Prefabrication of a ribosomal protein subcomplex essential for eukaryotic ribosome formation
Figures
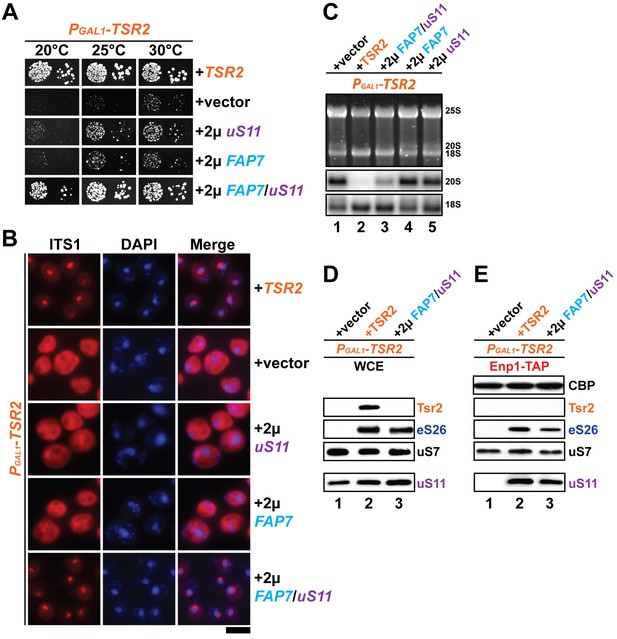
Co-overexpression of Fap7 and the r-protein uS11 bypasses in vivo requirement for Tsr2.
(A) Co-overexpression of FAP7 and uS11 rescues slow growth of Tsr2-depleted cells. The PGAL1-TSR2 strain was transformed with indicated plasmids and spotted in 10-fold dilutions on selective and repressive glucose-containing plates and grown at indicated temperatures for 3–7 days. (B) Co-overexpression of FAP7 and uS11 rescues 20S pre-rRNA processing defect of Tsr2-depleted cells. Indicated strains were grown to mid-log phase at 25°C in selective glucose-containing medium. 20S pre-rRNA was localized by FISH using a Cy3-labeled oligonucleotide complementary to the 5’ portion of ITS1 (red). Nuclear and and mitochondrial DNA was stained with DAPI (blue). Scale bar = 5 µm. (C) Indicated strains were grown as in above for extraction of total RNA and then analyzed by GelRed straining and Northern blotting using probes against 20S and 18S rRNAs. (D) Co-overexpression of FAP7 and uS11 restores protein levels of eS26 in Tsr2-depleted cells. Whole cell extracts (WCE) were prepared from indicated strains and subjected to Western analysis. (E) Co-overexpression of FAP7 and uS11 restores co-enrichment of eS26 with Enp1-TAP in Tsr2-depleted cells. Enp1-TAP was isolated from indicated strains and subjected to Western analysis. uS7 and CBP (TAP-tag) protein levels served as loading controls.
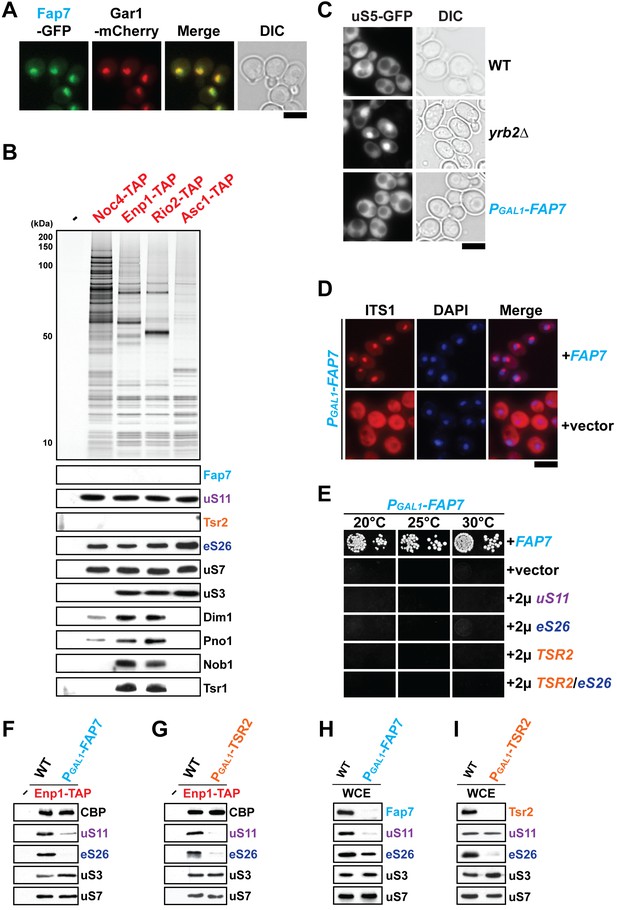
uS11 and eS26 depend upon each other for their incorporation into pre-ribosomes.
