Toddler signaling regulates mesodermal cell migration downstream of Nodal signaling
Figures
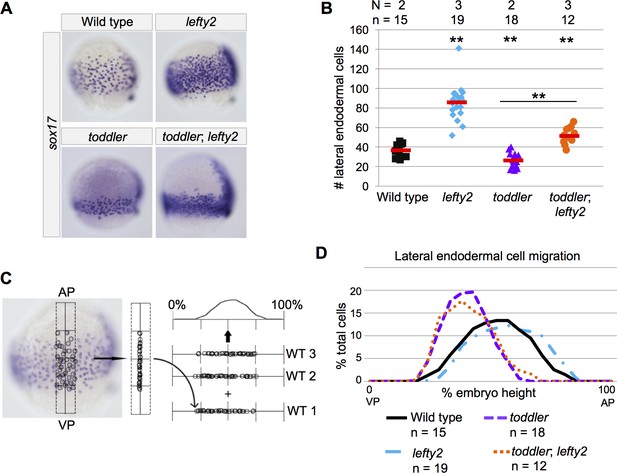
Increased endodermal specification does not rescue endodermal cell migration in toddler;lefty2 double mutants.
(A) Representative images of embryos analyzed in B and D. In situ hybridization with a sox17 probe at 75% epiboly; dorsal to the right. Endoderm in toddler;lefty2 double mutants resembles toddler single mutants. (B) The number of lateral endodermal cells is increased in toddler;lefty2 double mutants compared to wild-type and toddler single mutant embryos. Each point represents a single embryo. Red bars are averages. p-Values for pairwise comparison with wild type unless otherwise noted. **p<0.001; unpaired two-tailed t-test. N = number of independent experiments; n = number of embryos. (C) Schematic representation of experimental measurements shown in B and D. The locations of lateral endodermal cells in individual embryos are measured relative to the AP-VP axis and then consolidated across embryos. (D) Measurement of frequency with which cells were found at a given location in an embryo of a certain genotype. A cell at the animal pole corresponds to 100% embryo height, while a cell at the vegetal pole corresponds to 0%. AP = Animal pole; VP = vegetal pole. The same embryos were measured in B and D.
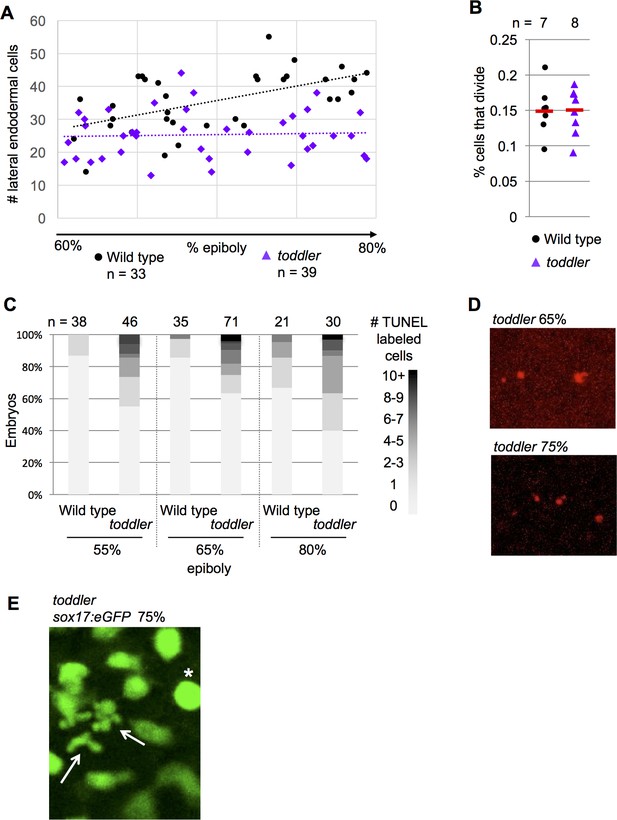
Initial specification of endoderm is normal in toddler mutants.
