CRIg, a tissue-resident macrophage specific immune checkpoint molecule, promotes immunological tolerance in NOD mice, via a dual role in effector and regulatory T cells
Figures
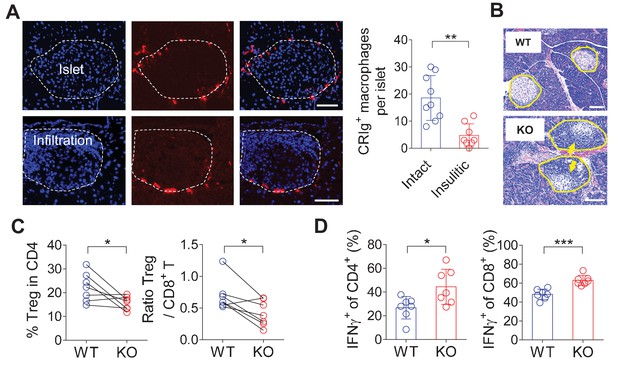
CRIg+TRMs form a protective barrier to prevent tissue autoimmune infiltration and activation.
(A) The distribution of CRIg+ TRMs within the pancreas. Immunostaining of pancreatic frozen sections of 10-week-old NOD mice. (left) Representative images depicting an intact (upper) and an insulitic (lower) islet, respectively. Red: CRIg; Blue: DAPI. The border of the islet was marked by dotted lines; immune infiltration was highlighted by higher density of DAPI+ dots (n = 9 for intact and n = 8 for insulitic). Bar, 50 um. (right) The numbers of CRIg+ TRMs per islet counted from immunostaining sections of pancreas as in (A). Islets were categorized into intact (free from infiltration) and insulitic groups. (B) The severity of insulitis in 10-week-old NOD and NOD/CRIg KO mice. Hematoxylin and eosin staining of pancreatic paraffin sections. Arrowheads depict massive immune infiltration. Bar, 50 um. (C) Flow cytometric analyses of digested pancreases of NOD/CRIg KO mice and littermate controls (n = 7 in each group). (Left) The percentages of pancreatic Treg cells in age-matched NOD and NOD/CRIg KO mice (mixed of both females and males). (Right) The ratio between Treg cells and CD8+ T cells (n = 7 in each group). (D) The production of IFN-γ in CD4+ Tconv and CD8+ T cells. Pancreatic digestions were the same as in (C). Data are representative of three (A, B) or four (C, D) experiments. Student’s t-test was used. *p<0.05; **p<0.01; ***p<0.001.
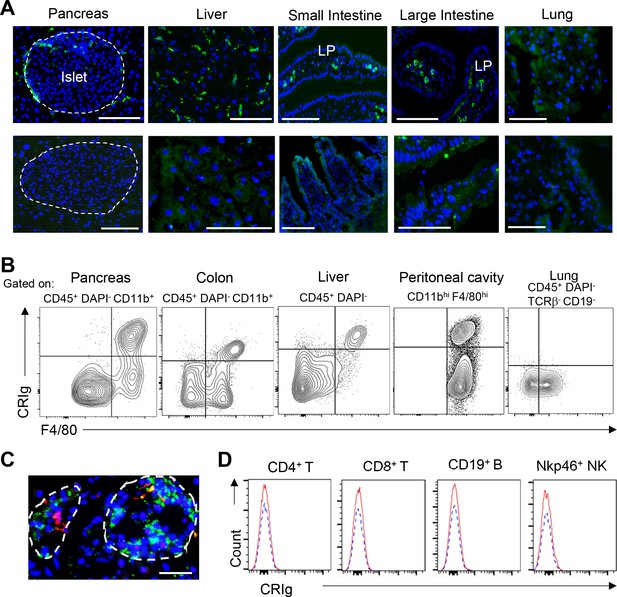
Tissue-distribution of CRIg+ TRMs.
(A) Immunostaining for CRIg on frozen sections of the pancreas, liver, small and large intestines and lung from 7-week-old B6 mice. lower panel: second Ab only. LP, lamina propria. Bar: 50 um. (B) Flow cytometric analyses of CRIg expression in different tissues. The strain and age of the mice were the same as in (A). Bar: 50 um. Data are representative of at least three experiments. (C) The expression of CRIg in human pancreas. Red, CRIg; green, insulin. Dotted lines depicted islets. Data are representative of four human pancreases. (D) Flow cytometric analyses of CRIg in lymphoid-lineage cells. None of these cells expresses CRIg.
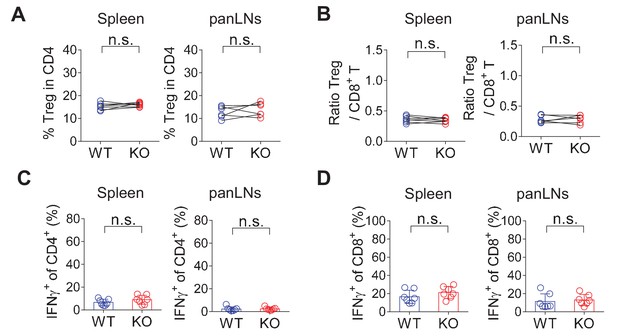
CRIg deficiency does not affect T cells in lymphoid organs.
(A) The percentages of Treg cells in age-matched wildtype and CRIg KO mice (mixed of both females and males) in spleen and pancreatic draining LNs (panLNs). (B) The ratios between Treg cells and CD8+ T cells in age-matched wildtype and CRIg KO mice (mixed of both females and males) in spleen and panLNs. (C) The production of IFN-γ in CD4+ Tconv from spleen or panLNs of wildtype and CRIg KO mice. (D) The production of IFN-γ in CD8+ T cells from spleen or panLNs of wildtype and CRIg KO mice. Data are representative of four experiments. Student’s t-test was used. n.s., non-significant. WT, wildtype; KO, CRIg knockout.
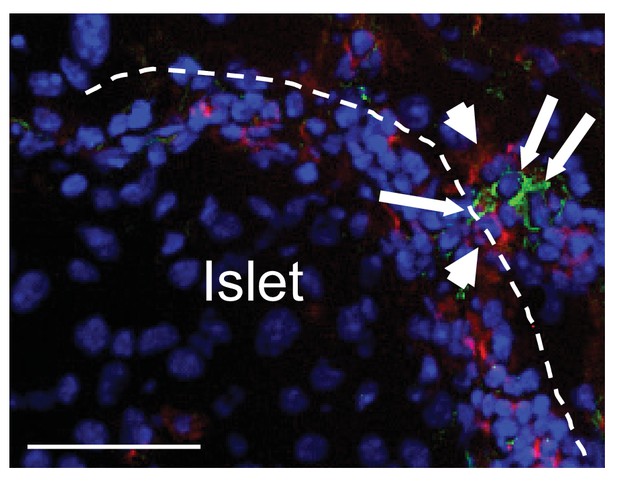
Cell-cell contact between CRIg+TRMs and T cells in pancreas.
(A) Cell-cell contact between CRIg+ macrophages (green) and islet-infiltrating CD4 T cells (red). Arrowheads, CD4+ T cells; long arrows, CRIg+ TRMs. Bar, 50 um.
