Nuclear export receptor CRM1 recognizes diverse conformations in nuclear export signals
Figures
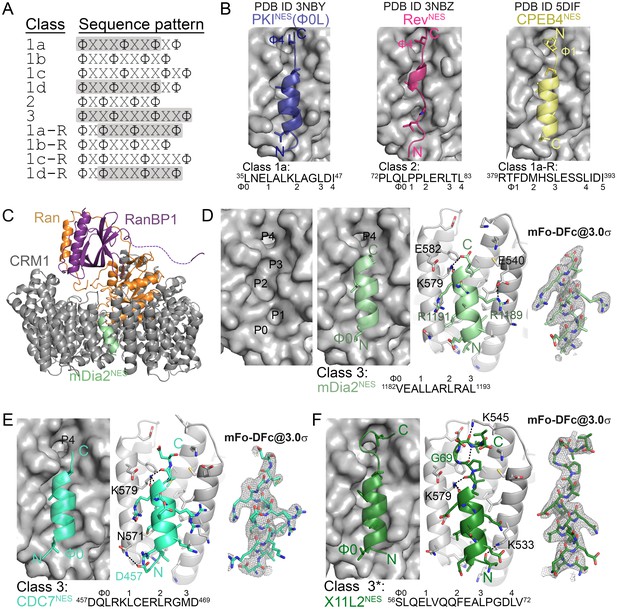
Structures of CRM1-bound NESs that match the potentially all-helical class 3 pattern.
(A) Current NES sequence patterns (Φ is Leu, Val, Ile, Phe or Met and X is any amino acid). Potential amphipathic α-helices, predicted with hydrophobic patterns of i, i + 4, i + 7 or i, i + 3, i + 7 or i, i + 3, i + 7, i + 10, are shaded grey. (B) Structure of PKINES(Ф0L) (dark blue, PDB ID: 3NBY), RevNES (pink, 3NBZ) and CPEB4NES (yellow, 5DIF) bound to the NES-binding groove of CRM1 (grey surface). NESs are shown in cartoon representations with their Φ side chains shown as sticks. (C) The overall structure of the engineered ScCRM1 (grey)-Ran•GTP (orange)-RanBP1 (purple)-mDia2NES (pale green) complex. The structure of (D) mDia2NES (pale green), (E) CDC7NES (green-cyan) and (F) X11L2NES (forest) bound to the NES-binding groove of ScCRM1 in the engineered ScCRM1-Ran-RanBP1 complex. *The X11L2NES sequence matches the class 3 pattern, but binds CRM1 according to the new hydrophobic pattern Φ0XXΦ1XXXΦ2XXΦ3XXXΦ4 that we termed the class 4 pattern. mDia2NES is not shown in the leftmost panel of (D) to view the five hydrophobic pockets (P0–P4) of the CRM1 groove. Rightmost panels of (D–F): overlays of 3.0σ positive densities of kick OMIT mFo-DFc maps (calculated with peptides omitted) and final coordinates of the NES peptides. Middle panels of (D–F): black dashes show CRM1-NES hydrogen bonds and polar contacts.
-
Figure 1—source data 1
Data collection and refinement statistics.
- https://doi.org/10.7554/eLife.23961.003
-
Figure 1—source data 2
Data collection and refinement statistics (cont.).
- https://doi.org/10.7554/eLife.23961.004
-
Figure 1—source data 3
Crystallization conditions of CRM1-NES complexes.
