Impaired respiration elicits SrrAB-dependent programmed cell lysis and biofilm formation in Staphylococcus aureus
Figures
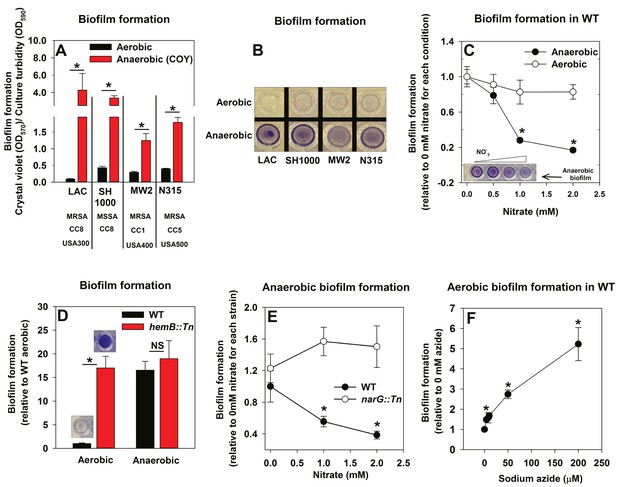
Oxygen impacts biofilm formation in its capacity as a terminal electron acceptor.
Panels A and B; Anaerobic growth elicits increased biofilm formation in multiple S. aureus isolates. Biofilm formation of the LAC (JMB1100; hereafter wild-type (WT)), SH1000 (JMB 1323), MW2 (JMB1324) and N315 (JMB 7570) isolates following aerobic or anaerobic growth is displayed. MRSA denotes methicillin resistance, MSSA denotes methicillin sensitivity, CC denotes clonal complex type and the USA number denotes the pulsed-field gel electrophoeresis type. Panel C; Supplementing growth media with the alternate terminal electron acceptor nitrate results in decreased biofilm formation during anaerobic growth. Biofilm formation for WT following aerobic or anaerobic growth and in media containing between 0–2 mM sodium nitrate is displayed. Panel D; A strain incapable of respiration upon oxygen forms increased biofilms when cultured aerobically, but not fermentatively. Biofilm formation for the WT and hemB::Tn (JMB6037) strains following aerobic or anaerobic growth is displayed. Panel E; Nitrate supplementation does not decrease anaerobic biofilm formation in a nitrate reductase mutant. Biofilm formation for the WT and narG::Tn (JMB7277) strains following anaerobic growth and in media containing between 0–2 mM sodium nitrate. Panel F; Chemical inhibition of respiration elicits increased biofilm formation during aerobic growth. Biofilm formation for the WT following aerobic growth in media supplemented with 0–250 µM sodium azide. The data represent the average values of eight wells (Panels A, C-E) or quadruplicates (Panel F) and error bars represent standard deviations. Representative photographs of biofilms formed upon the surface of a 96-well microtiter plate and stained with crystal violet are displayed in Panel B or insets in Panel C and D. Error bars are displayed for all data, but on occasion may be too small to see. Statistical significance was calculated using a two-tail Student's t-test and p-values>0.05 were considered to be not significant while * indicates p-value of <0.05.
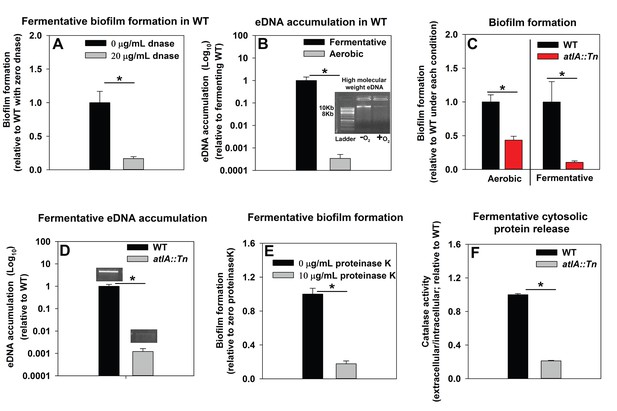
Impaired respiration results in AtlA-dependent release of high-molecular weight DNA, cytoplasmic proteins and an increase in biofilm formation.
Panel A; Fermentative biofilm formation is attenuated upon supplementation of growth medium with DNase. Biofilm formation of the WT (JMB 1100) following fermentative growth in media with or without 20 µg/mL DNase is displayed. Panel B; High-molecular weight DNA (eDNA) accumulation is increased in the biofilm matrix of fermenting cells. Biofilms of the WT were cultured aerobically or fermentatively, eDNA was extracted, and analyzed using agarose gel electrophoeresis (inset photograph). The data were normalized to the viable cell count, and thereafter, to eDNA accumulation in fermenting WT. Panel C; Fermentative biofilm formation is dependent upon the AtlA murein hydrolase. Biofilm formation for the WT and the atlA::Tn (JMB 6625) strains cultured aerobically or fermentatively is displayed. Panel D; eDNA accumulation in fermenting biofilms is dependent upon AtlA. Biofilms of the WT and atlA::Tn strains were cultured fermentatively and eDNA accumulation assessed. The data were normalized to the viable cell count, and thereafter, to eDNA accumulation in WT. Panel E; Fermentative biofilm formation is attenuated upon supplementation of growth medium with Proteinase K. Biofilm formation for the WT following fermentative growth in media with or without 10 µg/mL Proteinase K is displayed. Panel F; Fermentative growth results in AtlA-dependent release of a cytosolic protein into the extracellular milleu. Biofilms of the WT and atlA::Tn strains were cultured fermentatively and the activity of the cytosolic protein catalase (Kat) was measured in the spent media supernatant. The data were normalized to intracellular Kat activity, and thereafter to WT levels. The data represent the average values of eight wells (Panels A, C and E), sextuplets (Panel B) or triplicates (Panels D and F) and error bars represent standard deviations. Representative photographs of high-molecular weight eDNA are displayed in Panel B or inset in Panel D. Error bars are displayed for all data, but might be too small to see on occasion. Statistical significance was calculated using a two-tail Student's t-test and p-values>0.05 were considered to be not significant while * indicates p-value of <0.05.
