Tuning of in vivo cognate B-T cell interactions by Intersectin 2 is required for effective anti-viral B cell immunity
Figures
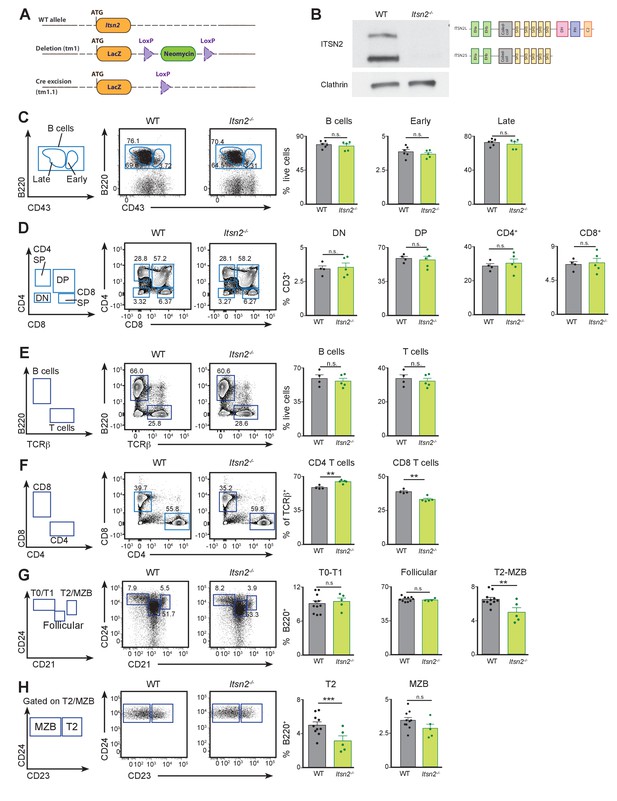
Lymphocyte development is not compromised by ITSN2 deletion.
(A) Genetic approach used to delete ITSN2. A LacZ cassette was inserted in the Itsn2 locus to disrupt protein expression. A neomycin resistance cassette flanked by two loxP sites was used as a selection marker, and subsequently excised by Cre-mediated recombination (KOMP allele tm1.1). (B) Naïve B cells were purified from the spleens of WT and Itsn2-/- mice and protein expression of ITSN2 (upper immunoblot, two isoforms as illustrated on the right) and clathrin heavy chain (lower immunoblot) were detected by Western blot. (C) Bone marrow from WT and Itsn2-/- littermates was analysed by flow cytometry using the gating strategy shown on the left. B cell progenitors (B220+) were divided into immature (CD43+) and mature (CD43-) cells on the basis of CD43 expression. Data quantified in the panels on the right show the percentage of live cells in the indicated gates. (D) Thymus from WT and Itsn2-/- littermates were analysed by flow cytometry. Gating strategy is shown on the left. T cell progenitors (CD3+) were divided into double negative (CD4- CD8-), double positive (CD4+ CD8+), CD4 single positive (CD4+ CD8-) and CD4 single positive (CD4- CD8+) cells. Data quantified in the panels on the right show the percentage of CD3+ cells in the indicated gates. (E–H) Spleens from WT and Itsn2-/- were analysed by flow cytometry. (E) Identification and quantification of B cells (B220+ TCRβ−) and T cells (B220- TCRβ+). (F) T cells were subdivided into CD4 (CD4+ CD8-) and CD8 (CD4- CD8+) T cells and quantified. (G–H) B cells were divided on the basis of CD21, CD23 and CD24 expression into T0/T1 cells (CD21-CD24hi, G), follicular B cells (CD21loCD24lo, G), T2 cells (CD21hi CD24hi CD23-, H), MZB cells (CD21hi CD24hi CD23+, H) and quantified (right panels). For all flow cytometry experiments (C–G), data are from 1 out of 3 representative experiment with 4 or more animals in each group, and each dot represents an individual mouse. Student’s t-test, ns p>0.05, *p<0.05, **p<0.01, ***p<0.001.
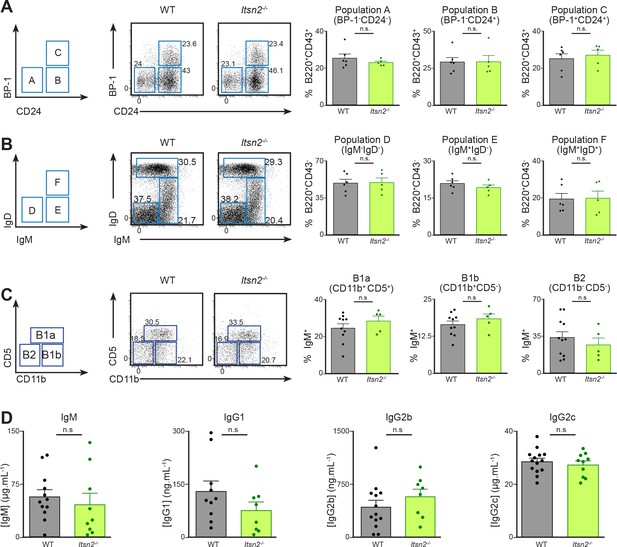
Lymphocyte development is not compromised by ITSN2 deletion.