- https://doi.org/10.7554/eLife.23961.005
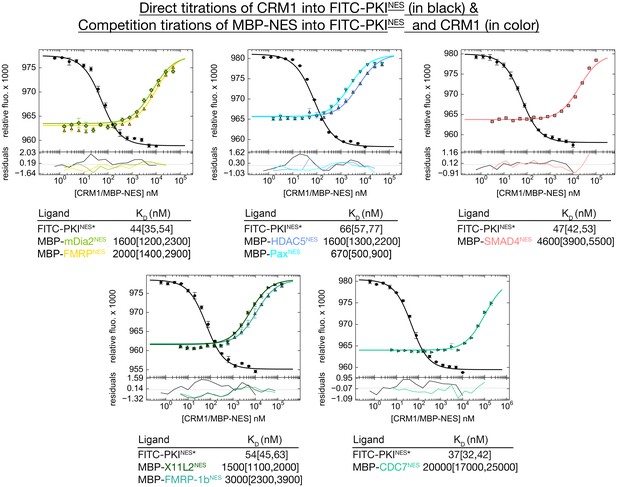
Binding affinities of NES to CRM1 measured by differential bleaching experiments.
Data from triplicate experiments of direct (titration of CRM1 to FITC-PKINES in the presence of excess RanGTP) or competition titrations (titration of MBP-NES to FITC-PKINES and CRM1 with excess RanGTP) are plotted as mean ± SD and fitted binding curves are overlaid on top. MBP-NES competes with FITC-PKINES for CRM1. Residuals are plotted at the bottom. Dissociation constants are reported below the graphs with ranges in brackets representing the 68.3% confidence interval. *Experiments done on different days were fitted using a corresponding triplicate set of direct titrations (traces in black).
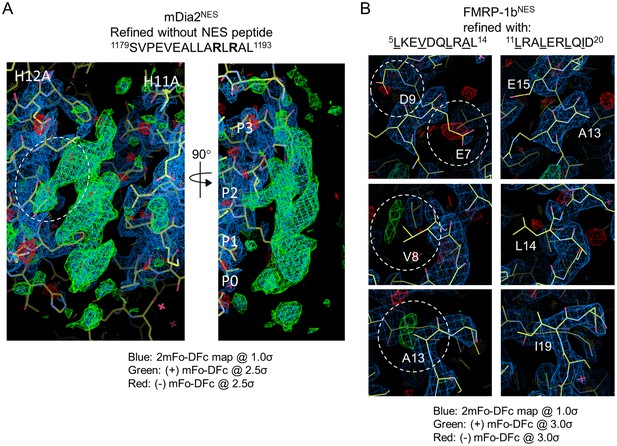
Modeling strategy for NESs with weak Φ side chain densities.
(A) The difference densities (mFo-DFc contoured at 2.5σ) for mDia2NES when the dataset was refined without the NES peptide. White circle on the left panel shows good side chain density for an arginine side chain. The image in the right panel shows the structure rotated 90 degrees to show strong side chain density for a hydrophobic residue pointing into the P2 pocket of CRM1. (B) Left panels: difference densities (mFo-DFc contoured at 3.0σ; shown in dashed white circles) for the incorrect model of FMRP-1bNES. Right panels: difference densities (mFo-DFc contoured at 3.0σ) for the correct model of FMRP-1bNES are shown at the same positions as the left panels. The absence of difference densities at side chain positions provides support for assignment of 11LRALERLQID20 to the NES peptide structure.
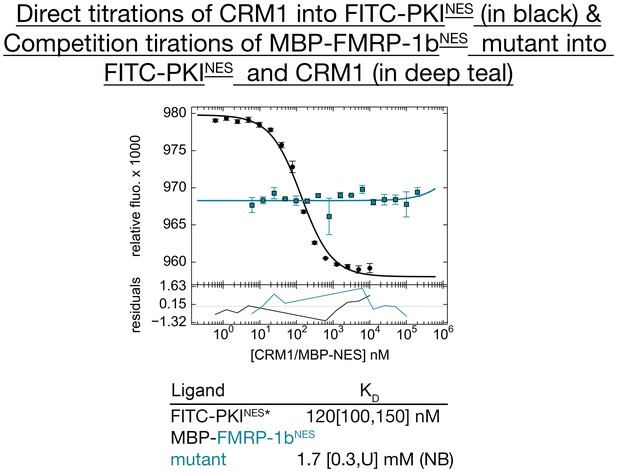
Mutagenesis of FMRP-1bNES to validate the sequence assignment of the NES.
