Stress-induced Cdk5 activity enhances cytoprotective basal autophagy in Drosophila melanogaster by phosphorylating acinus at serine437
Figures
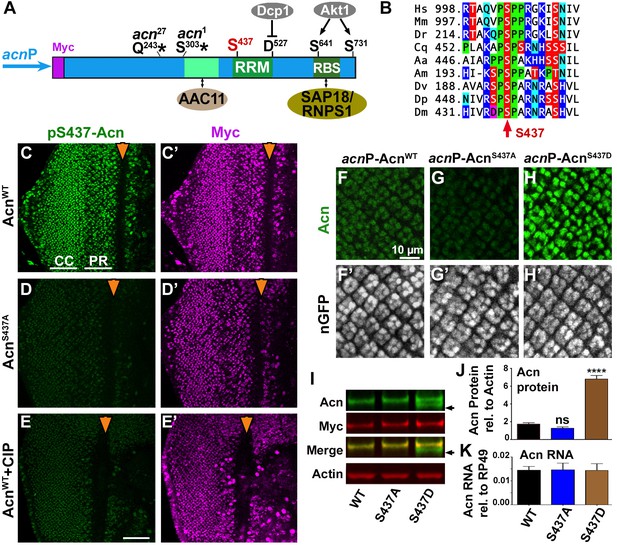
Acn protein is stabilized by phosphorylation at Serine 437.
(A) Cartoon shows regulatory elements of Acn. Genomic Acn transgenes are Myc-tagged and expressed from the endogenous acnP promoter in the background of acn1 and acn27 null alleles. Green boxes indicate the RRM domain and regions that bind to AAC11 (Rigou et al., 2009) or SAP18/RNPS1 (Murachelli et al., 2012). Phosphorylation at the AKT1 target sites S641 and S731 reduces Dcp-1-mediated cleavage at D527 (Nandi et al., 2014). (B) Serine S437 is conserved from humans to insects. Sequences shown: Homo sapiens NP_055792.1; Mus musculus AAF89661.1; Danio rerio AAI16537.1; Culex quinquefasciatus XP_001846490.1; Aedes aegypti XP_001664312.1; Apis mellifera XP_006570961.1; Drosophila virilis XP_015028002.1; Drosophila persimilis XP_002014510.1; Drosophila melanogaster NP_609935.1. (C–E) Projections of confocal micrographs of eye discs from AcnWT (C,E) or AcnS437A (D) larvae stained for pS437-Acn and Myc. Strong staining for pS437-Acn is dominated by cone cells (CC) in posterior regions of eye discs, and by photoreceptors cells (PR) closer to the furrow (arrowhead). Staining for pS437-Acn, but not Myc, is reduced to background level in AcnS437A eye discs (D) or after calf intestinal phosphatase (CIP) treatment of AcnWT (E). (F–H) Micrographs focusing on early photoreceptor clusters in eye disc stained for Acn and DNA. Compared to AcnWT (F), Acn levels are reduced for AcnS437A and increased for the phosphomimetic AcnS437D. (I,J) Western blots of larval lysates show elevated levels of AcnS437D compared to AcnWT or AcnS437A. An Acn cleavage product (arrow) removing the N-terminal Myc tag is stable in AcnS437D, but not AcnWT or AcnS437A. (J) Acn levels relative to the Actin loading control are quantified from three blots from biological repeats. (K) RT-qPCR reveals equal expression for Acn transgenes controlled by the endogenous acnP promoter relative to ribosomal protein RP49. Bar graphs show mean ±SD. ns, not significant; ****p<0.0001; each compared to wild-type Acn control. Scale bar in E is 50 µm in C-E, scale bar in F is 10 µm in F-H. Detailed genotypes are listed in Supplementary file 3.
-
Figure 1—source data 1
Quantification of Acn proteins in Western blots.
