Cyclin A/Cdk1 modulates Plk1 activity in prometaphase to regulate kinetochore-microtubule attachment stability
Figures
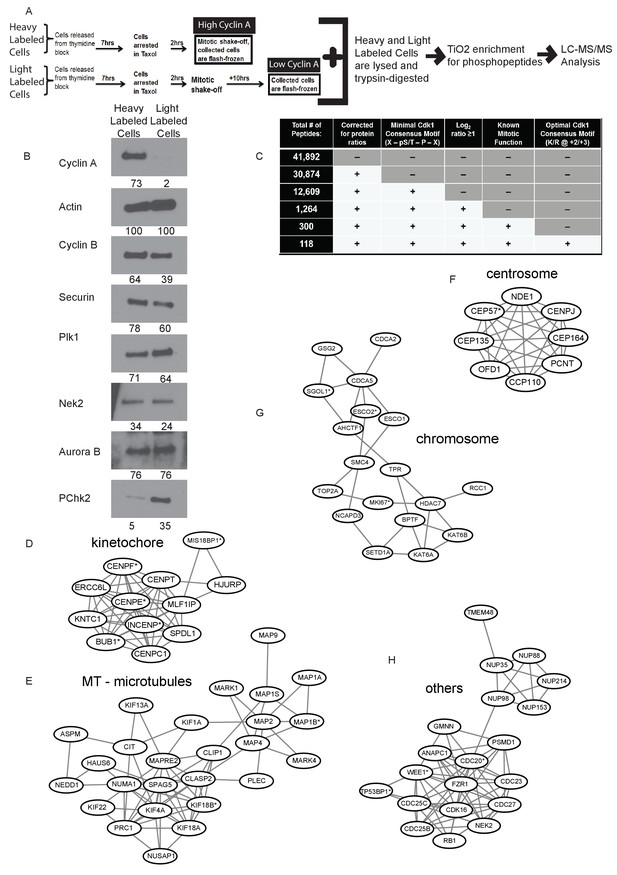
Identification of Cyclin A/Cdk1 substrates in prometaphase cells using SILAC-MS/MS.
(A) Experimental design for SILAC approach to generate two independently labeled prometaphase cell populations with differential Cyclin A levels for analysis of phosphorylation changes. (B) Western blots using whole cell lysates from the independently labeled prometaphase cell populations for the indicated protein targets. Numbers indicate protein levels relative to actin loading control in each column. (C) Summary of phosphopeptides identified by this proteomic approach. (D) –(H) Protein interactions within different mitotic categories based on STRING database analysis and drawn using Cytoscape. *indicates p<0.1 for at least one phosphopeptide on the corresponding protein.
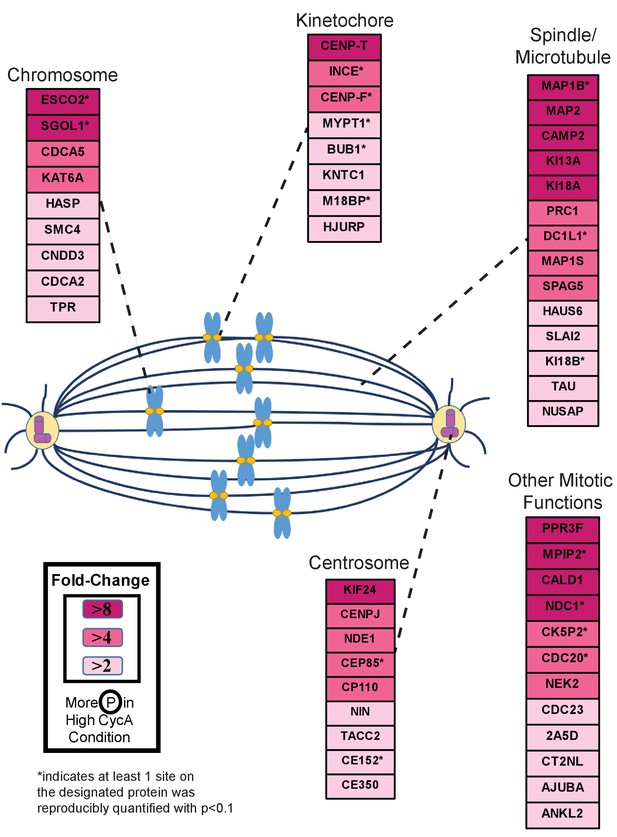
Triaged Cyclin A/Cdk1 Prometaphase Substrates by Category.
Cartoon spindle depicting examples of Cdk1 substrates that were preferentially phosphorylated in the presence of Cyclin A. Substrates are separated by mitotic function. Color intensity represents fold-change in phosphorylation; color scheme is based upon maximal difference observed in any given trial. *indicates at least one site on corresponding protein was reproducibly quantified with p<0.1 across the four trials.