Differential bleaching experiments of FMRP-1bNES mutant YLKEVDQLRALER show that it does not bind CRM1, thus providing support that assignment of 11LRALERLQID20 to the FMRP-1bNES is most likely correct. Triplicate direct tirations of CRM1 into FITC-PKINES in the presence of excess RanGTP is shown in black, and triplicate competition titrations of MBP-FMRP-1bNES mutant to CRM1 and FITC-PKINES is shown in teal. The KD values obtained are shown below the graph and numbers in brackets represent 68.3% confidence level (U=underteremined, NB=not binding).
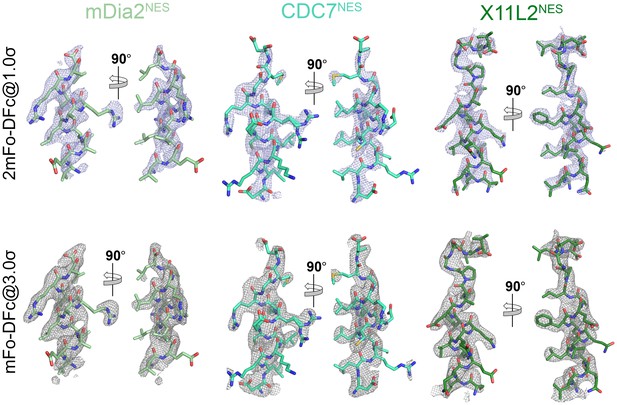
Electron densities of the NES peptides.
2mFo-DFc map meshes contoured at 1.0σ and kicked OMIT map meshes contoured at 3.0σ are overlaid on the final coordinates of the indicated structures with NES peptides shown as sticks.
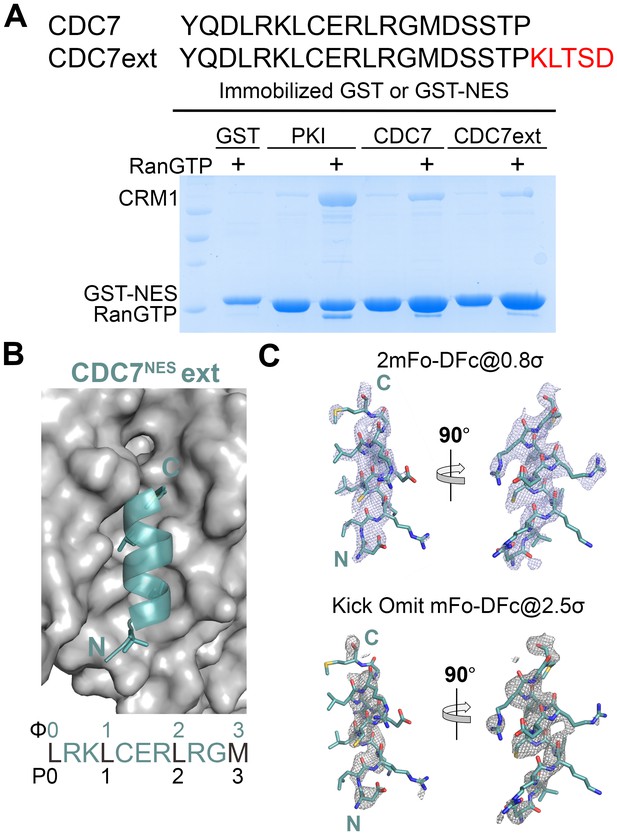
A longer CDC7NES peptide binds the CRM1 groove without interacting with the P4 pocket.
