A conformational switch regulates the ubiquitin ligase HUWE1
Figures
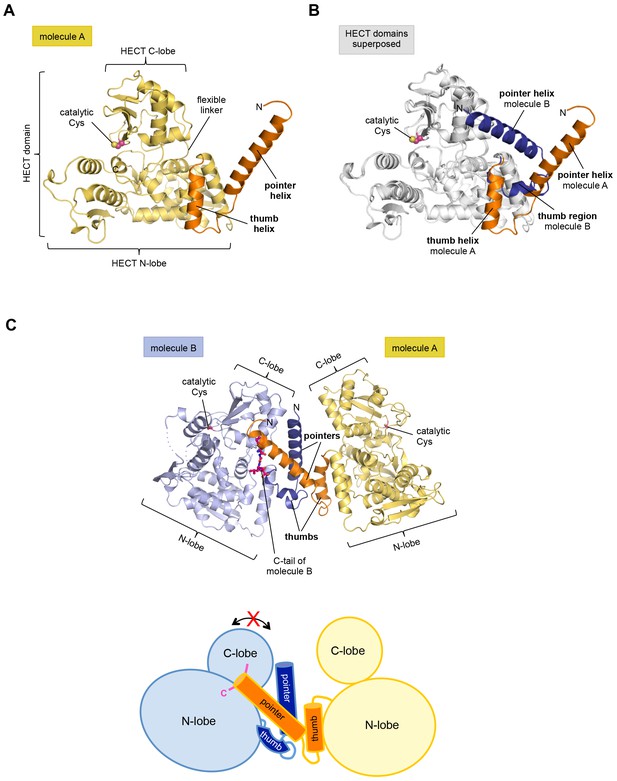
The crystal structure of the C-terminal region of HUWE1 reveals an asymmetric dimer.
(A) Structural organization of HUWE1 (3951–4374), shown for molecule A. The bi-lobal catalytic HECT domain is colored yellow, the previously uncharacterized region orange. (B) Superposition of the HECT domains of molecules A and B (grey). The N-terminal regions are highlighted. (C) Asymmetric dimer formed of molecules A and B in the crystal. The C-terminal residues (4370–4374, ‘C-tail’) of molecule B are highlighted in magenta. Two loop regions in molecule B, distant from the dimer interface, are disordered (dotted lines) (top). Cartoon model of the asymmetric dimer (bottom). The C-lobe and the C-tail (magenta) of molecule B are in a locked conformation (see Figure 2C,D). In all structural figures, the protein backbones are rendered as cartoons and the side chains of relevant residues as ball-and-stick models.

Crystallographic B-factor analysis.
(A) Isotropic Cα B-factors are plotted onto the crystal structure of the HUWE1 dimer. The table lists the mean isotropic B-factors per molecule and region, as calculated by Phenix (Adams et al., 2010). The dimerization regions of molecules A and B show B-factors that are below average overall. For molecule B that has higher B-factors than molecule A, the dimerization region has B-factors below average; for molecule A, the opposite is true. Importantly, however, both dimerization regions are well defined in the electron density. Residues at the N-terminus of the pointer helices that belong to the expression tag, including a disconnected stretch of residues, whose chain identity could not be assigned (see Figure 3—figure supplement 2) are characterized by high B-factors. While the B-factors for molecule B are higher than for molecule A, they fall into an acceptable range of the Wilson B factor for this dataset, as validated by Phenix (Adams et al., 2010). (B) Lattice environment of the asymmetric HUWE1 dimer. Surrounding symmetry mates are shown in ribbon representation (grey). The difference in the B-factors for the two molecules may originate from the different environments of two molecules in the context of the crystal lattice. A significant portion of molecule B – the one that is characterized by relatively high B-factors – faces a large water channel. In contrast, molecule A is more tightly embedded into lattice contacts and, consistently, displays lower B-factors.
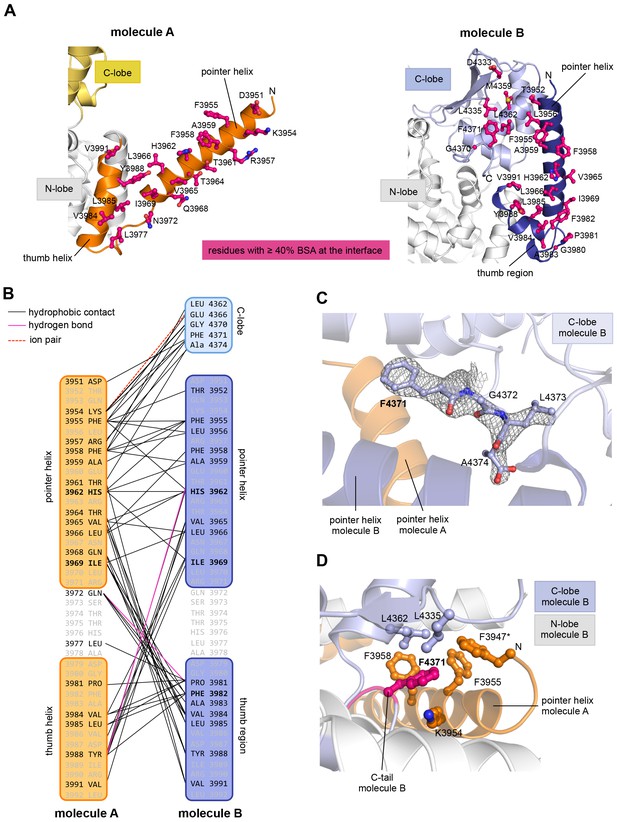
The dimer interface in HUWE1 is extraordinarily hydrophobic and locks the C-lobe of one subunit conformationally.