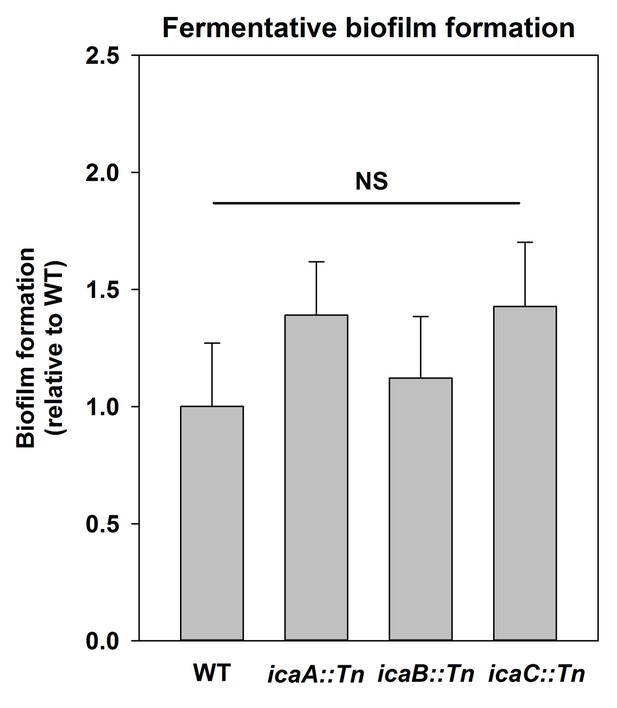
Polysaccharide intercellular adhesin (PIA) is dispensable for fermentative biofilm formation.
Biofilm formation of the WT (JMB 1100), icaA::Tn (JMB 5577), icaB::Tn (JMB 5579), and icaC::Tn (JMB 5578) strains following fermentative growth is displayed. Data represent the average value of eight wells and error bars represent standard deviation. Statistical significance was calculated using a two-tail Student's t-test and p-values>0.05 were considered to be not significant.
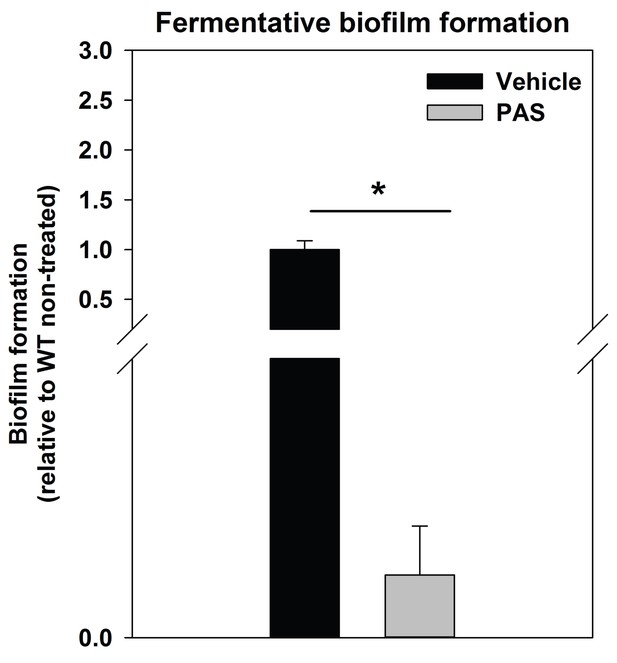
Supplementing growth media with the autolysis inhibitor polyanethole sulfonate (PAS) attenuates fermentative biofilm formation.
Biofilm formation of the WT (JMB 1100) cultured fermentatively in the presence of vehicle or 300 µg/mL PAS is displayed. Data represent the average value of eight wells and error bars represent standard deviation. Statistical significance was calculated using a two-tail Student's t-test and * indicates p-value of <0.05.
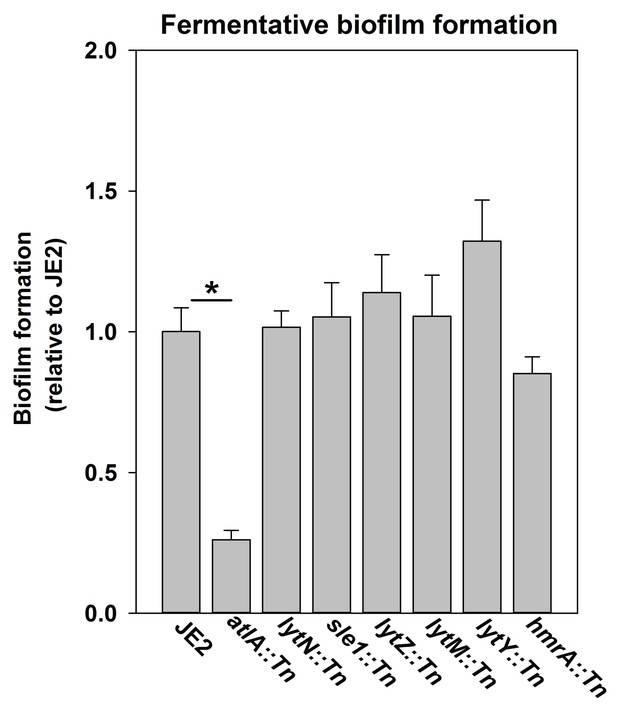
Fermentative biofilm formation is dependent on the AtlA murein hydrolase.
Biofilm formation of the WT (JMB 2977), atlA::Tn (JMB 6625), lytN::Tn (JMB 7265), sle1::Tn (JMB 7266), lytZ::Tn (JMB 7269), lytM::Tn (JMB 7271), lytY::Tn (JMB 7268), and hmrA::Tn (JMB 7270) strains cultured fermentatively is displayed. Data represent the average value of eight wells and error bars represent standard deviation. Statistical significance was calculated using a two-tail Student's t-test and * indicates p-value of <0.05.