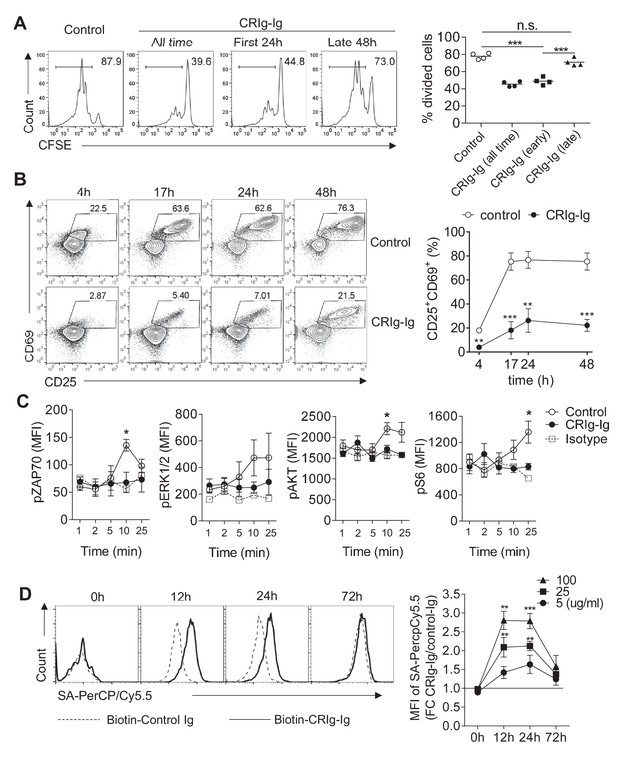
CRIg suppresses T cell activation.
(A) CFSE-labeled CD4+ CD25- Tconv cells were stimulated with anti-CD3/CD28. Plate-bound CRIg-Ig, or control Ig, was added either all time during T cell culture (3 d), or only the first 24 hr, or the late 48 hr. T cell proliferation was measured by CFSE dilution. (B) The expression of early T cell activation markers-CD69 and CD25 in cultured Tconv cells with plate-bound control Ig, or CRIg-Ig. (C) The phosphorylation of early T cell activation cascade proteins. CD4+ CD25- Tconv cells were activated in vitro with anti-CD3/CD28 in the presence of plate-bound control Ig, or CRIg-Ig. (D) The binding of CRIg to activated T cells. CD4+ CD25- Tconv cells were activated in vitro by anti-CD3/CD28 for various lengths of time (12 hr, 24 hr and 72 hr). At each time-point of T cell activation, various concentrations of biotinylated control Ig or CRIg-Ig were incubated with T cells at 37°C for 1 hr. The binding of biotin-labeled Ig proteins was detected using streptavidin-conjugated antibody. (left) Representative histograms of SA-PercpCy5.5 MFIs depicting the binding intensities of biotinylated proteins (25 ug/ml) bound to T cells. (right) Statistical data showing fold-changes of SA-PercpCy5.5 MFIs between CRIg-Ig and control Ig at each time-point and each concentration. P values were calculated by comparing the binding intensities between biotin-CRIg-Ig and biotin-control-Ig. The data are representative from five (A), three (B), and four (C, D) experiments. Student’s t-test was used. *p<0.05; **p<0.01; ***p<0.001, n.s., non-significant.
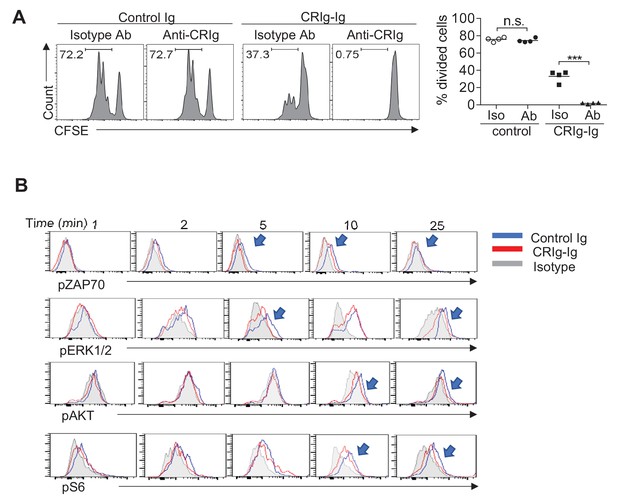
The suppressive effect of CRIg on T cells is complement-independent.
(A) CFSE-labeled CD4+ Foxp3(GFP)- Tconv cells were stimulated with anti-CD3/CD28, in the presence of plate-coated control Ig, or CRIg-Ig. Soluble anti-CRIg mAb (clone 14G8) was added. T cell proliferation was analyzed by CFSE dilutions after 3 days. (B) CRIg can attenuate TCR signaling in serum-free condition (deprived of complements). Arrow heads depict the suppression of phosphorylation of each respective TCR signaling cascade proteins. Data are representative of four (A) and three (B) experiments. Student’s t-test was used. n.s., non-significant; ***p<0.001.
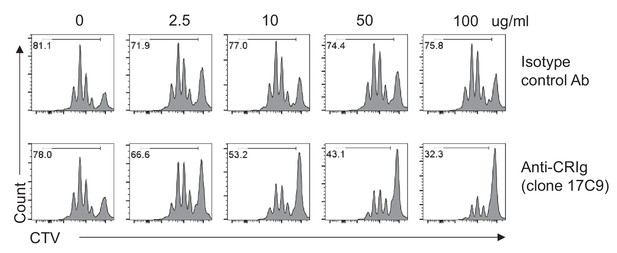
Plate-bound anti-CRIg mAb augments the effect of CRIg-Ig in T cells.
CTV-labeled CD4+ Foxp3(GFP)- Tconv cells were stimulated with anti-CD3/CD28, in the presence of plate-coated isotype-matched control mAb, or anti-CRIg-Ig (17C9) (with various concentrations as indicated). Soluble CRIg-Ig was added (5 ug/ml) into the culture medium. After 3 days of culture, cell proliferation was measured by CTV dilution. Numbers depicted proportions of divided cells. Data are representative of three independent experiments.
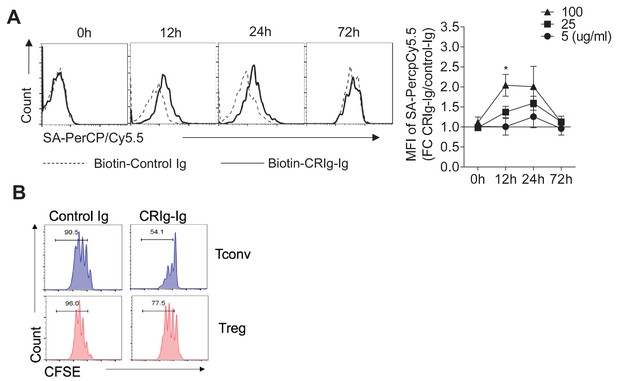
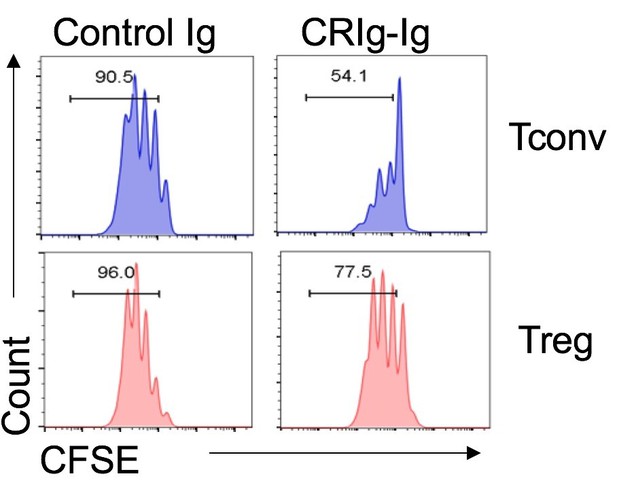
The binding of CRIg to activated Treg cells and differential suppression of CRIg-Ig for Tconv and Treg cells.