(A) Fap7 is a nuclear localized protein. Fap7 and the nucleolar protein Gar1 were endogenously tagged with GFP and mCherry, respectively, and then strains expressing them were grown to mid-log phase at 25°C. Localization of Fap7-GFP and Gar1-mCherry was visualized by fluorescence microscopy. Scale bar = 5 µm. (B) uS11 and eS26, but not Fap7 or Tsr2 co-enrich with pre-ribosomal particles along the 40S maturation pathway. Pre-ribosomal particles in the 40S maturation pathway were purified using the indicated TAP-tagged baits. Calmodulin-eluates were analyzed by Silver staining, and Western analyses was performed using the indicated antibodies. The r-protein uS7 served as loading control for the TAPs. (C) Fap7-depletion does not impair pre-40S subunit nuclear export. The indicated strains expressing uS5-GFP were grown in repressive glucose-containing medium to mid-log phase at 25°C. Localization of uS5-GFP was monitored by fluorescence microscopy. Scale bar = 5 µm. (D) Fap7-depleted cells accumulate immature 20S pre-rRNA in the cytoplasm. Indicated strains were grown to mid-log phase at 25°C in selective glucose-containing medium. 20S pre-rRNA was localized by FISH using a Cy3-labeled oligonucleotide complementary to the 5’ portion of ITS1 (red). Nuclear and mitochondrial DNA was stained with DAPI (blue). Scale bar = 5 µm. (E) Slow growth of Fap7-depleted cells cannot be rescued by either (co-)overexpression of uS11, eS26 or TSR2. The PGAL1-FAP7 strain was transformed with indicated plasmids and spotted in 10-fold dilutions on selective and repressive glucose-containing plates and grown at indicated temperatures for 3–7 days. (F) Efficient recruitment of uS11 and eS26 to the 90S requires Fap7. Enp1-TAP was isolated from indicated strains and subjected to Western analysis. (G) Efficient recruitment of uS11 and eS26 to the 90S requires Tsr2. Enp1-TAP was isolated from indicated strains and subjected to Western analysis. Protein levels of uS7 and CBP (TAP-tag) served as loading control. (H) uS11, but not eS26 levels are strongly reduced in Fap7-depleted cells. (I) eS26, but not uS11 levels are strongly reduced in Tsr2-depleted cells. Whole cell extracts (WCE) were prepared from indicated strains and subjected to Western analysis. uS7 protein levels served as a loading control.
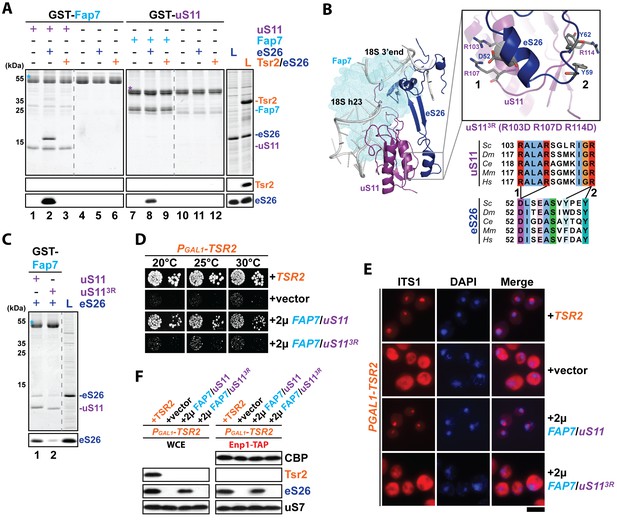
The Fap7:uS11 complex recruits eS26.
