Structure-based analysis of CysZ-mediated cellular uptake of sulfate
Figures
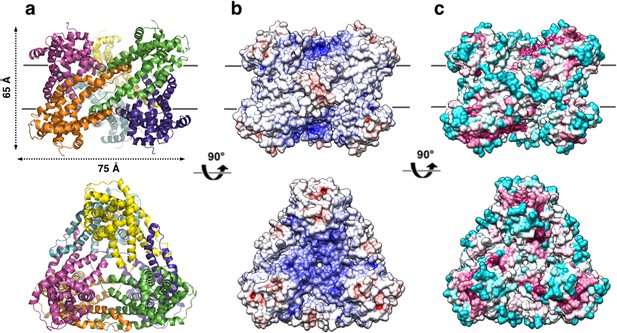
Overall structure of the P. denitrificans CysZ (PdCysZ) hexamer.
(a) Side and top views of the hexamer as a ribbon diagram with each protomer chain colored differently. The approximate dimensions of the hexamer marked in Å. (b) Side and top views represented by surface electrostatics as calculated by APBS, with negative and positive surface potential represented in red and blue respectively. (c) Side and top views representing conservation of residues as calculated by ConSurf, with maroon being most conserved to cyan being least conserved.
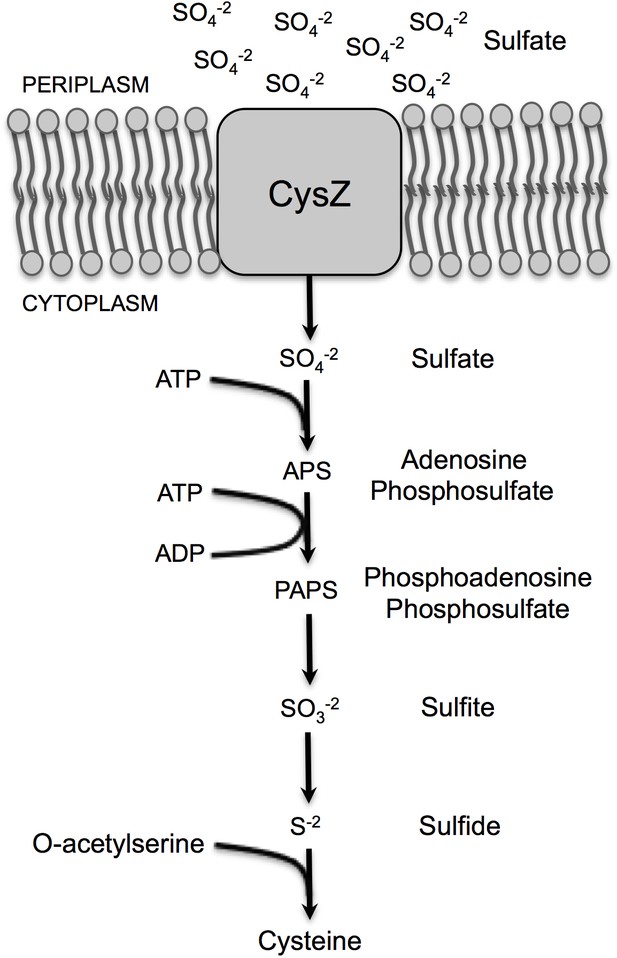
Schematic of assimilatory sulfate reduction in bacteria for cysteine biosynthesis.
Assimilatory reduction is energetically dispendious as sulfate is extremely stable. In this pathway, sulfate ions enter the cell via a transporter or channel, like CysZ; this is followed by its activation by ATP-sulfurylase, which utilizes ATP to form adensosine phosphosulfate (APS) and inorganic pyrophosphate (PPi) as a by-product. APS is then further activated by the addition of a second phosphate forming phosphoadenosine phosphosulfate (PAPS) and this high-energy intermediate product is poised for sulfur removal to form sulfite (SO32-) by the thioredoxin enzyme, which is then further reduced to sulfide ions (S-). Sulfide, the final reduced sulfur product, can then be used in a variety of ways by the cell, such as the incorporation into O-acetylserine to synthesize cysteine.
