SENP8 limits aberrant neddylation of NEDD8 pathway components to promote cullin-RING ubiquitin ligase function
Figures
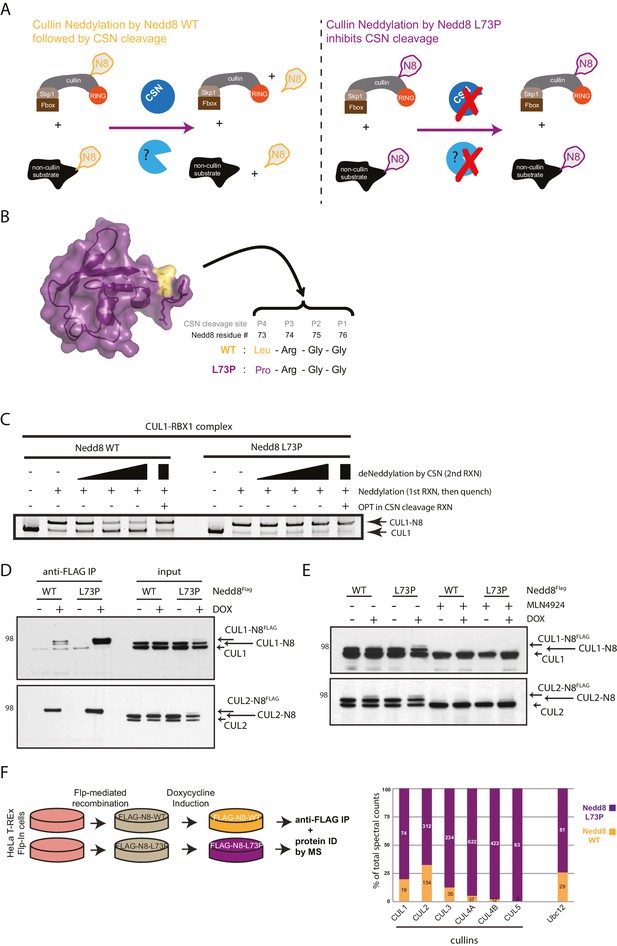
Expression of a deconjugation-resistant NEDD8 mutant (L73P) stabilizes neddylation of cullins and other non-cullin substrates.
(A) Schematics of the regulation of NEDD8 substrates by modification with either WT- (left panel) or L73P-Nedd8 (right panel), and deneddylation by NEDD8-specific proteases. CSN is the deneddylase responsible for deconjugating NEDD8 from cullin substrates, but proteases regulating deneddylation of non-cullin substrates are largely uncharacterized. (B) Surface representation of NEDD8 (pdb: 1NDD) and details of its C-terminal tail, showing its proteolytic cleavage site and location of the L73P mutation. (C) Recombinant CRL1/Rbx1 was in vitro neddylated by purified His-NEDD8-WT or His-NEDD8-L73P, in the presence of E1 and E2 enzymes and ATP. Reactions were quenched, and recombinant CSN was added at increasing concentrations to monitor the ability of each NEDD8 moiety to be deconjugated from CUL1. OPT (1,10-orthophenatroline, 1 mM) was added to samples containing the highest concentration of CSN (last lane) to completely inhibit CSN activity. (D) FLAG-NEDD8-WT or FLAG-NEDD8-L73P was induced in HeLa-FlpIn-N8 cells using 1 ug/mL doxycycline for 48 hr prior to collection. Whole-cell lysates of untreated or Dox-treated cells were incubated with anti-FLAG beads to purify FLAG-NEDD8-conjugates. Immunoblots of input and IP samples were analyzed for FLAG-NEDD8-modified CUL1 and CUL2. (E) HeLa-FlpIn-N8 cells were treated with or without Dox as in D to induce FLAG-NEDD8-WT or FLAG-NEDD8-L73P, and subsequently incubated with or without the of the CRL inhibitor MLN4924 (5 µM for 4 hr) before harvesting. Whole-cell extracts were analyzed for FLAG-NEDD8-conjugated CUL1 and CUL2. (F) (left panel) Workflow for expression and purification of FLAG-NEDD8-WT and FLAG-NEDD8-L73P for MS analysis. (right panel) Percentages of total spectral counts detected in FLAG-IPs from cells expressing either FLAG-NEDD8-WT (orange bars) or FLAG-NEDD8-L73P (purple bars). The numbers in the columns indicate actual spectral counts. The IPs were performed on lysates from the same number of cells.
-
Figure 1—source data 1
NEDD8- modified peptides identified by MS analysis of FLAG-NEDD8 IP samples.