(A) The Fap7:uS11 complex directly binds eS26 but not Tsr2:eS26 in vitro. Recombinant GST-tagged Fap7 and uS11 complexes were immobilized on Glutathione Sepharose beads before incubation with E. coli lysate containing recombinant eS26 or Tsr2:eS26. After washing away unbound proteins, beads were eluted and analyzed by SDS-PAGE followed by Coomassie Blue staining and Western blotting. L = 10% input. GST-baits are indicated with asterisks. (B) Conserved protein-protein interactions between uS11 and eS26 on the mature 40S subunit. uS11 (purple) binds to 18S rRNA helix 23 (grey) with its C-terminus embedded into the 18S rRNA 3’end. eS26 (blue) forms major contacts with uS11 via (1) a salt-bridge between uS11 R103 / R107 and eS26 D52 and (2) a hydrophobic side-chain interaction between uS11 R114 and eS26 Y59 / Y62. Yeast Fap7 which acts as a RNA mimic for helix 23 (light blue) was modeled into the structure by superposition of the Fap7:uS11 complex from Pyrococcus abyssi (PDB: 4CVN; Loc'h et al., 2014) onto yeast uS11 from the mature 40S ribosome (PDB: 4 V88; Ben-Shem et al., 2011). For the uS113R mutant, R103 / R107 / R114 were each mutated to aspartates. The sequences for the following organisms were aligned: Saccharomyces cerevisiae (Sc), Drosophila melanogaster (Dm), Caenorhabditis elegans (Ce), Mus musculus (Mm) and Homo sapiens (Hm) (C) The uS113R mutant is impaired in recruiting eS26 in vitro. Recombinant GST-Fap7:uS11 or GST-Fap7:uS113R was incubated with eS26 and analyzed by SDS-PAGE followed by Coomassie Blue staining and Western blotting. L = 100% input. GST-baits are indicated with asterisks. (D) Co-overexpression of FAP7 and uS113R does not rescue the slow growth of Tsr2-depleted cells. The PGAL1-TSR2 strain was transformed with indicated plasmids and spotted in 10-fold dilutions on selective and repressive glucose-containing plates and grown at indicated temperatures for 3–7 days. (E) Co-overexpression of FAP7 and uS113R does not rescue the 20S pre-rRNA processing defect of Tsr2-depleted cells. Indicated strains were grown to mid-log phase at 25°C in selective glucose-containing medium. 20S pre-rRNA was localized by FISH using a Cy3-labeled oligonucleotide complementary to the 5’ portion of ITS1 (red). Nuclear and and mitochondrial DNA was stained with DAPI (blue). Scale bar = 5 µm. (F) Co-overexpression of FAP7 and uS113R does not restore protein levels of eS26 and neither allows co-enrichment with Enp1-TAP in Tsr2-depleted cells. Whole cell extracts (WCE, left panel) and Enp1-TAP (right panel) were prepared and isolated, respectively, from indicated strains and subjected to Western analysis. uS7 and CBP (TAP-tag) protein levels served as loading controls.
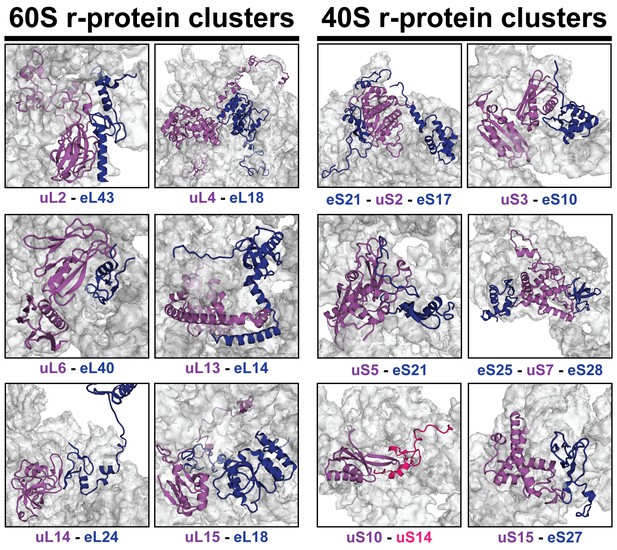
r-proteins cluster on the eukaryotic ribosome via tertiary contacts.
Eukaryotic r-proteins interacting via tertiary protein-protein interactions are each highlighted in purple/red (universally conserved) or blue colors (archaea or eukaryote-specific) on the surface of ribosomal RNA (grey). Figures were constructed using the structure of the mature S. cerevisiae 80S ribosome (PDB: 4 V88; Ben-Shem et al., 2011).