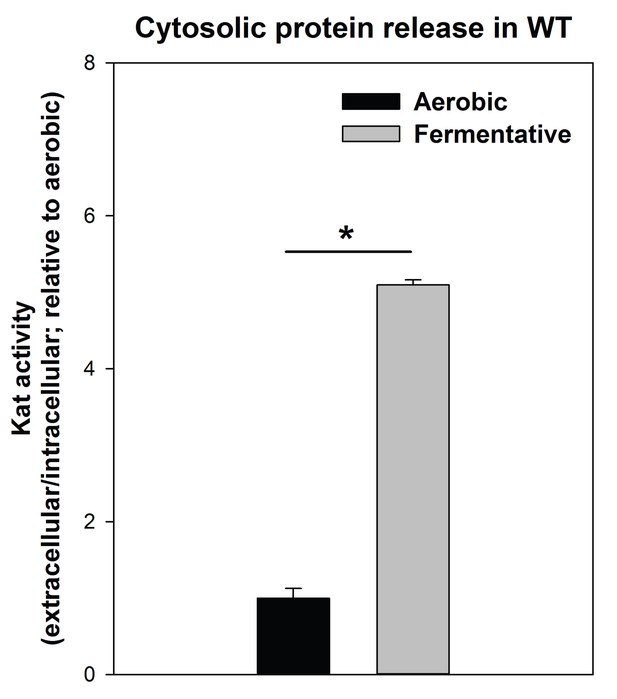
Cytosolic protein release is increased upon fermentative growth.
Biofilms of the WT (JMB 1100) were cultured aerobically or fermentatively and the activity of the cytosolic protein catalase (Kat) was measured in the spent media supernatant. Extracellular Kat activity was normalized to intracellular Kat activity and thereafter to levels under aerobic growth. Statistical significance was calculated using a two-tail Student's t-test and * indicates p-value of <0.05.
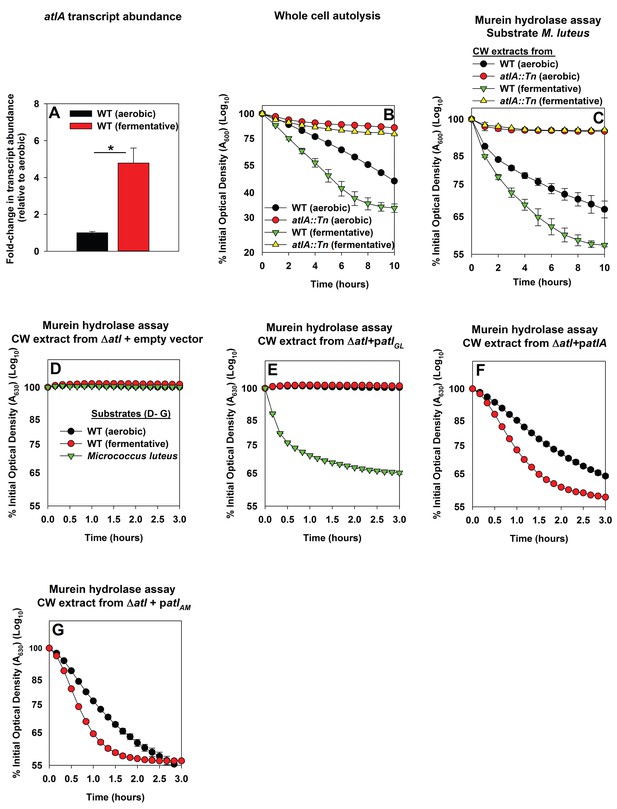
Impaired respiration elicits increased expression of AtlA and alterations that make cells more amenable to cleavage by AtlA.
Panel A; The atlA transcript is increased upon fermentative growth. Biofilms of the WT (JMB 1100) were cultured aerobically or fermentatively, mRNA was extracted, and the abundance of the atlA transcript was quantified. The data were normalized to 16S rRNA levels, and thereafter, to levels observed aerobically. Panel B; Fermenting cells undergo increased autolysis in an AtlA-dependent manner. The WT and atlA::Tn (JMB 6625) strains were cultured aerobically or fermentatively and autolysis was examined in intact whole cells. Panel C; AtlA-dependent bacteriolytic activity is increased in fermenting cells. Murein-hydrolase activity in cell-wall associated proteins (CW-extracts) detached from the WT or atlA::Tn strains cultured aerobically or fermentatively is displayed (pH of 7.5). Heat-killed Micrococcus luteus was used as a substrate. Panel D-G; Fermenting cells are more amenable to AtlA and N-acetylmuramyl-L-alanine amidase (AM)-dependent cleavage. Murein-hydrolase activity using CW-extracts detached from a ΔatlA strain (KB 5000) carrying plasmids encoding for empty vector control (Panel D), GL only (patlAGL) (Panel E), full-length AtlA (patlA) (Panel F), or AM only (patlAAM) (Panel G) upon heat-killed cells of the WT cultured aerobically or fermentatively or M. luteus as substrates is displayed (pH of 7.5). The data in Panel A represent the average values of triplicates. Statistical significance was calculated using a two-tail Student's t-test and * indicates p-value of <0.05. The data in Panels B-G represent the average value of technical duplicates from one set of substrate preparation, autolysis experiments, or CW extract preparations. Autolysis experiments or the preparation of heat-killed substrates or CW-extracts were conducted on least three separate occasions and similar results were obtained. Error bars in all panels represent standard deviations. Error bars are displayed for all data, but might be too small to see on occasion.