(A) CD4+ CD25+ Treg cells were activated in vitro by anti-CD3/CD28 for various lengths of time (12 hr, 24 hr and 72 hr). At each time-point of activation, various concentrations of biotinylated control Ig or CRIg-Ig were incubated with T cells at 37°C for 1 hr. The binding of biotin-labeled Ig proteins to Foxp3(GFP)+ Treg cells was detected using streptavidin-conjugated antibody. (left) Representative histograms of SA-PercpCy5.5 MFIs depicting the binding intensities of biotinylated proteins (25 ug/ml) bound to Treg cells. (right) Statistical data showing fold-changes of SA-PercpCy5.5 MFIs between CRIg-Ig and control Ig at each time-point and each concentration. P values were calculated by comparing the binding intensities of Biotin-CRIg-Ig to Biotin-control-Ig. *p<0.05. (B) Tconv and Treg cells were labeled with CFSE and cultured under the same condition (anti-CD3/CD28, IL-2 with the presence of plate-coated CRIg-Ig or control Ig. Cell proliferation was measured after 3 days of culture. Data were representative of three experiments.
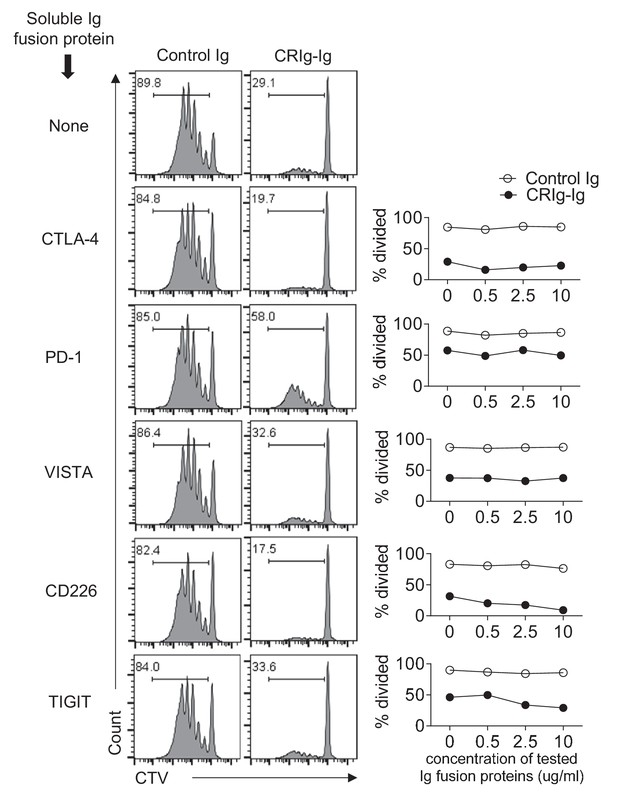
Ig fusion proteins of CTLA4, PD-1, VISTA, CD226 and TIGIT do not abolish the suppression of CRIg-Ig in T cells.
CTV-labeled CD4+ Foxp3(GFP)- Tconv cells were stimulated with anti-CD3/CD28, in the presence of plate-coated control Ig, or CRIg-Ig (5 ug/ml). Soluble Ig fusion proteins for CTLA-4, PD-1, VISTA, CD226 and TIGIT (from R and D) were added at indicated concentrations. T cell proliferation was analyzed by CTV dilutions at day 3. Shown are representative of three experiments.
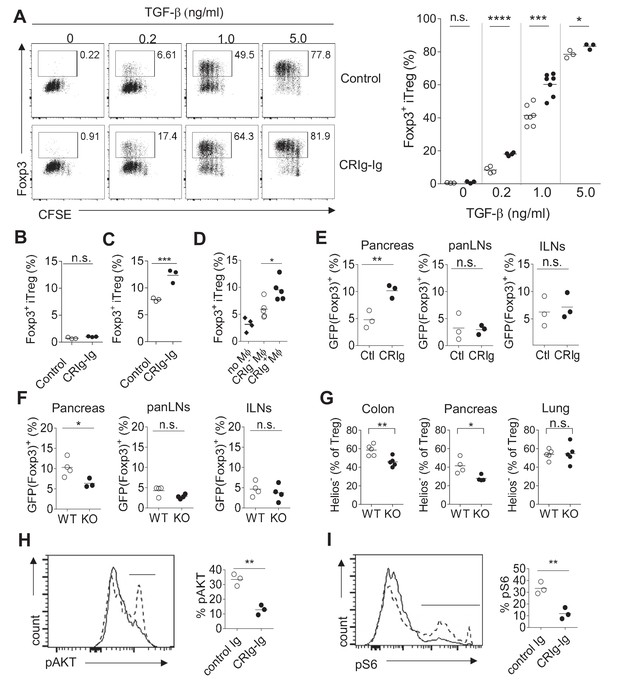
CRIg promotes iTreg generation in vitro.
(A) (left) Representative FACS plots depicting the generation of iTreg cells in the presence of CRIg-Ig, or control Ig, under various concentrations of TGF-β. (right) Statistics of multiple experiments. (B) Tconv cells were cultured in the condition of anti-CD3/CD28 and IL-2, anti-TGF-β neutralizing antibody (clone 1D11) and either control Ig, or CRIg-Ig. (C) Total splenocytes from 8-week-old NOD/BDC2.5/Thy1.1 mice were labeled with CTV and cultured with BDC2.5 mimotope (100 ng/ml) for 3 days. The generation of Treg cells were analyzed by intracellular Foxp3 staining. (D) iTreg generation in vitro as in (C) with the inclusion of CRIg+ or CRIg- macrophages, sorted from peritoneal cavity. (E) Purified CD4+Foxp3 (GFP)- T cells from NOD/BDC2.5/Foxp3GFP/Thy1.1 mice were transferred into 4-week-old NOD mice, followed by i.p. injection of CRIg-Ig, or control Ig every other day for 2 weeks. (F) Purified CD4+Foxp3 (GFP)- T cells from NOD/BDC2.5/Foxp3GFP/Th1.1 mice were transferred into 4-week-old NOD or NOD/CRIg KO mice. The generation of Foxp3(GFP)+ cells in pancreatic islets, panLNs and inguinal LNs(ILNs) was analyzed 2 weeks after the transfer. (G) Flow cytometric analyses of Helios- Treg cells from pancreas, colon and lung of NOD/CRIgKO and wildtype controls. (H, I) Purified CD4+ Foxp3(GFP)- T cells were cultured with either control Ig or CRIg-Ig for 18 hr and analyzed for the phosphorylation of AKT (H) and ribosomal protein S6 (I). Dotted line, control Ig; solid line, CRIg-Ig. Data are representative of five (A), three (B–I) experiments. Student’s t-test was used. n.s., non-significant. *p<0.05; ***p<0.001; ****p<0.0001. Ctl, control; KO, CRIg knockout.