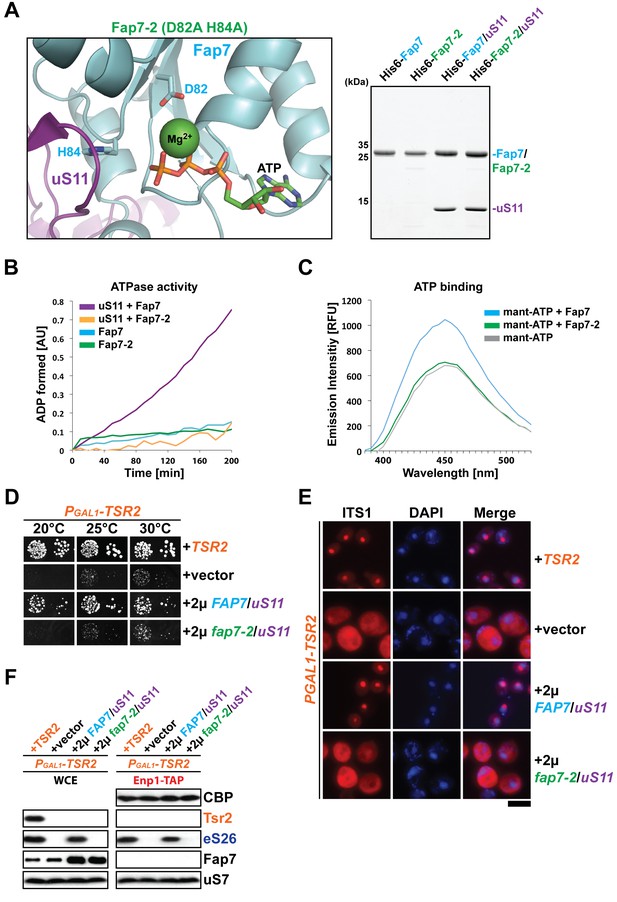
Fap7 ATPase activity is required to bypass Tsr2 requirement.
(A) To generate a catalytically inactive ATPase mutant for in vitro assays, the conserved D82 and H84 residues in the Walker B motif of Fap7 were each mutated to alanine (left panel), and expressed and purified from E. coli (right panel). The structure shows the Fap7:uS11 complex from P. abyssi in complex with ATP and Mg2+ (PDB: 4CW7; Loc'h et al., 2014). (B) Fap7-2 is deficient in ATP hydrolysis. The ATPase activity of indicated Fap7 and uS11 complexes was monitored by an indirect enzyme assay that detects formation of ADP, which is then coupled to β-NADH oxidation via the action of pyruvate kinase (PK) and lactate dehydrogenase (LDH). The decrease in β-NADH absorbance over time was measured at 340 nm in arbitrary units (AU). (C) Fap7-2 is deficient in ATP binding. Fluorescent mant-ATP was mixed with purified Fap7 or Fap7-2 and the change in emission spectra upon nucleotide binding was measured in relative fluorescence units (RFUs). (D) Co-overexpression of fap7-2 and uS11 does not rescue the slow growth of Tsr2-depleted cells. The PGAL1-TSR2 strain was transformed with indicated plasmids and spotted in 10-fold dilutions on selective and repressive glucose-containing plates and grown at indicated temperatures for 3–7 days. (E) Co-overexpression of fap7-2 and uS11 does not rescue the 20S pre-rRNA processing defect of Tsr2-depleted cells. Indicated strains were grown to mid-log phase at 25°C in selective glucose-containing medium. Localization of 20S pre-rRNA was analyzed by FISH using a Cy3-labeled oligonucleotide complementary to the 5’ portion of ITS1 (red). Nuclear and and mitochondrial DNA was stained with DAPI (blue). Scale bar = 5 µm. (F) Co-overexpression of fap7-2 and uS11 does not restore protein levels of eS26 and neither allows co-enrichment with Enp1-TAP in Tsr2-depleted cells. Whole cell extracts (WCE, left panel) and Enp1-TAP (right panel) were prepared and isolated, respectively, from indicated strains and subjected to Western analysis. uS7 and CBP (TAP-tag) protein levels served as loading controls.
