Conserved RNA-binding specificity of polycomb repressive complex 2 is achieved by dispersed amino acid patches in EZH2
Figures
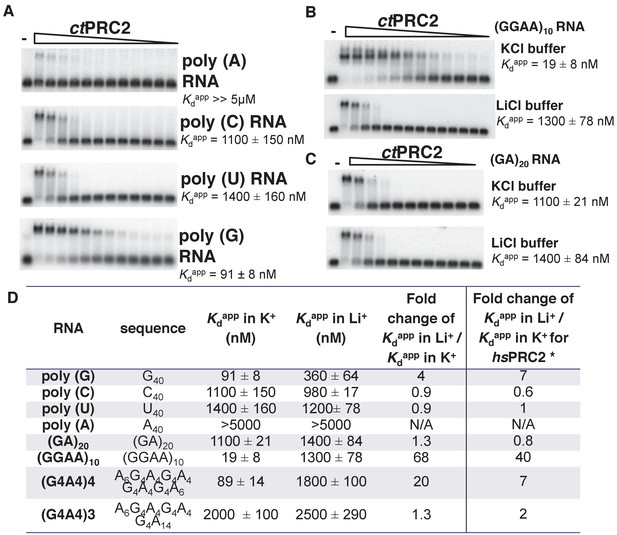
ctPRC2 prefers binding to G-rich RNAs and G-quadruplexes, and shares RNA-binding specificity with hsPRC2.
(A) Binding of ctPRC2 to four 40-mer homopolymeric RNAs was tested using EMSA with standard 100 mM KCl-binding buffer. (B) Binding of ctPRC2 to a G-quadruplex-forming (GGAA)10 RNA was tested in 100 mM KCl and 100 mM LiCl-binding buffers. (C) Binding of ctPRC2 to a control RNA (GA)20 was tested in 100 mM KCl and 100 mM LiCl-binding buffers. (D) Comprehensive analysis of binding affinities of ctPRC2 to a variety of 40-nt RNAs. * Results of this column were from Wang et al. (2017a). For (A), (B) and (C), ctPRC2 was used at successive threefold dilutions starting at 5 µM concentration or 4 µM for (GGAA)10 RNA in KCl buffer. Kdapp values and errors are mean and standard derivation of at least three binding experiments performed on different days and often by a different person.
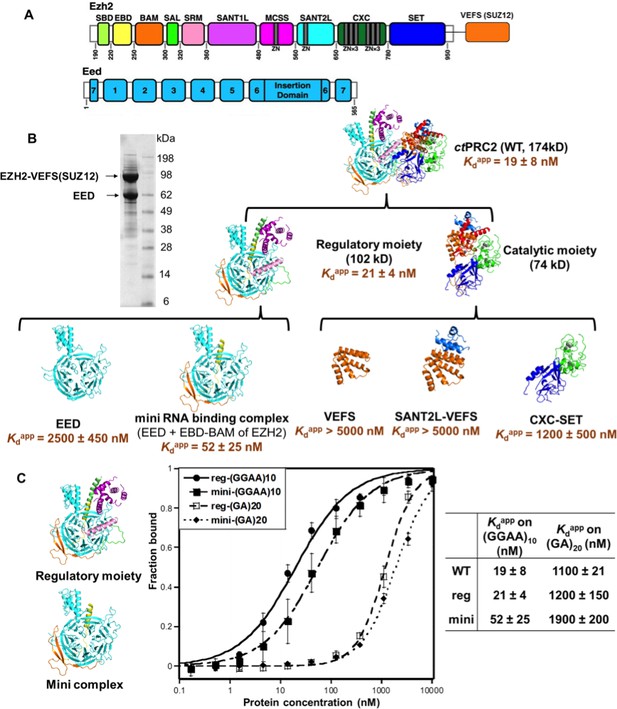
The minimal RNA-binding complex of ctPRC2 includes EED and EBD-BAM domains of EZH2, and additional affinity might be provided by the CXC-SET domains.