(A) Pull-down binding assay (~10 µg immobilized GST-CDC7NES and the longer GST-CDC7NESext, 2.5 µM HsCRM1 and 7.5 µM ScRanGTP in a reaction volume of 200 µL) show that both peptides binding to CRM1. (B) Structure of the longer CDC7NESext peptide (deep teal) bound to the CRM1 groove (grey surface). (C) 2mFo-DFc map (mesh) contoured at 0.8σ and kick OMIT map (mesh) at 2.5σ overlaid onto the final coordinates of CDC7NESext peptide (sticks).
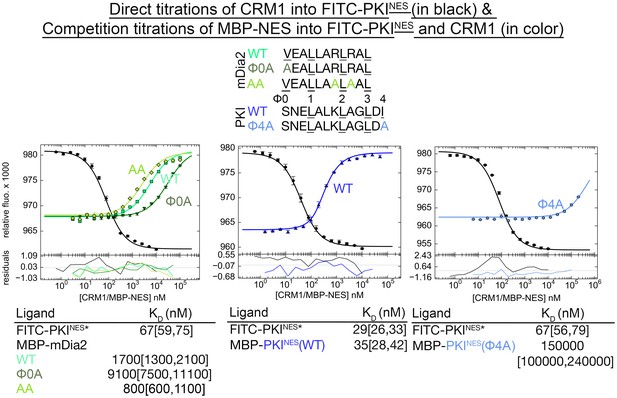
Variable number of Ф side chains are needed to have an active NES.
Sequences of wild type mDia2NES, the mDia2NES(Φ0A) and mDia2NES(AA) mutants are shown along with those for wild type PKINES and the PKINES(Φ4A) mutant. mDia2NES(Φ0A) and PKINES(Φ4A) were mutated to each have only 3 Ф side chains, leucines in positions Ф1, Ф2 and Ф3. Arg1189 and Arg1191 side chains of mDia2NES make electrostatic interactions with the CRM1 groove, and were both mutated to alanines to generate the mDia2NES(AA) mutant. Affinities of MBP-mDia2NES and PKINES mutants binding to HsCRM1 were measured using differential bleaching competition assays. Fitted binding curves are overlaid onto data points plotted as mean ± SD and residuals are plotted in the bottom. KD values are reported with ranges in brackets representing the 68.3% confidence interval. *Experiments done on different days were fitted using a corresponding triplicate set of direct titrations (in black). mDia2NES(Φ0A) and PKINES(Φ4A) bind CRM1 very differently even though they both have the same 3 Ф (Leu) side chains. mDia2NES(Φ0A) retains CRM1 binding (KD = 9.1 [7.5, 11.1] μM, vs wild type mDia2NESKD = 1.7 [1.3, 2.1] μM) but the PKINES(Φ4A) mutant no longer binds CRM1 (KD = 150 [100, 240] μM vs wild type PKI NESKD = 35 [26, 42] nM). mDia2NES(AA) binds CRM1 2-fold tighter (KD = 800 [600, 1100] nM), suggesting that electrostatic interactions involving Arg1189 and Arg1191 of the NES are not critical for CRM1 binding. Loss of the electrostatic interactions may be compensated by the gain of helical propensity, which may lead to stabilization of the mDia2NES helix.
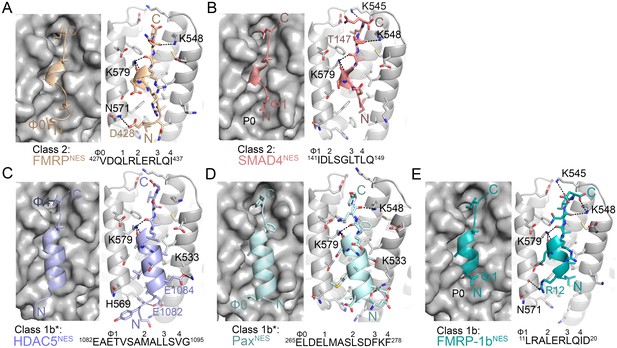
Structures of NESs with non-helical sequence patterns.