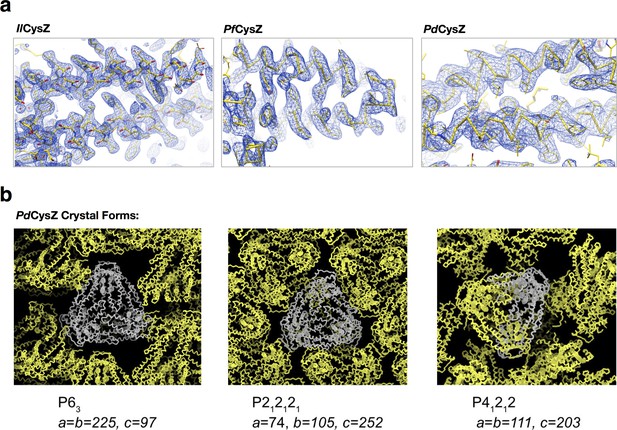
Representative electron density of the crystal structures of IlCysZ, PfCysZ and PdCysZ, and the different crystal forms observed for PdCysZ.
(a) Representative electron density in blue mesh for: IlCysZ, contoured at 1 x r.m.s.d., PfCysZ, contoured at 2 x r.m.s.d, with a negative B-factor of −100 Å2 applied to map; PdCysZ, contoured at 1 x r.m.s.d, with a negative B-factor of −100 Å2 applied to map. Final atomic models (Cα trace with side chains depicted) are shown as yellow stick representation. (b) Preliminary molecular replacement analyses of the three different crystal forms obtained for PdCysZ, showing the hexameric assembly of the molecule, seen in each case. The Cα trace of an asymmetric unit in each crystal form is depicted in grey, with their symmetry mates shown in yellow. Space group and unit cell dimensions (in Å) are listed below.
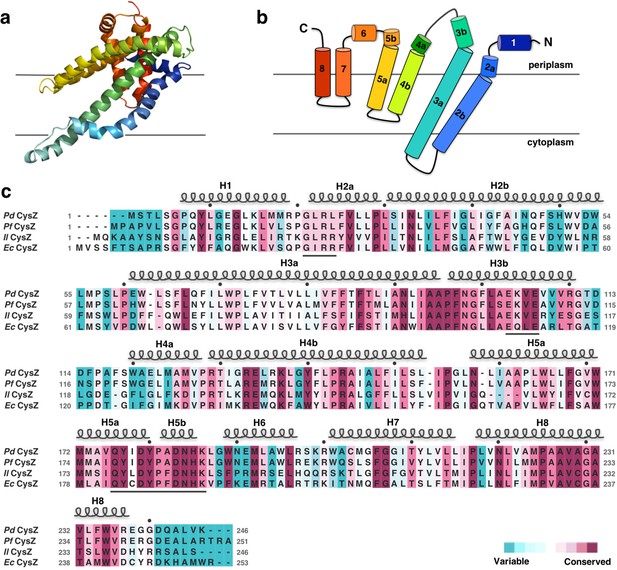
Structure and topology diagram of the PdCysZ protomer, with sequence alignment and conservation.
(a) Ribbon representation of the PdCysZ protomer colored in rainbow colors from N (blue) to C terminus (red), viewed from within the plane of the membrane, shown in the same orientation as the protomer drawn in green in Figure 1a. (b) Topology diagram of the PdCysZ protomer with helices marked from 1 to 8. Helices H2b and H3a are transmembrane helices, whereas helices H4-H5 and H7-H8 are hemi-penetrating helical hairpins, only partially inserted into the membrane. (c) Sequence alignment of E. coli CysZ (EcCysZ), P. denitrificans CysZ (PdCysZ), P. fragi CysZ (PfCysZ), and I. loihiensis CysZ (IlCysZ). Residues are colored based on conservation, with maroon being most conserved and cyan least conserved, as calculated by ConSurf using a sequence alignment of 150 non-redundant sequences from the CysZ family as input. Spirals above residues mark the extent of the helical segments based on the atomic structure of PdCysZ with helices numbered H1-H8; letters mark residue identities; black dots above identify every tenth residue (modulo 10) in the PdCysZ sequence and black underlines mark functionally relevant motifs discussed in the text.
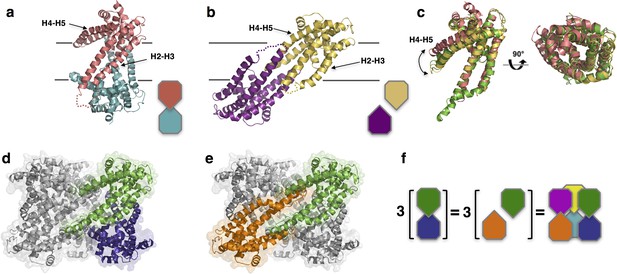
Structures of IlCysZ and PfCysZ, with comparison to PdCysZ.