(A) Purified ctPRC2 comprises two polypeptides: a 107 kDa peptide consisting of EZH2 fused with the VEFS domain of SUZ12, and a 67 kDa EED subunit. Individual domains are color coded as represented in structures of panel (B). (B) Purified ctPRC2 wild type was Coomassie stained on a SDS-PAGE gel (left). Sub-complexes of ctPRC2 were purified and tested for binding with (GGAA)10 RNA (right). Regulatory moiety contains EED and N-terminal half of EZH2 (ending at SANT1L domain), and catalytic moiety contains C-terminal half of EZH2 (starting at MCSS domain) fused with VEFS domain of SUZ12. Kdapp values and errors are mean and standard derivation of at least three binding experiments performed on different days. Minimal RNA-binding complex maintains the specificity of binding to G-quadruplex RNA. Regulatory moiety (reg) and minimal RNA-binding complex (mini; EED + EBD BAM of Ezh2) of ctPRC2. G-quadruplex-forming (GGAA)10 RNA and control (GA)20 RNA were tested for binding with the two complexes.
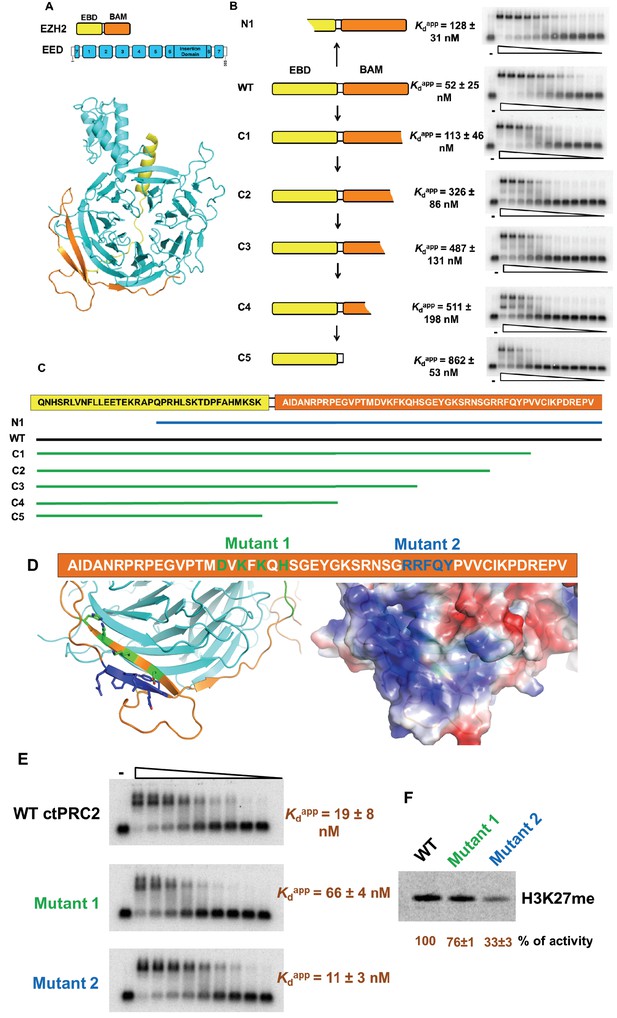
Four residues in BAM domain of ctPRC2 are important for RNA binding.
(A) The minimal RNA-binding complex consists of EED and EBD-BAM domains of EZH2. (B) Mutagenesis study was conducted on the minimal RNA-binding complex, by trimming the EZH2 peptide from the N and C termini. Purified complexes were tested for binding to (GGAA)10 RNA. Threefold titration of proteins was used in each EMSA experiment. Starting concentrations of each complex were: 10 µM for N1, C2, C3 and C4, 5 µM for wild type and C1, and 4 µM for C5. (C) Details of the EZH2 peptide sequence for wild type and the six mutants. Blue line indicates the N-terminal truncation mutant (N1), and green lines refer to the C-terminal truncation mutants (C1-5). (D) Two sets of residues (green and blue) in BAM domain were mutated in the context of the entire ctPRC2. Mutant 1 includes alanine substitution of four residues (shown in green), and mutant 2 includes mutation of five residues (shown in blue) to GGSGG sequence. These residues are also indicated in the structure and the electrostatic map. (E) Wild type and mutant ctPRC2 were tested for binding with (GGAA)10 RNA, and each protein was titrated threefold from 500 nM. (F) Methyltransferase activity on histone H3 was tested for wild type and the two mutants. 14C-labeled SAM and purified H3 were incubated with each protein for 30 min and then resolved on a SDS-PAGE gel. Methyltransferase activity is indicated by the intensity of the radiolabeled H3 band. Activity is normalized to that of wild type, and the mean ±S.D. of three independent experiments is shown.