(A) Open-book-view of the dimerization sites on molecules A and B of HUWE1 (3951–4374). The side chains of interfacing residues with ≥ 40% buried surface area (BSA) are displayed. (B) Schematic representation of the contact network between molecules A and B. The most important hydrophobic interactions are illustrated. Ion pairs and hydrogen bonds are sparse. The mutation sites, His 3962, Ile 3969, and Phe 3982, are marked bold (see Figure 4C–E). Residues that do not make contacts at the dimer interface are colored grey. For the C-lobe of molecule B, only those residues that form significant contacts and that have an interfacing BSA ≥ 40% are shown. (C) The C-tail of molecule B is buried at the dimer interface and well resolved in the electron density. An omit map countered at 1 σ is shown around residues 4370–4374. Phe 4371 corresponds to the conserved ‘−4 Phe’. For the C-tail of molecule A, see Figure 2—figure supplement 2. (D) Detailed view of the hydrophobic environment of −4 Phe (magenta) of molecule B at the dimer interface. The asterisk marks ‘F3947’ as not being part of HUWE1, but of an N-terminal expression tag (see Figure 3—figure supplement 2).
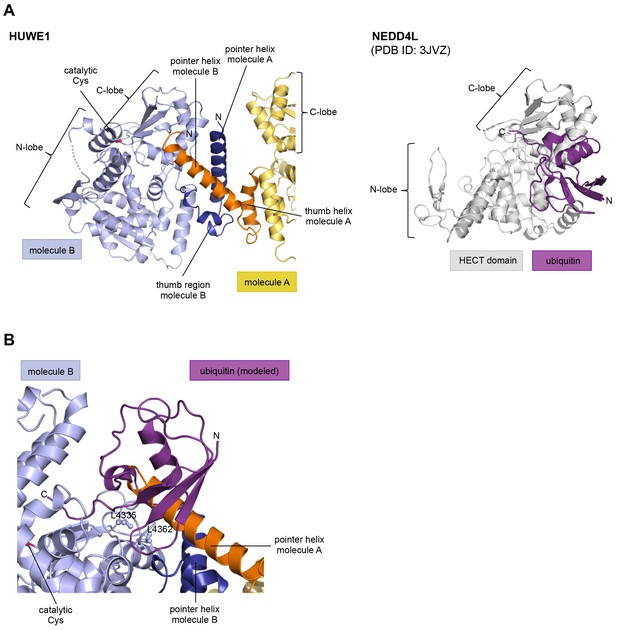
The HUWE1 dimer buries a hydrophobic region that mediates ubiquitin recognition in E3 enzymes of the NEDD4-subfamily.
(A) HUWE1 dimer (left) and NEDD4L-ubiquitin complex (as extracted from PDB ID: 3JVZ [Kamadurai et al., 2009]) (right). The C-lobes of molecule B of HUWE1 and of NEDD4L are shown in the same orientation, illustrating that the interactions of the C-lobe with the pointer helix of molecule A in HUWE1 and with ubiquitin, as seen in the NEDD4-family, are incompatible with each other. (B) Detailed view of the interface between the C-lobe of molecule B and the pointer helix of molecule A of HUWE1, including Leu 4335 and Leu 4362. The homologous residues in NEDD4L (Leu 916 and Leu 943) are important for the binding of ubiquitin (Maspero et al., 2013; Kamadurai et al., 2009, 2013). A ubiquitin molecule, modeled in the same orientation, in which it interacts with the C-lobe of NEDD4-subfamily enzymes clashes with the pointer helix of molecule A.
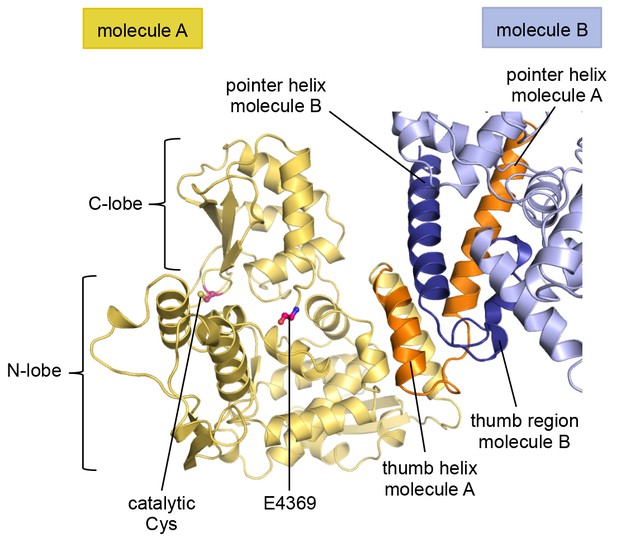
The C-tail of molecule A is disordered in the crystal structure.
In contrast to molecule B (Figure 2C,D), the C-tail of molecule A is disordered in the crystal and could not be modeled. However, the location of the most C-terminal residue that was built, Glu 4369, indicates that the C-tail of molecule A is removed from the dimer interface.
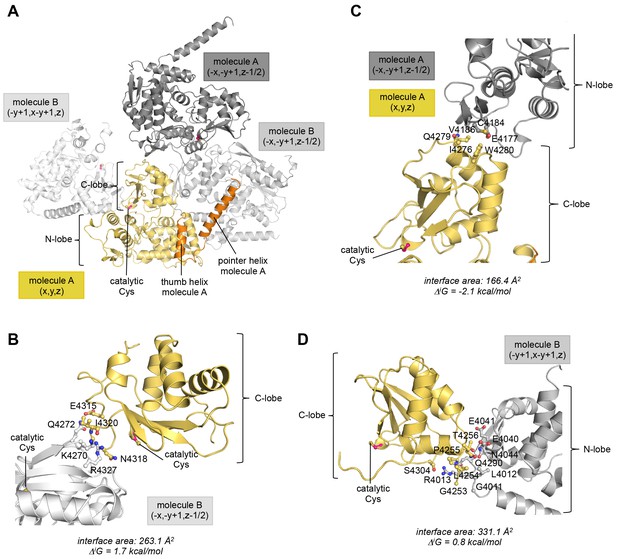
The position of the C-lobe of molecule A in the crystal is influenced by peripheral lattice contacts.