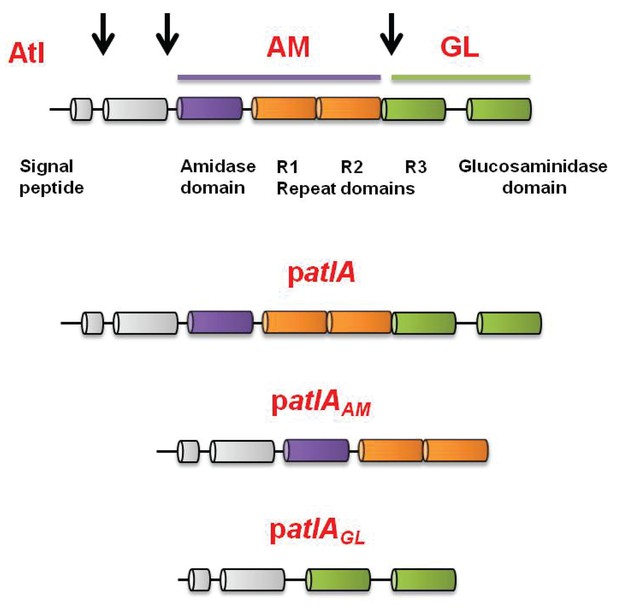
Representation of the full-length AtlA precursor protein and of the plasmid encoded variants used in this study.
S. aureus AtlA is a bifunctional protein with an amidase (AM) and a glucosaminidase (GM) domain. The schematic is a modification of previous illustrations (Bose et al., 2012; Götz et al., 2014). AtlA is post-translationally processed (indicated by arrows) between the propeptide and AM domain and between the repeat domains to free AM-R1-2 and R3-GM. The four allelic variants used in this study were constructed previously (Bose et al., 2012). The alleles are carried upon multi-copy plasmids that encode for full length AtlA (patlA), the amidase and repeat domains (AM-R1-R2) (patlAAM), catalytically inactivated amidase and repeat domains via a point mutation H263A (AMH263A-R1-R2) (patlAAMH263A) (not displayed), or glucosaminidase (R3-GM).
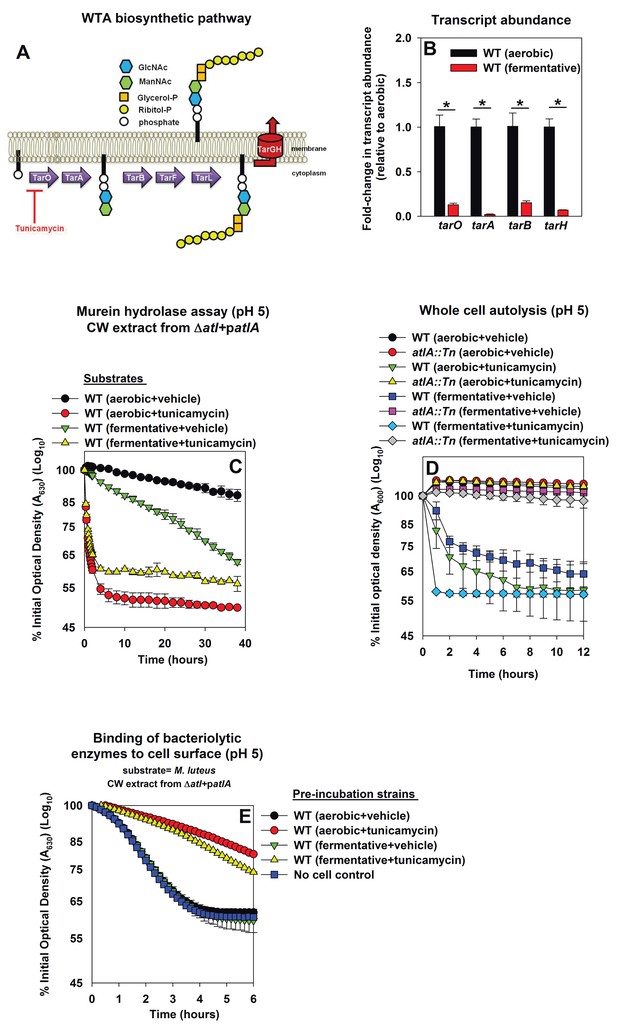
Decreased expression of wall-teichoic acids during fermentative growth makes S. aureus more amenable to cleavage by AtlA.
Panel A; Schematic of wall-teichoic acid (WTA) biosynthesis in S. aureus. The diagram displays select proteins involved in WTA biosynthesis and is redrawn as initially presented by Campbell et al. (2012). The initial transformations in the pathway catalyzed by TarO and TarA are non-essential, while the latter steps are essential. Tunicamycin inhibits TarO, as well as the 2-epimerase MnaA, which modulates the substrate levels for TarO (Campbell et al., 2011; Mann et al., 2016). MnaA is not displayed. Panel B; Transcript levels corresponding to genes encoding for WTA biosynthesis proteins are decreased upon fermentative growth. Biofilms of the WT (JMB 1100) were cultured aerobically or fermentatively, mRNA was extracted, and the abundances of the tarO, tarA, tarB, and tarH transcripts were quantified. The data were normalized to 16S rRNA levels, and thereafter to levels observed aerobically. Panel C; AtlA-dependent cleavage of heat-killed cells at a decreased pH is modulated via wall-teichoic acids. Murein-hydrolase activity at pH of 5 for cell-wall associated proteins (CW-extracts) detached from a ΔatlA strain (KB 5000) carrying patlA and incubated with heat-killed cells of the WT cultured aerobically or fermentatively in the presence or absence of 100 ng/mL tunicamycin as substrates is displayed. Panel D; AtlA-dependent autolysis of intact whole cells at decreased pH is modulated via wall-teichoic acids. The WT and atlA::Tn (JMB 6625) strains were cultured aerobically or fermentatively in the presence or absence of 100 ng/mL tunicamycin. Autolysis was examined in intact cells resuspended in a buffer with pH of 5. Panels E; Heat-killed aerobic or fermenting WT bind similar amounts of AtlA. CW-extract detached from a ΔatlA strain (KB 5000) carrying patlA was incubated at pH of 5 with heat-killed WT, cultured aerobically or fermentatively in the presence or absence of 100 ng/mL tunicamycin, or in the absence of cells (control) for 8 min. The cells were separated by centrifugation and bacteriolytic activity in the resultant supernatant was assessed upon heat-killed M. luteus as a substrate is displayed. Data in Panel B represents the average value of triplicates. Statistical significance was calculated using a two-tail Student's t-test and * indicates p-value of <0.05. Data in Panels C-E represent the average value of technical duplicates from one set of substrate preparation or autolysis assays. The heat-killed substrates were prepared or autolysis assays were conducted on least three separate occasions and similar results were obtained. Error bars in all panels represent standard deviations. Error bars are displayed for all data, but might be too small to see on occasion.