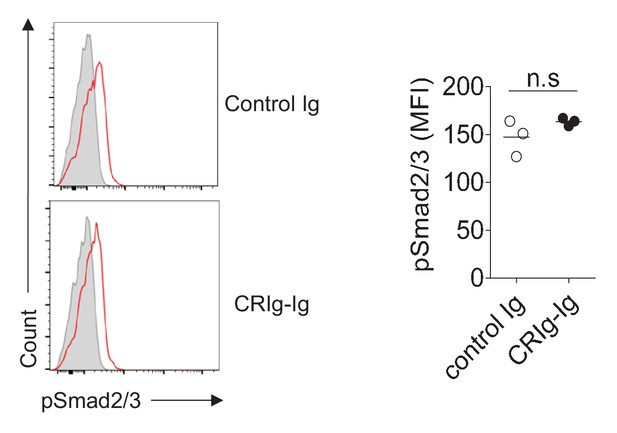
CRIg-Ig does not enhance TGF-β induced phosphorylation of Smad2/3.
CD4+ CD25- cells were sorted and cultured with 0.2% of fetal bovine serum under iTreg differentiation condition. Phosphorylation of Smad2/3 was detected 2 hr after TGF-β stimulation by flow cytometry. Data are representative of three experiments. n.s., non-significant.
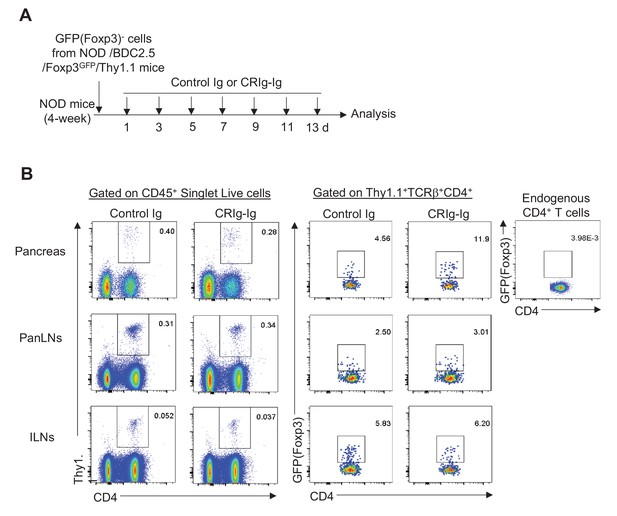
Experimental design and FACS profiles of in vivo pTreg generation promoted by CRIg-Ig.
(A) Schematic diagram depicting the experimental design. (B) Representative FACS profiles of gating strategy showing the engraftments of transferred T cells and the proportions of GFP+ cells. The top-right panel was negative control for GFP gating.
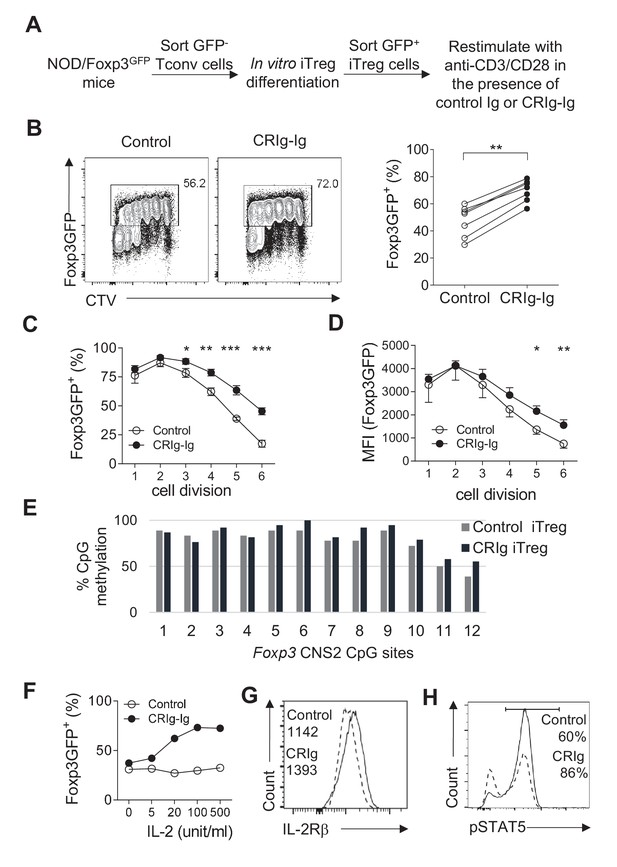
CRIg stabilizes Foxp3 expression in TGF−β induced iTreg cells.
(A) Experimental setting. (B) The fraction of Foxp3 positivity and the division of recultured iTreg cells. Left, representative FACS plots; right, the statistics of multiple experiments (n = 7). (C) The percentage of cells retaining Foxp3 expression in each cell division. (D) The MFI of Foxp3 protein in each generation of cell division. (E) The methylation percentage at each CpG motif in Foxp3 CNS2 of control iTreg (grey bars), or CRIg iTreg cells (black bars) (see Figure 4—source data 1) (F) In vitro differentiated iTreg cells were restimulated with anti-CD3/CD28, and various concentrations of IL-2, in the presence of CRIg-Ig, or control Ig. The fraction of cells retaining Foxp3 expression was analyzed after 3 days. (G) The expression of IL-2Rβ in control and CRIg iTreg cells after 3 days of culture. (H) The phosphorylation of STAT5 in control and CRIg iTreg cells. Data are representative of seven (B–D), two (E), and three (F–H) experiments, respectively. Student’s t-test was used. *p<0.05; **p<0.01; ***p<0.001.
-
Figure 4—source data 1
The methylation percentage at each CpG motif in Foxp3 CNS2 of control iTreg cells, CRIg iTreg cells and ex vivo Treg cells (associated with Figure 4E).
- https://doi.org/10.7554/eLife.29540.016
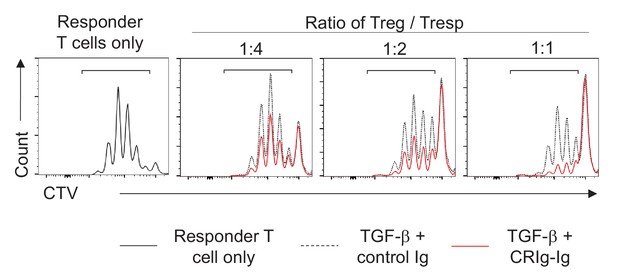
CRIg enhances iTreg suppressive function.
In an in vitro Treg suppression assay, responder T (Tresp) cells were labeled with CTV and cocultured with indicated ratios of control iTreg or CRIg-induced iTreg cells. The proliferation of responder T cells was analyzed after 3 days. Data are representative of three experiments.