- https://doi.org/10.7554/eLife.24325.003
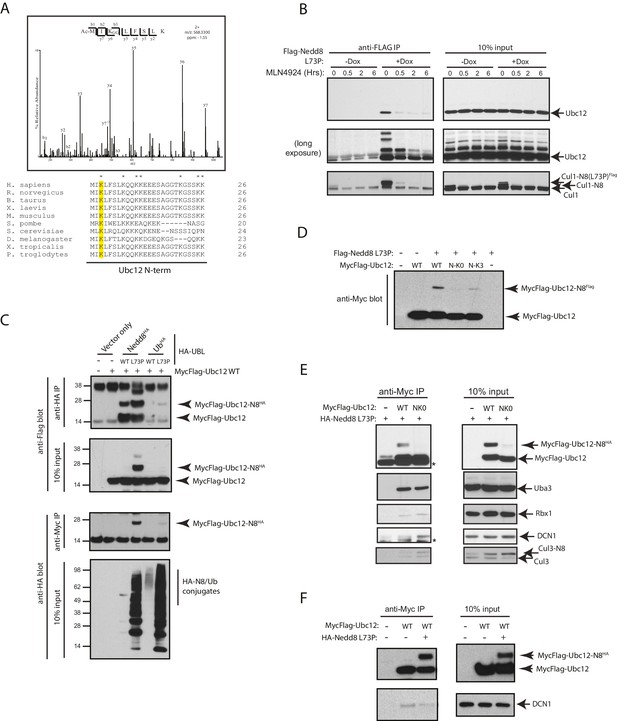
Nedd8 L73P is conjugated to Ubc12 in vitro and in cells.
(A) An HCD MS2 spectrum of the doubly charged Ubc12 N-terminal peptide, MIKLFSLK, N-terminally acetylated and K-ε-GG modified on Lys3 (top) and sequence alignment of Ubc12 N-termini from the indicated species (bottom). The conserved Lys3 residue in the Ubc12 N-terminus identified as a K-ε-GG-modified residue from the MS dataset is highlighted in yellow. Lys residues mutated to Arg in the Ubc12 NK0 mutant are indicated by asterisks. Sequence alignments were generated using Clustal Omega. (B) HeLa-Flp-In-NEDD8 cells were either left untreated or treated with Dox to induce FLAG-NEDD8-L73P expression, and subsequently treated with MLN4924 (5 µM) for the indicated times. Immunoblots of input and anti-FLAG IP samples were analyzed for modified Ubc12 and CUL1. (C) HA-tagged (WT or L73P) NEDD8 or Ub constructs were co-transfected with MycFlag-Ubc12 in HEK293T cells. Conjugation of either NEDD8 or Ub to Ubc12 was analyzed by reciprocal IPs using either anti-HA or anti-Myc antibodies and immunoblotting. (D) MycFlag-tagged WT, NK0 , or N-K3 (NK0 mutant with K3 restored) Ubc12 constructs were co-transfected with Flag-Nedd8-L73P in HEK293T cells. Modified Ubc12 was analyzed in whole-cell lysates by immunoblotting with anti-Myc antibody. (E) WT or NK0 MycFlag-Ubc12 was co-expressed in HEK293T cells with HA-NEDD8-L73P. Cell lysates were incubated with anti-Myc antibody to IP MycFlag-Ubc12, and input and IP samples were analyzed by immunoblotting with the indicated antibodies. Asterisks signify non-specific bands. (F) WT MycFlag-Ubc12 was transfected into HEK293T cells in the presence or absence of HA-Nedd8-L73P. Anti-Myc antibody was used to IP MycFlag-Ubc12 from cell extracts, and input and IP samples were analyzed by immunoblotting with the indicated antibodies.
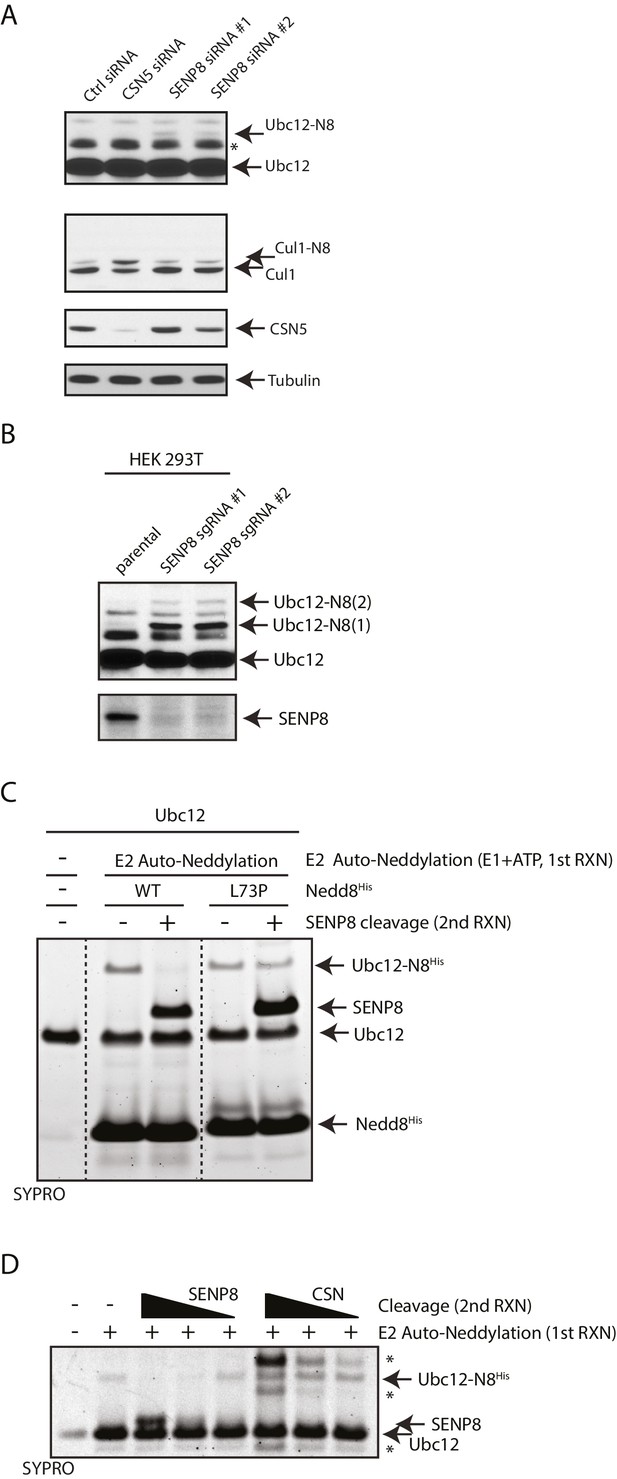
SENP8 regulates Ubc12 deneddylation.