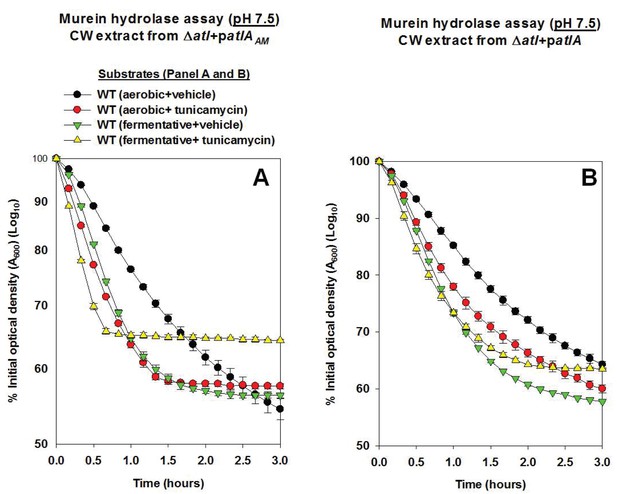
AtlA- and AM-dependent cleavage of heat-killed cells is modulated via altered expression of wall-teichoic acids.
Murein hydrolase assays were conducted using cell-wall associated proteins (CW-extracts) detached from a ΔatlA (KB 5000) strain carrying patlAAM (Panel A) or patlA (Panel B) and upon heat-killed cells of the WT (JMB 1100) cultured aerobically or fermentatively and in the presence or absence of 100 ng/mL tunicamycin as substrates (pH 7.5). Data represent the average value of technical duplicates from one set of substrate preparation. The heat-killed substrates were prepared on least three separate occasions and similar results were obtained. Error bars represent standard deviations.
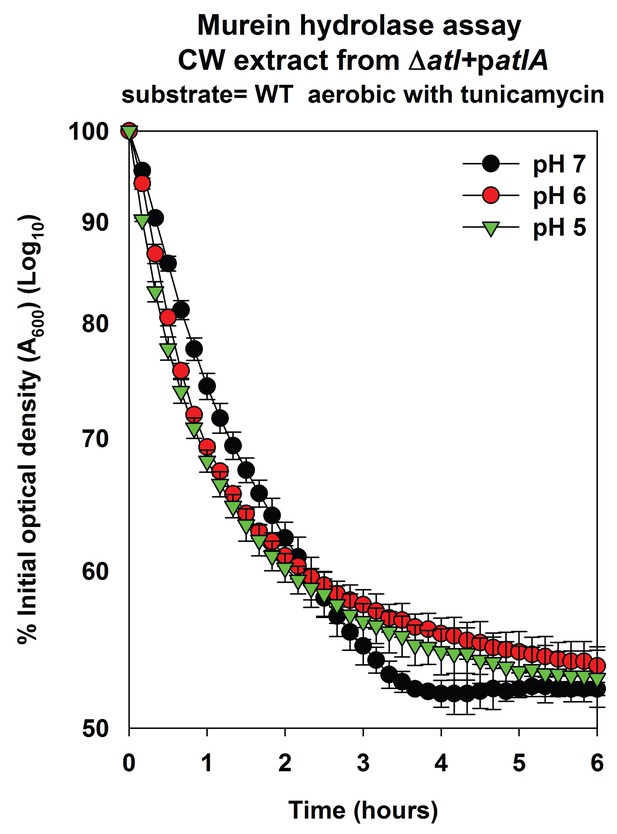
AtlA-dependent lysis rates of heat-killed tunicamycin treated cells are not altered upon alterations in the assay buffer pH.
Murein hydrolase assays were conducted at pH 7, pH 6, or pH 5 using cell-wall associated proteins (CW-extracts) detached from a ΔatlA (KB 5000) strain carrying patlA upon heat-killed cells of the WT cultured aerobically in the presence of 100 ng/mL tunicamycin as substrates. Data represent the average value of technical duplicates from one set of substrate preparation. The heat-killed substrates were prepared on least three separate occasions and similar results were obtained. Error bars represent standard deviations.
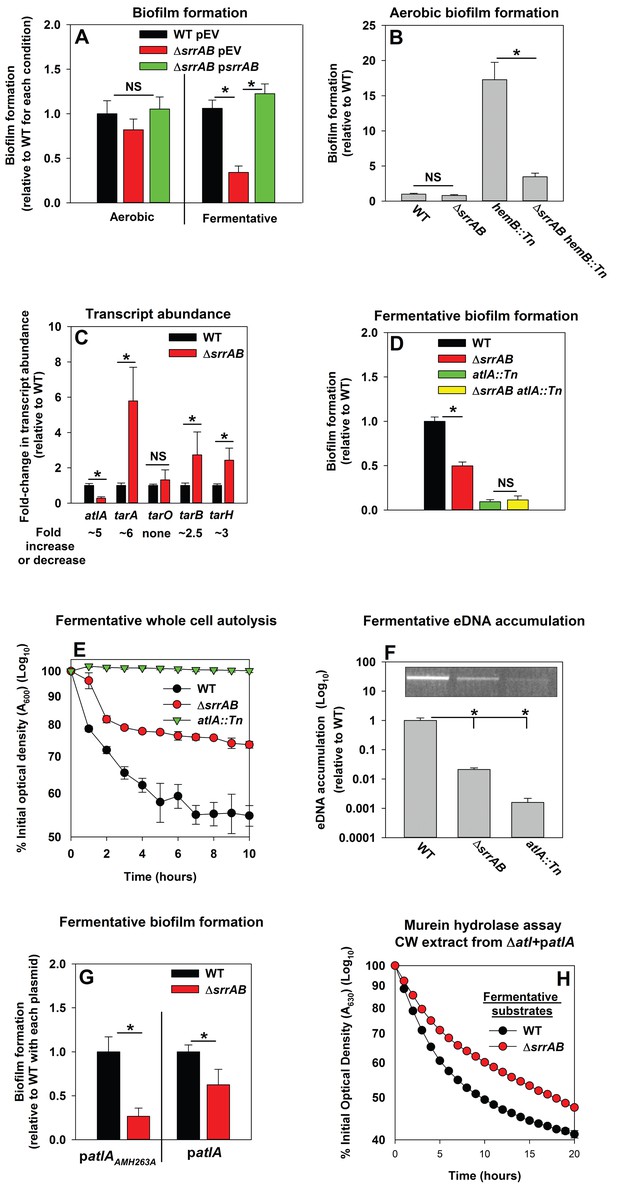
Programmed cell lysis and biofilm formation in fermenting cells are governed by the SrrAB two-component regulatory system.