(A–E) FMRPNES (light orange), SMAD4NES (salmon), HDACNES (slate), PaxNES (pale cyan) and FMRP-1bNES (deep teal) bound to the ScCRM1 groove. Black dashes show CRM1-NES hydrogen bonds and polar contacts, and unoccupied CRM1 hydrophobic pockets are labeled. *HDAC5NES and PaxNES sequences match the class 1b pattern, but both peptides bind CRM1 using Ф residues that match class 1a pattern. Average displacements of each Φ Cα in the eight NESs (including mDia2NES, CDC7NES, X11L2NES in Figure 1) from the equivalent Φ Cα of PKINES(Ф0L) are 1.3 ± 0.6 (Φ4), 0.8 ± 0.5 (Φ3), 0.7 ± 0.4 (Φ2), 0.9 ± 0.3 (Φ1) and 1.8 ± 0.9 (Φ0) Å.
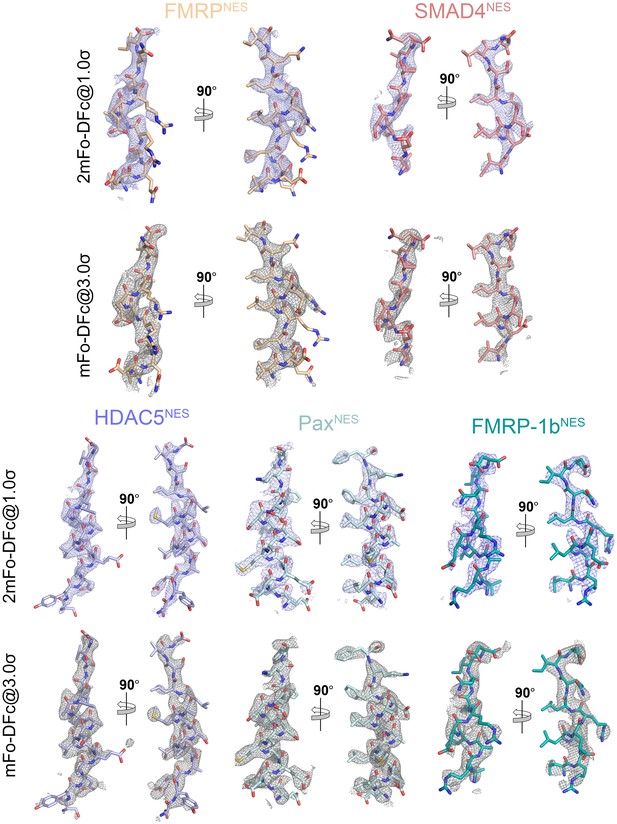

Electron densities of the NES peptides.
2mFo-DFc map meshes contoured at 1.0σ and kicked OMIT map meshes contoured at 3.0σ are overlaid on the final coordinates of the indicated structures with NES peptides shown as sticks.
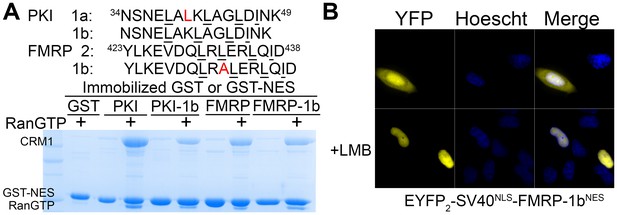
Engineering class 1b NESs.
(A) Sequences of class 1b NESs generated from PKINES and FMRPNES. The binding of mutants to CRM1 were tested in pull-down assay with ~7.5 µg immobilized GST-NESs, 2.5 µM HsCRM1 and 7.5 µM ScRanGTP in 200 µL reaction volumes. The reactions were extensively washed and visualized with SDS/PAGE and Coomassie staining. (B) Leptomycin B (LMB) sensitive nuclear export activity of FMRP-1bNES was verified by expressing EYFP2-SV40NLS-FMRP-1bNES in HeLa cells and imaged with spinning disk confocal microscope (60X).