- https://doi.org/10.7554/eLife.30760.005
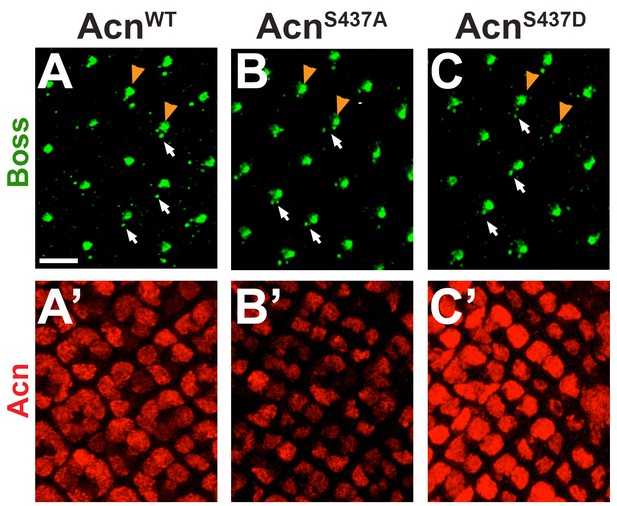
AcnS437A and AcnS437D support normal Sevenless-mediated Boss endocytosis.
Micrographs of AcnWT (A), AcnS437A (B), or AcnS437D (C) larval eye discs stained for Boss and Acn. In eye discs that completely lack acn function, the Boss ligand cannot be detected in R7 cells (Haberman et al., 2010). Similar to other regulatory mutants altering the level of Acn protein (Nandi et al., 2014), AcnS437A and AcnS437D mutants displayed no change in the uptake of the Boss ligand from the R8 surface (yellow arrowheads) into the stereotypical Boss-positive endosomes within R7 cells (arrows). Scale bar: 5 µm.
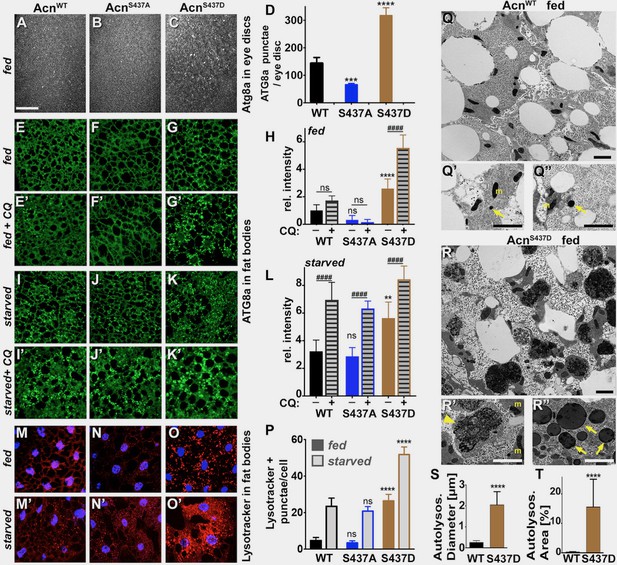
Stabilized AcnS437D enhances basal autophagy.
(A–C) Micrographs of fed AcnWT, AcnS437A or AcnS437D larval eye discs stained for Atg8a. Scale bar: 40 µm in A-O. (D) Quantification of Atg8a punctae in eye discs from five larvae. (E–G, I–K, M–O) Micrographs of AcnWT, AcnS437A or AcnS437D larval fat bodies encompassing 6 to 8 cells from 96 hr fed or starved size-matched larvae (as indicated) stained for ATG8a (E–G, I–K) or with Lysotracker (M–O). For panels E’-G’ and I’-K’ lysosomal degradation was inhibited with chloroquine to visualize autophagic flux. (H,L,P) Quantification of Atg8a intensity (H,L) or Lysotracker punctae (P) in fat bodies averaged from six to eight cells from four to five larvae from one representative experiment out of three repeats. (Q,R) TEMs of fed AcnWT or AcnS437D fat bodies. Smaller panels show higher magnification examples of dense lysosomes (arrowheads), membrane-enriched autolysosomes (arrows) and mitochondria (m). Scale bars are 2 µm. (S) Quantification of diameters of at least 100 lysosomes and autolysosomes per genotype. (T) Quantification of percentage of autolysosomal area averaged from 25 images per genotype. Bar graphs show mean ± SD. **p<0.01; ****p<0.0001; ns, not significant; each compared to corresponding fed or starved wild-type Acn control. For each genotype, starved and fed were significantly different (p<0.01). Significant differences between untreated and chloroquine-treated larvae are indicated (##p<0.01; ###p<0.001, ####p<0.0001). Detailed genotypes are listed in Supplementary file 3.