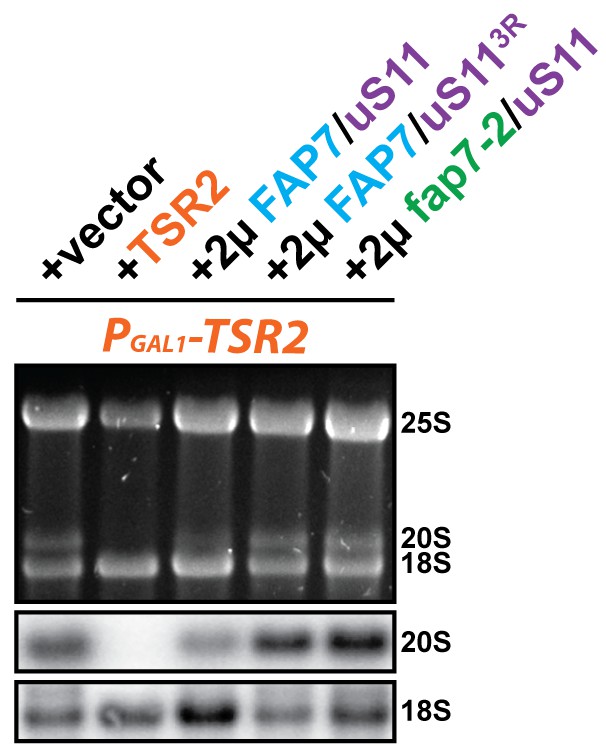
Co-overexpression of FAP7 and uS113R or fap7-2 and uS11 does not rescue the 20S pre-rRNA processing defect of Tsr2-depleted cells.
Indicated strains were grown to mid-log phase at 25°C in selective glucose-containing medium for extraction of total RNA and then analyzed by GelRed straining and Northern blotting using probes against 20S and 18S rRNAs.
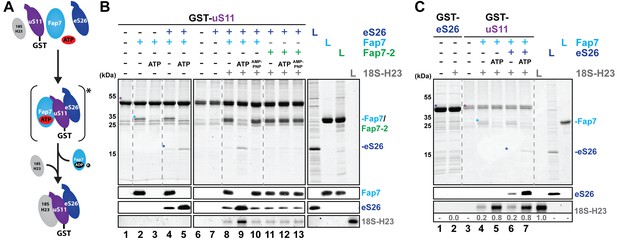
Fap7 ATPase activity organizes and recruits the uS11:eS26 subcomplex to helix 23 of 18S rRNA.
(A) Schematic for ATP-dependent loading of uS11:eS26 onto helix 23 of 18S rRNA by Fap7. Asterisk indicates a potential intermediate Fap7-ATP:uS11:eS26 complex. (B) Recombinant GST-uS11 was immobilized on Glutathione Sepharose beads before incubation with Fap7 or Fap7-2, eS26, ATP or non-hydrolysable ATP analog AMP-PNP and 18S helix 23 rRNA. (C) Recombinant GST-eS26 was immobilized on Glutathione Sepharose beads before incubation with 18S helix 23 rRNA. After washing away unbound proteins and RNA, beads were eluted and analyzed by SDS-PAGE followed by Coomassie Blue staining and Western blotting. For analysis of RNA, samples were Phenol-extracted and separated by denaturing PAGE followed by GelRed staining. RNA was quantified with respect to the input. L = 10% input. GST-baits and pulled-down proteins are indicated with asterisks.
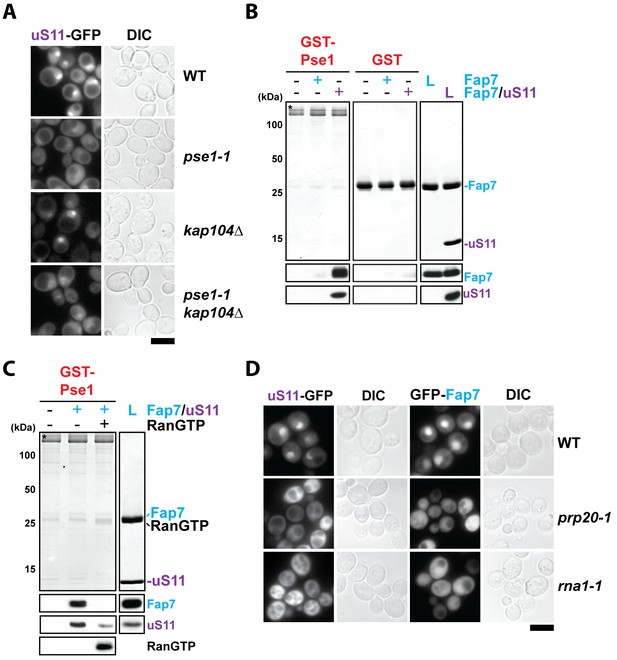
RanGTP-dependent nuclear import of uS11 and Fap7.