(A) Overview of all molecules that the C-lobe of molecule A contacts in the context of the crystal lattice. The symmetry operations that ought to be applied in order to obtain the respective interface (specified in fractional space relative to the structure position in the PDB file) are provided. (B–D) Details of each lattice interface that the C-lobe of molecule A is involved in. The interface area (difference in the total accessible surface areas of the isolated and interfacing structures divided by 2) and the solvation free energy gain upon formation of the interface, ΔiG, (difference in the total solvation energies of the isolated and the interfacing structures) were calculated with PISA (Krissinel and Henrick, 2007). In (B) residues that form hydrogen bonds at the interface are displayed. In (C) and (D) interfacing residues with >20% buried surface area are displayed. (B) Interface formed between the C-lobe of molecule A and the C-lobe of a neighboring molecule B. The positive ΔiG-value reflects the polar nature of this interface that is mainly made up of hydrogen bonds. These connect Ile 4320 and Lys 4270, Glu 4315 and Gln 4272, Asn 4318 and Arg 4327, and Ile 4320 and Arg 4327, of the C-lobes of molecules A and B, respectively. (C) Interface between the C-lobe of molecule A and the N-lobe of another copy of molecule A. This lattice interface is at least partially hydrophobic. Only few residues are buried on each molecule. These include Ile 4276, Gln 4279, and Trp 4280 of the C-lobe and Glu 4177, Cys 4184, and Val 4186 of the neighboring N-lobe, respectively. When considering additional contacts formed between the N-lobe of molecule A and this copy of molecule B (−x,−y + 1,z−1/2) (not shown), the ΔiG-value amounts to 2.6 kcal/mol, demonstrating that this intermolecular interface is rather polar overall. (D) Interface formed between residues in the C-lobe and linker region of molecule A and the N-lobe of a neighboring copy of molecule B. This interface is rather polar and includes a total of five hydrogen bonds that are formed between Leu 4254 and Gly 4011, Leu 4254 and Arg 4013, Ser 4304 and Arg 4013 (two hydrogen bonds), and Gly 4253 and Arg 4013 of the C-lobe and neighboring N-lobe, respectively. When considering additional contacts formed between the N-lobe of molecule A and this copy of molecule B (−y + 1,x−y +1,z) (not shown), the interaction has an even more positive ΔiG-value of 1.5 kcal/mol.

The asymmetric HUWE1 dimer is compatible, in principle, with the C-lobe of molecule A adopting catalytically competent orientations, as seen for E3 enzymes of the NEDD4-subfamily.
The N-lobe of molecule A in our structure was superposed with the N-lobes of two NEDD4-subfamily enzymes, RSP5 (PDB ID: 4LCD [Kamadurai et al.,2013] and NEDD4L (PDB ID: 3JVZ [Kamadurai et al., 2009]), respectively. The RSP5 structure has the C-lobe in a conformation that is adopted during substrate modification/isopeptide bond formation, while the NEDD4L structure represents a state that occurs during trans-thioesterification. For clarity, only the N-lobe of molecule A of HUWE1 is shown. The three C-lobes are rendered as surfaces in addition to the cartoon models. Note that in this view the C-lobe of RSP5 is located behind the C-lobe of NEDD4L and thus partially obscured.
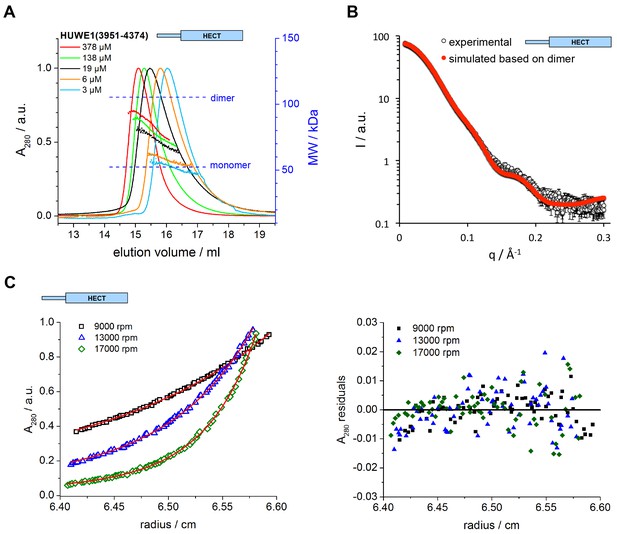
The C-terminal region of HUWE1 dimerizes in solution.
(A) SEC-MALS experiments with the crystallization construct, HUWE1 (3951–4374), at different concentrations. The heights of the absorbance peaks were normalized to a value of 1. The MALS-based MW profiles for each elution peak are shown, along with the calculated MWs of the monomer (53 kDa) and dimer (106 kDa), as references. The MALS-derived MWs in the order of increasing protein concentration are 54, 60, 76, 90, and 93 kDa. (B) SAXS experiment with HUWE1 (3951–4374) at 40 μM concentration. The experimental SAXS intensity, I, plotted over the momentum transfer, q, is compared to a simulated scattering curve, based on the crystal structure of the HUWE1 dimer. The corresponding fit score, χ, is 1.4. Analogous simulations of scattering curves, based on the structures of the two HUWE1 monomers seen in the crystal are shown in Figure 3—figure supplement 1. (C) AUC sedimentation equilibrium experiments with HUWE1 (3951–4374) at 10 μM concentration. The protein distribution across the cell, monitored by absorbance (λ = 280 nm), is shown for three rotation speeds (left). The data were fitted with Sedphat (Houtman et al., 2007) (red line), yielding a KD of 2.9 ± 1.5 μM. The residuals of the fit are shown (right).