-
Figure 2—source data 1
Relates to Figure 2D.
Quantification of Atg8a punctae in eye discs.
- https://doi.org/10.7554/eLife.30760.008
-
Figure 2—source data 2
Relates to Figure 2H and L.
Quantification of integrated intensity of Atg8a punctae in fat bodies from fed and starved larvae with or without Chloroquine treatment.
- https://doi.org/10.7554/eLife.30760.009
-
Figure 2—source data 3
Relates to Figure 2P.
Quantification of lysotracker punctae in fat bodies from fed and starved larvae.
- https://doi.org/10.7554/eLife.30760.010
-
Figure 2—source data 4
Quantification of autolysosome and lysosome diameters and relative area.
- https://doi.org/10.7554/eLife.30760.011
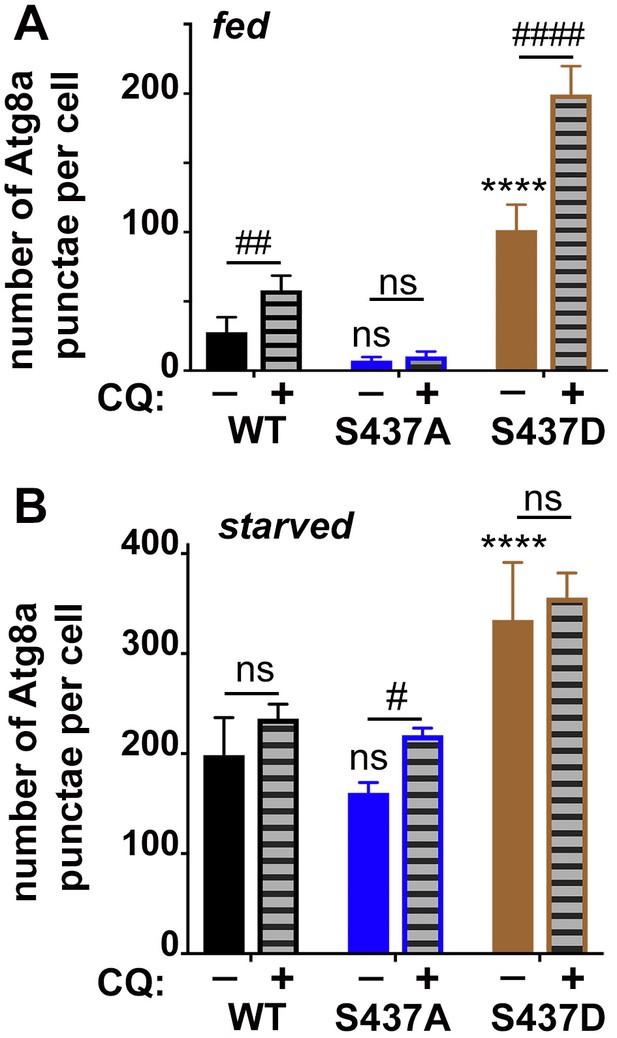
Quantification of Atg8a punctae in fat bodies.
Quantification of Atg8a punctae in fed (A) or starved (B) fat bodies averaged from six to eight cells from four to five larvae from one representative experiment out of three repeats.

Stabilized AcnS437D enhances life span.
