Tracking zoonotic pathogens using blood-sucking flies as 'flying syringes'
Figures
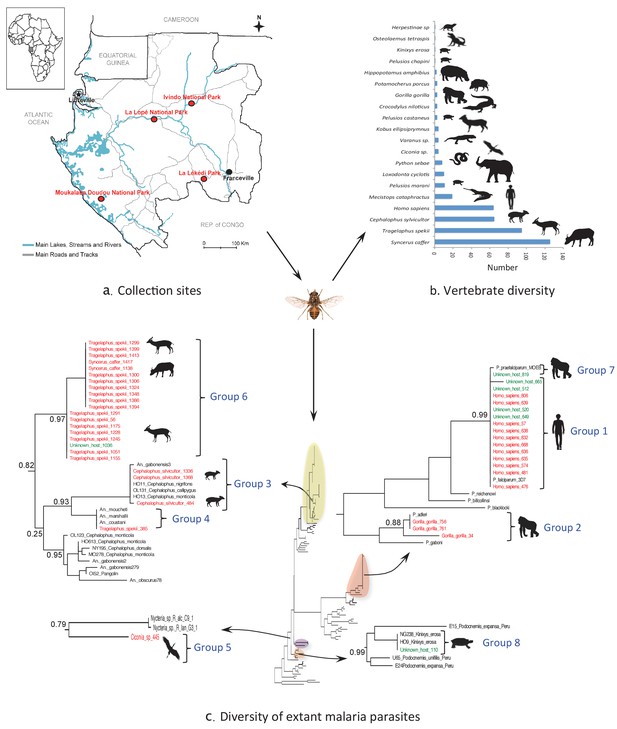
Monitoring vertebrate haemosporidian diversity using haematophagous flies.
(a) Localization of the sampling sites (red dots) in Gabon (Central Africa). (b) Number of blood meals originating from the different vertebrate species. (c) Position within the Cytb phylogeny of the haemosporidian Cytb sequences PCR-amplified from the blood meals of engorged flies with identified hosts (red isolates) and unidentified hosts (green isolates). Black isolates: references (Table 4). Bootstrap values at important nodes are shown.
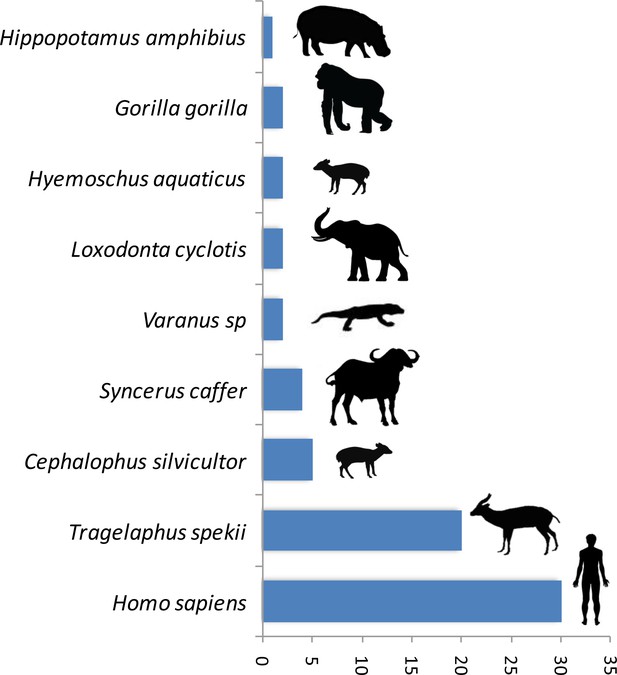
Number of blood meals identified using the shorter PCR system of Boessenkool et al. (2012) out of the previously unidentified 89 blood meals.
https://doi.org/10.7554/eLife.22069.007Tables
Number and proportion of specimens captured per fly species. The number of engorged specimens and blood meals identified in each fly species are also indicated.