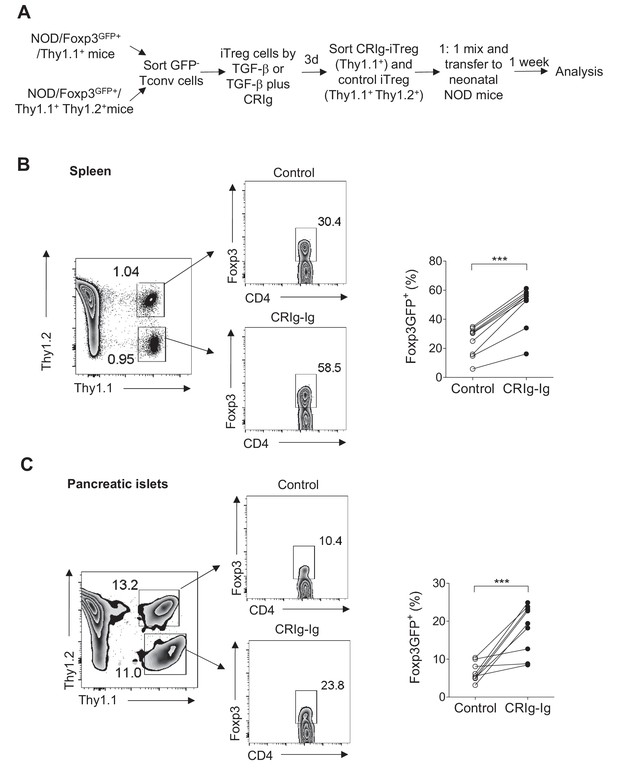
CRIg stabilizes adoptively transferred iTreg cells in vivo.
(A) Experimental design. (B, C) One week later, the transferred cells were isolated from spleen (B) and pancreatic islets (C) and were analyzed for the expression of Foxp3. Control iTreg cells, Thy1.1+ Thy1.2+; CRIg iTreg cells, Thy1.1+ Thy1.2-. Data are representative of two independent experiments with 9 mice in total. Student’s t-test was used. ***p<0.001.
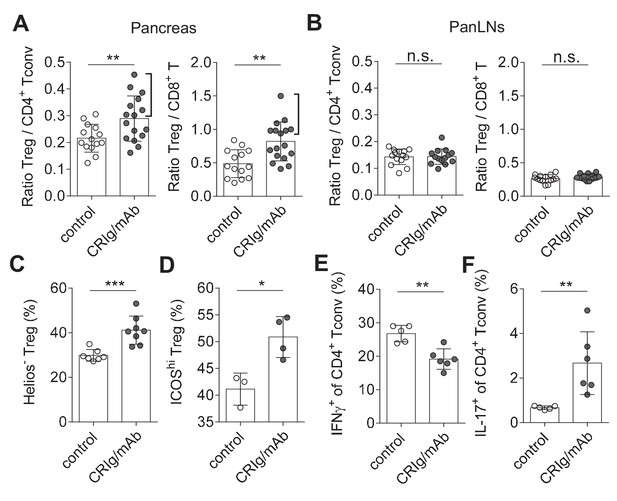
CRIg restores immune tolerance in pancreatic islets of NOD mice.
The ratios of Treg/Tconv cells, and Treg/CD8+ T cells in the pancreas (A) and panLNs (B) of control (n = 14) and CRIg-Ig/anti-CRIg treated mice (n = 16). The expression of Helios (C) and ICOS (D) in Treg cells from pancreatic islets of control (n = 7 in C, n=3 in D) and CRIg-Ig/anti-CRIg (labeled as CRIg/mAb in the figure panels, n = 8 in C, n=4 in D) treated mice. The production of IFN-γ (E) and IL-17 (F) in CD4+ Tconv cells from pancreatic islets in control Ig (n = 5) and CRIg-Ig/anti-CRIg (n = 6) treated mice. Data are representative of six (A, B), three (C), and two (D–F) experiments, respectively. Student’s t-test was used. n.s., non-significant; *p<0.05; **p<0.01; ***p<0.001.
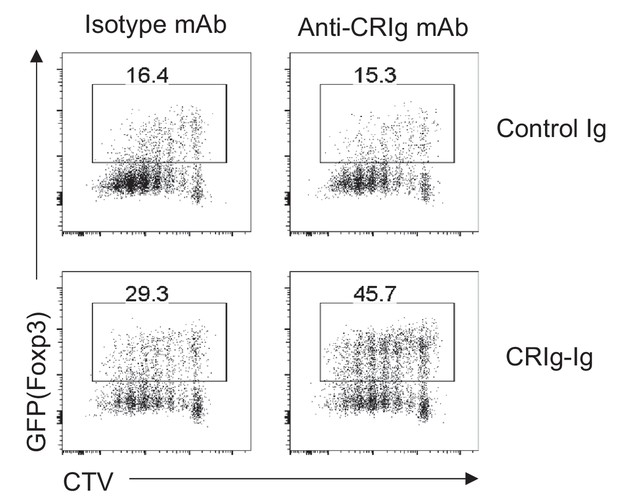
Cross-linking CRIg with anti-CRIg mAb enhances iTreg generation.
CD4+ GFP(Foxp3)- Tconv cells were sorted, labeled with CTV, and cultured in iTreg differentiation condition (anti-CD3/CD28, TGF-β and IL-2) with plate-bound control Ig or CRIg-Ig. Soluble isotype mAb or anti-CRIg mAb (17C9) was added. After 3 days, cultured cells were analyzed for their expression of Foxp3 (GFP) and the dilution of CTV. Data are representative of three experiments.
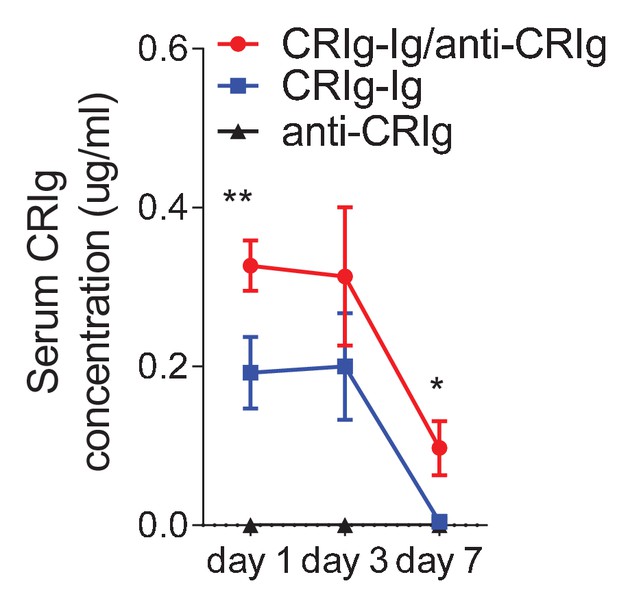
Anti-CRIg mAb prolongs in vivo half-life of CRIg-Ig.
ELISA of serum CRIg-Ig concentrations at different time-points. 10-week-old NOD mice were treated with one-dose of CRIg-Ig (3.5 mg/kg), or CRIg-Ig (3.5mg/kg) plus anti-CRIg (clone 17C9, 7 mg/kg). Small amounts of blood were collected at various time-points post treatment. CRIg-Ig concentrations were measured by ELISA using different isotypes of anti-CRIg mAbs that recognized different epitopes of CRIg. Data are representative of three independent experiments. Student’s t-test was used. *p<0.05; **p<0.01.