(A) Starvation-induced mortality of flies expressing the indicated Acn proteins. In parenthesis shown are numbers of initial flies in a single representative experiment out of three. (B) Survival curves of flies expressing the indicated Acn proteins. In parenthesis shown are numbers of initial flies in a single representative experiment out of three. Log-rank comparisons revealed significant differences between survival curves: *p<0.05; ****p<0.0001.
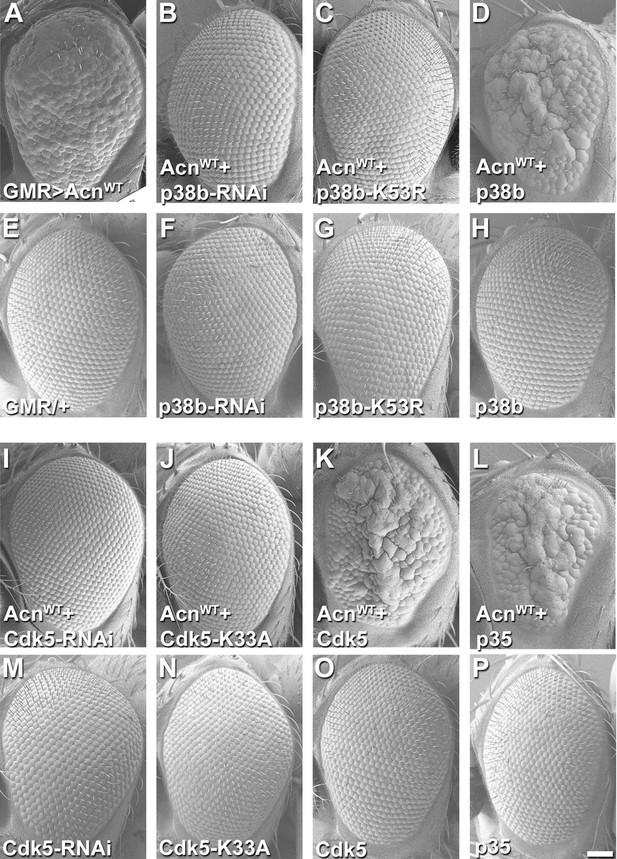

Acn genetically interacts with p38b MAP kinase and Cdk5/p35.
SEM images of eye expressing the indicated transgenes under GMR-Gal4 control at 28°C. (A) Expression of UAS-AcnWT causes a rough eye. This eye-roughness is suppressed by knockdown of p38b MAPK (B) or co-expression of dominant-negative p38b MAPKK53R (C). By contrast, co-expression of wild-type p38b MAPK enhances the roughness (D). Expression of Gal4 (E) or the indicated p38b MAPK transgenes in the absence of UAS-AcnWT (F–H) causes little or no changes in eye morphology. Eye-roughness induced by UAS-AcnWT is suppressed by knockdown of Cdk5 (I) or co-expression of dominant-negative Cdk5K33A (J). By contrast, co-expression of wild-type Cdk5 (K) or the required cofactor p35 (L) enhance AcnWT-induced roughness. Expression of the indicated Cdk5 and p35 transgene in the absence of UAS-AcnWT (M–P) causes little or no changes in eye morphology. Scale bar in A-P: 50 µm. Quantification of genetic interactions is shown in Supplementary files 1 and 2. Detailed genotypes are listed in Supplementary file 3.
-
Figure 4—source data 1
Relates to Figure 4—figure supplement 1.
Quantification of efficiency of RNAi-mediated Kinase knockdowns.
- https://doi.org/10.7554/eLife.30760.016
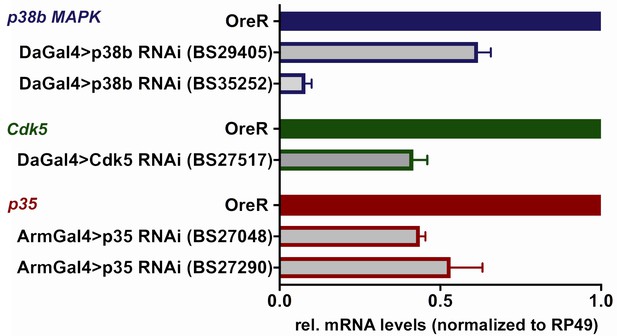
Knockdown efficiency of p38b MAPK, Cdk5 and p35 RNAi UAS-transgenes.