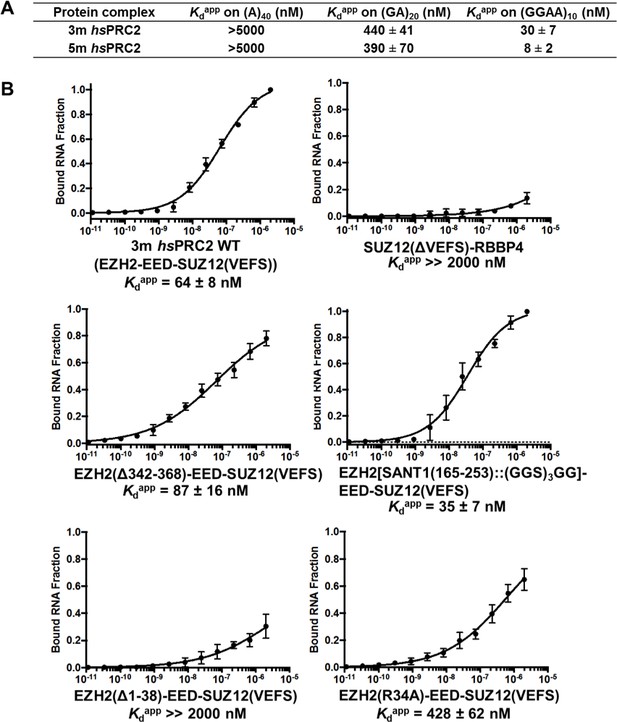
The crystallized 3 m hsPRC2 complex contains the major RNA-binding sites of hsPRC2, and the N-terminal helix of EZH2 plays an important role.
(A) Previously tested hsPRC2 5 m complex and the crystallized 3 m core complex were expressed in insect cells, purified and tested for binding with (GGAA)10, (GA)20 and (A)40 RNA. PRC2 proteins were titrated threefold starting at 5 µM concentration. (B) Mutagenesis analysis of the crystallized 3 m core complex uncovers the role of the N-terminal helix of EZH2 in RNA binding. Wild type and mutants were expressed using an Saccharomyces cerevisiae expression system. Wild type 3 m complex and the remaining part of the 4 m holoenzyme were tested for binding with (GGAA)10 RNA (top panel). Three large deletion mutants and one point mutant (R34A) were tested for RNA binding (middle and bottom panels). Kdapp values and errors are mean and standard derivation of at least three binding experiments performed at different days.
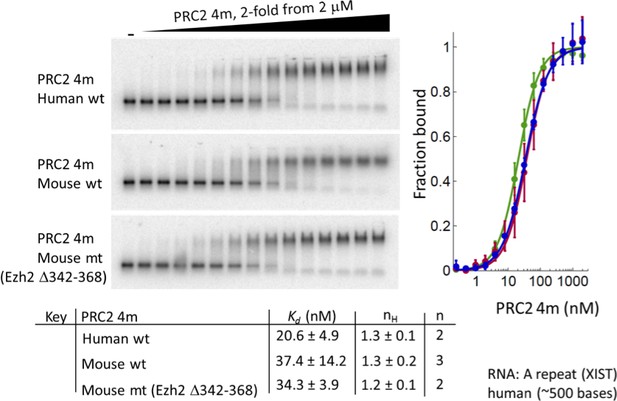
Deletion of residues 342–368 of EZH2 in mouse 4 m PRC2 does not affect binding to a RepA RNA.
Purified wild-type human 4 m PRC2, wild-type mouse 4 m PRC2 and mutant mouse 4 m PRC2 (Δ342–368) were tested with an in vitro transcribed human RepA RNA using EMSA assays. Kdapp values and errors are mean and standard derivation of two or three (as indicated) independent experiments.