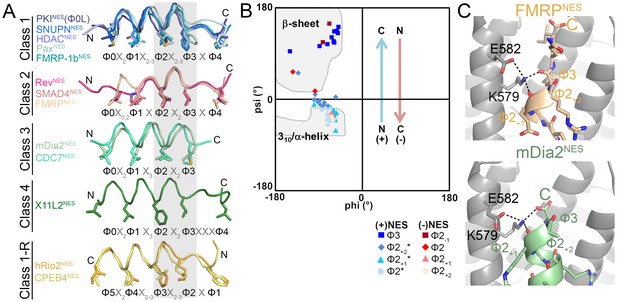
NESs adopt diverse conformations to bind CRM1, sharing only a small one turn of the helix structural element.
(A) Overlay of 13 different CRM1-bound NESs: mDia2NES, CDC7NES, X11L2NES, FMRPNES, SMAD4NES, HDACNES, PaxNES and FMRP-1bNES, SNUPNNES (3GB8), PKINES(Φ0L) (3NBY), RevNES (3NBZ), hRio2NES (5DHF), CPEB4NES (5DIF) (shown as ribbons with Φ residues in sticks; their CRM1 H12A helices were superimposed). A grey rectangle highlights the only secondary structural element shared by all 13 NESs: a single turn of helix. (B) Ramachandran plot of phi/psi angles of the four residues in each of the conserved one-turn of helix. Arrows indicate changes in psi angles along the polypeptide direction. For example, (+) HDAC5NES: Φ2 Ψ = −43.5°, Φ2+1Ψ = −21.7°, Φ2+2Ψ = −1.3°, Φ3 Ψ = 133.8° and (−) CPEB4NES: Φ2+2Ψ = −40.2°, Φ2+1Ψ = −31.0°, Φ2 Ψ = 16.5°, Φ2-1Ψ = 111.5°. *Residues in SNUPNNES plotted are Φ2+1, Φ2+2, Φ2+3 and Φ3. (C) Detailed view of niche motifs in (+) NESs, FMRPNES and mDia2NES.
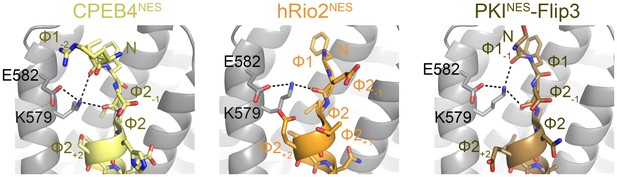
NES main chain hydrogen bonds with ScCRM1 Lys579 in (−) NESs.
Structures of CPEB4NES (5DIF), hRio2NES (5DHF) and PKINES-Flip3 mutant (5DH9) bound to ScCRM1, showing hydrogen bonds made by the Lys579 side chain of ScCRM1. Niche motifs observed in (+) NESs are absent here, but in CPEB4NES and PKINES-Flip3 mutant, Lys579 is still coordinated by two main chain carbonyls from Ф1-1 and Ф2-1.
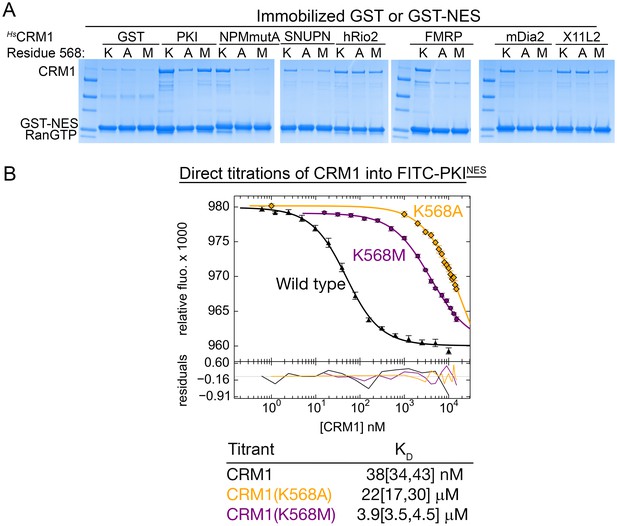
The Lys568 of HsCRM1 is important for NES binding.