Indicated UAS-RNAi transgenes were expressed using Da-Gal4 or Arm-Gal4 drivers in adult flies. Transcript levels for the indicated genes were determined using RT-qPCR relative to the ribosomal protein RP49 and normalized to wild-type OreR flies. Bar graphs show mean ±SD from three biological replicas.
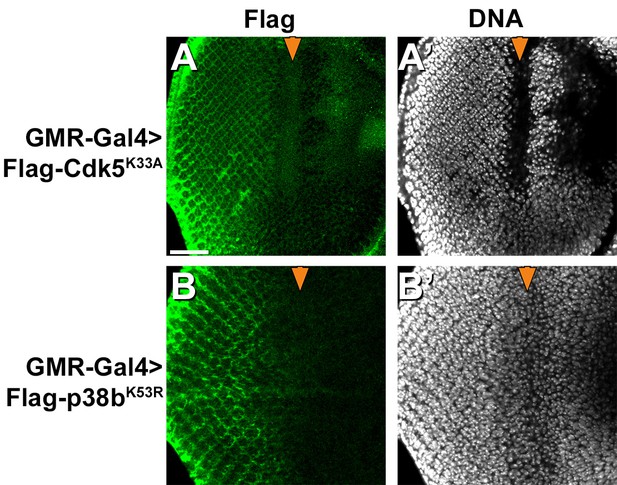
Expression of dominant-negative mutant Cdk5 and p38b MAP kinases.
FLAG-tagged dominant negative mutant Cdk5 or p38b kinases as used in Figure 4 were expressed in eye discs under control of the GMR-Gal4 driver and detected with anti-FLAG antibodies. Scale bar: 50 µm.
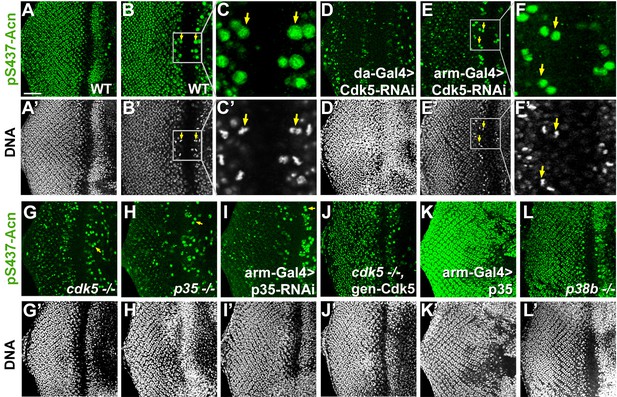
Cdk5/p35-mediates phosphorylation of Acinus.
Projections (A, D, G–L) or individual optical sections (B,C,D,E) of confocal micrographs of eye discs stained for pS437-Acn or DNA from wild-type controls (A–C), or from larvae with knockdown for Cdk5 (D–F), mutant for Cdk5 (G) or p35 (H), or with p35 knockdown (I), or Cdk5 mutant larvae rescued with a genomic Cdk5 transgene (J), from larvae overexpressing p35 (K) or mutant for p38b MAP kinase (L). Arrows indicate examples of mitotic cells with high pS437-Acn levels. These are best seen in individual optical sections of wild-type (B,C) or Cdk5-RNAi eye discs (E,F). Scale bar is 50 µm in A, D,E, G-L; 32 µm in B; 10 µm in C; and 15 µm in F.