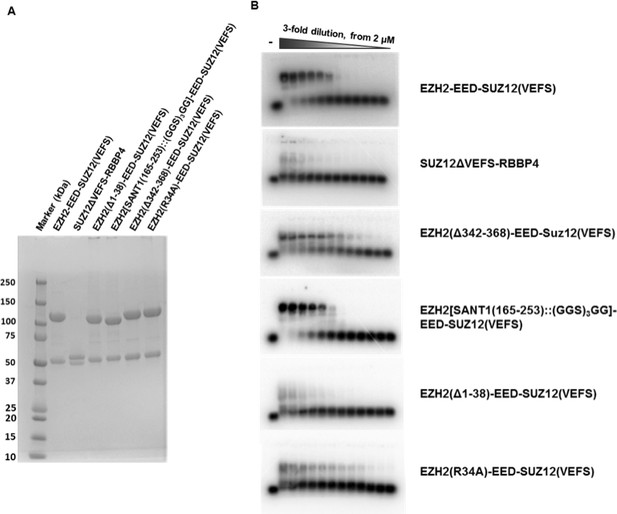
SDS-PAGE gels and EMSA gels of S.
cerevisiae expressed hsPRC2 complexes. S. cerevisiae expressed hsPRC2 complexes shown in Figure 4 are resolved on SDS-PAGE gels to illustrate two polypeptides in each protein complex (hsEZH-VEFS(hsSUZ12) and hsEED; or hsRBBP4 and hsSUZ12(ΔVEFS))(A). The EMSA gels used for calculation of the binding curves are shown in (B).
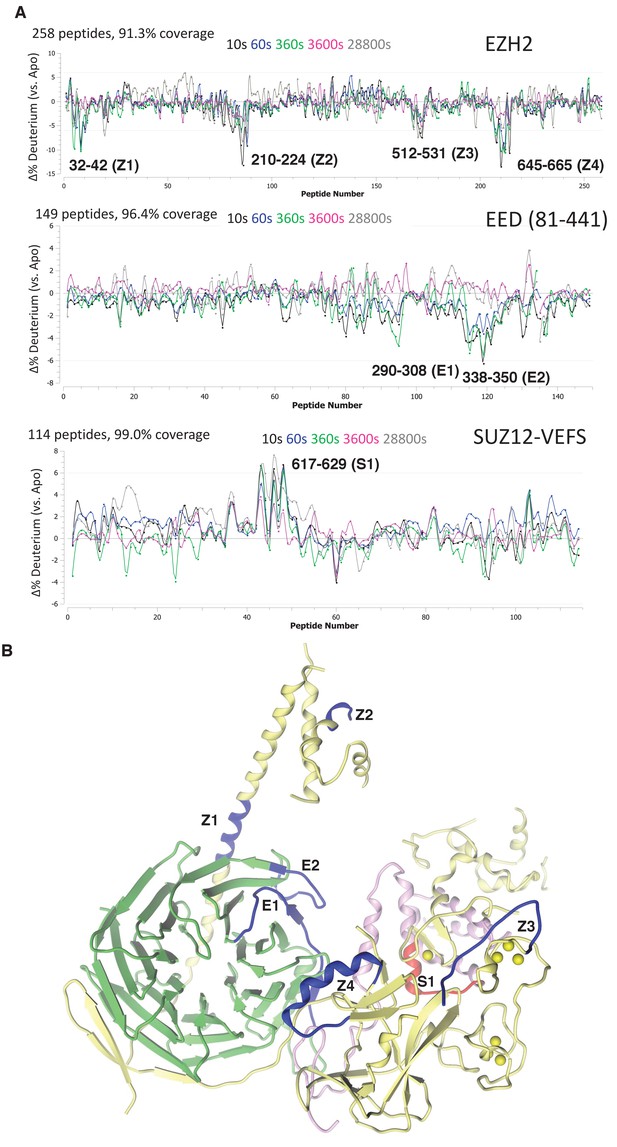
Regions protected by RNA binding identified using HDX-MS.