Panel A; Fermentative biofilm formation is dependent upon SrrAB. Biofilm formation is displayed following aerobic or fermentative growth in the WT (JMB 1100) carrying pLL39 (pEV) or the ΔsrrAB (JMB 1467) strains carrying either pLL39 (pEV) or pLL39_srrAB (psrrAB). Panel B; A hemB mutant forms SrrAB-dependent biofilms aerobically. Biofilm formation following aerobic growth is displayed for the WT, ΔsrrAB, hemB::Tn (JMB 6037), and ΔsrrAB hemB::Tn (JMB 6039) strains. Panel C; Transcript levels corresponding to genes involved in programmed cell lysis and biofilm formation are altered in a ΔsrrAB strain. Biofilms of the WT and ΔsrrAB strains were cultured fermentatively, mRNA was extracted, and the abundances of the atlA, tarO, tarA, tarB, and tarH transcripts were quantified. Data were normalized to 16S rRNA levels, and thereafter, to levels observed in the WT. Panel D; The fermentative biofilm formation phenotypes associated with the ΔsrrAB and atlA::Tn mutations are not additive. Biofilm formation is displayed following fermentative growth for the WT, ΔsrrAB, atlA::Tn (JMB 6625), and ΔsrrAB atlA::Tn (JMB 6624) strains. Panel E; Autolysis of fermenting S. aureus is decreased in a strain lacking SrrAB. The WT, ΔsrrAB, and atlA::Tn strains were cultured fermentatively and autolysis was examined (pH of 5). Panel F; eDNA accumulation is decreased in a strain lacking SrrAB. Biofilms of the WT, ΔsrrAB, and atlA::Tn strains were cultured fermentatively and eDNA was quantified. The data were normalized to the viable cell count and thereafter to the levels in the WT. Panel G; atlA in multicopy partially suppresses the biofilm formation defect of the ΔsrrAB strain. Fermentative biofilm formation is displayed for the WT and ΔsrrAB strains carrying either patlAAM H263A or patlA. Panel H; Heat-killed cells of a ΔsrrAB strain are less amenable towards AtlA-dependent lysis. Murein-hydrolase activity for cell-wall associated proteins (CW-extracts) detached from a ΔatlA strain (KB 5000) carrying patlA and combined with fermentatively cultured and heat-killed WT or ΔsrrAB strains as substrates are displayed. Data presented represent the average value of eight wells (Panels A, B, D-G) or biological triplicates (Panel C and F). Data in Panels E and H represent the average value of technical duplicates from one set of autolysis assays or substrate preparations. The heat-killed substrates were prepared or autolysis assays were conducted on least three separate occasions and similar results were obtained. Error bars in all panels represent standard deviations. Error bars are displayed for all data, but might be too small to see on occasion. Statistical significance was calculated using a two-tail Student's t-test and p-values>0.05 were considered to be not significant while * indicates p-value of <0.05.
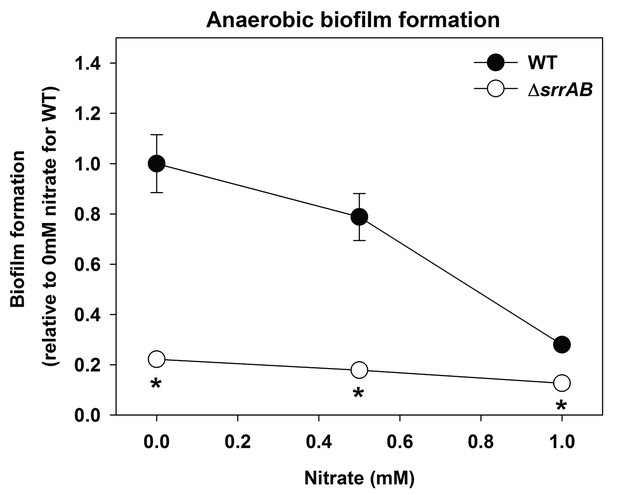
Biofilm formation of a ΔsrrAB strain is largely unaltered upon supplementing anaerobic biofilms with the alternate terminal electron acceptor nitrate.
Biofilm formation following anaerobic growth in the presence or absence of varying concentrations of sodium nitrate is displayed for the WT (JMB1100) and ΔsrrAB (JMB1467) strains. Data represent the average value of eight wells and error bars represent standard deviation. Statistical significance was calculated using a two-tail Student's t-test and p-values>0.05 were considered to be not significant while * indicates p-value of <0.05.