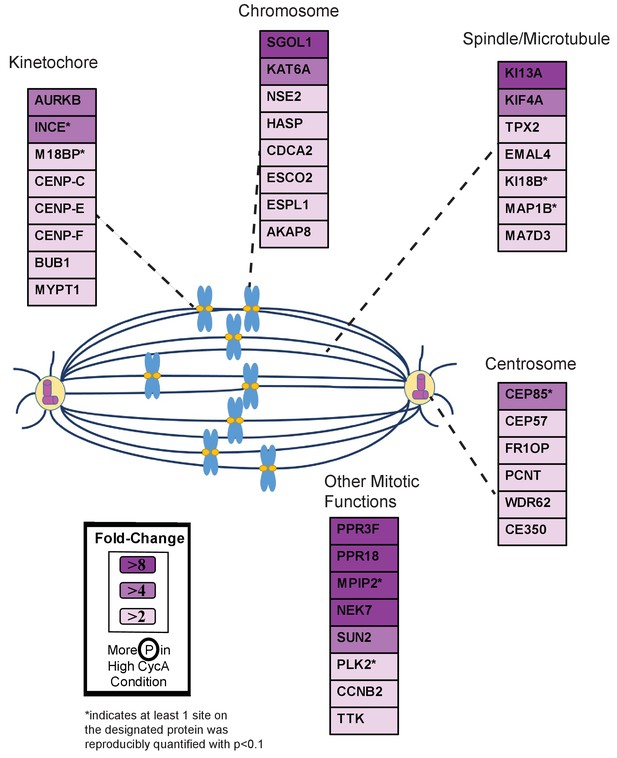
Triaged Aurora B Prometaphase Substrates by Category.
Cartoon spindle depicting examples of Aurora B kinase substrates that were preferentially phosphorylated in the presence of Cyclin A. Substrates are separated by mitotic function. Color intensity represents fold-change in phosphorylation; color scheme is based upon maximal difference observed in any given trial. *indicates at least one site on corresponding protein was reproducibly quantified with p<0.1 across the four trials.
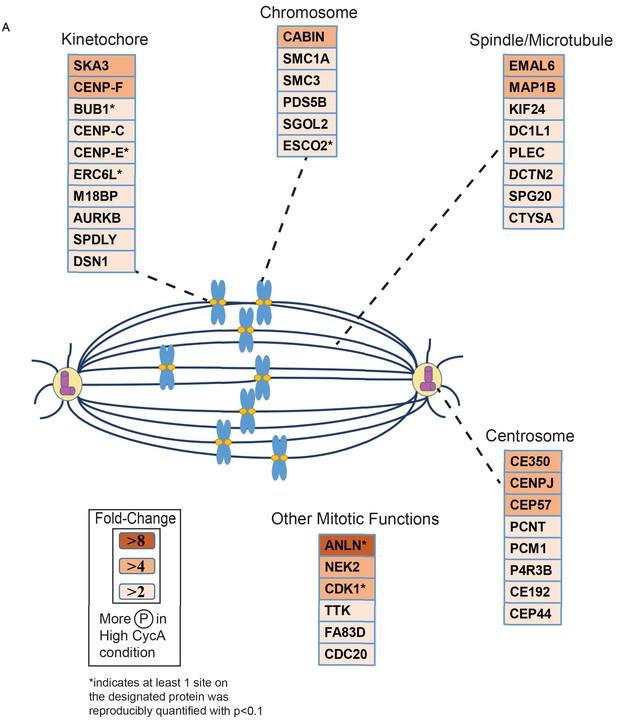
Triaged Plk1 Prometaphase Substrates by Category.
Cartoon spindle depicting examples of Plk1 substrates that were preferentially phosphorylated in the presence of Cyclin A. Substrates are separated by mitotic function. Color intensity represents fold-change in phosphorylation; color scheme is based upon maximal difference observed in any given trial. *indicates at least one site on corresponding protein was reproducibly quantified with p<0.1 across the four trials.