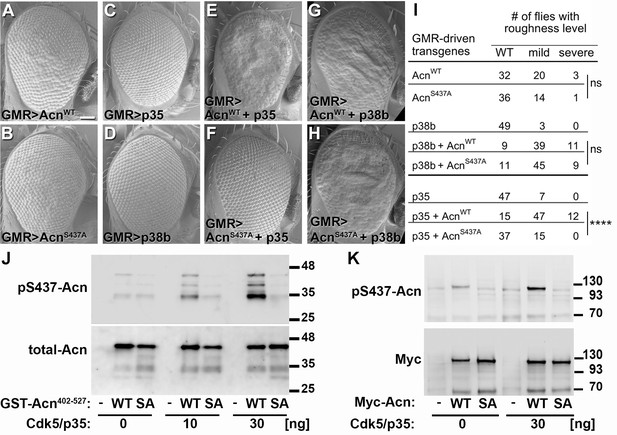
Acn-S437 is the critical target site for Cdk5/p35-mediated phosphorylation.
(A–H) SEM images of eyes expressing the indicated transgenes under UAS/GMR-Gal4 control at 25°. Under these conditions, expression of AcnWT (A) or AcnS437A (B) causes a mildly rough eye, but not expression of p35 (C) or p38b MAP kinase (D). Coexpression of p35 with AcnWT (E) causes a severely rough eye, but not p35 coexpression with AcnS437A (F). By contrast, p38b MAP kinase coexpression enhances roughness of both, AcnWT (G) and AcnS437A (H). Quantification of these genetic interactions is shown in panel I. Statistical significance was calculated by Chi square test (****p<0.0001; ns, not significant). (J–K) Western blots of purified wild-type (WT) or S437A mutant (SA) GST-Acn402-527 fusion proteins (J) or full-length Streptag-Myc-Acn proteins (K) that were incubated with the indicated amount of Cdk5/p35 kinase complex. Blots were developed with antibodies against pS437-Acn (1:2000), Myc (1:2000), or total Acn (1:2000) as described (Nandi et al., 2014). Note that the Acn antibody (Haberman et al., 2010) preferentially recognizes a C-terminal Acn epitope that is deleted in some of the partially degraded GST-fusion proteins, which however still contain the Cdk5 target site at S437. Detailed genotypes are listed in Supplementary file 3.
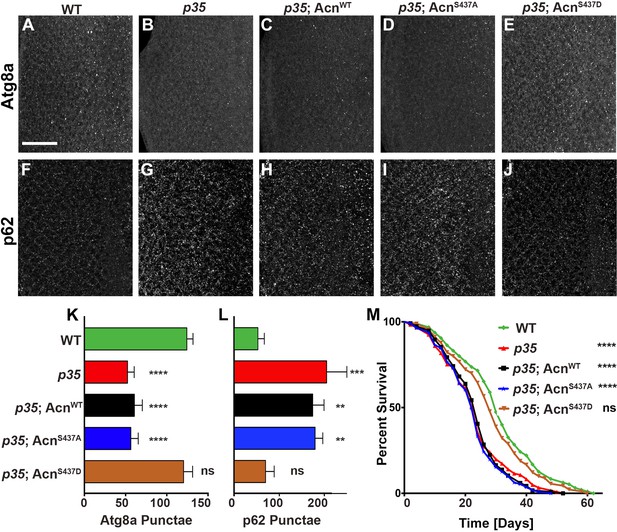
Cdk5/p35-mediated phosphorylation of Acn regulates basal autophagy and longevity.
Projections of confocal micrographs of eye discs stained for Atg8a (A–E) or p62 (F–J) from wild-type controls (A,F), p35 mutant larvae (B,G) or p35 mutant larvae expressing AcnWT (C,H), AcnS437A (D,I) or the phosphomimetic AcnS437D (E,J) under control of the acnP promoter. Scale bar in A is 40 µm in A-J. (K) Quantification of Atg8a punctae from three eye discs each with genotypes as shown in A-E. (L) Quantification of p62 punctae from three eye discs each with genotypes as shown in F-J. Bar graphs show mean ±SD. ns, not significant; **p<0.01; ***p<0.001; ***p<0.0001; each compared to wild-type Acn control. (M) Survival curves of WT controls or p35 mutants, or p35 mutants expressing the indicated Acn proteins. The initial numbers of flies were 95 (WT), 121 (p35), 116 (p35, AcnWT), 111 (p35, AcnS437A), 90 (p35, AcnS437D). Log rank tests compared to WT control: ns, not significant; ****p<0.0001. Detailed genotypes are listed in Supplementary file 3.