(A) Nuclear uptake of uS11-GFP is impaired in pse1-1 and pse1-1 kap104∆ mutants. Strains expressing uS11-GFP were grown in synthetic media at 25°C to mid-log phase. Ts-mutant strains were then shifted to 37°C for 4 hr and localization of uS11-GFP was analyzed by fluorescence microscopy. Scale bar = 5 µm. (B) Fap7:uS11, but not Fap7, interacts with Pse1. Recombinant GST-Pse1 or GST alone were immobilized on Glutathione Sepharose beads and incubated with buffer, purified Fap7 or Fap7:uS11 for 1 hr. After washing away unbound proteins, beads were eluted and analyzed by SDS-PAGE followed by Coomassie Blue staining and Western blotting. L = 10% input. GST-baits are indicated with asterisks. RanGTP (His6-Gsp1Q71L-GTP) efficiently releases Fap7:uS11 from Pse1. GST-Pse1:Fap7:uS11 complexes immobilized on Glutathione Sepharose were incubated with either buffer alone or with RanGTP for 10 min and then analyzed as described above. (D) Nuclear uptake of uS11-GFP and GFP-Fap7 is impaired in prp20-1 and rna1-1 mutants. Strains expressing uS11-GFP or GFP-Fap7 were grown in synthetic media at 25°C to mid-log phase. Ts-mutant strains were then shifted to 37°C for 4 hr and localization of uS11-GFP and GFP-Fap7 was analyzed by fluorescence microscopy. Scale bar = 5 µm.
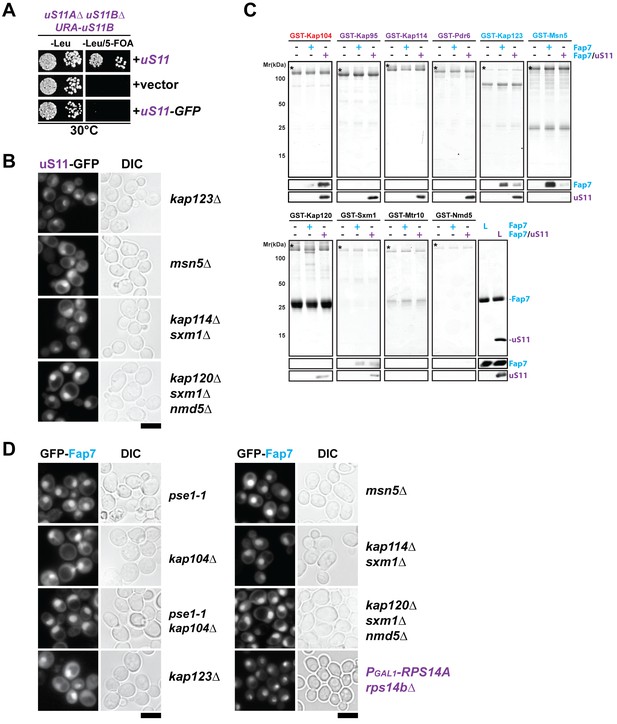
Nuclear import of uS11 and Fap7.