(A) HeLa cells were treated with control, CSN5, or SENP8 siRNAs for 48 hr prior to harvesting. Whole-cell lysates were analyzed by immunoblotting for the indicated proteins. (B) SENP8 knockout clones (#1 and #2) of HEK293T cells were derived as specified in Materials and methods. Cell lysates were analyzed by immunoblotting with the indicated antibodies. (C) Recombinant Ubc12 was auto-neddylated in vitro in the presence of E1 enzyme, ATP, and either WT or L73P His-tagged NEDD8. Reactions were quenched, and SENP8 was added to deneddylate Ubc12-NEDD8 conjugates. Deneddylation of recombinant Ubc12 was analyzed by SDS-PAGE and SYPRO staining. (D) Ubc12 was in vitro neddylated as in (C) in the presence of WT His-tagged NEDD8. Decreasing concentrations of either recombinant SENP8 or CSN were added to quenched reactions, and Ubc12 deneddylation was assessed by SDS-PAGE and SYPRO staining.
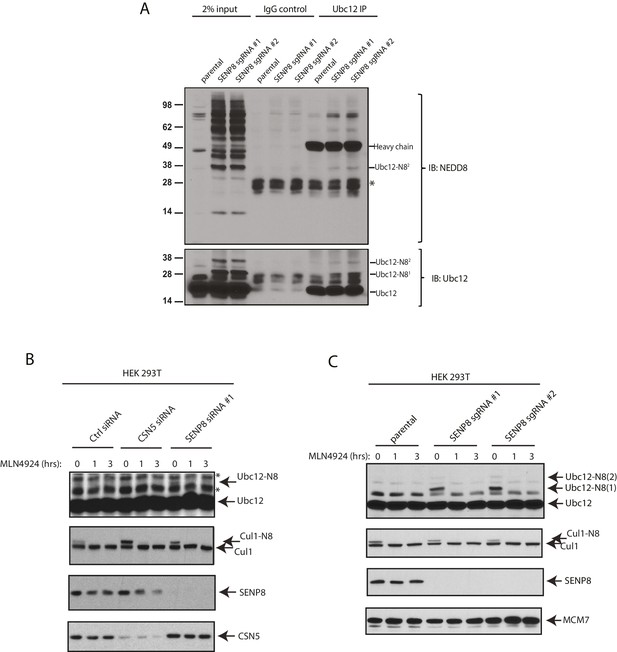
Detection of NEDD8-modified Ubc12 in SENP8-deficient cells.
(A) Whole-cell lysates of parental and SENP8-deficient HEK 293 T cells were incubated with or without Ubc12 antibody, and immune complexes were captured using Protein G beads. Input and IP samples were analyzed for NEDD8 or Ubc12 conjugates. Non-specific band is indicated by *. (B) HEK293T cells were treated with the indicated siRNAs for 48 hr and subsequently treated with or without 5 µM MLN4924. Cells were harvested for immunoblotting analysis at the indicated times following MLN4924 treatment. (C) CRISPR-generated SENP8 knockout cells were treated for the specified times with 5 µM MLN4924 and harvested for immunoblotting analysis.
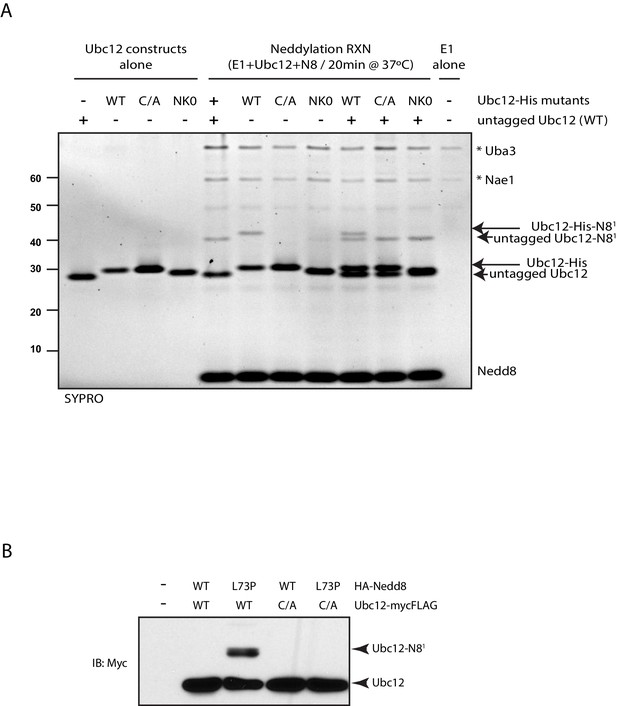
Modification of Ubc12 with NEDD8-L73P is dependent on Ubc12 catalytic activity.