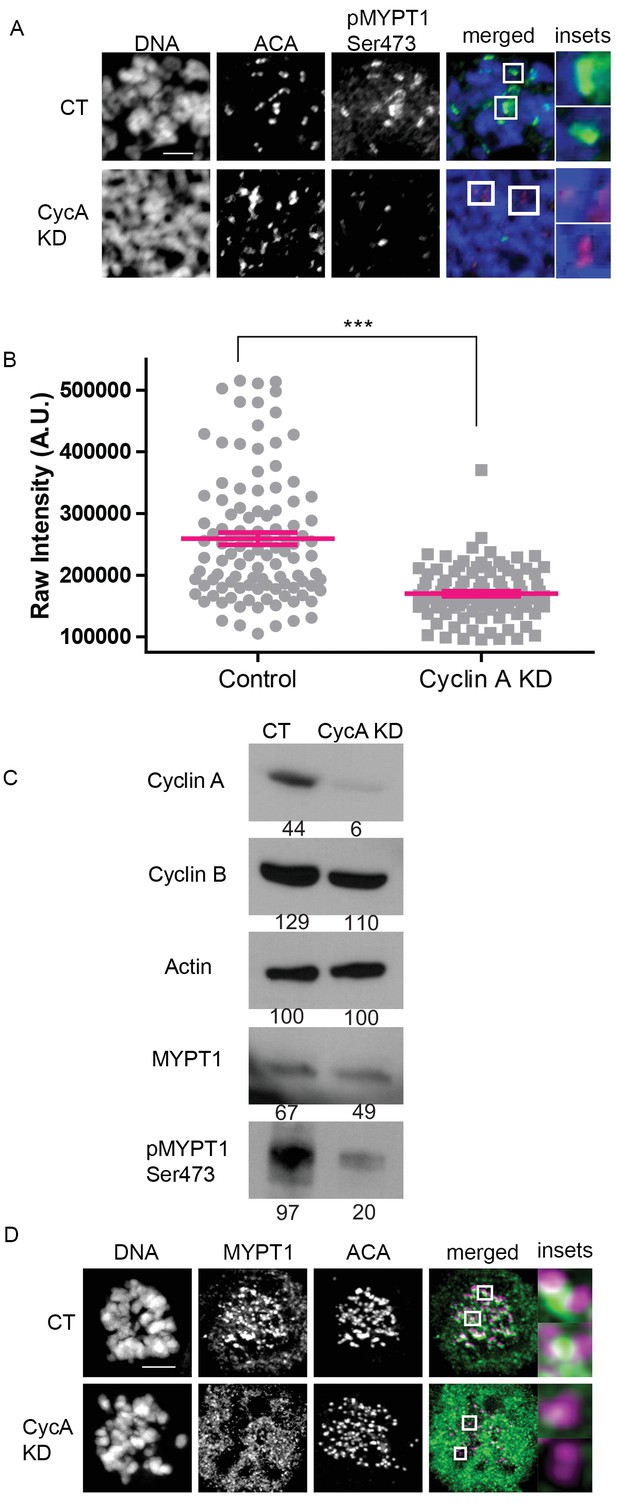
MYPT1 is a mitotic Cyclin A/Cdk1 substrate.
(A) Chromosome spreads from nocodazole-treated U2OS before (CT) and after si-RNA-mediated Cyclin A-knockdown (CycA KD) stained for DNA, centromere-specific human antisera (ACA), antibody specific to MYPT1 pSer473 (pMYPT1 Ser473). Scale bar, 5 µm. Insets highlight centromeres at 3X magnification. (B) Quantification of the intensity of centromere MYPT1 pSer473 staining on chromosome spreads (n ≥ 100 centromeres/condition. p<0.0001, unpaired, two-tailed t-test). (C) Western blots using whole cell lysates from U2OS before (CT) and after si-RNA-mediated Cyclin A-knockdown (CycA KD) for the indicated protein targets. Numbers indicate protein levels relative to actin loading control in each column. (D) U2OS before (CT) and after si-RNA-mediated Cyclin A-knockdown (CycA KD) stained for DNA, MYPT1, and centromere-specific human antisera (ACA). Scale bar, 5 µm. Insets highlight centromeres at 5X magnification.
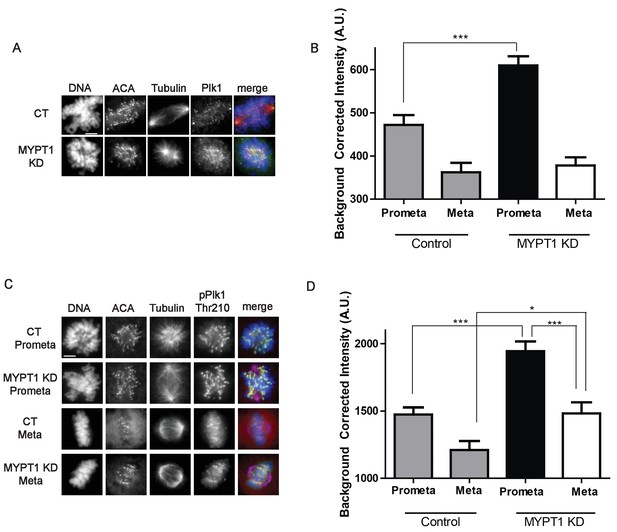
MYPT1 limits kinetochore Plk1 localization and phosphorylation.