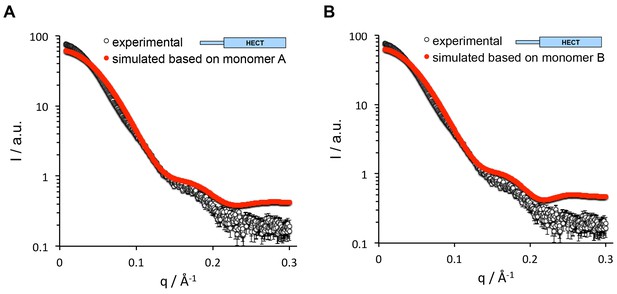
The SAXS data for HUWE1 (3951-4374) are inconsistent with a monomeric state.
The experimental SAXS intensity, I, of HUWE1 (3951–4374) at ~40 μM concentration, plotted over the momentum transfer, q, is compared with simulated scattering curves, based on the crystal structures of the HUWE1 monomers, molecules A (A) and B (B). The corresponding fit scores, χ, amount to 19.9 and 17.7, respectively.
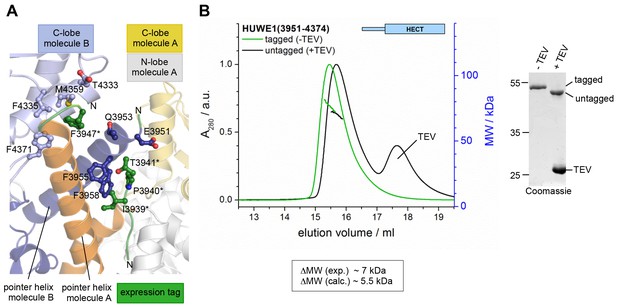
The N-terminal expression tag that was present in the crystallized construct does not induce the dimerization of HUWE1 in solution.
(A) Parts of the N-terminal expression tags of molecules A and B that could be built in the crystal structure are shown in green. The side chains of those residues that form inter- or intramolecular contacts with HUWE1 are shown as ball-and-stick models. Note that, due to a lack of connecting electron density, it could not be determined if the tag region 3936*−3941* belongs to molecule A or B. (B) SEC-MALS experiments with HUWE1 (3951–4374) before and after TEV protease-mediated cleavage of the expression tag (left). The samples were loaded at 75 μM concentration, and the peak heights were normalized to a value of 1. The MALS-derived MW difference between the tagged and untagged protein variants and the calculated MW difference, based on the MW of the expression tag and assuming a constant fraction of dimer, are shown. The completeness of the TEV protease-mediated cleavage reaction was confirmed by SDS PAGE (right). ~2 μg of HUWE1 were loaded after incubation with or without ~3 μg of TEV protease.
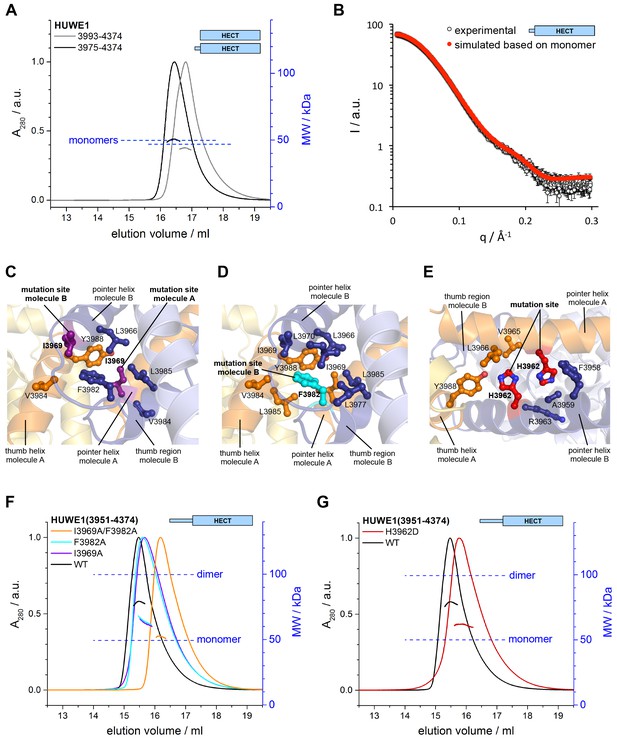
Key contacts in the crystallographic dimer interface mediate the dimerization of HUWE1 in solution.
(A) SEC-MALS experiments with a truncated construct, HUWE1 (3975–4374), that lacks the pointer helix and with the HECT domain (HUWE1 [3993–4374]), at 380 and 400 μM, respectively. The absorbance peak heights were normalized to a value of 1. The MALS-derived MWs are 51 kDa and 44 kDa; the calculated MWs for the monomers are 50 and 47 kDa, respectively. (B) Comparison of the experimental SAXS intensity profile, I(q), of HUWE1 (3975–4374) at 40 μM concentration with a simulated scattering curve, based on the crystal structure of a monomer, as extracted from the crystal structure of HUWE1 (3951–4374). The fit score, χ, is 1.3. (C–E) Detailed views of the hydrophobic side chain environments (within a radius of ~4 Å) of the mutation sites Ile 3969 (purple) of molecules A and B (C), Phe 3982 (cyan) of molecule B (D), and His 3962 (red) of molecules A and B (E). For the environment of Phe 3982 of molecule A, see Figure 4—figure supplement 1. (F) SEC-MALS experiments with WT and mutated variants of HUWE1 (3951–4374) at ~350 μM concentration. The MALS-derived MWs of I3969A, F3982A, and I3969A/F3982A are 63, 64, and 53 kDa, respectively, compared to 79 kDa for the WT. The calculated MW of each protein monomer is 50 kDa. (G) Analogous SEC-MALS experiment with the tumor-associated mutated variant H3962D of HUWE1 (3951–4374). The MALS-derived MW is 62 kDa; the calculated MW of a monomer is 50 kDa. For comparison, the WT profile is displayed (same as in panel F).

In the asymmetric dimer Phe 3982 of molecule A engages in intramolecular interactions.