(A) A C-terminally GFP-tagged uS11 cannot complement the lethality of a rps14a∆rps14b∆ double knockout strain. The rps14a∆rps14b∆ pURA-uS11B shuffle strain was transformed with a plasmid encoding C-terminally tagged uS11-GFP and spotted in 10-fold dilutions on selective Leucine-deficient media plates with or without FOA to select against pURA-uS11B and grown at 30°C for 3–4 days. (B) Nuclear uptake of uS11-GFP is not impaired in kap123∆, msn5∆, kap114∆ sxm1∆ and kap120∆ sxm1∆ nmd5∆ mutants. Strains expressing uS11-GFP were grown in synthetic media at 25°C to mid-log phase. Ts-mutant strains were then shifted to 37°C for 4 hr and localization of uS11-GFP was analyzed by fluorescence microscopy. Scale bar = 5 µm. (C) Fap7:uS11, but not Fap7, interacts with Kap104. uS11, but not Fap7 interacts with Kap95, Kap114, Pdr6. Fap7 and uS11 alone, but not the Fap7:uS11 dimer interact with Kap123. Only Fap7 interacts with Msn5. Recombinant GST-tagged importins were immobilized on Glutathione Sepharose beads and incubated with buffer, purified Fap7 or Fap7:uS11 for 1 hr. After washing away unbound proteins, beads were eluted and analyzed by SDS-PAGE followed by Coomassie Blue staining and Western blotting. L = 10% input. GST-baits are indicated with asterisks. (D) Nuclear uptake of GFP-Fap7 is not impaired in several importin mutants and upon depletion of uS11. Strains expressing GFP-Fap7 were grown in synthetic media at 25°C to mid-log phase. Ts-mutant strains were then shifted to 37°C for 4 hr while PGal1-RPS14A rps14b∆ was grown overnight at 25°C in repressive glucose medium. Localization of GFP-Fap7 was analyzed by fluorescence microscopy. Scale bar = 5 µm.

Fap7:uS11 facilitates RanGTP-dependent release of eS26 from Pse1.
(A) eS26 and Fap7:uS11 interact with Pse1. Recombinant GST-Pse1 was immobilized on Glutathione Sepharose beads and incubated with buffer or eS26 alone. After washing away unbound proteins, beads were further incubated in presence of buffer or Fap7:uS11 for 1 hr and washed again. Beads were then eluted and analyzed by SDS-PAGE followed by Coomassie Blue staining and Western blotting. L = 10% input. GST-baits are indicated with asterisks. (B) Fap7:uS11 facilitates RanGTP-dependent release of eS26. Immobilized GST-Pse1 was incubated first with eS26 alone (left panel). After washing away unbound proteins, beads were further incubated with RanGTP alone or RanGTP together with either Fap7:uS11 or Fap7:uS113R for indicated time points and washed again. Beads were then eluted and analyzed as described above. Right panel shows a schematic for how Fap7:uS11 facilitates RanGTP-dependent release of eS26 from Pse1.
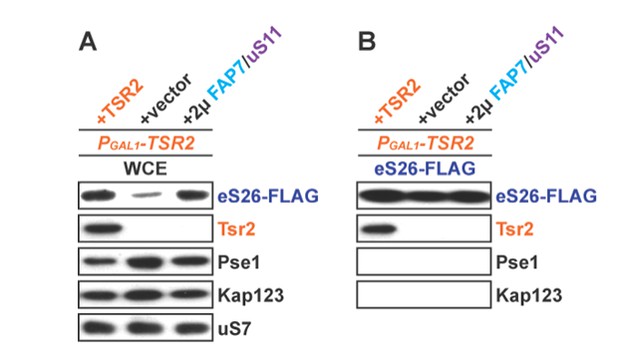
(A) Whole cell extracts (WCE) were prepared from indicated strains and subjected to Western analysis using antibodies directed against Pse1 and Kap123. uS7 served as loading control. (B) eS26-FLAG was isolated from indicated strains. Equal levels of eS26-FLAG were loaded and separated on a SDS-gel and subjected to Western analysis using the indicated antibodies.
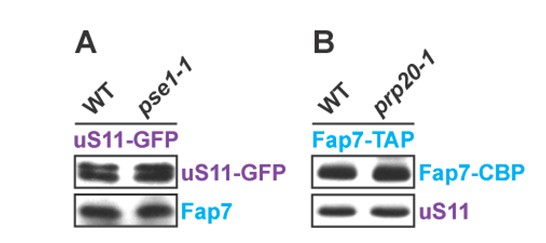
(A) Wild-type and pse1-1 strains expressing uS11-GFP were grown in synthetic media at 25°C to mid-log phase and then shifted to 37°C for 4 h. uS11-GFP was then immuno-precipitated and subjected to Western analysis.(B) Wild-type and prp20-1 strains expressing Fap7-TAP were grown in synthetic media at 25°C to mid-log phase and then shifted to 37°C for 4 h. Fap7-TAP was then purified and subjected to Western analysis.
Additional files
-
Supplementary file 1
Yeast strains and plasmids used in this study.
(A) Yeast strains used in this study. (B) Plasmids used in this study.
- https://doi.org/10.7554/eLife.21755.012