(A–B) Bone marrow from WT and Itsn2-/- littermates was analysed by flow cytometry. Gating strategy is shown on the left. B cell progenitors (B220+) were divided into early (CD43+) and late (CD43-) cells on the basis of CD43 expression. (A) Early (CD43+) B cell progenitors were subdivided according to CD24 and BP-1 expression into Hardy populations A (CD24-BP-1-), B (CD24+BP-1-) and C (CD24+BP-1+). Data quantified in the panels on the right show the percentage of CD43+ cells in the indicated gates. (B) Late (CD43-) B cell progenitors were subdivided according to IgM and IgD expression into Hardy populations D (IgM-IgD-), E (IgM+ IgDint) and F (IgM+IgD+). The data quantified in the panels on the right show the percentage of CD43- cells in the indicated gates. (C) Peritoneal lavage was performed on WT and Itsn2-/- littermates, and the exudate was analysed by flow cytometry. B cells (IgM+) were divided into B2 (CD11b-CD5-), B1b (CD11b+CD5-) and B1a (CD11b+CD5+). Data quantified in the panels on the right show the percentage of IgM+ cells in the indicated gates. Data are from 1 out of 3 experiments with 5 or more animals in each group. Student’s t-test, ns – p>0.05. (D) Serum from naïve age-matched WT and Itsn2-/- animals was collected and baseline antibody titres were measured by sandwich ELISA. Data are pooled from two independent experiments with more than four animals in each group. Student’s t-test, ns p>0.05.
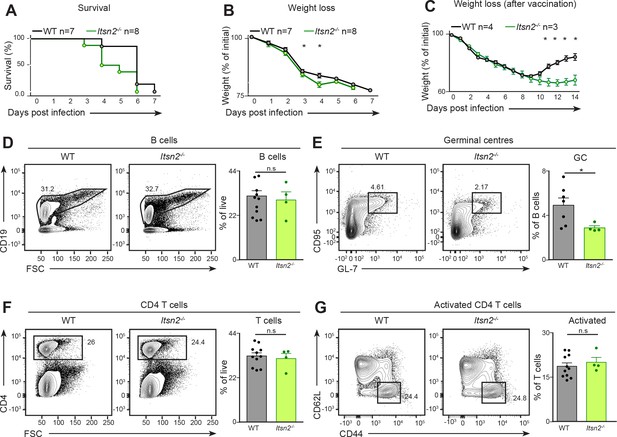
Itsn2-/- animals are impaired in responding to viral infection.
(A–B) Age-matched WT and Itsn2-/- mice were infected intranasally with 3 × 104 PFU of Influenza virus (PR8 strain). Animals were weighed daily and culled when exhibiting 20% weight loss compared to the weight at Day 0. Survival curve (A) and weight loss (B) as a function of days post infection are shown. Survival analysis, p=0.0345. (C) Age-matched WT and Itsn2-/-mice were vaccinated by i.v infection of Hemagglutinin trimer with Sigma Adjuvant. After one month, mice were infected with 1.5.105 PFU Influenza PR8 and weighed daily for the whole duration of the experiment. (D–G) WT and Itsn2-/- littermates were infected with 104 PFU Vaccinia virus (Western Reserve), and draining popliteal lymph nodes were analysed by flow cytometry 7 days post-infection. B cells (CD19+, D), GC (B220+ GL-7+ CD95+, E), CD4 T cells (CD4+, F) and activated T cells (CD4+CD44+CD62L-, G) are shown. Student’s t-test, ns p>0.05, *p<0.05.
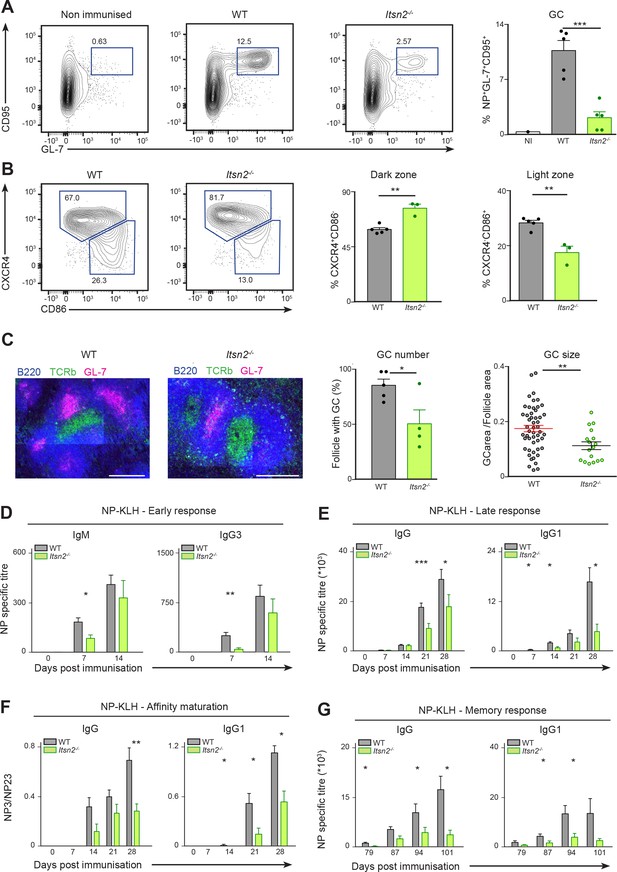
Defective humoral responses in Itsn2-/- animals.