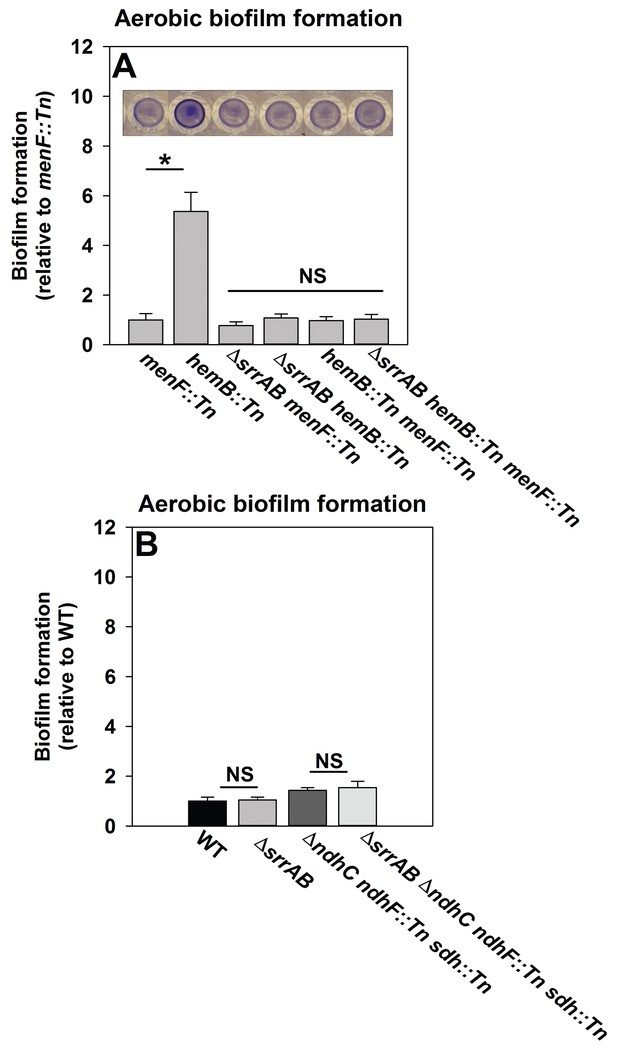
SrrAB-dependent biofilm formation is responsive to the oxidation state of the cellular menaquinone pool.
Panel A; SrrAB-dependent biofilm formation is inactivated in strains lacking the ability to synthesize menaquinone. Biofilm formation following aerobic growth is displayed for the menF::Tn (JMB6219), hemB::Tn (JMB6037), ΔsrrAB menF::Tn (JMB6221), ΔsrrAB hemB::Tn (JMB6039), hemB::Tn menF::Tn (JMB6217), and ΔsrrAB hemB::Tn menF::Tn (JMB6673) strains. Panel B; SrrAB-dependent biofilm formation is not stimulated in strains enriched for oxidized menaquinone. Biofilm formation following aerobic growth is displayed for the WT (JMB 1100), ΔsrrAB (JMB 1467), ΔndhC ndhF::Tn sdh:Tn (JMB 6613), and ΔsrrAB ΔndhC ndhF::Tn sdh:Tn (JMB 6614) strains. Data in both panels represent the average value of eight wells and the errors bars represent standard deviation. Statistical significance was calculated using a two-tail Student's t-test and p-values>0.05 were considered to be not significant while * indicates p-value of <0.05.
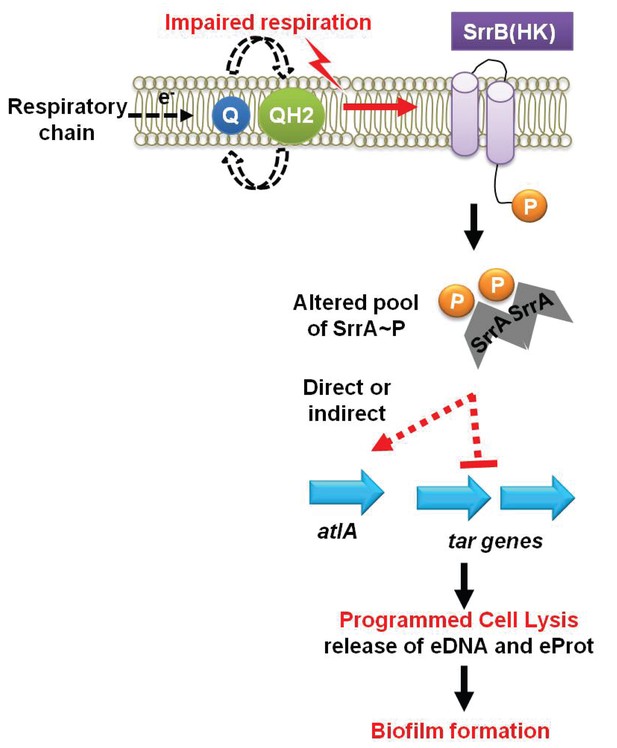
A working model for the influence of respiration upon autolysis and biofilm formation in S. aureus.
A decreased capacity to respire results in an enrichment of reduced menaquinone effecting altered activity of the SrrAB two-component regulatory system. Altered SrrAB activity leads to increased transcription of atlA and decreased transcription of genes (tar) encoding for wall-teichoic acid (WTA) biosynthesis. The consequent decrease in WTA expression and increase in AtlA expression results in the release of DNA and proteins, cell lysis and biofilm formation. Since cell lysis is effected via regulatory tuning of two divergent processes we term this mechanism as programmed cell lysis (PCL).
Tables
Strains and plasmids used in this study.