-
Figure 7—source data 1
Quantification of Atg8a and p62 punctae in eye discs.
- https://doi.org/10.7554/eLife.30760.020

Protein-aggregate-induced phosphorylation of Acn S437 depends on Cdk5/p35.
Projections of confocal micrographs of eye discs stained for pS437-Acn (A–E) or polyQ proteins (G–K) and DNA. Compared to a GMR-Gal4 control (A), GMR-Gal4-driven UAS-Htt.Q93 (B) or GMR-Htt.Q120 (D) expression induced elevated S437 phosphorylation, which was suppressed in eye discs with p35 knockdown (C) or mutant for p35 (E). GMR-directed expression initiates at the furrow (arrowhead) and is more developed toward the posterior. (F) Quantification of S437 phosphorylation (F) averaged constant areas containing about 50 ommatidial clusters located at least 6–8 rows posterior to the furrow. Bar graphs show mean ±SD of integrated densities. Values were normalized to GMR-Gal4 controls and were from one representative experiment out of three repeats. ns, not significant; **p<0.01; ***p<0.001; ***p<0.0001; for indicated comparisons. Early polyQ accumulation was compared between GMR-Gal4 eye discs as controls (G), and eye discs expressing GMR-Htt.Q120 and carrying a copy of the indicated genomic Acn transgene (H–J) or the p35 null allele (K). Compared to AcnWT (H) and AcnS437A (I) eye discs, in AcnS437D (J) polyQ accumulation was reduced until some 7 to 8 rows of ommatidia posterior to the furrow (arrowhead) and enhanced in p35 mutants (H). Scale bar in A: 40 µm in A-N. Posterior is to the left. (L) Quantification of PolyQ accumulation averaged constant areas containing about 100 ommatidial clusters located at least 2–3 rows posterior to the furrow. Bar graphs show mean ± SD of integrated densities. Values were normalized to AcnWT/GMR-Htt.Q120 (L) and were from one representative experiment out of three repeats. ns, not significant; **p<0.01; ***p<0.001; ***p<0.0001; compared to AcnWT/GMR-Htt.Q120. (M) Quantification of dot blots measuring aggregated polyQ protein in fly heads expressing GMR-Htt.Q120 and the indicated genomic Acn transgenes or in a p35 mutant background. Control OreR flies (+/+) indicate background level of dot blot measurements (n = 3 biological repeats). The scatter plot shows mean and standard deviation from three separate experiments. Detailed genotypes are listed in Supplementary file 3.
-
Figure 8—source data 1
Quantification of pS437 and polyQ-levels in eye discs.
- https://doi.org/10.7554/eLife.30760.022
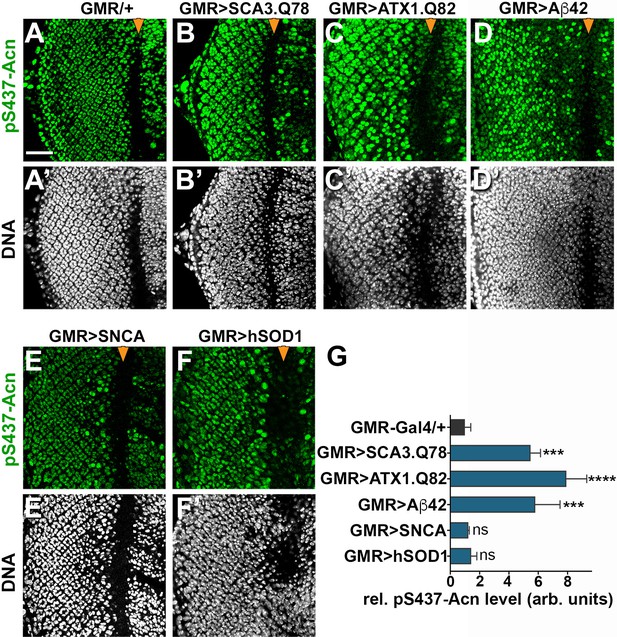
Increased Acn-S437 phosphorylation is specific for a subset of neurodegeneration models.