(A) In vitro neddylation reactions were performed with WT, NK0, or catalytically inactive (C111A) Ubc12 in the presence of E1, WT NEDD8, and ATP. As controls, reactions were performed with Ubc12 constructs only, E1 only, and untagged Ubc12 constructs. NEDD8 modification of Ubc12 was assessed by SDS-PAGE and SYPRO staining. (B) HA-tagged WT or L73P NEDD8 constructs were co-expressed with MycFLAG-tagged WT or C111A Ubc12 constructs in HEK293T cells. Ubc12 conjugates were analyzed by immunoblotting with anti-Myc antibody.
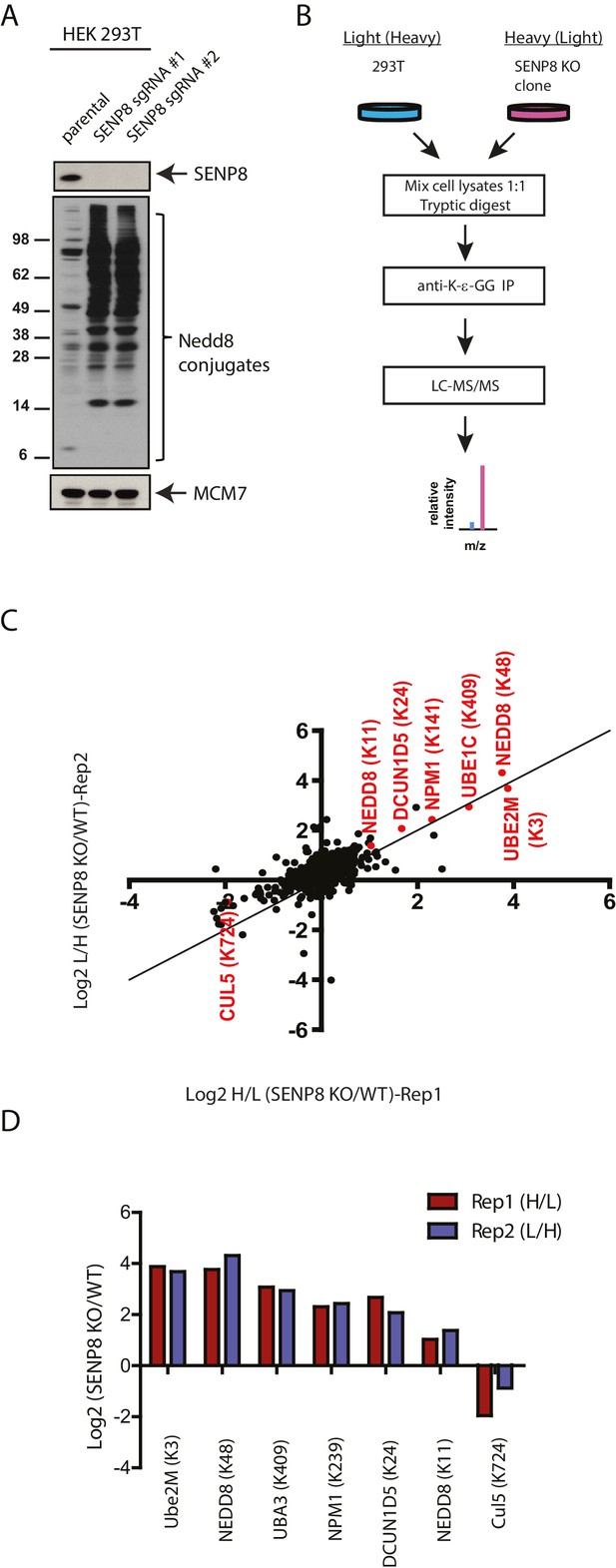
Identification of endogenous neddylation substrates in SENP8-deficient cells.
(A) Lysates from parental and CRISPR-generated SENP8 knockout HEK293T cells were immunoblotted for the indicated proteins. For this figure and all subsequent figures, MCM7 serves as a loading control. (B) Workflow for identification of K-ε-GG-remnant containing peptides in WT and SENP8 knockout cells by antibody-based enrichment and quantitative SILAC mass spectrometry. (C) Scatter plot showing Log2 ratios of all K-ε-GG sites identified in both the forward and reverse SENP8 KO:WT SILAC samples, prepared as in B. K-ε-GG sites of interest identified in both replicates are highlighted in red. Line of identity with slope = 1 is shown for reference. (D) Log2 ratios of K-ε-GG modification sites highlighted in C. Error bars represent SEM.