WT and Itsn2-/- littermates were immunised with NP-KLH precipitated in Alum, and B cell responses analysed 13d after immunisation. (A, B and C) Spleens of immunised animals were analysed by flow cytometry. Antigen-specific GC B cells (NP+ GL-7+ CD95+, A), Dark zone GC B cells (GL-7+ CD95+ CD86loCXCR4hi, B), light zone GC B cells (GL-7+ CD95+ CD86hiCXCR4lo, B). Panels on the right show the show percentage of cells in the indicated gates (each dot represents a different animal). (C) Tiled images of spleen frozen sections acquired by confocal microscopy. GC cells (GL7+, magenta), B cell area (B220+, blue) and T cell area (TCRβ+, green) are shown. Scale bars – 200 µm. Quantification in the two right panels indicate the percentage of B cell follicles with a germinal centre (each dot corresponds to one section), and the ratio between GC area and total follicle area (each dot represents an individual B cell follicle). (D–G) WT and Itsn2-/- littermates were immunised with NP-KLH precipitated in Alum. (D) Serum was collected from WT and Itsn2-/- mice and titres of NP23-specific IgM and IgG3 measured by ELISA. (E) Serum from WT and Itsn2-/- mice was collected and titres of NP23-specific or total IgG and IgG1 were measured by ELISA. (F) ELISA analysis showing affinity maturation (expressed as the ratio of NP3 to NP23 titres) of total IgG and IgG1 isoform after immunisation. (G) Immunised WT and Itsn2-/- animals were submitted to a secondary challenge at day 80 with NP-KLH in PBS, and titres of NP23-specific total IgG and IgG1 were measured by ELISA. Data are representative of at least 2 independent experiments with more than three animals in each group. Student’s t-test, ns p>0.05, *p<0.05, *p<0.01, ***p<0.0001.
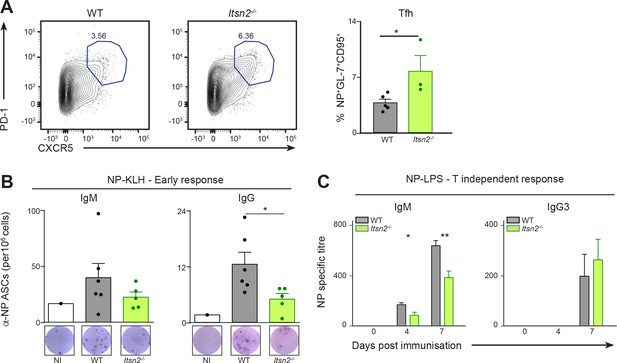
Defective humoral responses in Itsn2-/- animals
(A) WT and Itsn2-/- littermates were immunised with NP-KLH precipitated in Alum, and spleens of immunised animals analysed by flow cytometry 13d after immunisation. Tfh cells (CD4+CXCR5+PD-1+). Panels on the right show the show percentage of cells in the indicated gates (each dot represents a different animal). (B) WT and Itsn2-/- littermates were immunised with NP-KLH precipitated in Alum, and 13 days post-immunisation, NP-specific antibody secreting cells were detected in the spleen by ELISPOT. (F) WT and Itsn2-/- littermates were immunised with NP-LPS and titres of NP23-specific IgM and IgG3 were measured by ELISA. Data are representative of at least 2 independent experiments with more than three animals in each group. Student’s t-test, ns p>0.05, *p<0.05, * p<0.01, *** p<0.0001.
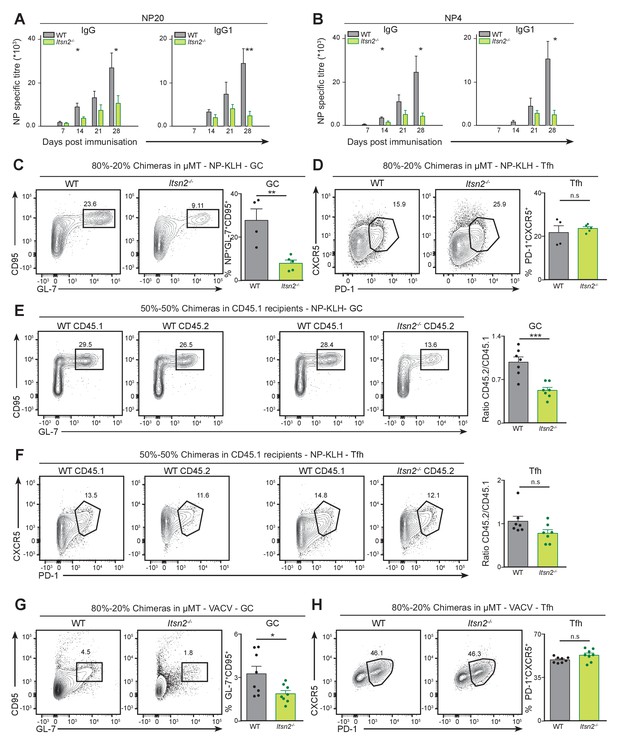
Itsn2-/- B cells are intrinsically impaired in responding to immunisation.