| Fly species | Number of collected specimens | Proportion (%) | Number of engorged specimens | Number of identified blood meals |
|---|---|---|---|---|
| Glossinidae | 2252 | 54.94 | 1218 | 423 |
| Glossina caliginea | 144 | 3.51 | 87 | 33 |
| G. fusca congolensis | 210 | 5.12 | 104 | 42 |
| G. fuscipes fuscipes | 290 | 7.07 | 214 | 93 |
| G. pallicera newsteadi | 157 | 3.83 | 97 | 37 |
| G. palpalis palpalis | 1372 | 33.47 | 662 | 218 |
| G. tabaniformis | 79 | 1.93 | 54 | 0 |
| Muscidae | 1362 | 33.23 | 9 | 4 |
| Stomoxys calcitrans | 245 | 5.98 | 5 | 2 |
| S. inornatus | 334 | 8.14 | 0 | 0 |
| S. niger niger | 253 | 6.17 | 4 | 2 |
| S. niger bilineatus | 224 | 5.46 | 0 | 0 |
| S. omega omega | 197 | 4.81 | 0 | 0 |
| S. transvittatus | 109 | 2.66 | 0 | 0 |
| Tabanidae | 485 | 11.83 | 3 | 1 |
| Ancala sp | 41 | 1 | 0 | 0 |
| Atylotus sp | 104 | 2.53 | 0 | 0 |
| Chrysops sp | 156 | 3.81 | 3 | 1 |
| Haematopota sp | 13 | 0.31 | 0 | 0 |
| Tabanus par | 52 | 1.27 | 0 | 0 |
| Tabanus taeniola | 120 | 2.93 | 0 | 0 |
| Total | 4099 | 100 | 1230 | 428 |
Number and origin of blood meals according to the fly species (Fsp), park and climatic season.
| Number of identified blood meals by fly species (Fsp) | |||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Moukalaba-Doudou | Lopé | ||||||||||||||||||||||||
| Rainy season | Dry season | Rainy season | Dry season | ||||||||||||||||||||||
| Taxonomic group/Order/Family | Host species | N° Identified | Fsp1 | Fsp2 | Fsp3 | Fsp4 | Fsp5 | Fsp1 | Fsp2 | Fsp3 | Fsp4 | Fsp5 | Fsp8 | Fsp1 | Fsp2 | Fsp3 | Fsp4 | Fsp5 | Fsp1 | Fsp2 | Fsp3 | Fsp4 | Fsp5 | Fsp6 | Fsp7 |
| Mammals | |||||||||||||||||||||||||
| Artiodactyla | 295 | 1 | 4 | 3 | 8 | 1 | 1 | 7 | 1 | 14 | 2 | 1 | 2 | 3 | 3 | 4 | 1 | 3 | |||||||
| Bovidae | Cephalophus silvicultor | 65 | |||||||||||||||||||||||
| kobus ellipsiprymnus | 4 | 3 | 1 | ||||||||||||||||||||||
| Syncerus caffer | 126 | 3 | 5 | 7 | 1 | 1 | 3 | 2 | 9 | 2 | 1 | 10 | 5 | 7 | 1 | 2 | 8 | 1 | 6 | 1 | |||||
| Tragelaphus spekii | 95 | 1 | 6 | 4 | 1 | 5 | 1 | 3 | 7 | 6 | 9 | 4 | 12 | 1 | |||||||||||
| Hippopotamidae | Hippopotamus amphibius | 2 | 1 | 1 | |||||||||||||||||||||
| Suidae | Potamochoerus porcus | 3 | 1 | 2 | |||||||||||||||||||||
| Carnivora | 1 | ||||||||||||||||||||||||
| Herpestidae | Herpestinae sp | 1 | 1 | ||||||||||||||||||||||
| Primates | 67 | ||||||||||||||||||||||||
| Hominidae | Gorilla gorilla | 3 | 2 | 1 | |||||||||||||||||||||
| Homo sapiens | 64 | 1 | 1 | 13 | 2 | 22 | 1 | 3 | 1 | 2 | 1 | 4 | 1 | ||||||||||||
| Proboscidae | 10 | ||||||||||||||||||||||||
| Elephantidae | Loxodonta cyclotis | 10 | 7 | 1 | 2 | ||||||||||||||||||||