(a) Ribbon diagram of the structure of CysZ from I. loihiensis (IlCysZ) at 2.3 Å. Protomers of the dimer are colored in salmon pink and teal blue, arranged in a head-to-tail conformation in the membrane, with helical hairpins H2-H3 and H4-H5 labeled for clarity. The dimer interface of IlCysZ involves helices H2-H3. (b) Ribbon diagram of the structure of CysZ from P. fragi (PfCysZ) at 3.2 Å. The protomers of the dual topology dimer in the membrane are colored gold and purple, with helical hairpins H2-H3 and H4-H5 again labeled. The dimer interface here involves the interaction of helices H4-H5 of each protomer. (c) Side and top views of the superposition of the three different protomers from PdCysZ (green), IlCysZ (pink) and PfCysZ (yellow) after aligning helices H1-H3, with a curved arrow indicating the flexible nature of helices H4-H5. d, e. The same dimer interfaces observed in IlCysZ (a) and PfCysZ (b) observed in the hexameric assembly of PdCysZ, as highlighted in green and blue (d) and green and orange (e). (f) Schematic representation showing how three copies of the dimeric protomers of IlCysZ (green and blue, left) and of PfCysZ (orange and green, center), can coexist in and each recapitulate the hexameric assembly of PdCysZ.
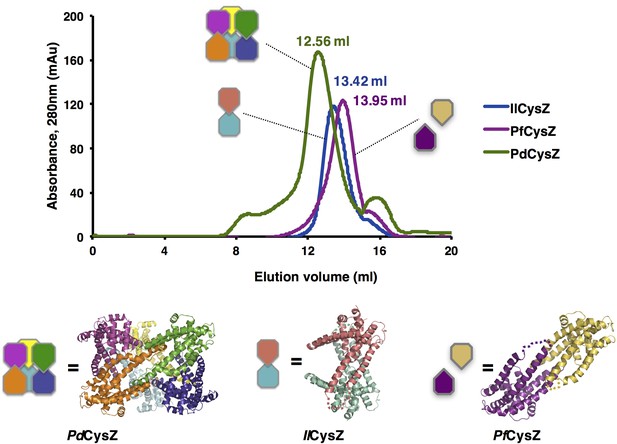
Size-exclusion chromatography of CysZ shows a mono-disperse elution profile for each of the three species purified - PdCysZ, PfCyZ and IlCysZ.
All three proteins were solubilized from isolated membrane fractions purified in the presence of 0.2% decyl maltopyranoside (DeM) and exchanged into buffer containing 1% β-octyl glucopyranoside (β-OG) on the size-exclusion column (Superdex 200 10/30 HR). Schematic of the oligomeric states observed in the three crystal structures of CysZ are shown below the elution profiles. Crystallization trials were set-up directly after the size-exclusion chromatography step. A shift of ~1 ml was observed for the elution of PdCysZ (peak at 12.56 ml), with respect to PfCysZ and IlCysZ, indicating that the size (volume) and shape of PdCysZ is significantly larger than the other two species. IlCysZ elutes at 13.42 ml, 0.5 ml ahead of PfCysZ, which could be explained by the shape of IlCysZ, occupying more volume as compared to PfCysZ, with its helices (H4–H5) pointing outward, away from the body of the dimer.
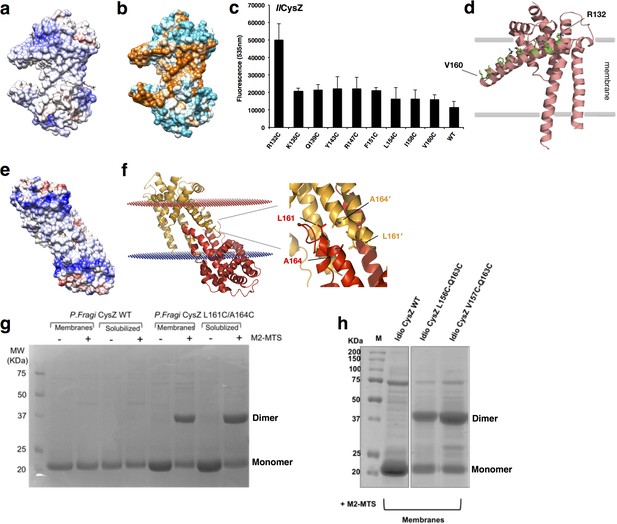
Surface electrostatics, hydrophobicity and site-directed fluorescence labeling of cysteine mutants located on H4 of IlCysZ.