(A–C) n = number of embryos (A) Number of lateral endodermal cells in wild-type and toddler mutant embryos during gastrulation. (B) Rate of endodermal cell division in wild-type and toddler mutant embryos. 40–60 cells were tracked per embryo and the percentage of those cells that divided is shown. Red bars are averages. (C) Percentage of embryos with a given number of TUNEL +cells on one side of the embryo. (D) Representative images of TUNEL staining at two time points in toddler mutants. (E) Live tracking of endoderm using sox17:GFP transgenic fish captures two endodermal cells undergoing cell death during gastrulation in toddler mutants (arrows). A cell that is about to divide is also present (asterisk).
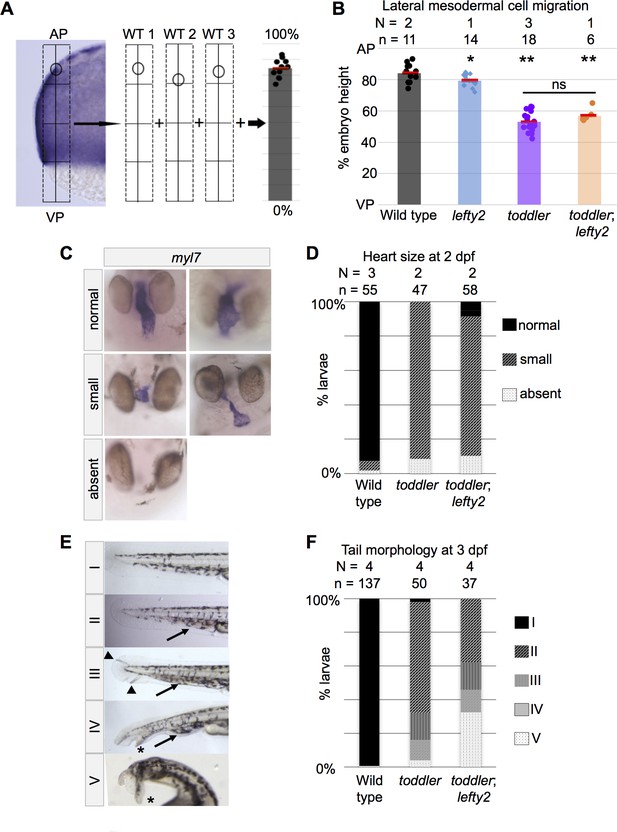
Mesodermal cell migration and larval phenotypes are not rescued by increased numbers of endodermal cells.
(A) Schematic representation of experimental measurements shown in B. Densely packed, fn1a + lateral mesodermal cells are viewed by cross section. Location of the most animally migrated lateral cells of individual embryos are measured relative to AP-VP axis, then consolidated across embryos. AP = Animal pole; VP = vegetal pole. (B) Defects in animal-pole directed migration of mesodermal cells are still present in toddler;lefty2 double mutants. Each point represents a single embryo. Red bars are averages. p-Values for pairwise comparison with wild type unless indicated otherwise. *p=0.04, **p<0.001; ns: p=0.08; unpaired two-tailed t-test. (C–D) toddler;lefty2 double mutants resemble toddler single mutant siblings in respect to heart phenotypes at 2 days post fertilization (dpf). Hearts were classified as small if shortened by more than half of normal length and/or excessively narrow or thin. Hearts that were neither thin nor short but appeared to have looping defects or other patterning defects were classified as normal. Heart phenotypes were scored blind to genotype for toddler and toddler;lefty2 mutants. (C) Representative images of phenotypic classes after in situ hybridization for myl7. (D) Quantification of C. (E–F) toddler;lefty2 double mutants are more poorly patterned than toddler single mutant siblings in respect to tail phenotypes at 3 dpf. (E) Representative images of phenotypic classes. Phenotype classes II-V lack circulation. Arrow: accumulated blood. Arrowhead: defects in mesenchyme. * Duplicated tail tip. (F) Quantification of E. (B,D,F) N = number of independent experiments; n = number of embryos.