(A) U2OS before (CT) and after si-RNA-mediated MYPT1-knockdown (MYPT1 KD) stained for DNA, centromere-specific human antisera (ACA), tubulin, and Plk1 as indicated. Scale bar, 5 µm. (B) Quantification of the intensity of kinetochore Plk1 staining in U2OS before (Control) and after si-RNA-mediated MYPT1-knockdown (MYPT1 KD) in prometaphase (Prometa) and metaphase (Meta) cells (n ≥ 100 kinetochores per condition. p<0.0001, unpaired, two-tailed t-test). (C) U2OS before (CT) and after si-RNA-mediated MYPT1-knockdown (MYPT1 KD) in either prometaphase (Prometa) or metaphase (Meta) stained for DNA, centromere-specific human antisera (ACA), tubulin, and pPlk1-Thr210 as indicated. Scale bar, 5 µm. (D) Quantification of the intensity of kinetochore pPlk1-Thr210 staining in U2OS cells before (Control) and after si-RNA-mediated MYPT1-knockdown (MYPT1 KD) in prometaphase (Prometa) and metaphase (Meta) cells (n ≥ 100 kinetochores per condition. p<0.0001, MYPT1KD prometaphase vs metaphase, p<0.0001, CT vs MYPT1 KD prometaphase; p=0.01 CT vs MYPT1 KD metaphase; unpaired, two-tailed t-test).
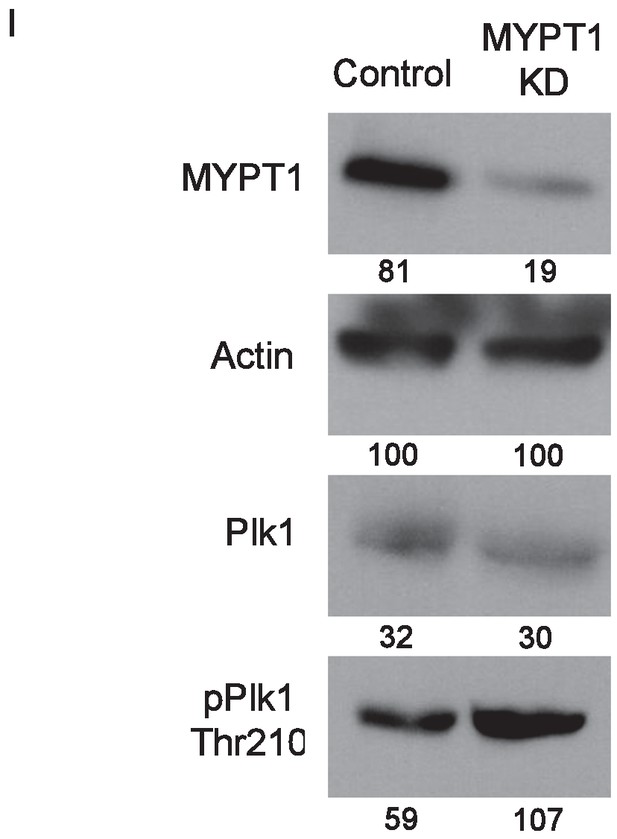
Whole cell lysates compared for changes in protein expression.
Western blots using whole cell lysates from U2OS before (Control) and after si-RNA-mediated MYPT1 knockdown (MYPT1 KD) for the indicated protein targets. Numbers indicate protein levels relative to actin loading control in each column.
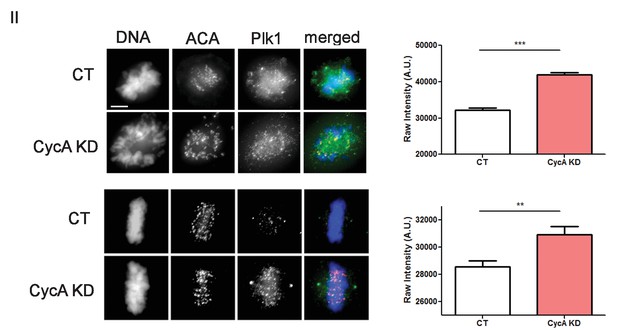
Analysis of Plk1 levels.
U2OS before (CT) and after si-RNA-mediated Cyclin A-knockdown (CycA KD) stained for DNA, centromere-specific human antisera (ACA), and Plk1 as indicated. Scale bar, 5 µm. Quantified intensity levels. (n ≥ 100 kinetochores/condition. p<0.0001 CT vs CycA KD prometaphase cells; p=0.0012 CT vs CycAKD metaphase cells).
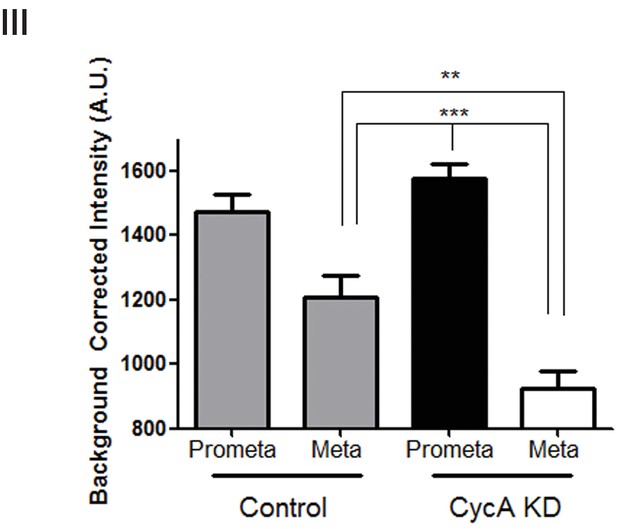
Analysis of pPlk1 levels.