(A–C) Lethally irradiated μMT hosts were reconstituted for 10 weeks with mixtures of 80% μMT BM and 20% WT or Itsn2-/- BM and immunised with NP-KLH in Alum. NP20 (A) and NP4 (B)-specific titres were determined by ELISA. (C–D) Spleens of immunised animals were analysed by flow cytometry 13 days after immunisation. Antigen-specific GC B cells (NP+ GL-7+ CD95+, C) and Tfh (CD4+CXCR5+PD-1+, D) are shown. Quantification shown on the right represent the percentage of cells in the indicated gate. (E–F) Lethally irradiated C57BL6-CD45.1 hosts were reconstituted for 10 weeks with mixtures of 50% WT-CD45.1 BM and 50% WT or Itsn2-/- BM, immunised with NP-KLH in Alum, and spleens of immunised animals were analysed by flow cytometry at day 13. CD45.1+ and CD45.2+ NP-specific GC B cells (NP+ GL-7+ CD95+, E) and Tfh (CD4+CXCR5+PD-1+, F) are shown. Graphs on the right represent the CD45.2/CD45.1 ratio calculated from the percentages of cells in the indicated gates. (G) 80–20% mixed chimeras in µMT recipients were infected with 104 PFU of Vaccinia virus via intra-footpad injections, and popliteal lymph nodes were analysed by flow cytometry at d7. GC (GL-7+ CD95+, G) and Tfh cells (GL-7+ CD95+, H) are shown. Data are representative of 2–3 independent experiments with at least 4 mice in each group. Student’s t-test, ns p>0.05, *p<0.05, *p<0.01, ***p<0.0001.
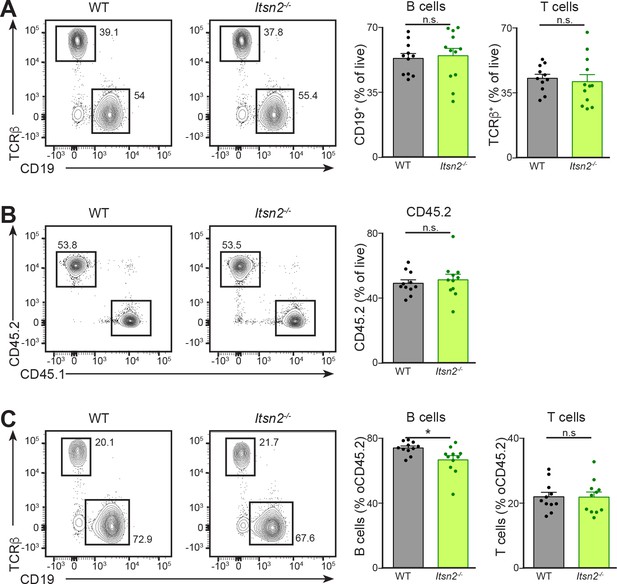
Immune compartment reconstitution by Itsn2-/-cells
(A) Lethally irradiated μMT hosts were reconstituted with mixtures of 80% μMT BM and 20% WT or Itsn2-/- BM. After 10 weeks, animals were bled and circulating B (CD19+) and T (TCRβ+) lymphocytes characterised by flow cytometry. (B–C) Lethally irradiated B. CD45.1 hosts were reconstituted with mixtures of 50% B. CD45.1 and 50% WT or Itsn2-/- BM. After 10 weeks, animals were bled and circulating cells were characterised by flow cytometry. (B) Circulating cells were divided into CD45.1+ (WT) and CD45.2+ (WT or Itsn2-/-). (C) CD45.2 cells were subdivided into B (CD19+) and T (TCRβ) lymphocytes. For all panels in this figure, data are quantified on the right panels and indicate percentage of live (A and B) or CD45.2 (C) cells in the indicated gates. Data are from 1 out of at least 3 experiments with 5 animals or more in each group. Student’s t-test, ns p>0.05, *p<0.05.
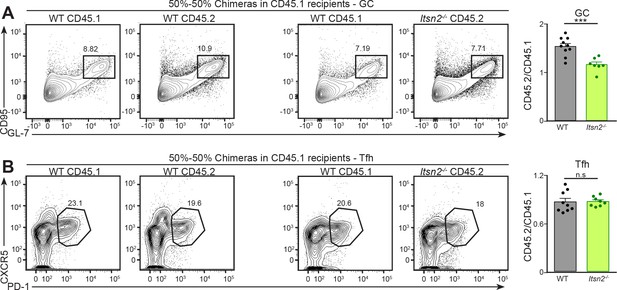
Itsn2-/- B cells are intrinsically impaired in responding to immunisation
(A–B) 50–50% mixed chimeras in CD45.1 recipients were infected with 104 PFU of Vaccinia virus via intra-footpad injections, and popliteal lymph nodes were analysed by flow cytometry at d7. CD45.1+ and CD45.2+ GC B cells (GL-7+ CD95+, (A) and Tfh cells (CD4+CXCR5+PD-1+, (B) are shown. Graphs on the right represent the CD45.2/CD45.1 ratio calculated from the percentages of cells in the indicated gates.