(a) A depiction of the surface electrostatic potentials of the IlCysZ dimer with negative surface potential represented in red, and positive potential in blue as calculated by APBS. (b) Surface of IlCysZ rendered by level of hydrophicity, using the Kyte-Doolitle hydrophobicity scale with cyan being most polar, to orange being most hydrophobic, reveals a hydrophobic belt along the center of the dimer, suggesting the orientation of the dimer in the lipid bilayer. (c, d) Single cysteine mutants were designed to be located along the length of helix H4 of CysZ, to gauge the extent of its membrane insertion. The labeled protein was extracted from the membrane and purified after quenching the labeling reaction. Fluorescence intensity was measured and quantified by a Tecan fluorescence plate reader at an excitation of 485 nm and emission of 535 nm the results of which were plotted on the left, with error bars representing the standard deviation from the mean for n = 3. The results show that of all the residues on the helix, only R132C (the top-most residue) was accessible to the fluorophore and hence exposed out of the membrane. (d) Locations of the mutated residues on helix H4 are marked on the model of IlCysZ in green. Crosslinking of L161C-A164C cysteine mutant of PfCysZ and IlCysZ exhibits dimer formation. (e) A depiction of the surface electrostatic potentials of the PfCysZ dimer with negative surface potential represented in red, and positive potential in blue as calculated by APBS, highlights hydrophobic belt marking the orientation of the PfCysZ dimer in the membrane. (f) Membrane orientation of PfCysZ predicted by OPM/PPM server shows its 31-degree tilt to the perpendicular, with a zoomed in view of the location of the cysteine mutants at the dimer interface, used for crosslinking of the dimer. (g, h) Dimer formation by the crosslinking of PfCysZ (g) and of IlCysZ (h) with introduced structure-based cysteine substitution mutations in the helices H4-H5 dimer interface. Analogous pairs of cysteine mutants from each species are shown here: PfCysZ L161C-A164C and IlCysZ L156C-Q163C and V157C-Q163C. Sulfydryl specific Bis-MTS crosslinkers of a certain spacer length (in this case 5.2 Å) were used at 0.5 mM for 1 hr at room temperature to crosslink the protein. Experiment was performed on protein in the membrane, as well as on solubilized protein. WT protein (cysteine-less) was used as a control. Proteins were purified after stopping the reaction, and run on an SDS PAGE with reducing dye, and stained with Coomassie blue.
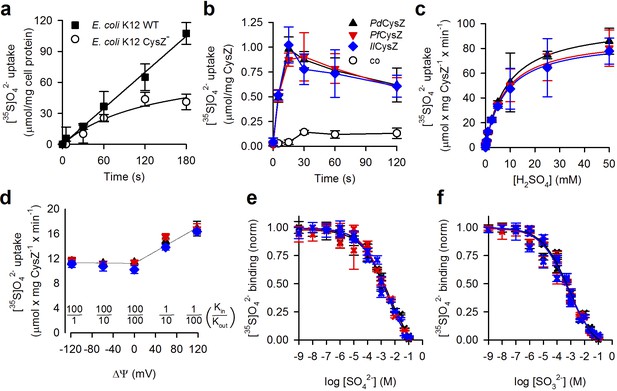
Functional characterization of CysZ.