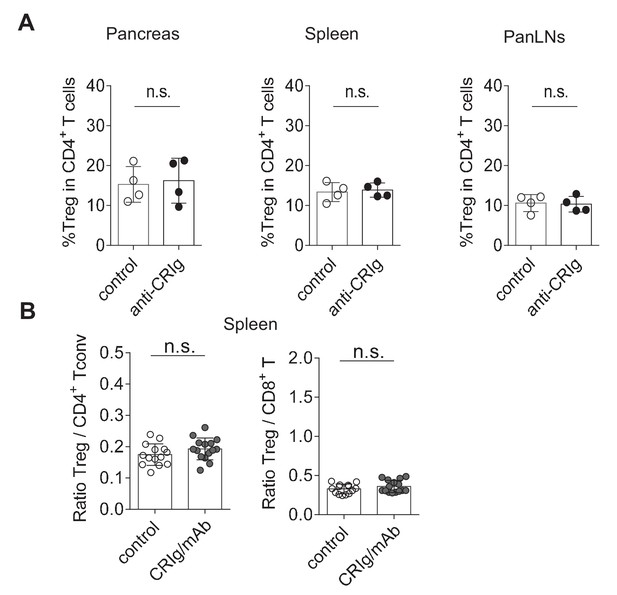
In vivo modulation of CRIg in NOD mice.
(A) anti-CRIg mAb treatment does not affect Treg cell abundance in NOD mice. 10-week-old prediabetic female NOD mice from the same litter were randomly grouped and i.p. injected with either control isotype mAb or anti-CRIg mAb twice a week for two weeks. Treg cells isolated from spleen, panLNs and pancreatic islets were analyzed. (B) The ratios of Treg cells to Tconv cells and Treg cells to CD8+ T cells in the spleen of control and CRIg-Ig/anti-CRIg treated mice. Data are representative of two (a) and six (b) experiments. Student’s t-test was used. n.s., non-significant.
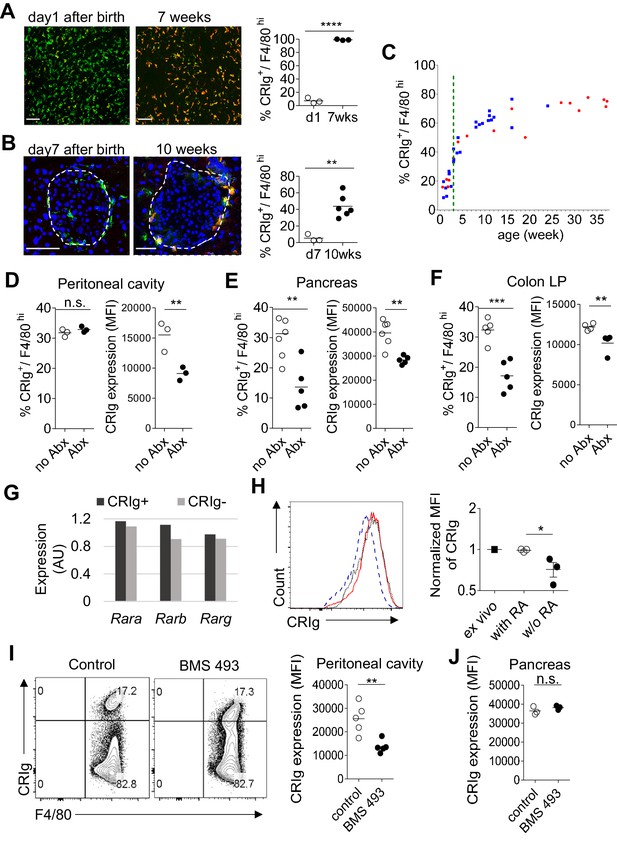
The expression of CRIg in TRMs is influenced by environmental factors.
(A) The expression of CRIg in liver Kupffer cells of adult (7 weeks of age) and neonatal (day one post birth) B6 mice. Green, F4/80; Red, CRIg. (B) The expression of CRIg in pancreatic islets of adult (10 weeks of age) and neonatal (day seven post birth) NOD mice. Green, F4/80; Red, CRIg. (C) Longitudinal analysis of CRIg expression in peritoneal F4/80hi TRMs of NOD mice. Red: females; Blue, males. Dotted green line depicts the time of weaning. (D–F) The percentages and the MFI of CRIg expression in TRMs from peritoneal cavity (D), pancreatic islets (E) and colon (F) of control and antibiotics treated B6 mice, respectively. (G) The expression of retinoic acid receptors in CRIg+ and CRIg- TRMs isolated from peritoneal cavity of 7 weeks old B6 mice. (H) CRIg+ macrophages were isolated from peritoneal cavity and cultured in vitro in the presence or absence of ATRA. The expression of CRIg in cultured cells were analyzed 3 days later. blue line, no ATRA; red line, with ATRA. (I, J) In vivo treatment of 7 weeks old B6 mice with an inverse pan-RA agonist BMS 493 to block RA signaling. The expression of CRIg in TRMs from peritoneal cavity (I) and pancreatic islets (J) was analyzed 3 days later. Data are representative of three (A, B, D–J), and more than five (C) experiments, respectively. Student’s t-test was used. n.s., non-significant; *p<0.05; **p<0.01; ***p<0.001; ****p<0.0001. w/o, without.
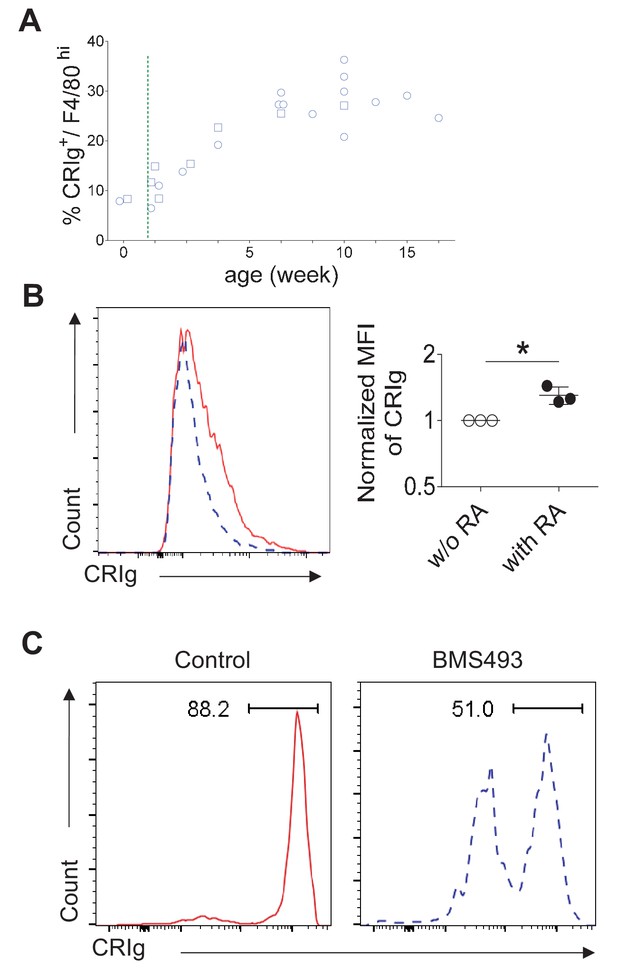
RA signaling is involved in CRIg expression in TRMs.
(A) Longitudinal analysis of CRIg expression in peritoneal resting TRMs from B6 mice. circle: females; square, males. Dotted green line depicts the time of weaning. (B) CRIg- peritoneal resting macrophages were sorted and cultured in vitro with ATRA and the expression of CRIg was analyzed after 3 days. blue line, no ATRA; red line, with ATRA. (C) CRIg+ macrophages (3 × 104) were sorted from peritoneal cavity of CD45.1+ B6 mice and transferred into 7 weeks old B6 mice. The recipient mice were treated with control PBS or an inverse pan-RA agonist BMS 493 to block RA signaling. The expression of CRIg was analyzed 3 days later. Blue, BMS 493; Red, DMSO.