Quantification of the intensity of kinetochore pPlk1-Thr210 staining in U2OS cells before (Control) and after si-RNA-mediated Cyclin A knockdown (Cyc A KD) in prometaphase (Prometa) and metaphase (Meta) cells (n ≥ 100 kinetochores/condition. p<0.0001 for CycA KD prometaphase vs metaphase, p<0.0001 for CT metaphase vs CycA KD prometaphase; p=0.001 for CT vs CycAKD metaphase).
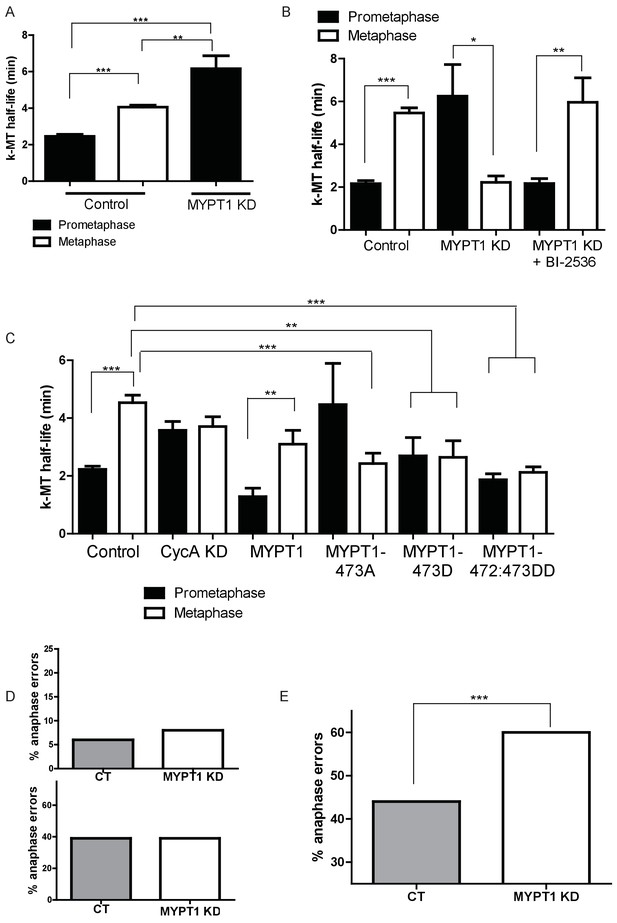
MYPT1 promotes efficient error correction by regulating Plk1.
(A) Microtubule turnover rates were measured and k-MT half-life was calculated from RPE1 cells before (Control) and after si-RNA-mediated MYPT1-knockdown (MYPT1 KD) in prometaphase and metaphase cells as indicated (n ≥ 10 cells/condition from ≥2 independent experiments. ***Indicates p<0.0001, **indicates p=0.0096, unpaired, two-tailed t-test. (B) Microtubule turnover rates were measured and k-MT half-life was calculated from U2OS cells before (Control) and after si-RNA-mediated MYPT1-knockdown (MYPT1 KD) and with the addition of the Plk1-specific inhibitor Bi-2536 in prometaphase and metaphase cells as indicated (n ≥ 15 cells/condition from ≥2 independent experiments. CT cells p<0.0001; MYPT1 KD cells p=0.04; MYPT1KD +BI-2536 p=0.0038; unpaired, two-tailed t-test). (C) Microtubule turnover rates were measured and k-MT half-life was calculated from U2OS cells before (Control) and after si-RNA-mediated Cyclin A2-knockdown (CycAKD) or transfection with plasmid containing either full-length MYPT1 (MYPT1), phosphonull (MYPT1-473A), phosphomimetic (MYPT1-473D) or tandem phosphomimetic (MYPT1-472:473DD) in prometaphase and metaphase cells as indicated (n ≥ 10 cells/condition in all conditions except MYPT1-473A metaphase where n = 9 cells). ***Indicates p<0.0001, **indicates p=0.004, unpaired, two-tailed t-test. (D) Fraction of anaphase cells displaying lagging chromosomes in RPE1 (Top) and USOS (Bottom) cells before (CT) and after si-RNA-mediated MYPT1-knockdown (MYPT1 KD) (n ≥ 300 anaphases/condition. p-values n.s., unpaired two-tailed t-test). (E) Fraction of anaphase cells displaying lagging chromosomes following release from monastrol treatment in U2OS cells before (Control) and after si-RNA-mediated MYPT1-knockdown (MYPT1 KD) (n ≥ 800 anaphases/condition from two independent experiments. p<0.0001, Chi square contingency analysis).