In contrast to Phe 3982 of molecule B (Figure 4D), Phe 3982 of molecule A does not participate in the dimer interface and is, instead, oriented toward the N-lobe of molecule A. The side chains of key contacting residues of Phe 3982 (within a radius ~4 Å) are shown as ball-and-stick models. Phe 3982 of each molecule is shown in cyan.
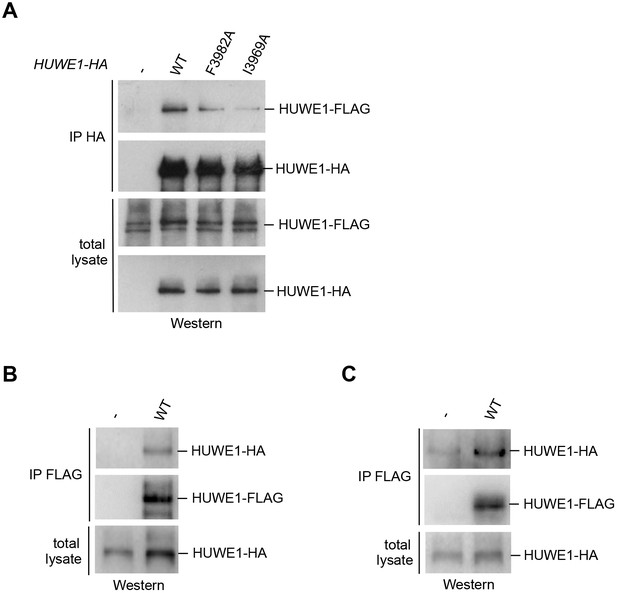
Self-association of HUWE1 occurs in cells and is mediated by the same key contacts as the dimerization observed in vitro.
(A) Co-IP experiments from HeLa cells upon transient transfection with HA- and FLAG-tagged HUWE1 (2474–4374) WT, I3969A, and F3982A. (B) Co-IP experiments from HeLa cells stably expressing HA- and FLAG-tagged HUWE1 (2474–4374) WT that were generated by lentiviral transduction. (C) Co-IP experiments from HeLa cells stably expressing HA- and FLAG-tagged HUWE1 (2474–4374) WT that were generated by transposon-mediated gene delivery.

Comparison of the HUWE1 expression levels in stable cell lines and upon transient transfection.
Expression levels of HUWE1 construct in stable cell lines, generated using lentiviruses or transposons, and upon transient expression (left). The expression is particularly low in the cell lines generated using transposon vectors. To visualize the protein in this system, a second blot is shown, where more cell lysate was loaded (right).
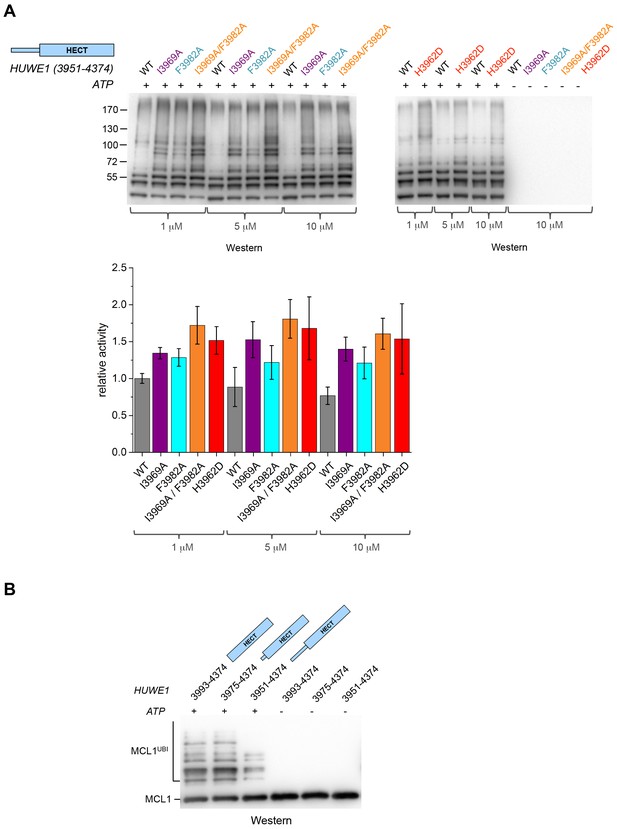
Disruption of the dimer interface enhances the activity of HUWE1 in vitro.
(A) Comparison of the auto-ubiquitination activities of HUWE1 (3951–4374) WT to dimerization-deficient mutated variants at three different concentrations (1, 5, and 10 μM) in vitro, monitored by anti-ubiquitin Western blotting. The reactions contain HUWE1, E2 enzyme (UBCH7), E1 enzyme (UBA1), ubiquitin, and ATP/Mg2+ (for details see Materials and methods). Negative controls were performed without ATP. The band intensities for HUWE1 species modified with at least four ubiquitin molecules were quantified with ImageJ (Abràmoff and Magalhães, 2004) and normalized by the intensity observed in the presence of the lowest concentration of HUWE1 WT. The error bars result from three independent reaction replicates. (B) Comparison of the ubiquitination activities of HUWE1 (3951–4374), HUWE1 (3975–4374) (pointer helix missing), and HUWE1 (3993–4374) (HECT domain only) at 5 μM concentration toward the substrate MCL1, monitored by anti-MCL1 Western blotting (for details, see Materials and methods).
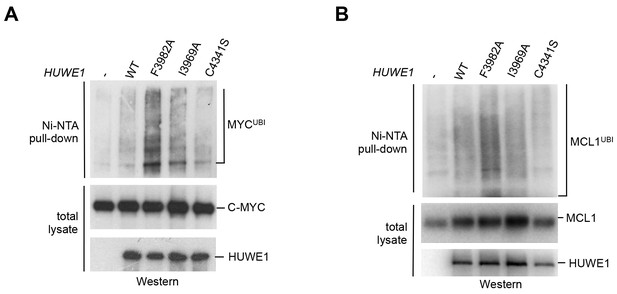
Mutations in the dimer interface enhance the activity of HUWE1 in cells.