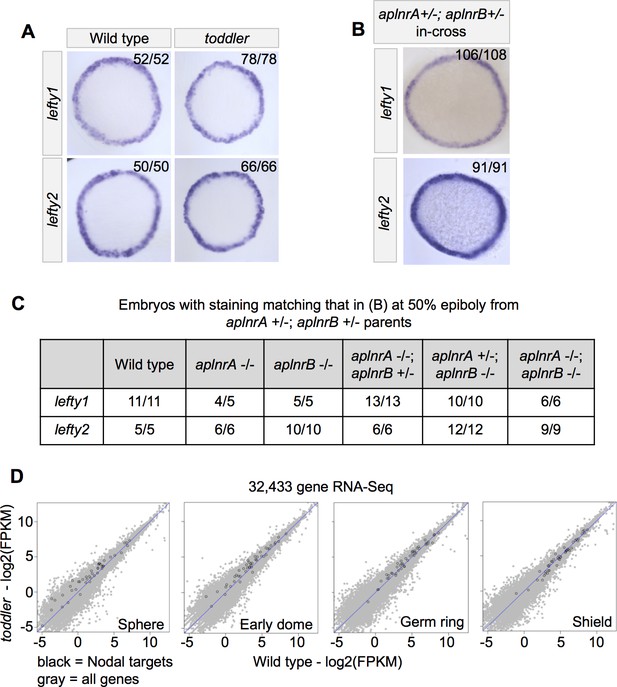
Wild-type and toddler mutant embryos have similar levels of Nodal signaling.
(A) Representative images of Nodal target gene expression in wild-type and toddler mutant embryos based on in situ hybridization at 50% epiboly. Animal pole facing up. Three biological replicates. (B–C) aplnrA;aplnrB double heterozygote parents were incrossed and their progeny examined by in situ hybridization at 50% epiboly. (B) Representative images of Nodal target gene expression in offspring from aplnrA;aplnrB double heterozygote parents. Animal pole facing up. (C) All embryos from B were genotyped. Results for relevant genotypes are shown. Expression of lefty1 and lefty2 in aplnrA;aplnrB double mutant embryos was indistinguishable from wild type. (D) RNA-sequencing before and during early gastrulation reveals no differential transcription of Nodal targets (black dots) between wild-type and toddler mutant embryos. All other genes are shaded in light gray. See also Figure 3—Source data 1.
-
Figure 3—source data 1
Gene information for variable genes in RNA-Seq data set.
Analysis with Cufflinks identified 47 genes with significantly different expression at one or more stages in the time course. Spreadsheet contains gene information and FPKM expression values for the variable genes in wild-type and toddler mutant embryos for each stage.
- https://doi.org/10.7554/eLife.22626.006
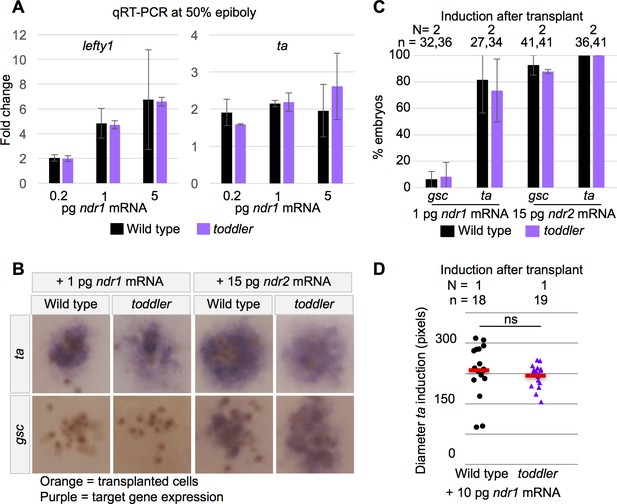
Wild-type and toddler mutant embryos respond similarly to exogenous Nodal sources.