(A) hsPRC2-(GGAA)10 complex dynamics. Differential hsPRC2 vs hsPRC2-(GGAA)10 (5 µM hsPRC2: 40 µM (GGAA)10) in deuterium uptake plots of EZH2, EED (81-441) and Suz12-VEFS (545-726; S583D). Deuterium incubation times: 10 s, 1 min, 6 min, 1 hr, 8 hr. Peptides with statistically significant changes in deuterium uptake are labeled based on human protein numbering. (B) Mapping of regions with altered deuterium uptake in presence of (GGAA)10 quadruplex onto the ac/hsPRC2 crystal structure.
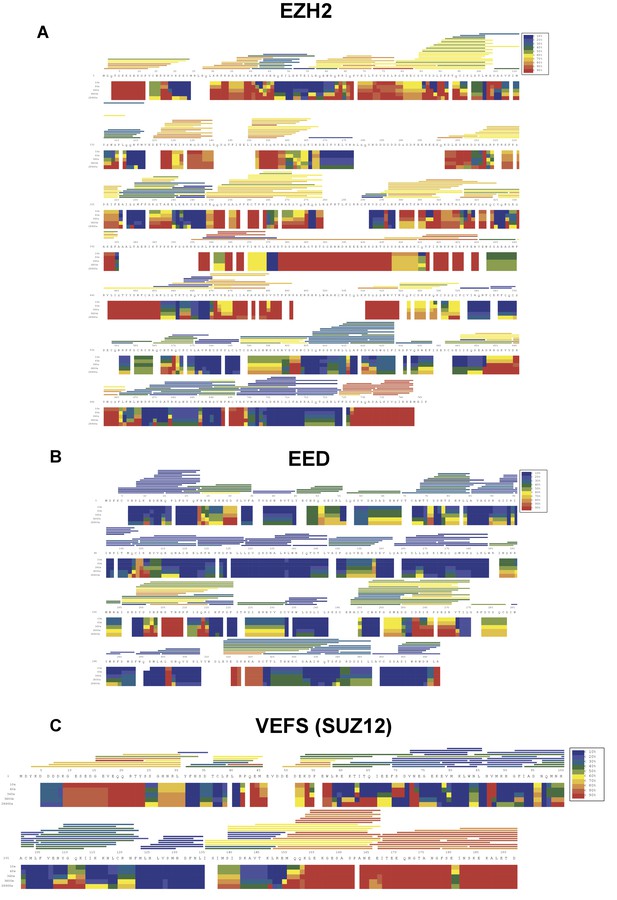
HDX-MS deuterium uptake heat map in the absence of RNA.
HDX-MS heat map represents the solution dynamics of full-length hsEZH2 (A), hsEED (B) and hsVEFS(SUZ12) (C) in the absence of RNA: red regions indicate protein exposed to solvent and readily available for backbone exchange. Blue regions are blocked from exchange because they are: (1) buried in protein (2) involved in hydrogen bonding (secondary structure) or (3) blocked by protein–protein interactions. The horizontal bars at the top of the heat map represent average deuteration for individual peptides.
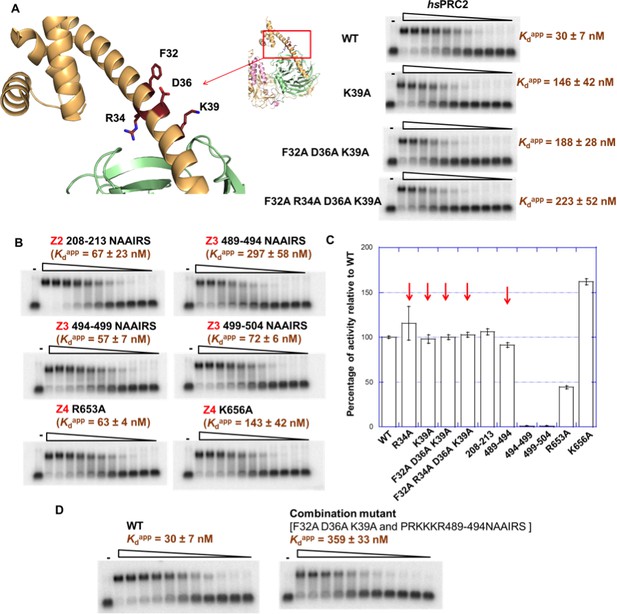
Mutagenesis results support RNA-binding roles of regions Z1, Z3 and Z4 in HDX data, and key residues are identified.