-
Figure 4—source data 1
SILAC analysis of K-ε-GG remnant-containing peptides detected in untreated parental and SENP8 knockout cell lysates.
- https://doi.org/10.7554/eLife.24325.009
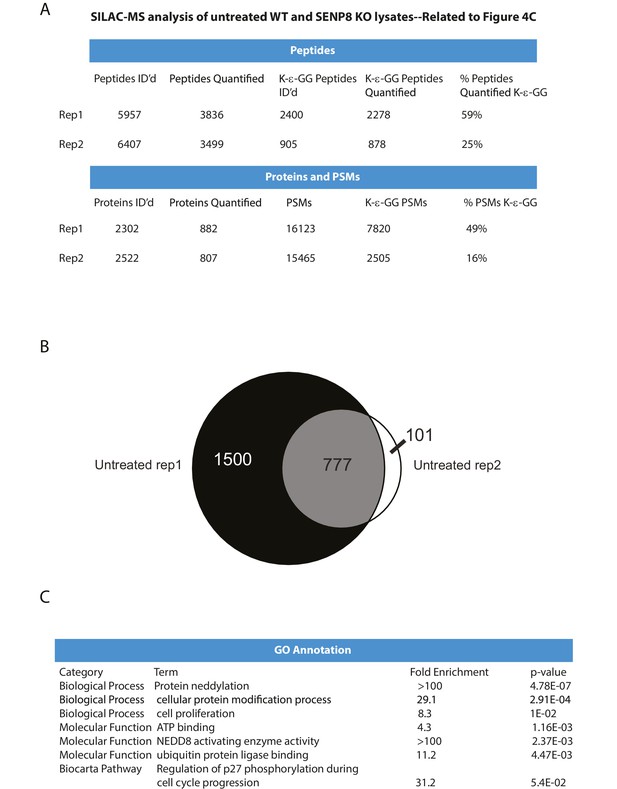
Statistical information related to K-ε-GG MS screen comparing untreated parental and SENP8 knockout cell lysates (Figure 4B).
(A) Quantification of K-ε-GG sites, peptides, proteins, and peptide spectrum matches (PSMs) in replicates 1 and 2 of Figure 4B. (B) Venn diagram showing degree of overlap between K-ε-GG remnant peptides quantified in replicates 1 and 2. (C) Analysis of Gene Ontology (GO) annotations of K-ε-GG remnant peptides with a SILAC ratio (SENP8 KO/WT) of >2 from replicates 1 and 2 of Figure 4B. GO term analysis was performed using the DAVID Gene Functional Classification Tool (Huang et al., 2009b, 2009c).
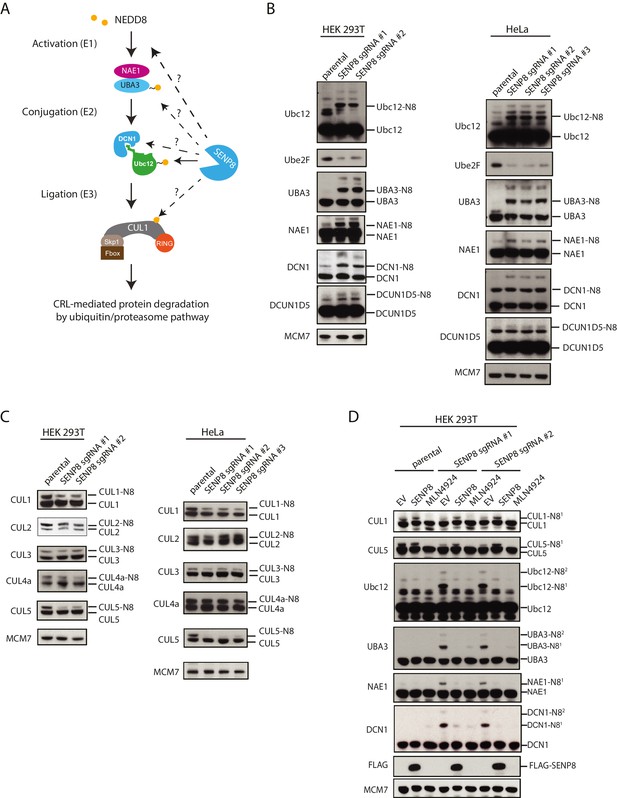
Loss of SENP8 leads to aberrant neddylation of NEDD8 pathway components.
(A) Schematic of the NEDD8 conjugation pathway. Dotted arrows represent potential points of regulation by SENP8-mediated deneddylation. Orange circle represents NEDD8. (B and C) Cellular extracts from parental and CRISPR-generated SENP8 knockout HEK293T (left) and HeLa (right) cells were subjected to immunoblot analysis for (B) NEDD8 pathway components and (C) cullin subunits. NEDD8-conjugated substrates are indicated. MCM7 serves as a loading control. (D) Parental or SENP8 knockout HEK293T cells were transfected with empty vector (EV) or a FLAG-SENP8-WT construct and analyzed for NEDD8-conjugated proteins by immunoblotting. As a control, samples from each cell line were treated with 0.3 µM MLN4924 for 24 hr to completely inhibit neddylation of the indicated proteins.
