. CDR at 100 µM Tc is indicated with brackets at the right. (D) Stable cells characterized in (C) were cultured in 100 µM Tc for 72 hr. Tc was then withdrawn and the cells were cultured for an additional 72 hr. Cell culture supernatants were collected and replaced with fresh medium every 12 hr. Secreted GLuc in the supernatants was measured by luminescence assay. Luciferase expression levels at each time point were normalized to those observed before Tc was added. CDR observed at 72 hr for each aptazyme variant is indicated. (E) Assays similar to those in (C) except that a destabilized enhanced green fluorescent protein (GFP) was used as a reporter, and flow cytometry was used to measure expression. CDR at 100 µM Tc is indicated with brackets. Data shown are representative of two or three independent experiments. Data points in panels B–D represent mean ± S.D. of three biological replicates. Data points in panel E are representative of three biological replicates.
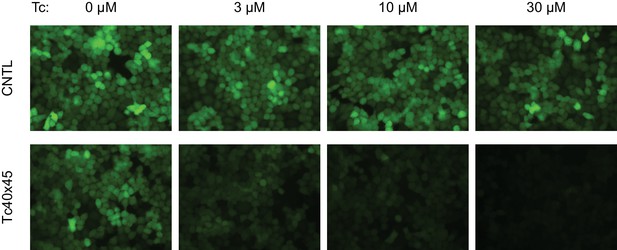
Control of retroviral vectored transgene expression by rationally designed aptazymes.
HeLa cells were transduced with MLV vectors expressing a destabilized enhanced green fluorescent protein (GFP) gene regulated by Tc40×45 or without any aptazymes (CNTL), and selected by puromycin. Stably transduced cells were cultured with the indicated concentration of Tc for 2 days. GFP expression was monitored by fluorescence microscopy at 400× magnification.
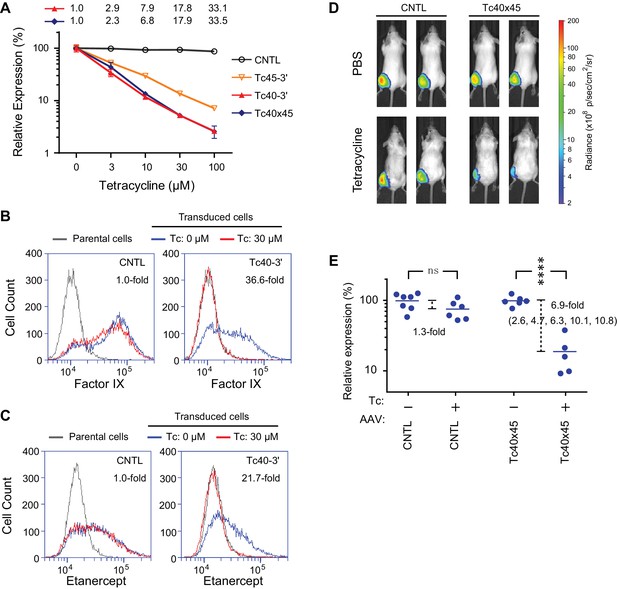
Aptazyme-mediated control of adeno-associated virus (AAV) vectored transgene expression.
(A) HeLa cells were transduced with AAV vectors expressing GLuc regulated by the indicated aptazymes and were cultured with the indicated concentrations of Tc for two days post-transduction. Luciferase expression level was normalized to that observed without Tc. CDR for Tc40-3′ and Tc40×45 is indicated above the figure. (B) HeLa cells were transduced with an AAV vector expressing human coagulation factor IX regulated (right panel) or not (left panel) by Tc40-3′. Cells were then incubated in presence (red) or absence (blue) of 30 µM Tc for two days and analyzed by flow cytometry using intracellular staining with an anti-human factor IX antibody. Grey indicates parental HeLa cells stained with the same antibody. CDR is indicated in the upper right corner of the panels. (C) An experiment similar to that in (B) except that cells were transduced with an AAV vector expressing etanercept (Enbrel), and stained with an anti-human IgG1 Fc antibody. (D) Fourteen 9-week old female BALB/c mice were injected with 1011 genome copies of AAV particles carrying GLuc gene regulated (Tc40×45, seven mice) or not (CNTL, seven mice) by Tc40×45 into the left gastrocnemius muscle. All mice were treated for 3 days with Tc (250 mg/kg) or phosphate-buffered saline (PBS) two weeks post-injection, and imaged for luciferase expression using the Xenogen IVIS In-Vivo Imager. (E) Bioluminescent photon outputs of all the imaged mice were analyzed using the Living Image Software. Relative luciferase expression was calculated as the percentage of bioluminescent photon output relative to the average photon output of PBS-treated mice injected with same AAV vector. Each point represents one mouse, and bars indicate the average expression level for each treatment group. Average fold changes of the luciferase expression between Tc- and PBS-treated groups are indicated. Numbers in the parentheses are the fold regulation of individual Tc-treated mice injected with Tc40×45-regulated vector. Differences between Tc- and PBS-treated groups were significant (two-sided t-test, p<0.0001) in mice injected with the Tc40×45-regulated vector, whereas differences in the control mice were not significant (two-sided t-test, p>0.05). Data shown in (A–D) are representative of two or three independent experiments. Data points in panel A represent mean ± S.D. of three biological replicates. Data points in panel B and C are representative of three biological replicates.
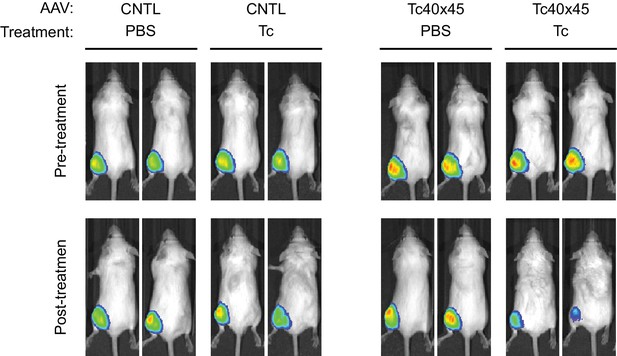
Control of AAV vectored transgene expression by rationally designed aptazymes.
Male 5-week old BALB/c mice were injected into the left gastrocnemius muscle with AAV particles carrying GLuc gene regulated (right panel, 1011 genome copies per mouse) or not (left panel, 3 × 1010 genome copies per mouse) by Tc40×45. Ten days post injection, mice were imaged for luciferase expression using the Xenogen IVIS In-Vivo Imager. Two weeks after injection, mice were treated for 4 days with tetracycline or phosphate-buffered saline (PBS) as indicated. Mice were then imaged for luciferase expression using the In-Vivo Imager.