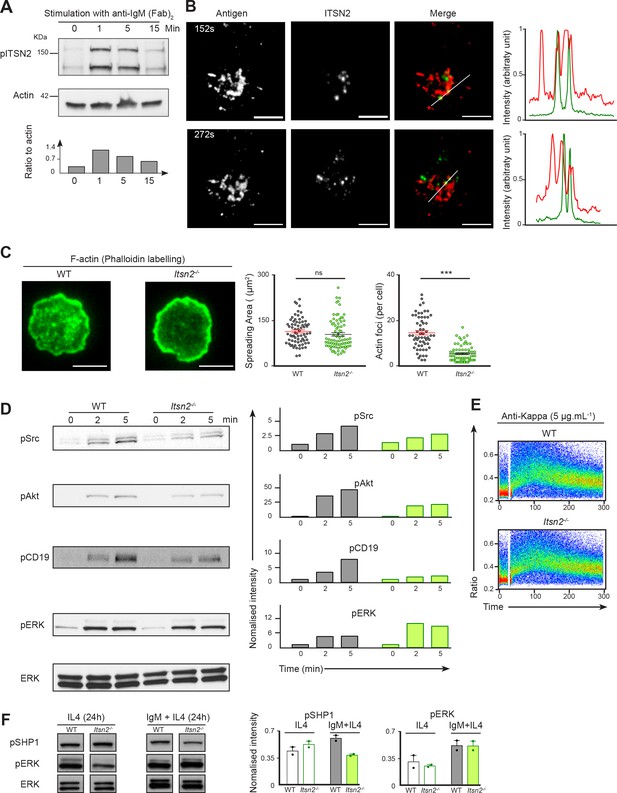
ITSN2 participates in the BCR signalling cascade.
(A) Purified naïve B cells were stimulated with anti-IgM F(ab’)2 (10 µg mL−1) for 0, 1, 5 or 15 min, the total phosphotyrosine fraction was immunoprecipitated, and phosphorylated ITSN2 detected by Western blot. Quantification below shows phosphorylated ITSN2 intensity related to β-Actin. Data are representative of 3 independent experiments. (B) A20 cells with a HEL-specific BCR were transfected with a plasmid encoding for ITSN2-L GFP, settled onto fluorescently labelled antigen-loaded lipid bilayers and imaged with a TIRF microscope. Localisation of ITSN2 (green) and antigen (red) were compared over time. Scale bars – 3 μm. Data are representative of 3 independent experiments. (C) Representative TIRF images of WT and Itsn2-/- B cells spread on anti-kappa coated glass coverslips and stained with fluorescently-labelled phalloidin. Scale bars – 3 µm. The quantification of spreading area and number of detected actin foci are shown graphically. Each dot represents an individual cell. Data are representative of 3 independent experiments where > 100 cells of each genotype were analysed. (D) WT and Itsn2-/- B cells were stimulated with 2 µg mL−1 anti-IgM F(ab’)2 for 0, 2 or 5 min, and phosphorylation of Src, CD19, Akt (S473) and ERK were detected by Western blotting. Graphs next to each lane indicate the relative intensity of each band relative to ERK at t0. (E) Intracellular calcium influx in purified WT or Itsn2-/- B cells after stimulation with 5 µg mL−1 anti-kappa. (F) Naïve purified WT and Itsn2-/- B cells were cultured in the presence of IL-4 alone or IL-4 and anti-IgM for 24 hr, and phosphorylation ERK and SHP1 measured by Western Blot. Student’s t-test, ns p>0.05, *p<0.05, **p<0.01, ***p<0.001.
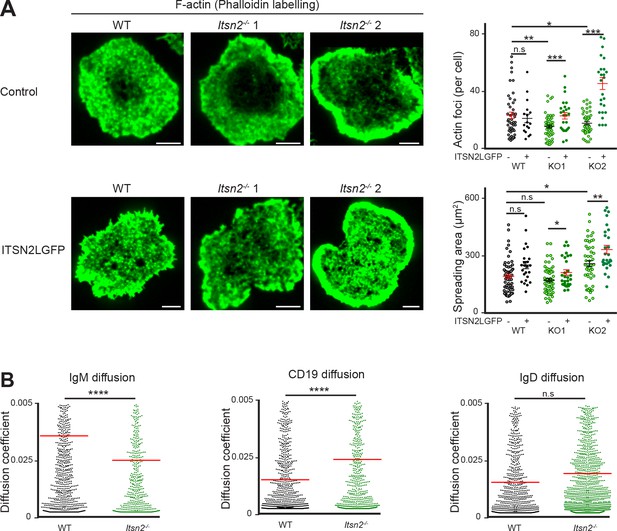
ITSN2 participates in the BCR signalling cascade.
(A) ITSN2 was ablated in A20 cells expressing a HEL-specific BCR by CRISPR-Cas9 mutagenesis. Representative TIRF images of WT and Itsn2-/- (two independent clones) A20 cells without (upper row) or after transfection with GFP-ITSN2-L (lower row) spread on anti-kappa coated glass coverslips and stained with fluorescently-labelled phalloidin. Scale bars – 5 µm. The quantification of spreading area and number of detected actin foci are shown graphically. Each dot represents an individual cell. Data are representative of 2 independent experiments where > 40 cells of each group were analysed. (B) Single-particle tracking (SPT) of CD19, IgM and IgD on WT or Itsn2-/- B cells settled on nonstimulatory coverslips. Shown is the diffusion coefficient (D) in µm2/s. The median of 300 representative cells is given as value and indicated by red bars. 1,000–2000 tracks were analysed with a minimum of 20 cells from two experiments.