(A) Pull-down binding assays with ~5 µg immobilized GST-NESs, 500 nM wild type HsCRM1, the HsCRM1(K568A) or HsCRM1(K568M) mutant and 1.5 µM ScRanGTP in 1 mL reaction volumes. Most of the NESs showed decreased binding to the K568A and K568M mutants compared to wild type CRM1. Because of the limited dynamic range of detection in pull-down assays, it is unclear if binding of hRio2NES and X11L2NES to mutant versus wild type CRM1 proteins are different. (B) Binding affinity of FITC-PKINES to HsCRM1, HsCRM1(K568A) and HsCRM1(K568M) measured using differential bleaching. Different concentration regimes were used for different mutants to obtain meaningful data points and at least three titrations were performed for each CRM1. Fitted binding curves are overlaid onto data points plotted as mean ± SD and residuals are plotted in the bottom. KD values are reported with ranges in brackets representing the 68.3% confidence interval.
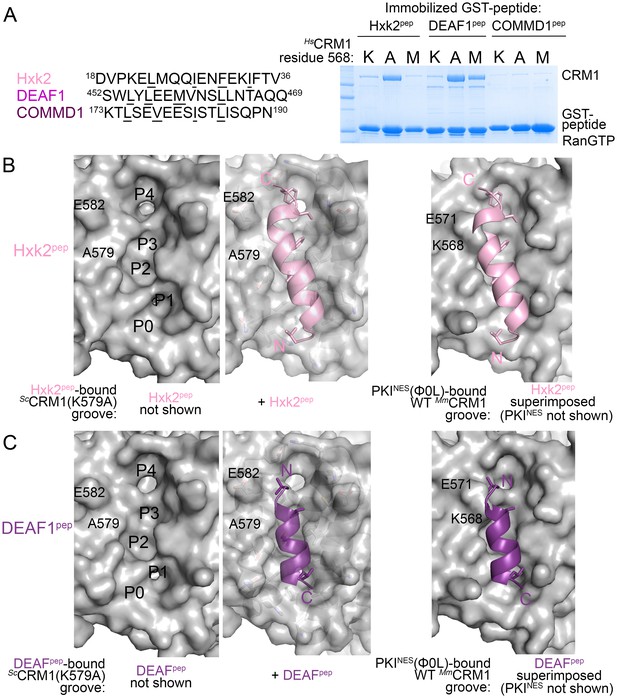
HsCRM1 Lys568 is a selectivity filter.
(A) False positive NES sequences with Φ residues of consensus matches underlined. Pull-down binding assay of ~5 µg immobilized GST-NESs and 7.5 µM ScRanGTP with 2.5 µM of either wild type HsCRM1 or mutant HsCRM1(K568A) or HsCRM1(K568M) in 200 µL reactions. (C–D) Structures of Hxk2pep (pink) (C) and DEAF1pep (purple) (D) bound to ScCRM1(K579A). Left panels, peptides were removed to show the surface of the mutant ScCRM1(K579A) groove. Middle panels, peptides bound to the mutant ScCRM1(K579A). Right panels, CRM1(K579A)-bound Hxk2pep and DEAF1pep superimposed onto the PKINES(Ф0L)-bound MmCRM1 groove (3NBY; CRM1 H12A helices were aligned; PKINES not shown) to show steric clash of the Hxk2pep and DEAF1pep peptides with the MmCRM1 Lys568 side chain.
-
Figure 4—source data 1
Data collection and refinement statistics and crystallization conditions.