(A) qRT-PCR at 50% epiboly on embryos injected at the one-cell stage with water or increasing levels of Nodal mRNA. Fold change in expression is relative to water control. Two biological replicates. (B–D) Clones of cells expressing exogenous Nodal mRNA and GFP were transplanted into host embryos at sphere stage and collected for in situ hybridization 1.5 hr later. Donors and hosts were always of matching genotype. (B) Representative images of quantifications in C. Orange staining marks anti-GFP labeled transplanted cells. Purple staining is from in situ hybridization for a Nodal target gene. (C) Percentage of embryos for which Nodal target gene expression was visible via in situ hybridization. N = number of independent experiments. n = number of embryos. (D) Diameter of induction of ta expression around the clone of transplanted cells. Each point represents a single embryo. Red bars are averages; ns: p=0.44; unpaired two-tailed t-test. (A and C) Means ±SEM.
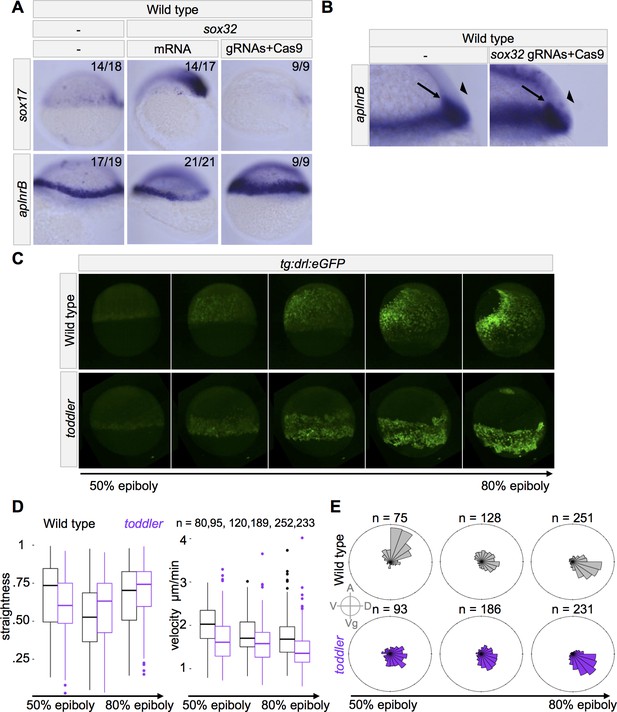
toddler mutant mesodermal cells have diminished animal pole-directed migration during gastrulation.
(A–B) Cas9-mediated mutagenesis was used to generate sox32 mutants. Embryos were injected at the one-cell stage either with sox32 mRNA or with Cas9 + sox32 gRNAs. in situ hybridization for sox17 and aplnrB at shield stage; dorsal to the right. (A) Expression of aplnrB decreases in the presence of excess endoderm and increases in the absence of endoderm. (B) Cross-section of shield stage embryo. The majority of cells expressing aplnrB in embryos lacking endoderm are internalized and unlikely to be ectodermal. Arrow: internalized cells. Arrowhead: ectoderm. (C–E) Mesodermal cell migration was tracked during gastrulation using a drl:eGFP transgene and lightsheet microscopy. drl:eGFP labels ventrolateral mesoderm during gastrulation. toddler mutant measurements represent three embryos. Wild-type measurements represent two wild-type and one heterozygous embryos. (C) Representative still frames of maximum intensity projections from a wild-type and toddler mutant drl:eGFP transgenic embryo. See also Video 1. (D–E) Movies spanning 50% to 85% epiboly were aligned at the onset of internalization and binned into 45 min windows. (D) Measurement of straightness and velocity of mesodermal cells in wild-type and toddler mutants. Straightness is defined as the difference between the total movement of a cell in all directions divided by the net displacement (the actual distance between the cell’s beginning and ending location). (E) toddler mutant cells have diminished migration animally during gastrulation. Each bin represents the proportion of cells moving in a given direction weighted by the total distance traveled in that direction. n = number of cells. A = animal, D = dorsal, Vg = vegetal, V = ventral.