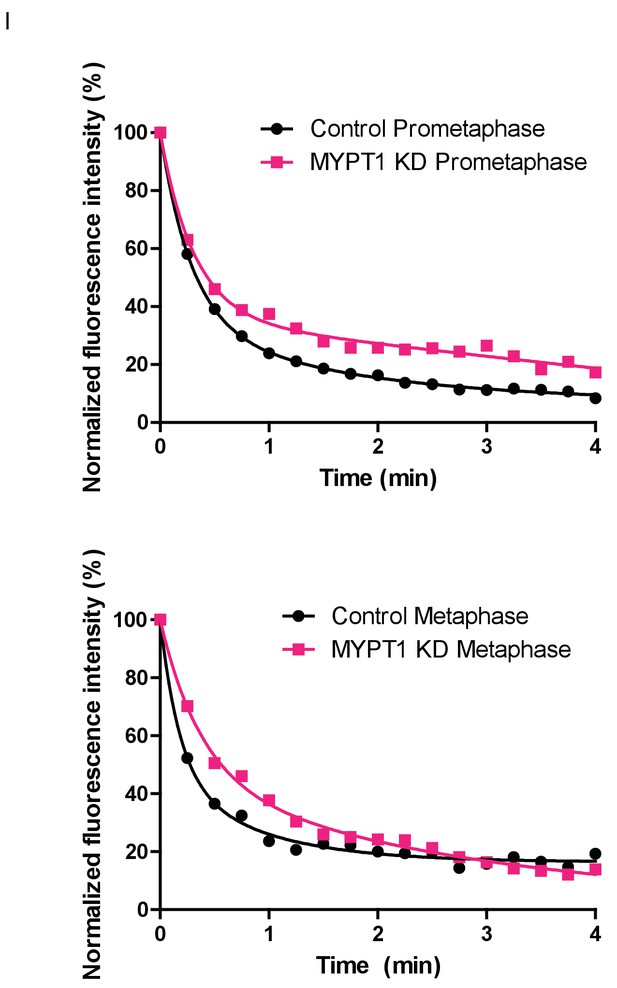
Fluorescence Dissipation After Photoactivation (FDAPA) Representative Curves From Microtubule Stability Measurements.
(Top) Examples of normalized fluorescence dissipation after photoactivation (FDAPA) in prometaphase for untreated (Control) and MYPT1-KD single cells that are representative of the mean. These data are then used to determine the rate of fluorescence dissipation over time, from which a microtubule half-life is calculated. (Bottom) Examples of normalized fluorescence dissipation after photoactivation (FDAPA) in metaphase for untreated (Control) and MYPT1-KD single cells that are representative of the mean. These data are then used to determine the rate of fluorescence dissipation over time, from which a microtubule half-life is calculated.
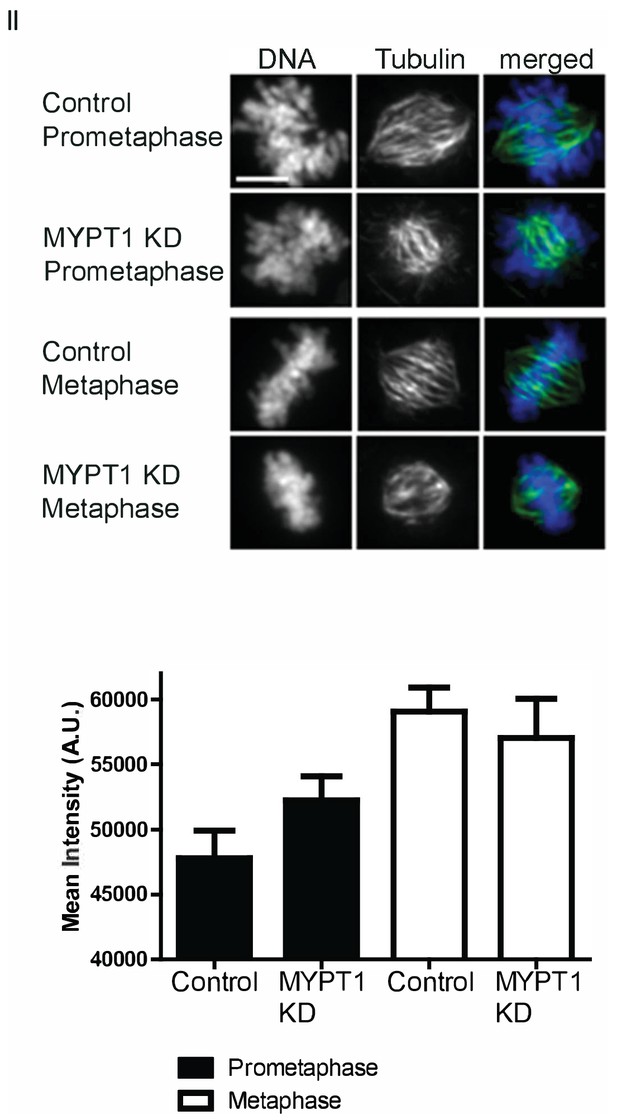
Calcium Stabilization Assay.