(A) Identification of key residues in Z1. Positions of F32, R34, D36 and K39 in N-terminal helix of EZH2 are indicated in the ac/hsPRC2 crystal structure. K39A point mutant, triple mutant [F32A D36A K39A] and quadruple mutant [F32A R34A D36A K39A] were purified and tested for binding with (GGAA)10 RNA, together with the wild type 3 m hsPRC2. All proteins were used at successive threefold dilutions starting at 2.5 µM concentration. (B) Mutagenesis in Z2, Z3 and Z4 leads to identification of important residues in Z3 and Z4. All proteins were used at successive threefold dilutions starting at 5 µM concentration, and Kdapp values and errors are mean and standard derivation of three binding experiments performed at different days. (C) Histone methyltransferase activity assays for RNA-binding mutants, normalized to that of wild type and error calculated from three independent experiments. Red arrows indicate the mutants that disrupt RNA binding while maintaining normal catalytic activity. (D) Combining identified mutations into a single mutant [F32A D36A K39A and PRKKKR489-494NAAIRS] causes more severe binding defect for (GGAA)10 RNA. Both proteins were used at successive threefold dilutions starting at 5 µM concentration, and Kdapp values and errors are mean and standard derivation of three independent experiments.
-
Figure 6—source data 1
Raw data in fRIP experiment.
- https://doi.org/10.7554/eLife.31558.014
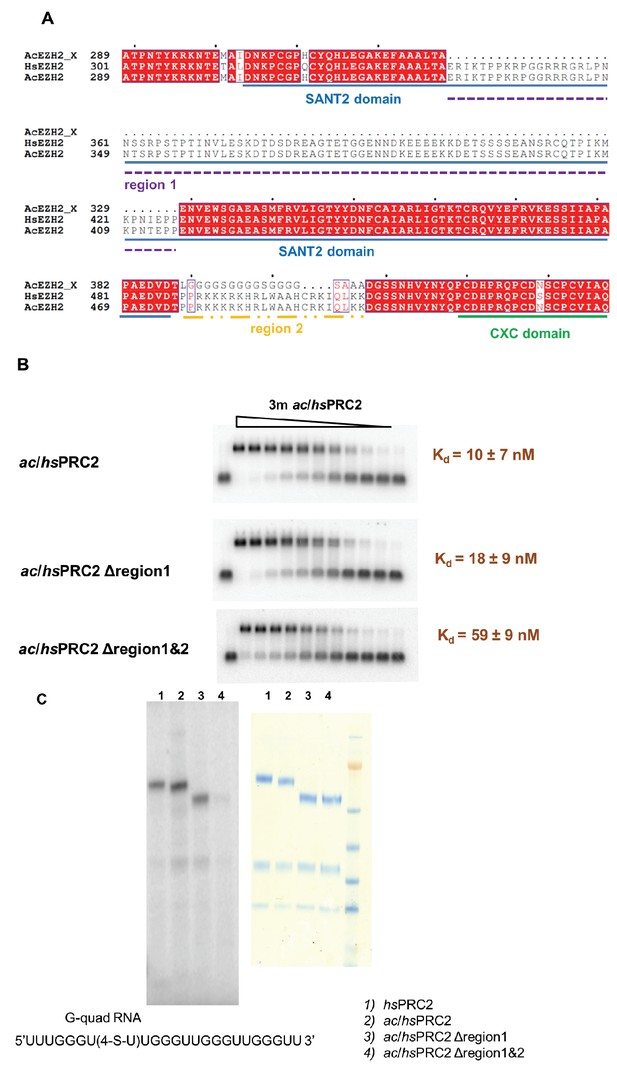
A basic region (adjacent to and include Z3 in HDX-MS result) of EZH2 is important for RNA binding.