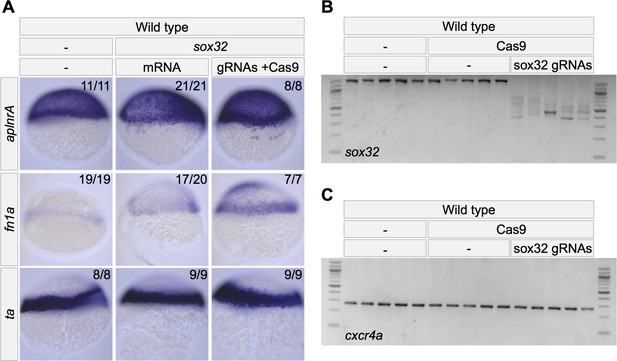
Apelin receptor A is expressed in mesodermal cells.
(A) Embryos injected at the one-cell stage with mRNA or Cas9 + gRNAs for endoderm master regulator sox32. in situ hybridization at shield stage, dorsal to the right. Expression of aplnrA, fn1a and ta decrease slightly or do not change in the presence of excess endoderm and increase in the absence of endoderm. (B–C) PCR of a genomic locus (B: sox32; C: cxcr4a) in wild-type embryos, wild-type embryos injected with Cas9 or wild-type embryos injected with Cas9 and sox32 gRNAs. Each lane represents an individual 2-day-old embryo that was injected at the one-cell stage. (B) Evidence of various large deletions in the genomic sox32 locus in response to injection of Cas9 and sox32 gRNAs. (C) The locus of a non-targeted gene, cxcr4a, is unaffected by injection of Cas9 and sox32 gRNAs.
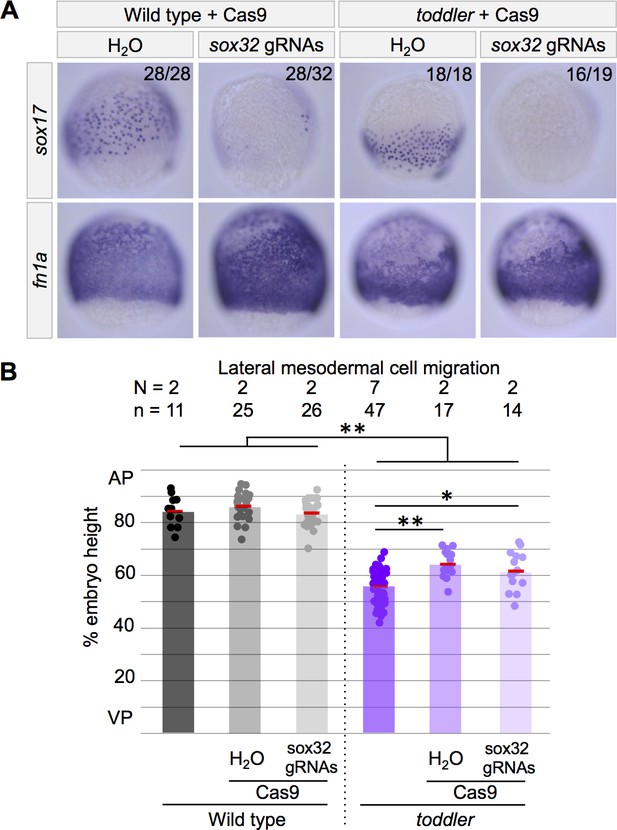
Toddler’s role in mesodermal cell migration is independent of endoderm.