(a) Time course of [35S]O42- uptake (320 μM) by CysZ+ (strain BW25113) or CysZ- (strain JW2406-1)E. coli K-12 cells (n = 3). (b) Time course of [35S]O42- uptake (500 μM) by CysZ-containing proteoliposomes (▲, PdCysZ; , PfCysZ; , IlCysZ) or by control liposomes (š). (c) Kinetics of [35S]O42- uptake by CysZ. Initial rates of [35S]O42- uptake were measured for 5 s with [35S]O42- concentrations ranging from 0.5 to 50 mM. (d) Effect of the membrane potential (ΔΨ) on the initial rate of 1 mM [35S]O42- uptake measured for a period of 5 s. ΔΨ of different polarity was imposed with the K+-ionophore valinomycin in CysZ-containing proteoliposomes that were pre-loaded with different concentrations of KCl and assayed in external buffer of different KCl concentrations (equiosmolar cis/trans conditions) as indicated. The valinomycin-generated K+ flux produced a KCl in/out concentration difference-dependent ΔΨ that was calculated using the Nernst equation (). Data in panel (b–d) depict the means ± S.E.M. of 3 independent measurements performed in triplicate. (e, f) Equilibrium binding of [35S]O42- measured with the SPA. (e) Isotopic dilution of 100 μM [35S]O42- with non-labeled SO42- (sodium salt) or (f) competition of 100 μM [35S]O42- with non-labeled SO32- (sodium salt) reveals the EC50 or IC50 (concentration yielding half-maximum displacement or inhibition, respectively). See text for kinetic constants. Data in (e and f) are shown as mean ± S.E.M. of ≥6 measurements and subjected to global fitting in Prism seven and kinetic constants reflect the mean ± S.E.M. of the fit. The use of color for symbols and bars in panel (b–f) is consistent.
Sulfite inhibition, effects of dissipated ion gradients, and ion conductance activities of CysZ.
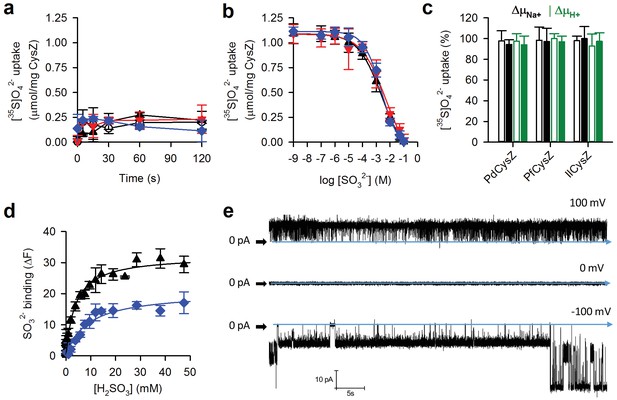
(a) Effect of SO32- on [35S]O42- accumulation in CysZ-containing proteoliposomes. Time course of [35S]O42- uptake (500 μM) by CysZ-containing proteoliposomes (▲, PdCysZ; , PfCysZ; , IlCysZ) or by control liposomes (š) in the presence of 500 μM Na2SO3. (b) Inhibition of [35S]O42- by SO32-. Initial rates of 500 μM [35S]O42- uptake were measured for 5 s in the presence of non-labeled SO32- at concentrations ranging from 1 nM – 100 mM. Data (means ± S.E.M. of ≥6 independent measurements) were subjected to nonlinear regression fitting (log inhibitor versus response – variable slope) in Prism 7, revealing an IC50 (SO32- concentration at which [35S]O42- uptake is inhibited by 50%) of 2.73 ± 0.42 mM for PdCysZ, 4.99 ± 0.46 mM for PfCysZ, and 2.31 ± 0.76 mM for IlCysZ. (c) Effect of the sodium motive force (ΔμNa+) or proton motive force (ΔμH+) on CysZ-mediated SO42- uptake. [35S]O42- (1 mM final concentration) uptake was measured for a period of 5 s in CysZ-containing proteoliposomes prepared in 200 mM Tris/Mes, pH 7.5 (inside buffer composition) in assay buffer composed of either (i) 100 mM Tris/Mes, pH 7.5/100 mM NaCl (generation of ΔμNa+; black bars) or (ii) 200 mM Tris/Mes, pH 5.5 (generation of ΔμH+; green bars) in the presence (solid bars) or absence (open bars) of 25 μg gramicidin/mL. Addition of the ionophore gramicidin did not markedly decrease the [35S]O42- uptake activity by CysZ as would be expected for Na+ or H+-coupled SO42- transporters. (d) SO32- binding by CysZ. Saturation binding of SO32- was measured with the MST assay, yielding EC50s of 3.73 ± 0.79 mM for PdCysZ and 8.68 ± 1.88 mM for IlCysZ. (e) Electrophysiology of IlCysZ reconstituted in planar lipid bilayers. Sample traces of IlCysZ under 150 mM symmetrical Na2SO4 solutions at the indicated membrane potentials and pH = 5.4.