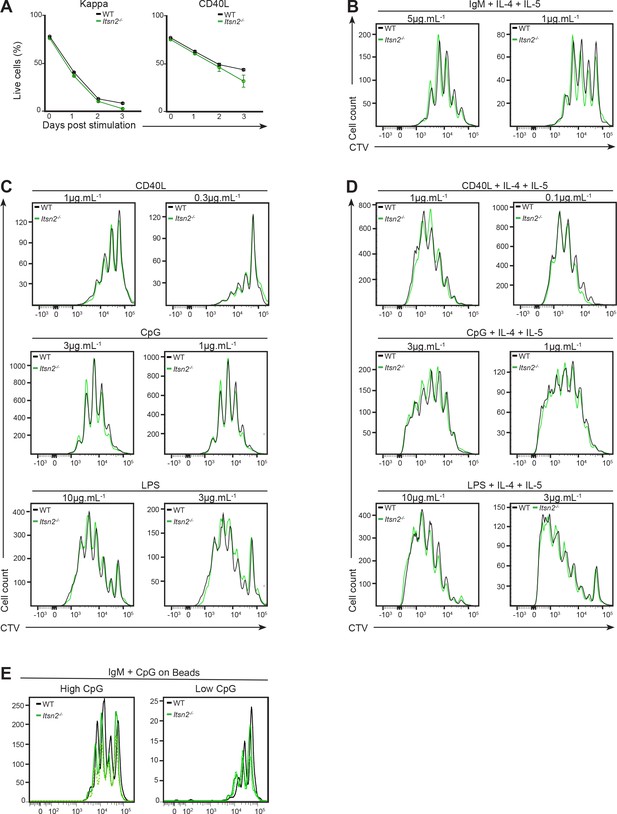
B cell activation ex vivo in response to BCR or TLR signals is not impaired in Itsn2-/- B cells.
(A) Naïve purified WT and Itsn2-/- B cells were cultured in the presence of CD40L (0.5 µg.mL−1) or anti-Kappa (1 µg.mL−1) for 3 days, and percentage of live cells measure daily. (B–D) CTV-labelled WT and Itsn2-/- B cells were cultured with anti-IgM (B), CD40L, CpG or LPS at the concentrations indicated on the graphs in presence (D) or absence (C) of IL-4 and IL-5. Proliferation profiles were analysed by flow cytometry after 3 (D) or 4 (C) days of culture. (E) CTV-labelled WT and Itsn2-/- B cells we loaded with particles coated with IgM and CpG, and proliferation measured after 3 days of culture.
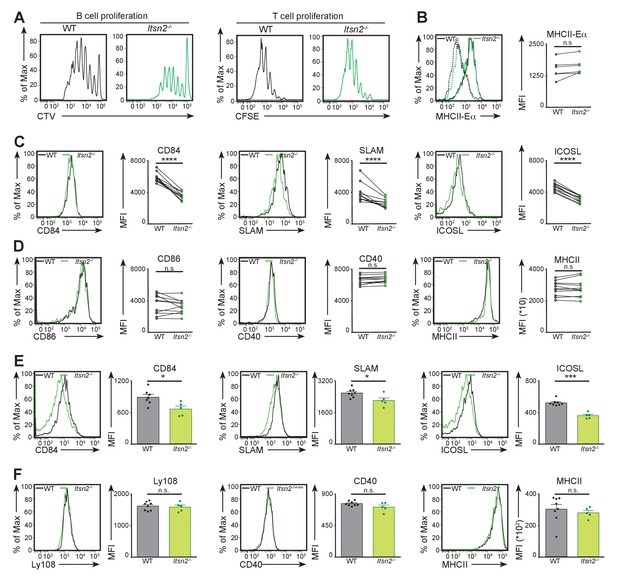
ITSN2 is required for B cell activation in vivo.
(A) CTV-labelled HEL-WT or HEL-Itsn2-/- B cells and CFSE-labelled OTII T cells were transferred to CD45.1 hosts, subsequently immunised with HEL and OVA coated microspheres. Proliferation of both cell types was measured by flow cytometry 3 days after immunisation. Data are representative of 4 independent experiments with 3–6 animals in each group. (B) CTV-labelled HEL-WT and CFSE-labelled HEL-Itsn2-/- B cells were transferred to WT recipients prior to intravenous immunisation with beads coated with HEL and Eα peptide. Expression of MHCII-Eαconjugates (quantified on the right as the geometric mean of the MHCII- Eα fluorescence signal) by both cell types was measured 24 hr after immunisation. Data are representative of 2 independent experiments with 5–10 mice. (C–F) CFSE-labelled HEL-WT and CTV-labelled HEL-Itsn2-/- B cells were transferred to CD45.1 hosts, and immunised with HEL and OVA coated microspheres. Expression of CD84 (C), SLAM (C), ICOSL (C), CD86 (D), CD40 (D) and MHCII (D) was measured by flow cytometry 24 hr after immunisation. Data are representative of 3 independent experiments with 5–10 mice. (E–F) WT and Itsn2-/- littermates were immunised with NP-KLH precipitated in Alum, and spleens analysed by flow cytometry 13 days after immunisation. Expression of CD84 (E), SLAM (E), ICOSL (E), Ly108 (F), CD40 (F) and MHCII (F) on germinal centre B cells was measured. Data are representative of 3 independent experiments with 5–10 mice in each group. Paired t-test (B, C, D) or Student’s t-test (E,F), ns p>0.05, *p<0.05, ***p<0.001, ****p<0.0001.