- https://doi.org/10.7554/eLife.23961.019
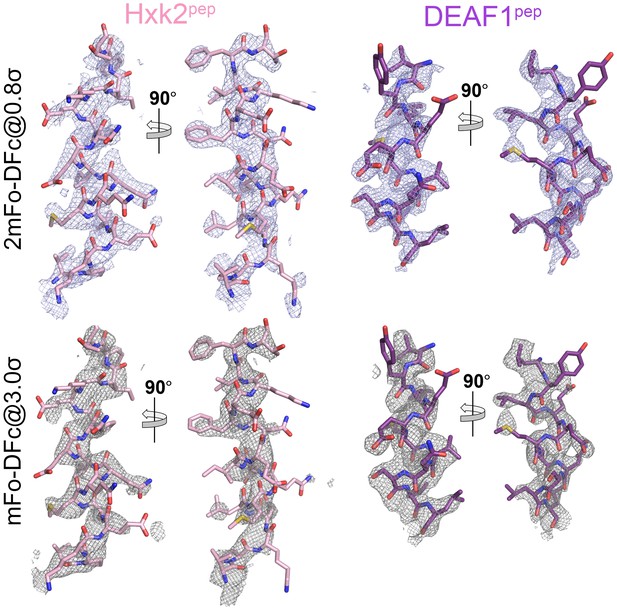
Electron densities of the HXK2pep and DEAFpep false positive NES peptides bound to the ScCRM1(K579A) mutant.
2mFo-DFc map meshes contoured at 0.8σ and kicked OMIT map meshes contoured at 3.0σ are overlaid on the final coordinates of the indicated structures with NES peptides shown as sticks.
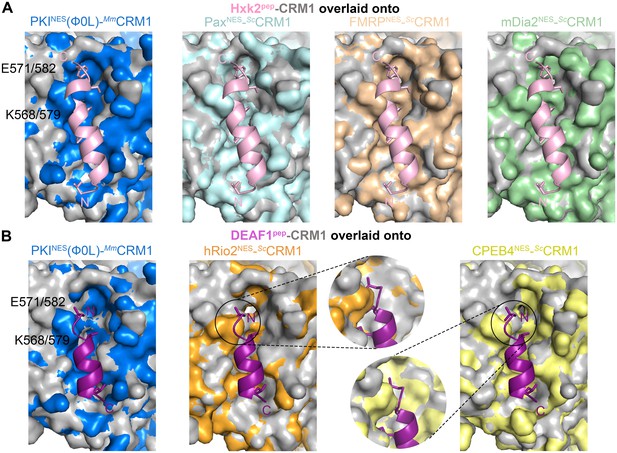
Additional overlays of Hxk2pep and DEAF1pep onto NES-bound wild type CRM1 grooves.
(A) Hxk2pep-ScCRM1(K579A) (Hxk2pep in pink cartoon and ScCRM1(K579A) shown as grey surface) superimposed onto the PKINES(Ф0L)-bound MmCRM1 groove (wild type CRM1, 3NBY; blue surface), the PaxNES- bound ScCRM1 groove (pale cyan), the FMRPNES-bound ScCRM1 (wheat) and the mDia2NES-bound ScCRM1 (pale green) by aligning their CRM1 H12A helices. The superpositions show steric clash of the Hxk2pep peptide with the MmCRM1 Lys568 or ScCRM1 Lys579 side chains. (B) DEAF1pep-ScCRM1(K579A) (DEAFpep in magenta cartoon and ScCRM1(K579A) shown as grey surface) superimposed onto the PKINES(Ф0L)-bound MmCRM1 groove (wild type CRM1, 3NBY; blue surface), the hRio2NES-bound ScCRM1 (orange, 5DHF) and the CPEB4NES-bound ScCRM1 (yellow, 5DIF). The N-terminus of DEAF1pep clashes with of MmCRM1 Lys568 or ScCRM1 Lys579 side chains. Zoomed in views are rotated to show proximity of DEAF1pep Ф1XФ2 to the ScCRM1 Lys579 side chain.