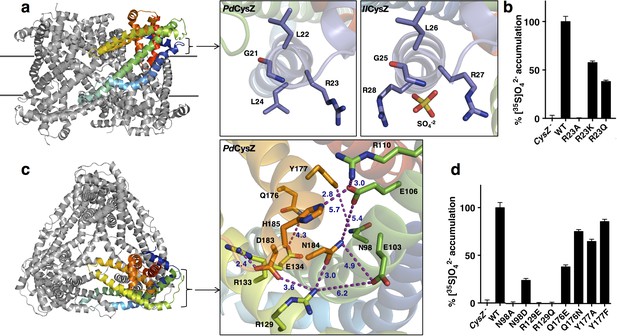
Functionally relevant features of the CysZ molecule.
(a) Side-view of a ribbon diagram of the PdCysZ hexamer, with one chain rainbow-colored from N- (blue) to C-termini (red). Insets show the sulfate-binding site located at the start of H2a in PdCysZ (left), with conserved residues G21, L22, R23 and L24 labeled, and the corresponding site in IlCysZ (right), with residues G25, L26, R27, and R28 labeled and showing the SO4−2 ion as bound in the crystal structure. (b) Sulfate uptake by sulfate-binding site mutants of PdCysZ. E. coli K-12 CysZ- cells transformed with the listed PdCysZ R23 mutants were used to measure 320 μM [35S]O4−2 uptake (n = 3). Sulfate uptake was abolished for R23A, and rescued to 50% and 40% of the wild type levels for the R23K and R23Q mutants respectively. (c) A top-view of the PdCysZ hexamer, colored as in a. The inset magnifies the central core of CysZ to show the associated network of hydrogen bonds (R129-N184, E106-H185), van der Waals interactions (E103-N184, E106-N184, Q176-E134) and salt bridges (R129-D183, R133-D183, R110-E106) between pairs of highly conserved residues. Interatomic contacts are shown as purple dotted lines with distances (in Å) marked in blue. (d) Sulfate uptake by central-core mutants of PdCysZ. E. coli K-12 CysZ- cells transformed with the listed PdCysZ mutants were used to measure 320 μM [35S]O4−2 uptake (n = 3). N98A and R129E and R129Q showed severely impaired sulfate uptake, whereas more conservative substitutions such as N98D, Q176E and Q176N had less of a negative effect on function. Y177A and Y177F do not show any impaired function.
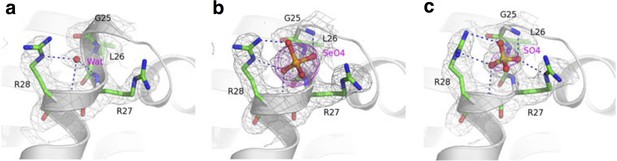
Substrate molecules bound at sulfate binding site of IlCysZ.
(a) Apo structure of IlCysZ showing a bound water (H2O) molecule, with no anomalous signal above noise level (collected at Fe edge). (b) IlCysZ in complex with selenate (SeO42-), anomalous Fourier map is shown at 5.0 sigma (collected at Se edge). (c) IlCysZ in complex with sulfate (SO42-), anomalous Fourier map is shown at 2.0 sigma (collected at Fe edge). 2Fo-Fc electron densities are in gray and anomalous Fouriers are in magenta.
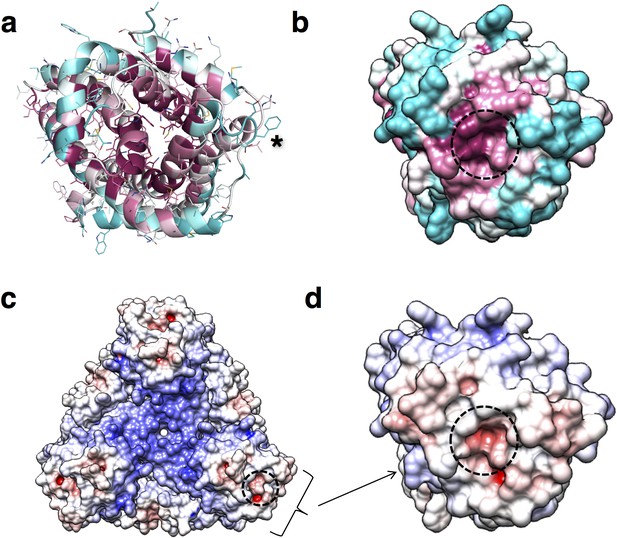
Conservation and entrance of putative pore of PdCysZ.