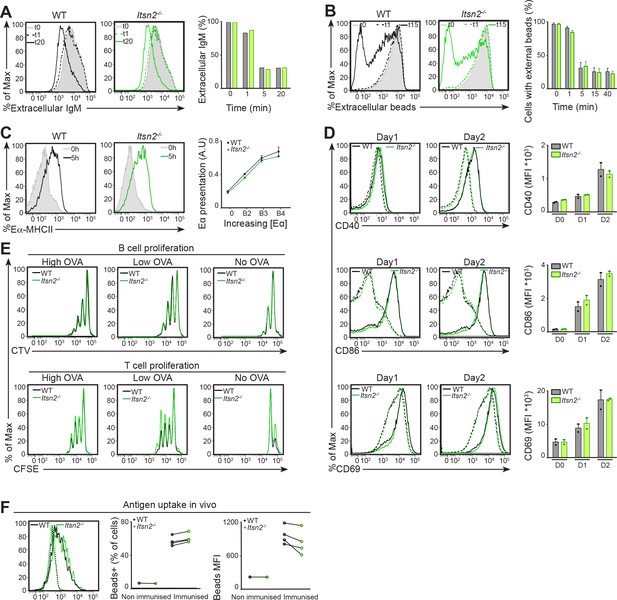
Antigen uptake and reception of T cell help ex vivois not impaired in Itsn2-/- B cells.
(A–B) Purified naïve WT or Itsn2-/- B cells were labelled on ice with soluble biotinylated anti-IgM (A) or anti-IgM coated particles (B), and incubated at 37°C for the indicated times, prior to fixation. Extracellular antigen was then detected by flow cytometry. Data are quantified on the right panel as percentage of initial MFI for each sample (A), or the percentage of the cells with external beads (B). Data are representative of more than 3 independent experiments with 2 mice in each group. (C) Purified naïve WT or Itsn2-/- B cells were stimulated with beads coated with anti-IgM and Eα peptide, and expression of MHCII-Eα conjugates measured by flow cytometry 5 hr after stimulation. Data are quantified on the right panel and show the Eα presentation for increasing amounts of Eα on the beads. (D–E) CTV-labelled WT or Itsn2-/- B cells were stimulated with beads coated with IgM and OVA, and co-cultured with CFSE-labelled OTII T cells. Expression of CD40, CD86 and CD69 was measured by flow cytometry at d1 and d2 (D). B and T cell proliferation rates according were characterised to CFSE and CTV dilution at d3 of the culture (E). (F) CFSE-labelled WT and CTV-labelled Itsn2-/- MD4 B cells were transferred adoptively into WT recipients, and after 24 hr, animals were injected i.v. with fluorescently-labelled HEL-coated particles. Beads internalisation was measured by flow cytometry 24 hr post-injection.
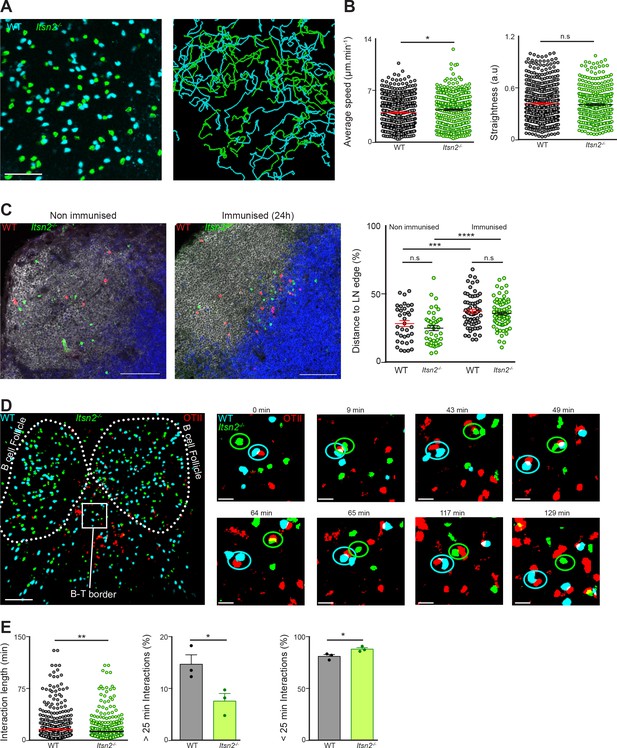
Itsn2-/- B cells are impaired in forming long-term conjugates with T cells.
(A–B) CTV-labelled HEL-WT and CFSE-labelled-Itsn2-/- B cells were transferred to WT recipients, and explanted popliteal lymph nodes were imaged with a multiphoton microscope 24 hr after adoptive transfer. A representative field of view as well as the associated tracks are shown. (B) Quantification representing the average speed and the straightness of the tracks. Data are representative of 2 independent experiments where 4–6 lymph nodes were imaged. Student-s t-test, n.s. p>0.05, *p<0.05. (C–E) CTV-labelled HEL-WT and CFSE-labelled HEL-Itsn2-/- B cells were transferred to WT recipients together with unlabelled (C) or SNARF-1-labelled (D–E) OTII T cells prior to intra-footpad immunisation with HEL and OVA coated microspheres. Popliteal lymph nodes were explanted 18–20 hr later imaged by confocal (C) or multiphoton microscopy (D–E). (C) Frozen sections were stained with antibodies against B220 and TCRβ to distinguish B and T cell areas. Quantified data on the right indicate the distance from each WT (red) or Itsn2-/- (green) B cell to the edge of the LN. Scale bars – 200 µm. Paired t-test, n.s. – p>0.05). (D) Representative images of a whole popliteal lymph node as well as higher magnification images showing typical interactions between WT B cells (cyan), Itsn2-/- B cells (green) and OTII T cells (red). Dotted lines were added for illustrative purposes and indicate the localisation of two B cell follicles. (Scale bar – 80 µm on the low magnification and 10 µm on the higher magnification.) (E) Quantification of the overall length of B-T interactions (left panel) and the fraction of interactions shorter (middle panel) or longer (right panel) than 25 min. Data were pooled from 3 independent experiments where 2–3 lymph nodes were imaged. Student’s t-test, ns p>0.05, *p<0.05, **p<0.01.