(A,B) Cell-based activity assays monitoring the substrate ubiquitination activities of HUWE1 (2474–4374) WT, the dimer interface mutants F3982A and I3969A, and the catalytically dead variant C4341S. HeLa cells were co-transfected with HUWE1 constructs, C-MYC, and MCL1, respectively. For details, see Materials and methods.
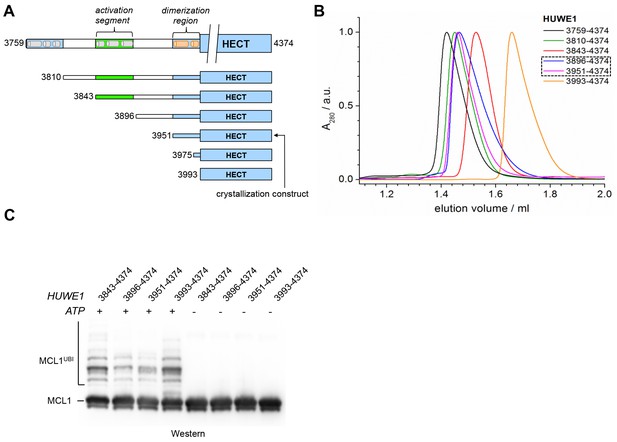
A segment located N-terminal to the dimerization region interferes with the dimerization of HUWE1 and stimulates its activity.
(A) The panel of HUWE1 constructs used in our in vitro studies. Helices >5 residues, as predicted by I-TASSER (Yang et al., 2015) in the structurally uncharacterized regions are colored grey. White boxes mark regions of low sequence complexity. (B) SEC experiments with six individual HUWE1 constructs at 120 μM concentration. The absorbance peak heights were normalized to a value of 1. HUWE1 (3896–4374) and (3951–4374) that dimerize are marked by a dashed box. For the calculated and MALS-derived MWs of the constructs, see Table 2. (C) Activities of different HUWE1 constructs at 5 μM concentration towards MCL1, monitored by anti-MCL1 Western blotting. For details, see Materials and methods.

SAXS data for C-terminal HUWE1 constructs.
SAXS intensity, I, plotted over the momentum transfer, q, and Guinier plots (insert) for HUWE1 (3759–4374) (A), (3810–4374) (B), (3843–4374) (C), (3896–4374) (D), (3951–4374) (E), and (3975–4374) (G) at 40 μM concentration each. The Guinier fits yield estimates of the radius of gyration, Rg (Table 2).
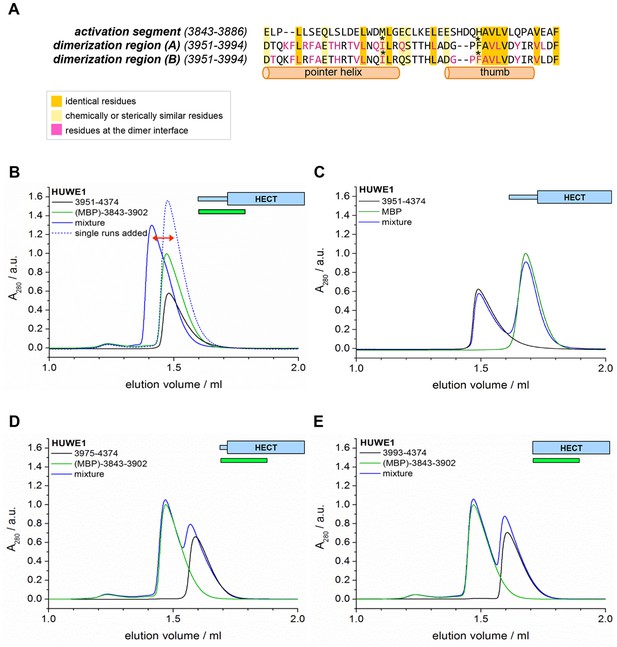
The activation segment displays sequence homology to the dimerization region and interacts with HUWE1 (3951–4374) specifically.
(A) Sequence alignment of the activation segment with the dimerization region of HUWE1, generated by EMBOSS stretcher (Rice et al., 2000). Residues that form contacts at the dimer interface are shown in magenta. The mutations sites, Ile 3969 and Phe 3982, are marked by asterisks. (B,D,E) SEC experiments with different HUWE1 constructs, monitoring interactions with the MBP-tagged activation segment (residues 3843–3902) in trans. SEC profiles of the individual proteins at 120 μM concentration are shown in black and green, respectively; their 1.5:1 molar mixture is shown in blue. The peak heights for the activation segment alone (green) were normalized to a value of 1. In (B) a dashed line represents the expected elution peak for a mixture of non-interacting proteins. A red arrow marks the offset between the calculated and experimental peaks. No significant complex formation is detected in (D) and (E). (C) Analogous SEC experiment using MBP as a negative control.

The MBP-tagged activation segment of HUWE1 is monomeric.
SEC-MALS experiments with the MBP-tagged activation segment (residues 3843–3902) of HUWE1 at 75 μM concentration. The height of the absorbance peak was normalized to a value of 1. The MALS-derived MW of 52 kDa matches the calculated MW of a monomer. A small amount of aggregate (void) and a higher MW contaminant (~200 kDa) are visible in the SEC profile. Experiments performed at protein concentrations of up to 400 μM showed similar results (data not shown).