(Top) Representative images of assay quantified in (D). Scale bar, 5 µm. (Bottom) Quantification of intensity of spindle tubulin staining in U2OS cells treated with calcium stabilization assay without (CT) and with si-RNA mediated MYPT1 KD in prometaphase or metaphase. n ≥ 20 cells/condition.
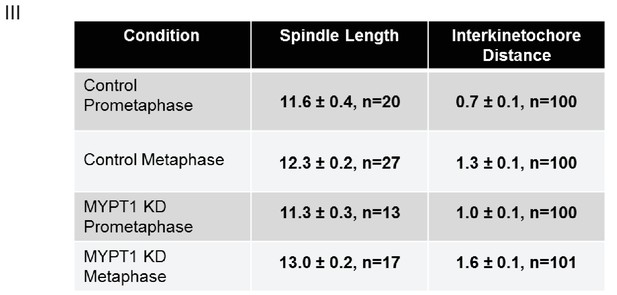
Quantifications of Other Mechanical Effects of MYPT1 in Mitotic Cells.
Table listing quantification of spindle length and inter-kinetochore distance ±SEM. For spindle length, n ≥ 13 cells per condition as listed; for inter-kinetochore distance, n ≥ 100 kinetochore pairs per condition as indicated.
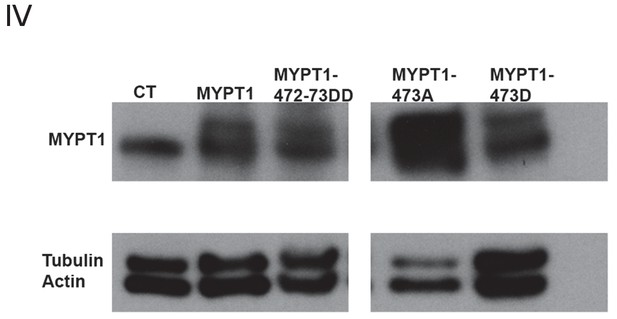
Analysis of Whole Cell Levels of Expression of Various MYPT1 Plasmids.
Western blots using whole-cell lysates from U2OS cells before (CT) and after lipofectamine-mediated transfection of either full-length MYPT1, MYPT1-472:473DD, MYPT1-473A, or MYPT1-473D mutant plasmids, for MYPT1 and loading controls (Tubulin and Actin).
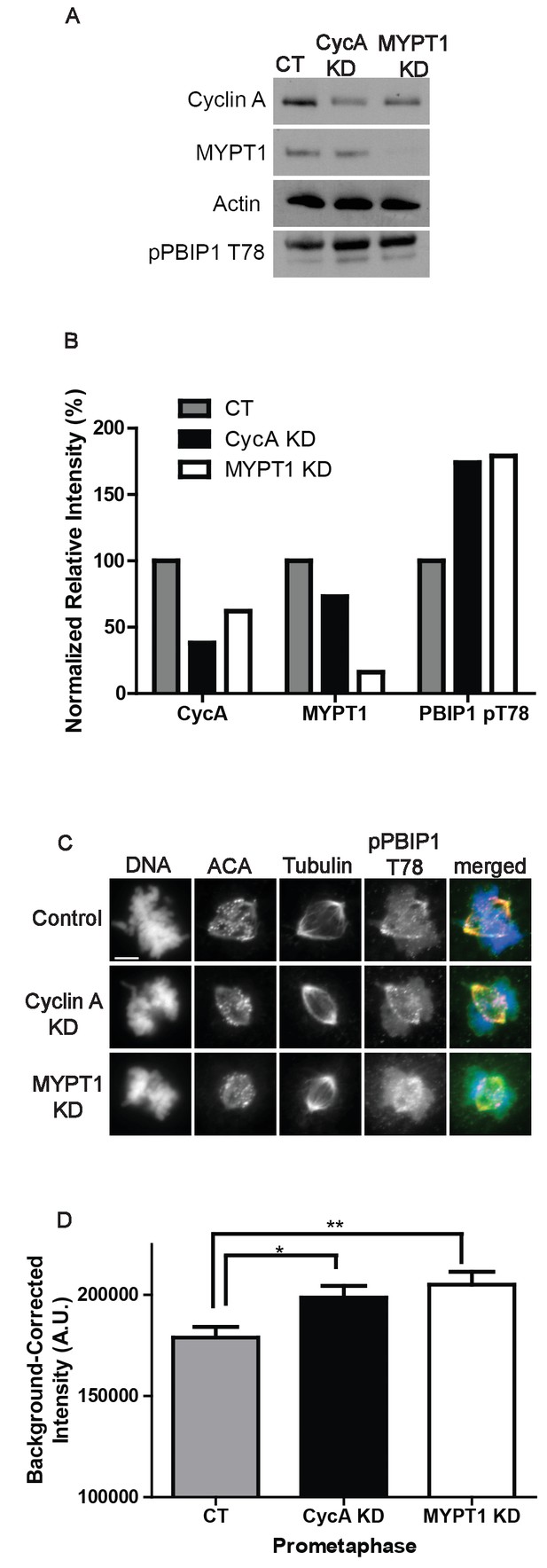
Plk1 self-priming of PBIP1 is influenced by Cyclin A/Cdk1 and MYPT1.