Tables
| Reagent type (species) or resources | Designation | Source or reference | Identifiers | Additional information |
|---|---|---|---|---|
| strain, strain background (Mus musculus) | C57BL/6 (B6) mice | The Jackson Laboratory | RRID:IMSR_JAX:000664 | |
| strain, strain background (Mus musculus) | NOD mice | Mathis-Benoist laboratory | RRID:IMSR_JAX:001976 | |
| strain, strain background (Mus musculus) | NOD/Foxp3GFP mice | The Jackson Laboratory | RRID:IMSR_JAX:025097 | |
| strain, strain background (Mus musculus) | B6/CD45.1 mice | Dr. Li-Fan Lu, UCSD | RRID:IMSR_JAX:002014 | |
| strain, strain background (Mus musculus) | NOD/BDC2.5 /Thy1.1 mice | Mathis-Benoist laboratory | ||
| strain, strain background (Mus musculus) | NOD/BDC2.5 /Foxp3GFP /Thy1.1 mice | this paper | NOD/BDC2.5/Foxp3GFP/ Thy1.1 mice were generated by crossing NOD/BDC2.5/Thy1.1 mice to NOD/ Foxp3GFP mice. | |
| strain, strain background (Mus musculus) | NOD/BDC2.5 /Foxp3GFP /Thy1.1/Thy1.2 mice | this paper | NOD/BDC2.5/Foxp3GFP/ Thy1.1/Thy1.2 mice were generated by crossing NOD/BDC2.5/Thy1.1 mice to NOD/Foxp3GFP mice. | |
| strain, strain background (Mus musculus) | B6/CRIg KO mice | Genentech | PMID: 16530040 | |
| strain, strain background (Mus musculus) | NOD/CRIg KO mice | this paper | NOD/CRIg-/- mice were generated by crossing B6/CRIg-/- mice onto a NOD background for more than 10 generations. | |
| biological sample (Mus musculus) | pancreas | other | prepared from NOD mice | |
| biological sample (Mus musculus) | colon lamina propria | other | prepared from NOD orB6 mice | |
| biological sample (Mus musculus) | peritoneal cavity cells | other | prepared from NOD or B6 mice | |
| biological sample (Mus musculus) | lung | other | prepared from NOD or B6 mice | |
| biological sample (Mus musculus) | liver | other | prepared from NOD or B6 mice | |
| biological sample (Mus musculus) | serum | other | prepared from NOD mice | |
| biological sample (Mus musculus) | panLNs | other | pancreatic draining lymph nodes from NOD mice | |
| biological sample (Mus musculus) | ILNs | other | Inguinal lymph nodes from NOD or B6 mice | |
| biological sample (Mus musculus) | spleen | other | prepared from NOD or B6 mice | |
| biological sample (Mus musculus) | macrophages | other | defined as F4/80+ or F4/80+ CD11b+ cells | |
| biological sample (Mus musculus) | Treg cells | other | defined as CD4+ Foxp3+ T cells | |
| biological sample (Mus musculus) | Tconv cells | other | defined as CD4+ Foxp3- T cells | |
| antibody | Ultra-LEAF Purified anti-mouse CD3ε (Armenian hamster monoclonal) | BioLegend | RRID:AB_11149115 | clone: 145–2C11 |
| antibody | Ultra-LEAF Purified anti-mouse CD28 (Syrian hamster monoclonal) | BioLegend | RRID:AB_11150408 | clone: 37.51 |
| antibody | anti-gp120 (mouse monoclonal) | Genentech | control Ig in this paper PMID: 16530040 | |
| antibody | anti-CRIg | Genentech | clone: 14G8 (mouse monoclonal; PMID: 19017980); clone: 17C9 (rat monoclonal; PMID: 16530040) | |
| antibody | anti-TGF-β 1, 2, 3 (mouse monoclonal) | R and D Systems | RRID:AB_357931 | clone: 1D11 |
| antibody | anti-CD16/CD32 (rat SD monoclonal) | BD Biosciences | RRID:AB_394656 | clone: 2.4G2 |
| antibody | anti-CRIg (mouse monoclonal) | this paper | Biotinylated anti-CRIg (clone: 14G8) prepared by our laboratory | |
| antibody | anti-CD45 (rat monoclonal) | BioLegend | RRID:AB_312981 | clone: 30-F11 |
| antibody | anti-CD45.1 (mouse monoclonal) | BioLegend | RRID:AB_893346 | clone: A20 |
| antibody | anti-CD45.2 (mouse monoclonal) | BioLegend | RRID:AB_389211 | clone: 104 |
| antibody | anti-TCRβ (Armenian hamster monoclonal) | BioLegend | RRID:AB_493344 | clone: H57-597 |
| antibody | anti-CD4 (rat monoclonal) | BioLegend | RRID:AB_312719; RRID:AB_312713; RRID:AB_312715 | clone: RM4-5 |
| antibody | anti-CD8α (rat monoclonal) | BioLegend | RRID:AB_312747; RRID:AB_312761 | clone: 53–6.7 |
| antibody | anti-Thy1.1 (mouse monoclonal) | BioLegend | RRID:AB_961437 | clone: OX-7 |
| antibody | anti-Thy1.2 (rat monoclonal) | BioLegend | RRID:AB_492888 | clone: 30-H12 |
| antibody | anti-Helios (Armenian hamster monoclonal) | BioLegend | RRID:AB_10660749 | clone: 22F6 |
| antibody | anti-ICOS (rat monoclonal) | ebioscience | RRID:AB_2573563 | clone: 7E.17G9 |
| antibody | anti-CD122 (rat monoclonal) | BioLegend | RRID:AB_313226 | clone: 5H4 |
| antibody | anti-CD25 (rat monoclonal) | BioLegend | RRID:AB_312857; RRID:AB_312865 | clone: PC61 |
| antibody | anti-CD69 (Armenian hamster monoclonal) | BioLegend | RRID:AB_2260065 | clone: H1.2F3 |
| antibody | anti-CDF4/80 (rat monoclonal) | BioLegend | RRID:AB_893481 | clone: BM8 |
| antibody | anti-CD11b (rat monoclonal) | BioLegend | RRID:AB_312791; RRID:AB_755986 | clone: M1/70 |
| antibody | anti-CD11c (Armenian hamster monoclonal) | BioLegend | RRID:AB_313777 | clone: N418 |
| antibody | anti-CD19 (rat monoclonal) | BioLegend | RRID:AB_313643 | clone: 6D5 |
| antibody | anti-NKp46 (rat monoclonal) | BioLegend | RRID:AB_2235755 | clone: 29A1.4 |
| antibody | anti-IL-17 (rat monoclonal) | BioLegend | RRID:AB_536018 | clone: TC11-18H10.1 |
| antibody | anti-IFN-γ (rat monoclonal) | BioLegend | RRID:AB_315402 | clone: XMG1.2 |
| antibody | anti-Foxp3 (rat monoclonal) | ebioscience | RRID:AB_1518812 | clone: FJK-16s |
| antibody | anti-phospho ZAP70/SykTyr319/Tyr352 (mouse monoclonal) | ebioscience | RRID:AB_2572664 | clone: n3kobu5 |