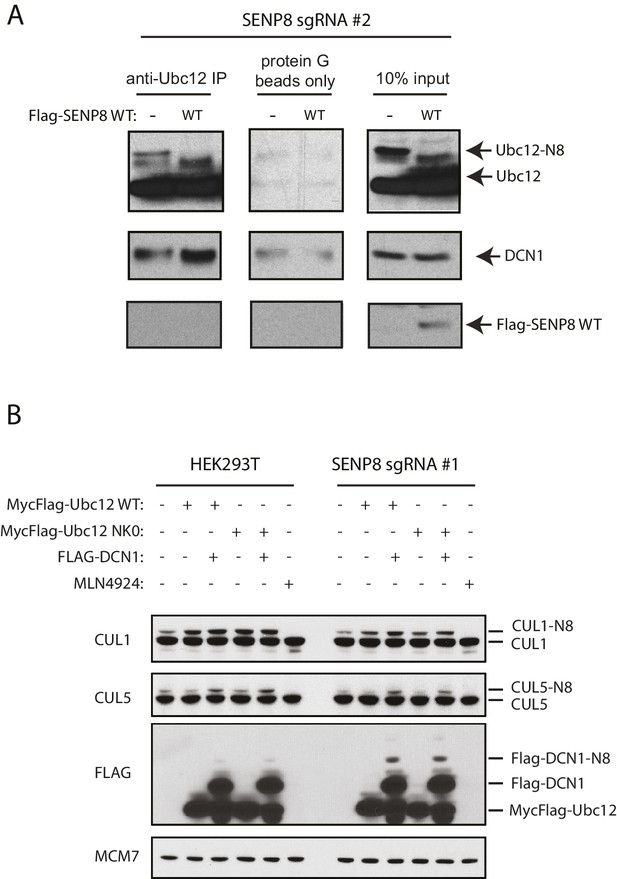
Correction of DCN1 binding and cullin neddylation defects in SENP8-deficient cells.
(A) SENP8-deficient HEK293T cells were transfected with either EV or SENP8 WT plasmids, harvested, and endogenous Ubc12 was immunoprecipitated from lysates using anti-Ubc12 antibody. Relative binding to endogenous DCN1 was assessed by immunoblotting. (B) Parental or SENP8 knockout HEK293T cells were transfected with Ubc12 WT, Ubc12 NK0, and DCN1 plasmids, or treated with 5 µM MLN4924 for 1 hr as indicated. Cell lysates were analyzed by immunoblotting with the antibodies shown.
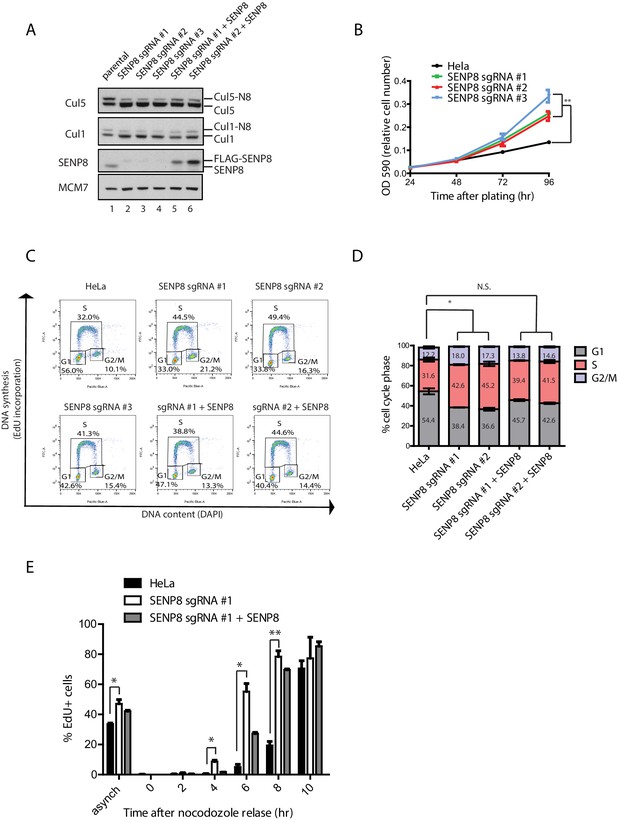
SENP8 depletion contributes to altered cell cycle progression and cell growth.