(a) Ribbon diagram colored by conservation with residues in maroon being most conserved to cyan being least conserved (calculated by ConSurf) to highlight the entrance to the putative pore; an asterisk (*) marks the location of the sulfate-binding site (GLR motif) at top of helix H2a. (b) Same view and coloring scheme as in a, but now shown in surface representation. (c) Electrostatic representation of hexameric PdCysZ as viewed from the top, with negative surface potential represented in red, and positive potential in blue as calculated by APBS, with location of the putative pore marked by a dashed circle. (d) Close-up view in electrostatic representation of the putative pore within a PdCysZ protomer, surface and orientation as in b.
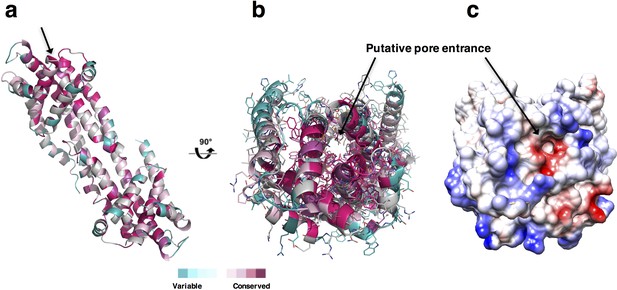
Visualization of putative pore of PfCysZ.
(a) The location of the putative sulfate translocation pore in the hydrophilic head of the PfCysZ transverse dimer is marked with an arrow. The cartoon is colored by conservation (maroon- highly conserved, cyan- more variable). (b) A top view (90° rotated) reveals the putative pore going all the way through the PfCysZ dimer. (c) A top view of the PfCysZ dimer in surface depiction, colored by surface electrostatics, shows the negatively charged nature of the entrance to the pore.
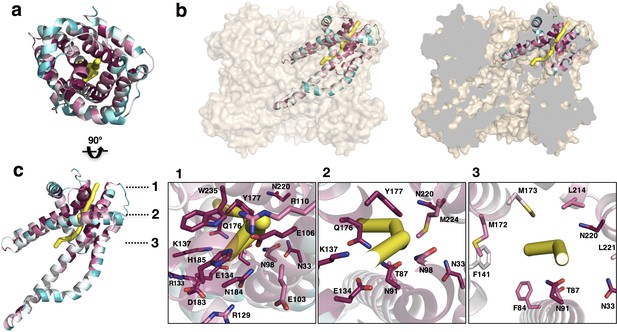
Putative ion conductance pathway of CysZ.
(a) A PdCysZ protomer looking into the incipient pore entrance. The polypeptide ribbon is oriented as in Figure 6a and colored by level of sequence conservation (calculated by ConSurf, with maroon being most conserved and cyan least). The putative pore and ion conduction pathway is shown as a yellow tube of 1 Å diameter, calculated by PoreWalker. (b) Surface representation of the PdCysZ hexamer, as side views, both semi-transparent (left) and with surface clipped (right, cut surfaces colored in grey) to allow for the internal visualization of the pathway leading to the central cavity. (c) Side view of a PdCysZ protomer left, viewed as in b and rotated 90° from a. Insets show magnified views of cross-sections along the pathway: At level 1, the entrance to the pore consists of a narrow constriction created by a network of highly conserved polar and charged residues (E106, R110, E134, N184, H185) tightly interacting with one another. At level 2, the pathway broadens and becomes less charged, but yet polar in nature as lined by conserved asparagine, tyrosine and threonine residues (N33, T87, N91, N98, Q176, Y177). From level 3, the pathway widens further and ultimately leads into the large central hydrophobic cavity.
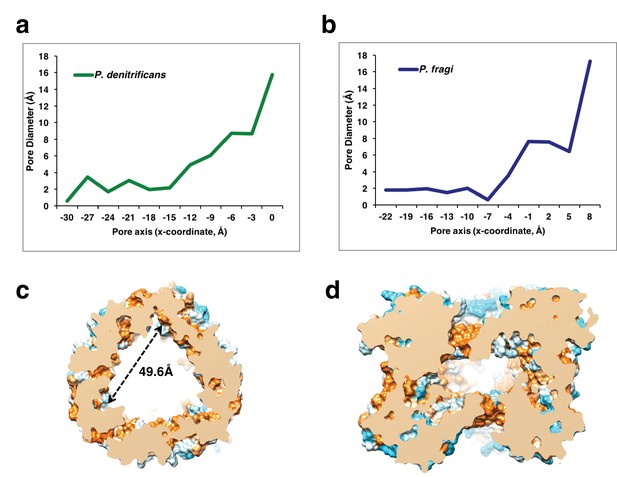
Plot of the pore diameter of the putative ion conduction pathway.