| Strains used in this study | |||
|---|---|---|---|
| S. aureus Strains | Genotype/Description | Genetic Background | Source/Reference |
| JMB1100 | Wild-type; USA300_LAC (erm sensitive); MRSA; CC8 | LAC | Boles et al. (2010) |
| RN4220 | Restriction minus; MSSA; CC8 | NCTC8325 | Kreiswirth et al. (1983) |
| JMB 1467 | ΔsrrAB (SAUSA300_1441–42) | LAC | Pang et al. (2014) |
| JMB 2047 | ΔsrrAB::tet | LAC | This work |
| JMB 2078 | katA::Tn (ermB) (SAUSA300_1232) | LAC | V. Torres |
| SH1000 | parent; MSSA; CC8 | SH1000 | Horsburgh et al. (2002) |
| JMB 1324 | parent, MRSA, USA400, CC1 | MW2 | Alex Horswill and Centers for Disease Control and Prevention (1999) |
| JMB 7570 | parent, MRSA, USA100; CC5 | N315 | Ann Stock and Kuroda et al. (2001) |
| JMB 1432 | Δfur::tetM | LAC | Horsburgh et al. (2001) |
| JMB 6231 | sdhA::Tn(ermB) | LAC | BEI resources and Fey et al. (2013) |
| JMB 6232 | ΔsrrAB sdhA::Tn(ermB) | LAC | This work |
| JMB 6384 | ndhF::Tn(ermB) (SAUSA300_0841) | LAC | This work; BEI resources and Fey et al. (2013) |
| JMB 2057 | ΔndhC::tet (SAUSA300_0844) | LAC | This work |
| JMB 6614 | ΔsrrAB sdhA::Tn(ermB) ΔndhC::tet ndhF::Tn(ermB) | LAC | This work |
| JMB 6613 | sdhA::Tn(ermB) ΔndhC::tet ndhF::Tn(ermB) | LAC | This work |
| JMB 6037 | hemB::Tn(ermB) | LAC | BEI resources and Fey et al. (2013) |
| JMB 6039 | ΔsrrAB hemB::Tn(ermB) | LAC | This work |
| JMB 6029 | menF::Tn(ermB) | LAC | BEI resources and Fey et al. (2013) |
| JMB 6033 | ΔsrrAB menF::Tn(ermB) | LAC | This work |
| JMB 6219 | menF::Tn(tet) | LAC | This work |
| JMB 6221 | ΔsrrAB menF::Tn(tet) | LAC | This work |
| JMB 6217 | hemB::Tn(ermB) menF::Tn(tet) | LAC | This work |
| JMB 6673 | ΔsrrAB hemB::Tn(ermB) menF::Tn(tet) | LAC | This work |
| JMB 6625 | atlA::Tn(ermB) | LAC | BEI resources and Fey et al. (2013) |
| KB5000 | ΔatlA | UAMS-1 | Bose et al. (2012) |
| JMB 6624 | ΔsrrAB atlA::Tn(ermB) | LAC | This work |
| JMB 5577 | icaA::Tn(ermB) | LAC | This work; BEI resources and Fey et al. (2013) |
| JMB 5579 | icaB::Tn(ermB) | LAC | This work; BEI resources and Fey et al. (2013) |
| JMB 5578 | icaC::Tn(ermB) | LAC | This work; BEI resources and Fey et al. (2013) |
| JMB 7270 | hmrA::Tn(ermB) | JE2 | BEI resources and Fey et al. (2013) |
| JMB 7265 | lytN::Tn(ermB) | JE2 | BEI resources and Fey et al. (2013) |
| JMB 7267 | lytX::Tn(ermB) | JE2 | BEI resources and Fey et al. (2013) |
| JMB 7266 | sle1::Tn(ermB) | JE2 | BEI resources and Fey et al. (2013) |
| JMB 7268 | lytY::Tn(ermB) | JE2 | BEI resources and Fey et al. (2013) |
| JMB 7269 | lytZ::Tn(ermB) | JE2 | BEI resources and Fey et al. (2013) |
| JMB 7271 | lytM::Tn(ermB) | JE2 | BEI resources and Fey et al. (2013) |
| JMB2977 | parent | JE2 | BEI resources and Fey et al. (2013) |
| JMB7277 | narG::Tn (ermB) | LAC | BEI resources and Fey et al. (2013) |
| JMB 1148 | ΔhptRS | LAC | Pang et al. (2014) |
| JMB 1357 | ΔlytSR | LAC | Pang et al. (2014) |
| JMB 1330 | graS::erm | LAC | Boles et al. (2010) |
| JMB 1335 | ΔsaePQRS::spec | LAC | Nygaard et al. (2010) |
| JMB 1219 | ΔSAUSA300_1219–1220 | LAC | Pang et al. (2014) |
| JMB 1383 | ΔarlSR | LAC | Pang et al. (2014) |
| JMB 1358 | ΔphoSR | LAC | Pang et al. (2014) |
| JMB 1241 | ΔairSR | LAC | Pang et al. (2014) |
| JMB 1377 | ΔvraSR | LAC | Pang et al. (2014) |
| JMB 1333 | Δagr::tetM | LAC | Kiedrowski et al. (2011) |
| JMB 1223 | ΔkdpSR | LAC | Pang et al. (2014) |
| JMB 1359 | ΔhssSR | LAC | Pang et al. (2014) |
| JMB 1145 | ΔnreSR | LAC | Pang et al. (2014) |
| JMB 1232 | ΔSAUSA300_2558–2559 | LAC | Pang et al. (2014) |
| Other Strains | |||
| Escherichia coli PX5 | |||
| Sacchromyces cerevisiae FY2 | |||
| Plasmids used in this study | |||
| Plasmid name | Insert Locus/function | Source/Reference | |
| pJB38 | Insertless vector for cloning chromosomal gene deletions | Bose et al. (2013) | |
| pJB38_srrAB::tet | Construction of srrAB::tet allele | This work | |
| pCM28 | Insertless cloning vector | A. Horswill | |
| pCM28_srrAB | srrAB complementing vector | Mashruwala and Boyd (2017) | |
| pLL39 | Insertless cloning vector for genetic complementation | Luong and Lee (2007) | |
| pLL39_srrAB | srrAB complementing vector | This work | |
| pJB141 | atlA complementing vector | Bose et al. (2012) | |
| pJB135 | atlAGL complementing vector | Bose et al. (2012) | |
| pJB122 | atlAAMH263A complementing vector | Bose et al. (2012) | |
| pJB128 | Insertless cloning vector | Bose et al. (2012) | |
| pJB111 | atlAAM complementing vector | Bose et al. (2012) | |
| pTnTet | Construction of menF::Tn (Tet) | Bose et al. (2013) | |
Additional files
-
Supplementary file 1
Oligonucleotides used in this study for real-time quantitative PCR and cloning.
- https://doi.org/10.7554/eLife.23845.019