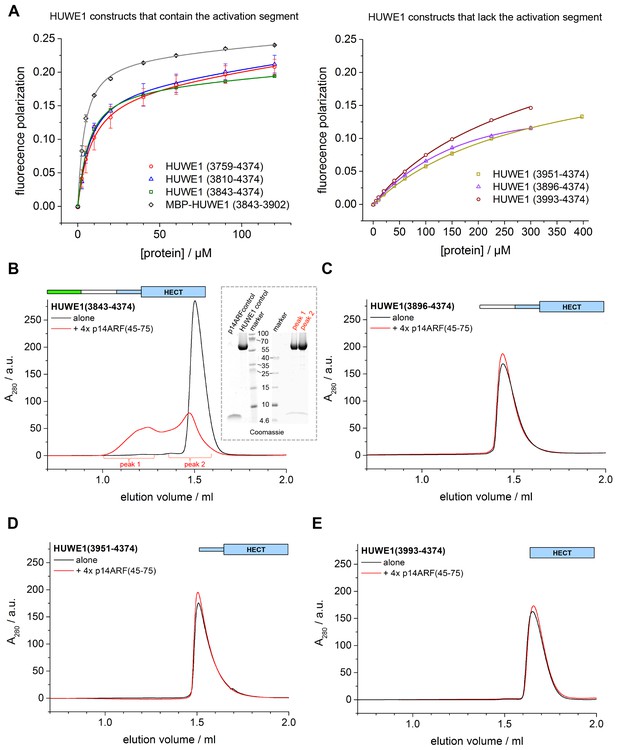
p14ARF interacts with the activation segment of HUWE1 and stimulates oligomerization of HUWE1.
(A) Interactions of a fluorophor-labeled peptide comprising residues 45–64 of p14ARF with HUWE1 constructs of different lengths, monitored by fluorescence polarization. A constant peptide concentration of 1 μM was titrated with different concentrations of protein, as indicated. Constructs containing the activation segment, including the MBP-tagged activation segment per se are shown on the left; constructs lacking the activation segment on the right. Note that the x-axes have different scales. The data points and errors reflect the means and standard deviations obtained from three independent experiment replicates. Data were fitted (lines), as described in the Materials and methods section. The resulting KDapp-values are summarized in Table 3. (B–E) SEC experiments with the specified constructs of HUWE1 at 120 μM concentration in the absence and presence of a four-fold molar excess of p14ARF (45-75). The p14ARF-derived peptide does not contain any aromatic residues and is, therefore, not detected by UV absorbance (λ = 280 nm). That HUWE1 (3843–4374) co-elutes with the p14ARF-derived peptide is demonstrated by SDS-PAGE, using 10–20% tricine gradient gels (insert). Two different molecular weight markers are shown. The HUWE1 bands look somewhat diffuse on this particular gel, since a prior pull-down using nonspecific protein-binding resin (Strataclean, Agilent) was performed in order to visualize the small amount of peptide.
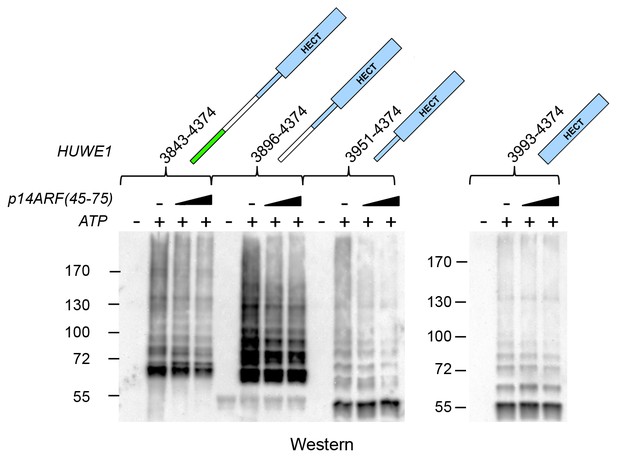
Effects of p14ARF (45-75) on the auto-ubiquitination activities of C-terminal HUWE1 constructs.
Auto-ubiquitination assays with various HUWE1 constructs in vitro, monitored by anti-ubiquitin Western blotting, in the absence and presence of p14ARF(45-75) (increasing concentrations of 15 and 75 μM, indicated by the triangle). Control reactions were performed without ATP. A separate blot is shown for HUWE1 (3993–4374), since the overall auto-ubiquitination activity of this construct is significantly lower than for the longer constructs.
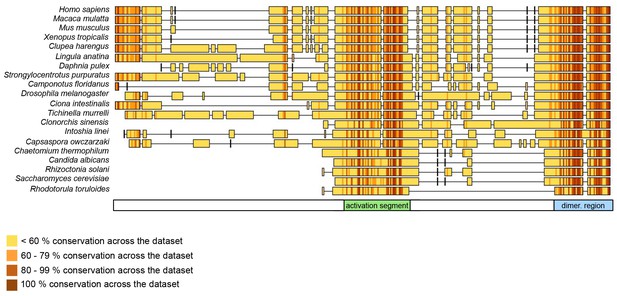
The dimerization region and the activation segment are disproportionally highly conserved across HUWE1 orthologues.
Sequence alignment of 20 selected HUWE1 orthologues, as output by Clustal Omega (RRID:SCR_001591; http://www.ebi.ac.uk/Tools/msa/clustalo/) and sorted by their degree of sequence identity with the HECT domain of the human protein (top row). The illustration was prepared with Geneious Basic (RRID:SCR_010519) (Kearse et al., 2012). The alignment contains regions homologous to residues 3759 to 3992 of human HUWE1, that is, the region preceding the HECT domain.