(A) SENP8 knockout (lanes 2–4) and SENP8-rescued (lanes 5–6) HeLa cell lines were generated as specified in Materials and methods. Whole-cell lysates were analyzed by immunoblotting with the indicated antibodies. (B) Parental and SENP8 knockout HeLa cells were plated and stained with crystal violet at the indicated times. OD590 was measured to calculate relative cell numbers at each time point for triplicate samples. Error bars represent SEM. p Values were calculated for the 96 hr time point by paired Student’s t-test. For all figures, asterisks represent either *p<0.05 or **p<0.01. (C and D) Parental and SENP8 knockout HeLa cells were pulse-labeled with 10 µM EdU for 30 min and analyzed by flow cytometry for DNA synthesis (EdU) and DAPI staining for DNA content (10,000 events per sample). Percentages of cells in G1, S, and G2/M phases are graphed for three biological replicates. Error bars indicate SEM. p Values were calculated for G1 populations by paired Student’s t test. (E) Parental, SENP8-deficient, and SENP8-rescued HeLa cells were synchronized in prometaphase by thymidine-nocodozole block, collected by mitotic shake-off, and released into fresh media for the indicated times. Percentages of EdU-positive cells from three biological replicates were analyzed at each time point following nocodozole release. Statistical significance was calculated using a paired Student’s t test. Error bars indicate SEM.
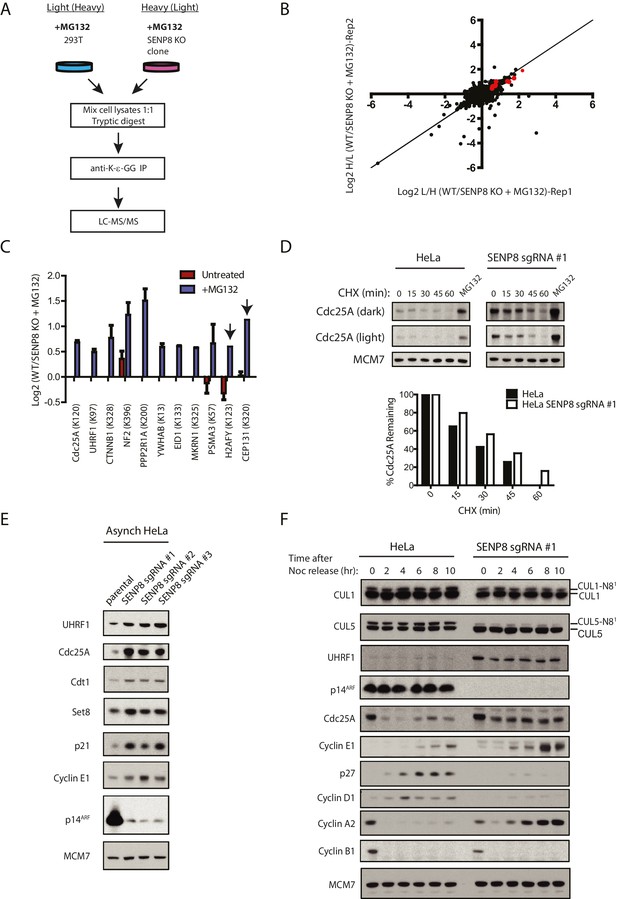
Loss of SENP8 contributes to decreased ubiquitylation and increased stability of CRL substrates.
(A) Workflow for identification of K-ε-GG-modified peptides in MG132-treated WT and SENP8 knockout cells by antibody-based enrichment and quantitative SILAC-MS. Light- and Heavy-labeled HEK293T cells were treated with 10 µM of the proteasome inhibitor MG132 for 2 hr prior to harvesting and further processing. K-ε-GG-remnant-containing peptides of interest in (C) are highlighted in red. (C) Scatter plot showing Log2 ratios of all K-ε-GG sites identified in both the forward and reverse WT: SENP8 KO MG132-treated SILAC samples, prepared as in A. Line of identity with slope = 1 is shown for reference. (D) Log2 ratios of selected K-ε-GG modification sites in untreated versus MG132-treated SILAC samples. Absence of bars indicates that a particular K-ε-GG modification site was not detected. K-ε-GG sites that were detected in only one of the two replicate samples are indicated by arrows. Error bars represent SEM. (C) Parental and SENP8 knockout HeLa cells were treated with 30 µg/ml cycloheximide (CHX) for the indicated times and subjected to immunoblot analysis for Cdc25A levels. As a control, cell lines were treated with 10 µM of the proteasome inhibitor MG132 for 1.5 hr. Percentages of Cdc25A remaining were quantified in the parental and CRISPR knockout cell lines by densitometry analysis using ImageJ software. (D) Relative levels of CRL substrates were analyzed in lysates from parental and SENP8-deficient HeLa cells by immunoblotting analysis. (E) Parental and SENP8 knockout HeLa cells were synchronized in prometaphase by sequential thymidine and nocodozole blocks, released into fresh media, and collected at the specified times. Cell lysates were subjected to immunoblotting analysis for the indicated proteins.
-
Figure 7—source data 1
SILAC analysis of K-ε-GG remnant-containing peptides detected in MG132-treated parental and SENP8 knockout cell lysates.