| Reptiles | |||||||||||||||||||||||||
| Crocodilia | 23 | 3 | |||||||||||||||||||||||
| Crocodylidae | Crocodylus niloticus | 3 | |||||||||||||||||||||||
| Mecistops cataphractus | 19 | 1 | 1 | 6 | |||||||||||||||||||||
| Osteolaemus tetraspis | 1 | 1 | |||||||||||||||||||||||
| Squamata | 12 | ||||||||||||||||||||||||
| Pythonidae | Python sebae | 8 | 2 | ||||||||||||||||||||||
| Varanidae | Varanus sp | 4 | 2 | 1 | 1 | ||||||||||||||||||||
| Testudines | 16 | ||||||||||||||||||||||||
| Testunidae | Kinixys erosa | 1 | 1 | ||||||||||||||||||||||
| Pelomedusidae | Pelusios castaneus | 3 | 1 | 1 | 1 | ||||||||||||||||||||
| Pelusios chapini | 1 | 1 | |||||||||||||||||||||||
| Pelusios marani | 11 | 3 | 8 | ||||||||||||||||||||||
| Birds | |||||||||||||||||||||||||
| Ciconiformes | 4 | ||||||||||||||||||||||||
| Ciconiidae | Ciconia sp | 4 | 1 | 2 | |||||||||||||||||||||
| 8 orders/12 families | 20 species | 428 | 3 | 1 | 11 | 4 | 41 | 2 | 2 | 16 | 4 | 89 | 1 | 6 | 7 | 18 | 7 | 16 | 11 | 11 | 22 | 6 | 25 | 1 | 2 |
-
Fsp1 = Glossina caliginea; Fsp2 = G. fusca congolensis; Fsp3 = G. fuscipes fuscipes; Fsp4 = G. pallicera newsteadi; Fsp5 = G. palpalis palpalis; Fsp6 = Stomoxys calcitrans; Fsp7 = S. niger niger; Fsp8 = Chrysops sp.
Number and origin of blood meals according to the fly species (Fsp), park and climatic season.
| Number of identified blood meals by fly species (Fsp) | ||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| La Lékédi | Ivindo | |||||||||||||||
| Rainy season | Dry season | Dry season | ||||||||||||||
| Taxonomic group/Order/Family | Host species | N° Identified | Fsp1 | Fsp3 | Fsp4 | Fsp5 | Fsp6 | Fsp1 | Fsp2 | Fsp3 | Fsp4 | Fsp5 | Fsp1 | Fsp2 | Fsp3 | Fsp5 |
| Mammals | ||||||||||||||||
| Artiodactyla | 295 | 2 | 1 | 3 | ||||||||||||
| Bovidae | Cephalophus silvicultor | 65 | ||||||||||||||
| kobus ellipsiprymnus | 4 | |||||||||||||||
| Syncerus caffer | 126 | 2 | 1 | 1 | 6 | 10 | 8 | 8 | 13 | 1 | 1 | |||||
| Tragelaphus spekii | 95 | 1 | 3 | 1 | 2 | 8 | 5 | 5 | 9 | 1 | ||||||
| Hippopotamidae | Hippopotamus amphibius | 2 | ||||||||||||||
| Suidae | Potamochoerus porcus | 3 | ||||||||||||||
| Carnivora | 1 | |||||||||||||||
| Herpestidae | Herpestinae sp | 1 | ||||||||||||||
| Primates | 67 | |||||||||||||||
| Hominidae | Gorilla gorilla | 3 | ||||||||||||||
| Homo sapiens | 64 | 4 | 1 | 5 | 1 | 1 | ||||||||||
| Proboscidae | 10 | |||||||||||||||
| Elephantidae | Loxodonta cyclotis | 10 | ||||||||||||||
| Reptiles | ||||||||||||||||
| Crocodilia | 23 | |||||||||||||||
| Crocodylidae | Crocodylus niloticus | 3 | ||||||||||||||