Representative movie of part of a popliteal lymph node, showing homeostatic migration of WT (gray) and Itsn2-/- (green) B cells.
https://doi.org/10.7554/eLife.26556.016Representative movie of part of a popliteal lymph node, showing long term interaction between a WT B cell (cyan) and OTII T cells (red), and shorter conjugates between Itsn2-/- B cells (green) and OTII T cells (red).
https://doi.org/10.7554/eLife.26556.017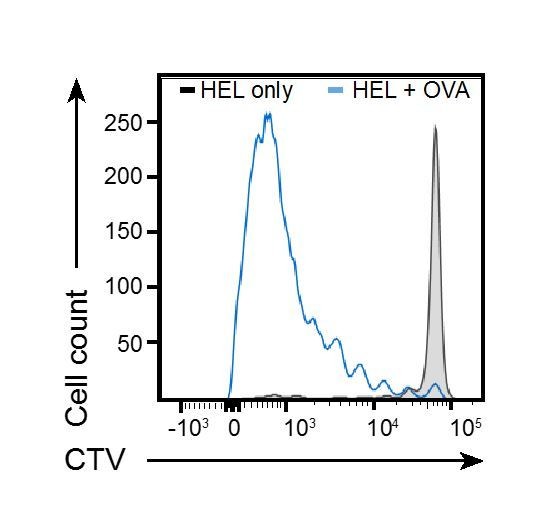
MD4 B cells and OTII T cells were transferred to WT recipients, and animals immunised with particles coated with HEL alone (black tinted line) or HEL and OVA (Blue line).
B cells proliferation was measured by FACS at d3.
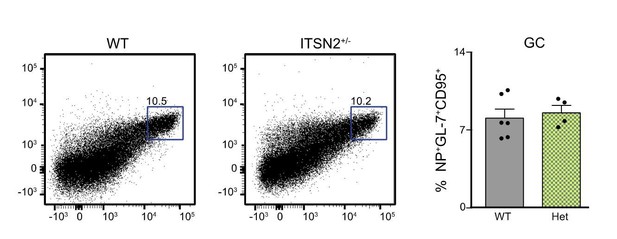
WT and ITSN2+/- littermates were infected with 104 PFU VACV by intra-footpad injection.
GC formation was measured in the popliteal LN 7 days after infection.
Tables
| Reagent type (species) or resource | Designation | Source or reference | Identifiers | Additional information |
|---|---|---|---|---|
| Gene (Mus musculus) | Itsn2 | NA | ENSMUSG00000020640; MGI:1338049 | |
| Genetic reagent (M. musculus) | Itsn2-/- | KOMP consortium | RRID:MGI:5631233 | Itsn2tm1.1(KOMP)Vlcg |
| Genetic reagent (M. musculus) | µMT | doi:10.1038/350423a0 | RRID:IMSR_HAR:1682 | (beta)-globin (mu)MT-/- |
| Genetic reagent (M. musculus) | MD4 | doi:10.1038/334676a0 | RRID:MGI:5006966 | Tg(IghelMD4)4Ccg |
| Genetic reagent (M. musculus) | OTII | doi:10.1046/j.1440–1711.1998.00709.x | RRID:IMSR_JAX:004194 | B6.Cg-Tg(TcraTcrb)425Cbn/J |
| Cell line (M. musculus) | A20 cells with HEL-specific D1.3 BCR | http://dx.doi.org/10.1016/S1074-7613(00)80580–4 | ||
| Transfected construct (plasmid) | ITSN2L GFP | Michael Way | pEGFP-C1 ITSN2L | |
| Antibody (rabbit polyclonal) | anti-ITSN2 antibody | Novus | NBP1-71833; RRID:AB_11038593 | 1:1000 in 5% Milk TBS-T |
| Sequence-based reagent | sgRNA1 | this paper | Forward - CACCGTAGCTATAGAGAACTCTTGC; Reverse - AAACGCAAGAGTTCTCTATAGCTAC | |
| Sequence-based reagent | sgRNA 2 | this paper | Forward - CACCGGGGGTTGTTTCATGATAGGA; Reverse - AAACTCCTATCATGAAACAACCCCC | |
| Peptide, recombinant protein | Eα peptide | The Francis Crick Institute peptide chemistry unit; doi:10.1038/353660a0 | Sequence: Biotin-GSGFAKFASFEAQGALANIAVDKA-COOH |
Additional files
-
Transparent reporting form
- https://doi.org/10.7554/eLife.26556.018