Projections of confocal micrographs of eye discs stained for pS437-Acn (A–F) or DNA (A’–F’). Compared to GMR-Gal4 controls (A), eye discs with GMR-Gal4-driven UAS-SCA3.Q78 (B) or UAS-ATX1.Q82 (C) or UAS-Aβ42 (D) display elevated levels of pS437-Acn staining in a subset of cells, in contrast to UAS-alpha-Synuclein (E) or UAS-hSOD1 (F) which appear unaltered compared to controls. GMR-directed expression of transgenes initiates at the furrow (arrowhead) and extends toward the posterior (left). Scale bar in A is 40 µm in A-N. Detailed genotypes are listed in Supplementary file 3. (G) Quantifications of S437 phosphorylation averaged constant areas containing about 50 ommatidial clusters located at least 6–8 rows posterior to the furrow. Bar graphs show mean ±SD of integrated densities of pS437 staining normalized to GMR-Gal4 controls. ns, not significant; **p<0.01; ***p<0.001; ***p<0.0001; compared to GMR-Gal4/+.
-
Figure 9—source data 1
Quantification of pS437-Acn in neurodegenerative model eye discs.
- https://doi.org/10.7554/eLife.30760.024
Additional files
-
Supplementary file 1
Suppression of Acn-induced eye roughness by kinase knock down.
The effect on eye morphology of GMR-Gal4-driven expression of UAS-RNAi transgenes with or without UAS-Acinus was scored and recorded as the number of flies with normal eyes (score = 1), eye with mild errors (score = 2), rough eyes (score = 3) or severely rough eyes (score = 4). The percentage suppression or enhancement was calculated from the average score for each RNAi transgene compared to UAS-Acn in the absence of RNAi transgenes. Positive or negative numbers indicate suppression and enhancement, respectively. Green or red colors highlight UAS-transgenes with more than 50% suppression or enhancement. Numbers in parenthesis indicated stock numbers of the Bloomington Drosophila stock center or, if starting with a ‘V..', from the Vienna Drosophila Resource Center. All flies were raised at 28°C.
- https://doi.org/10.7554/eLife.30760.025
-
Supplementary file 2
Genetic interactions of Acn with p38b and Cdk5/p35 loss and gain of function.
The effect on eye morphology of GMR-Gal4-driven expression of UAS-transgenes with or without UAS-Acinus was scored and recorded as the number of flies with normal eyes (score = 1), eye with mild errors (score = 2), rough eyes (score = 3) or severely rough eyes (score = 4). The percentage suppression or enhancement was calculated from the average score for each RNAi transgene compared to UAS-Acn in the absence of RNAi transgenes. Positive or negative numbers indicate suppression and enhancement, respectively. Green or red colors highlight UAS-transgenes with more than 50% suppression or enhancement. Numbers in parenthesis indicated stock numbers of the Bloomington Drosophila stock center. All flies were raised at 28°C.
- https://doi.org/10.7554/eLife.30760.026
-
Supplementary file 3
Genotypes of flies used for each figure.
Relevant genotypes are listed for all Figures.
- https://doi.org/10.7554/eLife.30760.027
-
Supplementary file 4
Key resources table.
- https://doi.org/10.7554/eLife.30760.028
-
Transparent reporting form
- https://doi.org/10.7554/eLife.30760.029