| Mecistops cataphractus | 19 | 2 | 4 | 2 | 3 | |||||||||||
| Osteolaemus tetraspis | 1 | |||||||||||||||
| Squamata | 12 | |||||||||||||||
| Pythonidae | Python sebae | 8 | 2 | 2 | 2 | |||||||||||
| Varanidae | Varanus sp | 4 | ||||||||||||||
| Testudines | 16 | |||||||||||||||
| Testunidae | Kinixys erosa | 1 | ||||||||||||||
| Pelomedusidae | Pelusios castaneus | 3 | ||||||||||||||
| Pelusios chapini | 1 | |||||||||||||||
| Pelusios marani | 11 | |||||||||||||||
| Birds | ||||||||||||||||
| Ciconiformes | 4 | |||||||||||||||
| Ciconiidae | Ciconia sp | 4 | 1 | |||||||||||||
| 8 orders/12 families | 20 species | 428 | 2 | 4 | 1 | 13 | 1 | 8 | 20 | 20 | 15 | 32 | 1 | 1 | 2 | 2 |
-
Fsp1 = Glossina caliginea; Fsp2 = G. fusca congolensis; Fsp3 = G. fuscipes fuscipes; Fsp4 = G. pallicera newsteadi; Fsp5 = G. palpalis palpalis; Fsp6 = Stomoxys calcitrans; Fsp7 = S. niger niger; Fsp8 = Chrysops sp.
Cytb sequences of parasites recovered in this study and of those used as references for phylogenetic analyses and their Genbank accession numbers.
| Isolate | Accession number |
|---|---|
| Anopheles coustani | KT367855 |
| An. gabonensis2 | KT367852 |
| An. gabonensis279 | KT367853 |
| An. gabonensis3 | KT367861 |
| An. marshallii | KT367857 |
| An. moucheti | KT367864 |
| An. obscurus2 | KT367846 |
| An. obscurus78 | KT367849 |
| Cephalophus_silvicultor_1336 | KY631949 |
| Cephalophus_silvicultor_1368 | KY631947 |
| Cephalophus_silvicultor_484 | KY631963 |
| Ciconia_sp_445 | KY631985 |
| E15_Podocnemis_expansa_Peru | KF049492 |
| E24_Podocnemis_expansa_Peru | KF049495 |
| Gorilla_gorilla_34 | KY631983 |
| Gorilla_gorilla_756 | KY631982 |
| Gorilla_gorilla_761 | KY631981 |
| P_sp._JA7_J725 | GU252027 |
| Haemoproteus_majoris | AY099045 |
| Haemoproteus_sp. | HM222472 |
| Haemoproteus_sp._GA02CI1 | HM222486 |
| Haemoproteus_sp._NA16K65 | HM222487 |
| Hepatocystis_sp. AA201_blike | JQ070951 |
| Hepatocystis_sp. | JQ070884 |
| Hepatocystis_sp._AA2012 | JQ070956 |
| HO11_Cephalophus_nigrofons | KT367819 |
| HO13_Cephalophus_monticola | KT367833 |
| HO613_Cephalophus_monticola | KT367836 |
| HO9_Kinixys_erosa | KT367843 |
| Homo_sapiens_476 | KY631978 |
| Homo_sapiens_481 | KY631977 |
| Homo_sapiens_57 | KY631979 |
| Homo_sapiens_574 | KY631976 |
| Homo_sapiens_635 | KY631975 |
| Homo_sapiens_636 | KY631974 |
| Homo_sapiens_638 | KY631973 |
| Homo_sapiens_639 | KY631972 |
| Homo_sapiens_668 | KY631969 |
| Homo_sapiens_806 | KY631968 |
| Homo_sapiens_832 | KY631967 |
| Leucocytozoon_caulleryi | AB302215 |
| Leucocytozoon_dubreuli | AY099063 |
| Leucocytozoon_majoris | FJ168563 |
| Leucocytozoon_sabrazesi | AB299369 |