- https://doi.org/10.7554/eLife.24325.015
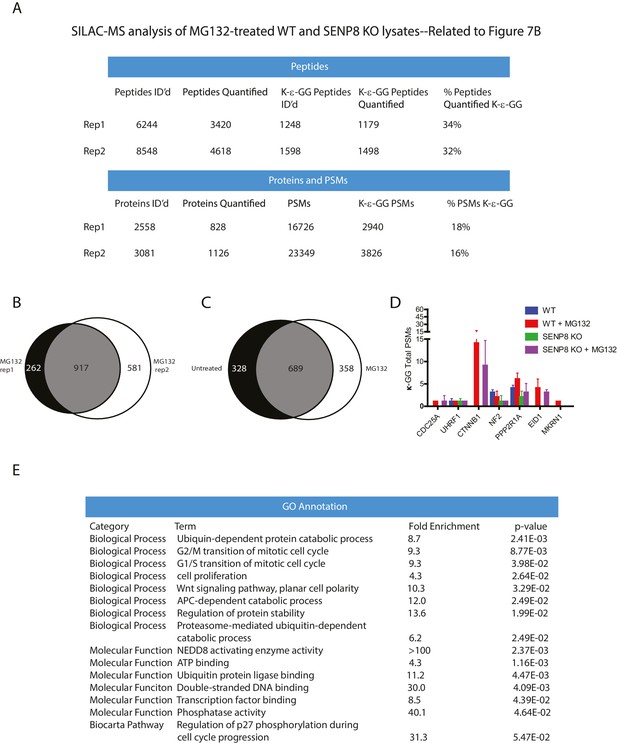
Statistical information related to K-ε-GG MS screen comparing MG132-treated parental and SENP8 knockout cell lysates (Figure 7B).
(A) Quantification of K-ε-GG sites, peptides, proteins, and PSMs in replicates 1 and 2 of Figure 7B. (B) Venn diagram showing degree of overlap between K-ε-GG remnant peptides quantified in replicates 1 and 2 (C) Venn diagram comparing overlap between all K-ε-GG-remnant containing peptides recovered from untreated (Figure 4B) and MG132-treated (Figure 7B) SILAC datasets. (D) Total PSMs detected from untreated (Figure 4B) and MG132-treated (Figure 7B) SILAC datasets for selected proteins of interest. Error bars represent SD. (E) Analysis of Gene Ontology (GO) terms of K-ε-GG remnant peptides with a SILAC ratio (WT/SENP8 KO) of >1.5 from replicates 1 and 2 of Figure 7B. GO term analysis was performed using the DAVID Gene Functional Classification Tool (Huang et al., 2009b, 2009c).
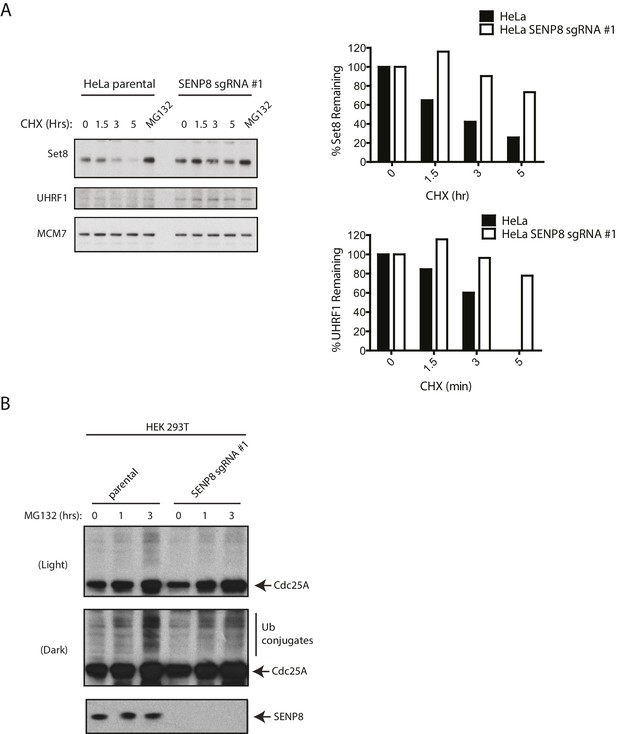
SENP8-deficient cells show increased stability and reduced ubiquitylation of CRL substrates.
(A) Parental and SENP8 knockout HeLa cells were treated with 30 µg/ml cycloheximide (CHX) for the indicated times and subjected to immunoblotting analysis for Set8 and UHRF1 levels. As a control, cell lines were treated with 10 µM of the proteasome inhibitor MG132 for 1.5 hr. Percentages of Set8 and UHRF1 remaining were quantified in the parental and CRISPR knockout cell lines by densitometry analysis using ImageJ software. (B) Parental and SENP8-deficient HEK293T cells were treated with 10 µM of MG132 for the specified times prior to harvesting for immunoblotting analysis.
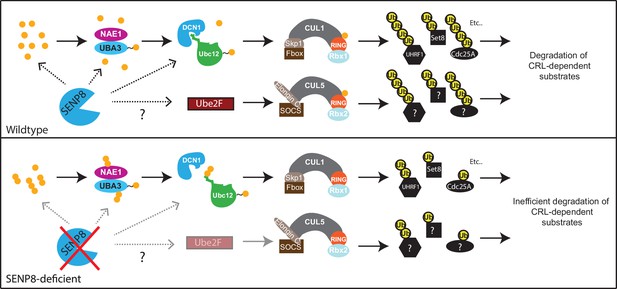
Model.
A model depicting the role of SENP8 in regulating reversible neddylation of NEDD8 conjugation pathway components.