(a) PdCysZ (b) PfCysZ, showing a very narrow constriction of under 2 Å in diameter near the entrance of the pore, which then widens as the ions move past the core of the protomer to the inner central cavity of the hexameric channel (as calculated by PoreWalker, measured at 3 Å steps along the x-axis) (c) Central hydrophobic cavity of PdCysZ hexamer, cross-section of central cavity of the hexameric PdCysZ (top view) and (d) side view, shows a triangular-shaped cavity with each side measuring 49.6 Å in length. The cavity is primarily hydrophobic, as depicted and colored with the Kyte-Doolitle hydrophobicity scale from most polar colored in cyan to most hydrophobic colored in orange.
Tables
Crystallographic data and refinement statistics.
https://doi.org/10.7554/eLife.27829.002| Data collection | Native IlCysZ | IlCysZ w/SeO42- (SAD) | SeMet PfCysZ (six crystals, refinement) | SeMet PfCysZ (15 crystals, SAD) | Native PdCysZ (MR-SAD) |
|---|---|---|---|---|---|
| Beamline | NSLS X4A | NSLS X4A/C | APS 24-ID-E | APS 24-ID-C | APS 24-ID-C |
| Space group | C2 | C2 | C2 | C2 | P63 |
| Cell dimensions: | |||||
| a, b, c (Å) | 128.9, 82.0, 100.4 | 128.9, 81.9, 100.3 | 172.29, 56.9, 96.17 | 172.35, 56.9, 96.31 | 225.13, 225.13, 96.62 |
| α, β, γ (°) | 90, 125.1, 90 | 90, 125.1, 90 | 90, 91.43, 90 | 90, 91.33, 90 | 90, 90, 120 |
| Za | 2 | 2 | 2 | 2 | 6 |
| Wavelength | 1.7432 | 0.96789 | 0.97890 | 0.97890 | 1.0230 |
| Bragg spacings (Å) | 30–2.30 | 50–2.10 | 40–3.20 | 40–3.50 | 86–3.40 |
| Rmerge | 0.047 (0.257) | 0.046 (0.417) | 0.136 (6.506) | 0.135 (2.13) | 0.095 (2.89) |
| I / σI | 29.4 (7.9) | 25.0 (1.9) | 12.3 (0.7) | 24.2 (2.6) | 22.4 (1.2) |
| Completeness (%) | 99.9 (99.9) | 99.9 (100.0) | 99.8 (99.3) | 99.9 (100.0) | 100.0 (100.0) |
| Multiplicity | 7.3 (7.2) | 7.6 (7.6) | 27.2 | 72.8 | 19.7 |
| Refinement: | |||||
| Resolution (Å) | 2.30 | 3.50 | 3.40 | ||
| No. of reflections | 38075 | 20741 | 37251 | ||
| Rwork/Rfree | 0.200/0.238 | 0.297/0.339 | 0.245/0.289 | ||
| No. of atoms: | |||||
| Protein | 3679 | 3428 | 11415 | ||
| Ligand/ion | 249 | 0 | 400 | ||
| Water | 162 | 0 | 0 | ||
| Average B-factors (Å2) Protein Ligand/Ion Water | 49.4 48.4 77.3 45.3 | 198.9 198.9 - - | 178.58 179.06 164.88 - | ||
| Bond Ideality (r.m.s.d.): | |||||
| Bond lengths (Å) | 0.006 | 0.006 | 0.010 | ||
| Bond angles (°) | 0.894 | 1.133 | 0.96 | ||
| Ramachandran Analysis: Favored (%) | 99.0 | 99.2 | 96.27 | ||
| PDB accession code | 3TX3 | 6D79 | 6D9Z |
-
Values in parentheses are from the highest resolution shell. Rfree was calculated using 5% of data excluded from refinement.
Additional files
-
Transparent reporting form
- https://doi.org/10.7554/eLife.27829.018