| M0278_Cephalophus_monticola | KT367834 |
| NG238_Kinixys_erosa | KT367844 |
| NG277_Ceratogymna_atrata | KT367825 |
| NY195_Cephalophus_dorsalis | KT367838 |
| Nycteria_sp._R_alc_C9_1 | KF159720 |
| Nycteria_sp._R_lan_G3_1 | KF159690 |
| OI52_Pangolin | KT367818 |
| OL123_Cephalophus_monticola | KT367822 |
| OL131_Cephalophus_callipygus | KT367830 |
| P_adleri | HM235081 |
| P_azurophilum | AY099055 |
| P_billcollinsi | KP875474 |
| P_blacklocki | HM235065 |
| P_cynomolgi | AB444126 |
| P_falciparum_3D7 | AF069605 |
| P_gaboni | JF895307 |
| P_gallinaceum | AF069612 |
| P_gonderi | JF923751 |
| P_knowlesi | JQ345504 |
| P_malariae | HM000110 |
| P_ovale | GU723548 |
| P_praefalciparum_MOEB | JF923761 |
| P_reichenowi | KP875479 |
| P_relictum | AY733090 |
| P_sp._DAJ | JF923753 |
| P_vivax | KF591834 |
| P_atheruri | AY099054 |
| P_giganteum | AY099053 |
| P_vinckei_isolate_1 | KJ700853 |
| P_vinckei_isolate_2 | KJ700854 |
| P_yoelii_killicki | DQ414658 |
| P. atheruri | HQ712051 |
| P. cyclopsi_Hip_cy_L4_1_Schaer | KF159674 |
| P. voltaicum_M_ang_G1_1_1 | KF159671 |
| Parahaemoproteus_sp._bird_sp.17 | GQ141581 |
| Parahaemoproteus_sp. _bird_sp.19 | GQ141585 |
| Parahaemoproteus_vireonis | FJ168561 |
| Plasmodium_sp._bird | GQ141574 |
| Plasmodium_sp._bird_sp._12 | HM222485 |
| Plasmodium_sp._GD2_GD201 | GU252012 |
| Plasmodium_sp._lineage_JA01 | KM598212 |
| Polychromophilus_melanipherus_haplotype_VIII | KJ131277 |
| Polychromophilus_murinus_haplotype_3 | HM055585 |
| Polychromophilus_sp._Min_vil_G3_2 | KF159699 |
| Polychromophilus_sp._Pip_gran_G3_1 | KF159714 |
| Polychromophilus_sp._Neo_cap_G3 | KF159700 |
| Syncerus_caffer_1138 | KY631953 |
| Syncerus_caffer_1417 | KY631942 |
| Tragelaphus_eurycerus_1324 | KY631950 |
| Tragelaphus_spekii_1051 | KY631961 |
| Tragelaphus_spekii_1155 | KY631959 |
| Tragelaphus_spekii_1175 | KY631958 |
| Tragelaphus_spekii_1228 | KY631957 |
| Tragelaphus_spekii_1245 | KY631956 |
| Tragelaphus_spekii_1291 | KY631955 |
| Tragelaphus_spekii_1299 | KY631954 |
| Tragelaphus_spekii_1300 | KY631952 |
| Tragelaphus_spekii_1306 | KY631951 |
| Tragelaphus_spekii_1348 | KY631948 |
| Tragelaphus_spekii_1386 | KY631946 |
| Tragelaphus_spekii_1394 | KY631945 |
| Tragelaphus_spekii_1399 | KY631944 |
| Tragelaphus_spekii_1413 | KY631943 |
| Tragelaphus_spekii_385 | KY631964 |
| Tragelaphus_spekii_56 | KY631965 |
| U65_Podocnemis_unifilis_Peru | KF049506 |
| Unknown_host_1036 | KY631960 |
| Unknown_host_110 | KY631966 |
| Unknown_host_512 | KY631962 |
| Unknown_host_520 | KY631980 |
| Unknown_host_649 | KY631971 |
| Unknown_host_665 | KY631970 |
| Unknown_host_819 | KY631984 |