(A) Western blots using whole cell lysates from U2OS before (CT) and after si-RNA-mediated Cyclin A-knockdown (CycA KD) of MYPT1 knockdown (MYPT1 KD) for the indicated protein targets. (B) Quantification of Western blots for knockdown efficiency and pPBIP1-Thr78 in the presence or absence of Cyclin A or MYPT1 (p<0.0001, Chi square contingency analysis). (C) U2OS before (Control) and after si-RNA-mediated Cyclin A knockdown (Cyclin A KD) or MYPT1-knockdown (MYPT1 KD) in prometaphase cells stained for DNA, centromere-specific human antisera (ACA), tubulin, and pPBIP1-Thr78 as indicated. Scale bar, 5 µm. (D) Quantification of the intensity of kinetochore pPBIP1-Thr78 staining in U2OS cells before (CT) and after si-RNA-mediated Cyclin A knockdown (CycA KD) or MYPT1-knockdown (MYPT1 KD) in prometaphase cells (n ≥ 100 kinetochores/condition. CT vs CycA KD p=0.01, CT vs MYPT1 KD p=0.001; unpaired, two-tailed t-test).
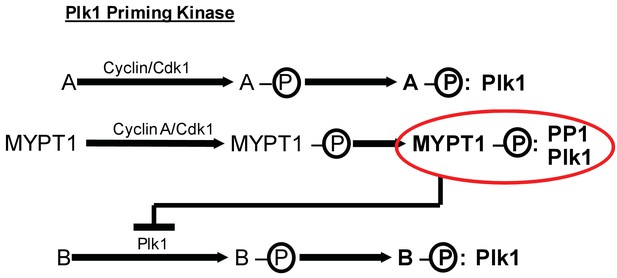
MYPT1 cross-regulates Cyclin A/Cdk1-dependent priming and self-priming of Plk1 substrates during mitosis.
Cyclin/Cdk1 has an established role in catalyzing the priming phosphorylation of substrates to promote Plk1 binding as depicted for hypothetical substrate ‘A’. MYPT1 is a specific substrate shown to be sensitive to Cyclin A/Cdk1 catalyzed priming phosphorylation for interaction with Plk1. In turn, MYPT1 recruits PP1 to dampen Plk1 which reduces the level of self-priming for hypothetical substrate ‘B’.
Additional files
-
Supplementary file 1
Mitotic phosphoproteome dataset unfiltered and filtered by Cdk1 consensus motif
Complete phosphoproteome dataset, along with dataset filtered by Cdk1 minimum consensus motif. Raw data were searched using COMET in high resolution mode against a target-decoy (reversed) version of the human proteome sequence database, filtered, and quantified, and phosphopeptide ratios were adjusted as described in the Methods section of the text. Significance of log2 phosphopeptide fold-change was determined by two-tailed Student’s t test assuming unequal variance. Dataset was also filtered by the Cdk1 minimum consensus motif (pS/pT-P; (Nigg, 1993; Moreno and Nurse, 1990; Songyang et al., 1994)), listed in the second tab of the file.
- https://doi.org/10.7554/eLife.29303.018
-
Supplementary file 2
Phosphoproteome dataset filtered by Aurora B consensus motif
Dataset filtered by the Aurora B kinase consensus motif (R/K – X – pS/pT) (Kim et al., 2010; Meraldi et al., 2004).
- https://doi.org/10.7554/eLife.29303.019
-
Supplementary file 3
Phosphoproteome dataset filtered by Plk1 consensus motif
Dataset filtered by the Plk1 kinase consensus motif (D/E-X-pS/pT) (Nakajima et al., 2003; Grosstessner-Hain et al., 2011).
- https://doi.org/10.7554/eLife.29303.020
-
Transparent reporting form
- https://doi.org/10.7554/eLife.29303.021