The presence or absence of endoderm does not affect lateral mesodermal cell migration in wild-type and toddler mutants. Cas9-mediated mutagenesis was used to generate sox32 mutants. Embryos were injected at the one-cell stage with Cas9 and sox32 gRNAs. In situ hybridization for sox17 and fn1a at 75% epiboly; dorsal to the right. (A) Representative images of embryos analyzed in B. (B) Quantification of mesodermal cell migration from embryos in A as described in Figure 2A. AP = Animal pole; VP = vegetal pole. Each point represents a single embryo. Red bars are averages; **p<1.5×10−6; *p<0.05; unpaired two-tailed t-test.
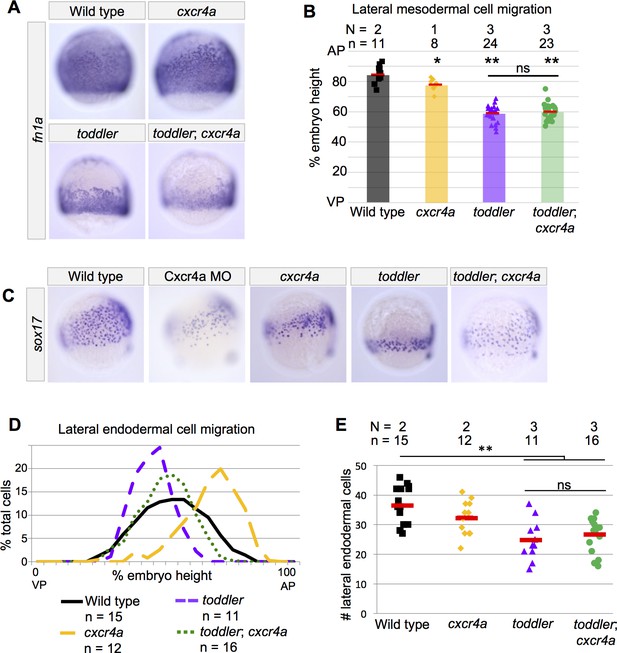
Endodermal cell migration does not require Toddler signaling in cxcr4a mutants.
(A–B) Analysis of lateral mesodermal cell migration defects in cxcr4a mutant embryos by in situ hybridization for fn1a. Mesodermal cell migration is normal in stable cxcr4a single mutants. toddler;cxcr4a double mutants show mesodermal cell migration defects that resemble toddler single mutant siblings. (C) Representative images of embryos analyzed in D and E. In situ hybridization for sox17 at 75% epiboly; dorsal to the right. Cxcr4a morphants and cxcr4a mutants have excessive animal pole-directed migration of vegetal endoderm. Endoderm patterning in toddler;cxcr4a double mutants resembles wild-type embryos more than toddler single mutant siblings. (D) Measurement of frequency with which cells were found at a given location in an embryo of a certain genotype. A cell at the animal pole corresponds to 100% embryo height, while a cell at the vegetal pole corresponds to 0%. AP = Animal pole=100%; VP = vegetal pole=0%. (E) The number of lateral endodermal cells is unchanged between toddler single and toddler;cxcr4a double mutants. Each point represents a single embryo. Red bars are averages. (B and E) ns: p>0.46; *p<0.05; **p<0.0005; unpaired two-tailed t-test. N = number of independent experiments; n = number of embryos.
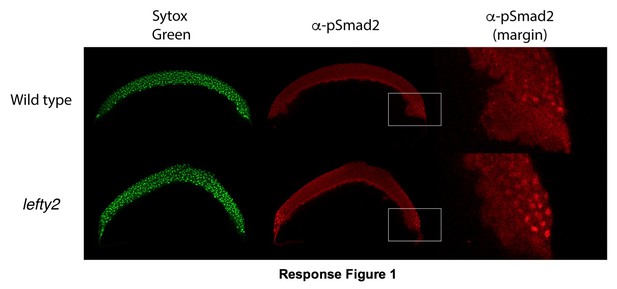
Nodal signaling is elevated in lefty2 mutants.