| antibody | anti-phospho ERK1/2Thr202/Tyr204 (mouse monoclonal) | BioLegend | RRID:AB_2629710 | clone: 6B8B69 |
| antibody | anti-phospho AKT1Ser473 (mouse monoclonal) | ebioscience | RRID:AB_2573309 | clone: SDRNR |
| antibody | anti-phospho S6Ser235, Ser236 (mouse monoclonal) | ebioscience | RRID:AB_2572666 | clone: cupk43k |
| antibody | anti-phospho STAT5 (mouse monoclonal) | BD Biosciences | RRID:AB_10894188 | clone: 47/Stat5(pY694) |
| antibody | anti-phospho Smad2 (pS465/pS467)/ Smad3 (pS423/pS425) (mouse monoclonal) | BD Biosciences | RRID:AB_2716578 | clone: O72-670 |
| antibody | PerCP/Cy5.5-streptavidin | BioLegend | RRID:AB_2716577 | |
| recombinant DNA reagent | pCR 2.1-TOPO (vector) | Invitrogen | CAT#: K204040 | |
| sequence-based reagent | Foxp3 Intron1 | Eton Bioscience | Forward: ATTTGAATTGGATATGGTTTGT; Reverse: AACCTTAAACCCCTCTAACATC | |
| sequence-based reagent | Foxp3 TSDR | Eton Bioscience | Forward: GTTTGTGTTTTTGAGATTTTAAAAT; Reverse: AACCAACTTCCTACACTATCTATTA | |
| sequence-based reagent | Rara | Eton Bioscience | Forward: CCAGTCAGTGGTTACAGCACA; Reverse: TAGTGGTAGCCGGATGATTTG | |
| sequence-based reagent | Rarb | Eton Bioscience | Forward: ACATGATCTACACTTGCCATCG; Reverse: TGAAGGCTCCTTCTTTTTCTTG | |
| sequence-based reagent | Rarg | Eton Bioscience | Forward: CATTTGAGATGCTGAGCCCTA; Reverse: GCTTATAGACCCGAGGAGGTG | |
| sequence-based reagent | Oligo(dT)12-18 Primer | Invitrogen | CAT#: 18418012 | |
| peptide, recombinant protein | CRIg-Ig | Genentech | PMID: 16530040 | |
| peptide, recombinant protein | Biotinylated CRIg-Ig | this paper | prepared by our laboratory | |
| peptide, recombinant protein | control Ig | Genentech | PMID: 16530040 | |
| peptide, recombinant protein | Biotinylated control Ig | this paper | prepared by our laboratory | |
| peptide, recombinant protein | CTLA-4 Ig | R and D Systems | CAT#: 434-CT-200/CF | |
| peptide, recombinant protein | PD-1 Ig | R and D Systems | CAT#: 1021-PD-100 | |
| peptide, recombinant protein | VISTA Ig | R and D Systems | CAT#: 7005-B7-050 | |
| peptide, recombinant protein | CD226 Ig | R and D Systems | CAT#: 4436-DN-050 | |
| peptide, recombinant protein | TIGIT Ig | R and D Systems | CAT#: 7267-TG-050 | |
| peptide, recombinant protein | BDC2.5 mimotope | AnaSpec | CAT#: AS-63774 | Sequence: RTRPLWVRME |
| peptide, recombinant protein | Recombinant Murine IL-2 | PeproTech | CAT#: 212–12 | |
| peptide, recombinant protein | Recombinant Human TGF-β1 | PeproTech | CAT#: 100–21 | |
| commercial assay or kit | ACK lysing buffer | Lonza | CAT#: 10-548E | |
| commercial assay or kit | Anti-PE microBeads | Miltenyi Biotec | CAT#: 130-048-801 | |
| commercial assay or kit | LIVE/DEAD fixable dead cell stain kits | Invitrogen | CAT#: L34972; CAT#: L34966 | |
| commercial assay or kit | Foxp3/Transcription factorstaining buffer set | ebioscience | CAT#: 00-5523-00 | |
| commercial assay or kit | Phosflow Lyse/Fix buffer | BD Biosciences | CAT#: 558049 | |
| commercial assay or kit | Phosflow Perm buffer III | BD Biosciences | CAT#: 558050 | |
| commercial assay or kit | Percoll | GE Healthcare Life Science | CAT#: 17-0891-01 | |
| commercial assay or kit | NucleoSpin Tissue XS | Macherey-Nagel | CAT#: 740901.50 | |
| commercial assay or kit | EZ DNA Methylaiton Kit | Zymo Research | CAT#: D5001 | |
| commercial assay or kit | HotStarTaq DNA Polymerase | QIAGEN | CAT#: 203203 | |
| commercial assay or kit | TOPO TA Cloning Kit | Invitrogen | CAT#: K204040 | |
| commercial assay or kit | TRIzol Reagent | Invitrogen | CAT#: 15596026 | |
| commercial assay or kit | SuperScript III Reverse Transcriptase | Invitrogen | CAT#: 18080044 | |
| commercial assay or kit | SYBR Green PCR Master Mix | Applied Biosystems | CAT#: 4309155 | |
| chemical compound, drug | Collagenase P | Roche | CAT#: 11249002001 | |
| chemical compound, drug | Collagenase D | Roche | CAT#: 11088882001 | |
| chemical compound, drug | DNase I | Sigma-Aldrich | CAT#: DN25-1G | |
| chemical compound, drug | ATRA | Sigma-Aldrich | CAT#: R2625-50MG | |
| chemical compound, drug | BMS 493 | Sigma-Aldrich | CAT#: B6688-5MG | |
| chemical compound, drug | Vancomycin | Acros Organics | CAT#: 296990010 | |
| chemical compound, drug | Metronidazole | Acros Organics | CAT#: 210340050 | |
| chemical compound, drug | Neomycin | Fisher Scientific | CAT#: BP266925 | |
| chemical compound, drug | Ampicillin | Sigma-Aldrich | CAT#: A0166-25G | |
| chemical compound, drug | PMA | Sigma-Aldrich | CAT#: P1585-1MG | |
| chemical compound, drug | Ionomycin | Sigma-Aldrich | CAT#: I0634-1MG | |
| chemical compound, drug | Brefeldin A solution | BioLegend | CAT#: 420601 | |
| chemical compound, drug | Fisher Healthcare Tissue-Plus O.C.T Compound | Fisher Scientific | CAT#: 23-730-571 | |
| chemical compound, drug | Avidin, HRP conjugate | Invitrogen | CAT#: 434423 | |
| chemical compound, drug | 1-Step Ultra TMB-ELISA Substrate Solution | Thermo Scientific | CAT#: 34028 | |
| chemical compound, drug | Stop Solution for TMB Substrates | Thermo Scientific | CAT#: N600 | |
| chemical compound, drug | DAPI | Invitrogen | CAT#: D1306 | |
| software, algorithm | FlowJo | FlowJo, LLC | RRID:SCR_008520 | |
| software, algorithm | ImageJ | NIH | RRID:SCR_003070 | |
| software, algorithm | BISMA software | other | RRID:SCR_000688 | public website, BDPC DNA methylation analysis platform |
Additional files
-
Transparent reporting form
- https://doi.org/10.7554/eLife.29540.024