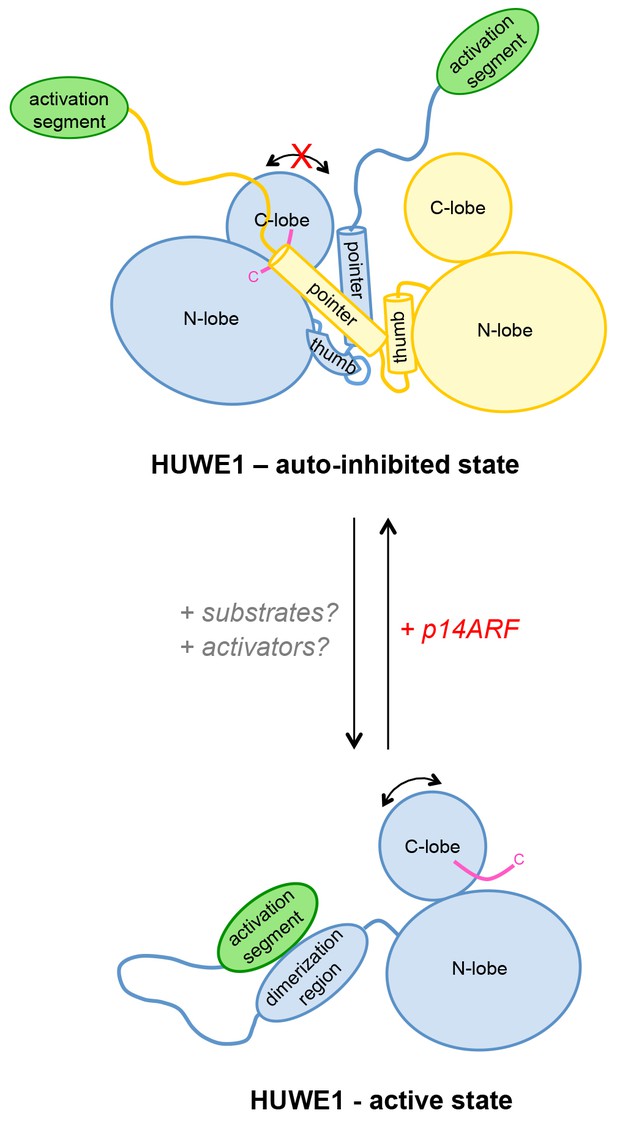
Model of the conformational regulation of HUWE1 and proposed mechanism of its inhibition by p14ARF.
Our studies suggest that the catalytic activity of HUWE1 is regulated conformationally by an intricate balance of inter- and intramolecular interactions. The thumb and pointer helices adjacent to the catalytic HECT domain can mediate the dimerization of HUWE1. The dimer locks the position of the C-lobe, buries the C-terminal tail (magenta), and occludes a putative ubiquitin binding site on the C-lobe of one subunit, hence representing an auto-inhibited state. Alternatively, the dimerization region of HUWE1 can associate with the activation segment in cis, which precludes dimer formation. The activation segment and the dimerization region are separated by a 55-residue linker that is presumably flexible, thus allowing for the re-positioning of the activation segment. In the monomeric state of HUWE1, the C-lobe is mobile with respect to the N-lobe (arrow), and the C-terminal tail may anchor the C-lobe on the N-lobe or interact with substrates, as required for catalytic activity.
The activation segment of HUWE1 presents a major interaction site for a physiological inhibitor of HUWE1, p14ARF. We propose that the binding of p14ARF to the activation segment releases the dimerization region from its intramolecular engagement, thus shifting the conformational equilibrium of HUWE1 toward the auto-inhibited, dimeric state. An additional/alternate mode of inhibition of HUWE1 by p14ARF is independent of the activation segment. We speculate that interactions of HUWE1 with its substrates and positive effectors favor the active state of HUWE1 through interactions that remain to be elucidated.
Tables
X-ray crystallographic data collection and refinement statistics for the structure of HUWE1 (3951–4374). Values in parentheses correspond to the highest resolution shell. For a detailed B-factor analysis, see Figure 1—figure supplement 1.
Data collection | |
|---|---|
Wavelength (Å) | 0.9677 |
Space group | P 63 |
Cell dimensions | |
a,b,c (Å) | 177.46 177.46 106.26 |
α, β, γ (°) | 90.00 90.00 120.00 |
Resolution (Å) | 46.15–2.70 |
Rmerge | 0.058 (0.349) |
I/σ(I) | 16.2 (2.6) |
CC1/2 | 0.998 (0.828) |
Completeness (%) | 99.4 (98.5) |
Redundancy | 4.2 (2.9) |
Wilson B factor | 51.1 |
Refinement | |
Resolution | 46.15–2.70 |
Reflections used | 52025 |
Rfree reflections | 2549 |
Rwork/Rfree | 0.197/0.227 |
No. of non-hydrogen atoms | 7015 |
Protein | 6983 |
Ligands | 32 |
Average B factors | 70.84 |
Protein | 70.97 |
Solvent | 44.19 |
RMSD from ideality | |
Bonds (Å) | 0.005 |
Angles (°) | 0.845 |
Ramachandran statistics | |
Favored (%) | 97.97% |
Disallowed (%) | 0.00% |
MolProbity clash score | 1.89 |
Summary of SEC-MALS-derived MWs and SAXS-derived Rg-values for C-terminal HUWE1 constructs. For SEC-MALS studies the proteins were injected at a concentration of ~375 μM. The SAXS experiments were performed at ~40 μM concentration (see Figure 8—figure supplement 1).
HUWE1 construct | MW (MALS) / kDa | MW (calc.) / kDa | Rg (SAXS) / Å |
|---|---|---|---|
3759–4374 | 68 | 74 | 35 |
3810–4374 | 61 | 68 | 35 |
3843–4374 | 60 | 65 | 31 |
3896–4374 | 92 | 60 | 39 |
3951–4374 | 93 | 53 | 36 |
3975–4374 | 51 | 50 | 27 |
3993–4374 | 44 | 47 | / |
Apparent dissociation constants, KDapp, of the interactions between p14ARF and C-terminal HUWE1 constructs, as derived from fluorescence polarization studies with a fluorophor-labeled p14ARF-derived peptide (residues 45–64) (see Figure 10A).
HUWE1 construct | KDapp/µM |
|---|---|
3759–4374 | 7.7 ± 0.4 |
3810–4374 | 6 ± 1 |
3843–4374 | 5.2 ± 0.2 |
3896–4374 | 340 ± 60 |
3951–4374 | 280 ± 40 |
3993–4374 | 230 ± 40 |
(MBP-) 3843–3902 | 3.6 ± 0.2 |