Smad2 is the intracellular effector of Nodal signaling and is phosphorylated when activated (Schier 2009). We measured phosphorylated Smad2 (pSmad2) by immunostaining as described in a forthcoming manuscript (Rogers et al. submitted). Representative cross-sections from pools of 3-6 embryos at 50% epiboly stage are shown. Left column: nuclei are labeled with Sytox green DNA stain. Middle column: α -pSmad2 immunostaining. Right column: magnified view of embryo margins highlighted in middle column with white boxes.
Videos
Toddler regulates animal-poled directed migration of mesodermal cells.
Embryo pair shown in Figure 5C. Embryos are transgenic for drl:eGFP, which labels ventrolateral mesoderm during gastrulation. Embryos are aligned at the onset of internalization. Maximum intensity projections from a light sheet microscope spanning 50% to 85% epiboly, time in hr:min.
Tables
Primer and target sequences.
https://doi.org/10.7554/eLife.22626.013| Primers | gRNA targets | ||
|---|---|---|---|
| lefty1 1 | CGTGGCTTTCATGTATCACCTTC | cxcr4a target 1 | G G A CATCGGAGC CAACTTTG |
| lefty1 1 | G CATTAG CCTATATG TTAACTTG CAC. | cxcr4a target 2 | GTAC CG TCTG CA C CT CTCAG |
| Iefty2 1 | GGGACACAAG CTTTG AAGG G | cxcr4a target 3 | CG C CTTC ATCA GTTTG G ACC |
| Iefty2 2 | TCCCTGTGTGAGTGAGATCG | cxcr4a target 4 | CGCCGCGCTCCTCACTGTGC |
| Iefty2 3 | CAGCTGTTCATTTTGACCACTCAC | cxcr4a target 5 | G GA CTC GTTTG TCACAT G G G |
| Iefty2 4 | AT G GAG CTTC AG C AT G G AC AG | sox32 target 1 | CGTTCTGATGTTG C AAATAGTGG |
| aplnrB 1 | TGTGTGAATATGATGAGTGGGAAC | sox32 target 2 | G G CTTAAT G G GCC CGACGCGGGG |
| aplnrB 2 | AGTGGTATCCCAGAGCAGTGTAG | sox32 target 3 | CC GC G TCG G G CCC ATTAAG CC CG |
| cxcr4a 1 | CGTCTTTGAAGATGATTTATCAGC | sox32 target 4 | G TT C ATCAT G TG G AC G AAAG AG G |
| cxcr4a 2 | CA C GTAAATG ATG CG G TT G G | sox32 target 5 | CGAAGTGGTATGATGAAGAGTGG |
| cxcr4a 3 | AG ACT G AA G GAG CTG G AG AAG | sox32 target 6 | AGTG GAAAC GT GTTTG AT G GT G G |
| qPCR ta F | G AAC CAC AG AG GTG CTC CATATC | sox32 target 7 | G ACTCT GA GTAAGC AG ACC G TG G |
| qPCR ta R | CTGGTGTTGGAGGTAGTGTTTGTG | sox32 target 8 | GTAGAGCTCCATGATAGGTGGGG |
| qPCR leftyl F | AAG CT CTAC AAG AAG G CCC C AC ACAAG | sox32 target 9 | G CGTGTGTG CTGTGTTTG G GTG G |
| qPCR leftyl R | TTCG TG AATG G G AATC AAC CTG G A A | ||
| qPCR actin F | ATCAGGGTGTCATGGTTGGT | ||
| qPCR actin R | CACGCAGCTCGTTGTAGAAG | ||
| sox32 F | AAACTTCTCACG CTT C AC ACC | ||
| sox32 R | CCATCCAGATTGCTG CTG ATTT | ||
Additional files
-
Transparent reporting form
- https://doi.org/10.7554/eLife.22626.014