(A) Two previously deleted regions (region 1 and 2) in the ac/hsPRC2 crystal structure are indicated with dashed underlines. (Brooun et al., 2016). (B) ac/hsPRC2 wild type and two mutants (Δregion 1 and Δregion1 and 2 < crystallization construct>) were tested with (GGAA)10 RNA. Kdapp values and errors are mean and standard derivation of three independent experiments. (C) Protein-RNA crosslinking experiment using a 5’-32P-labeled G-quadruplex RNA (UUUGGGU(4-thio-U)UGGGUUGGGUUGGGUU). [Left panel] Protein samples and radiolabeled RNA were crosslinked using 365 nM UV light, resolved using SDS-PAGE and exposed to a phosphorimaging plate. Radiolabeled bands indicate that RNA is crosslinked to these polypeptides. [Right panel] Coomassie staining of the four protein samples on a SDS-PAGE gel.
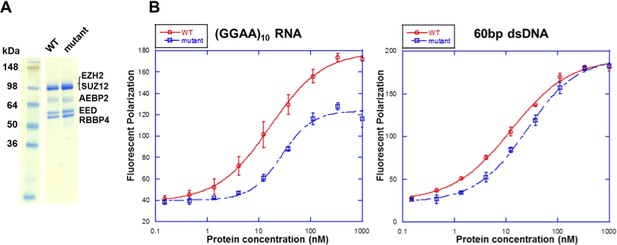
Mutations in the context of the 5 m hsPRC2 (holoenzyme) exhibit binding defect specifically for RNA relative to DNA.
(A) Wild type and mutant [F32A R34A D36A K39A PRKKKR489-494NAAIRS] EZH2 were expressed and purified in the context of the 5 m hsPRC2 (EZH2-SUZ12-EED-RBBP4-AEBP2). The recombinant 5 m hsPRC2 complexes reconstituted with the baculovirus system were resolved using a Novex 10–20% Tris-Glycine gel (WedgeWell format, ThermoFisher) and stained with Coomassie Blue for visualization. Individual polypeptides are marked. (B) Wild type and mutant 5 m hsPRC2 were tested for binding with a (GGAA)10 RNA and a 60 bp double-stranded DNA (produced by annealing two single-stranded (GC)30 DNA oligos) using fluorescent anisotropy. 5 nM of the Alexa488 fluorophore-attached RNA or DNA were folded and incubated as described for the EMSA experiments. RNA binding was measured at high-salt condition (100 mM KCl and 2.5 mM MgCl2) as used in the EMSA experiments, and DNA binding was measure in low-salt condition (10 mM KCl and 0.25 mM MgCl2) to enhance the binding. All proteins were used at successive 3-fold dilutions starting at 1 µM concentration, and all polarization values and errors are mean and standard derivation of three binding experiments performed on different days.
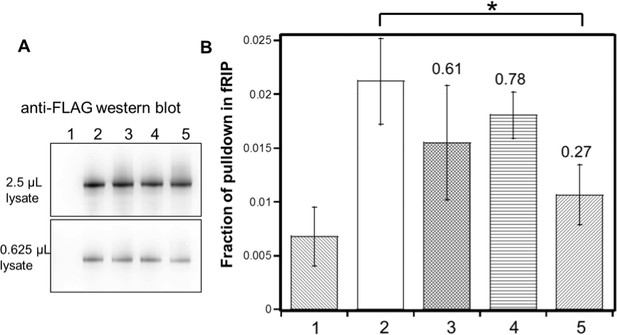
Identified residues are important for (GGAA)10 RNA interaction in vivo.
FLAG-tagged hsEZH2 (wild type or mutants) was co-expressed with (GGAA)10-MS2 RNA in HEK293T cells, and formaldehyde RNA immunoprecipitation (fRIP) was performed to evaluate the percentage of pull down of the RNA by each hsEZH2 variant. hsEZH2 variant used in each lane is: 1. no EZH2 (negative control); 2. wild type; 3. [K39A]; 4. [F32A R34A D36A K39A]; 5. [F32A R34A D36A K39A and PRKKKR489-494NAAIRS]. (A) Western Blot using anti-FLAG antibody was performed to compare the expression levels of exogenous hsEZH2 wild type and mutants. Two different amounts (2.5 or 0.625 µL) of lysate were used to better evaluate the expression levels. (B) Fraction of pulldown of (GGAA)10 RNA by each hsEZH2 variant is plotted in the bar graph, and the mean ± S.D. of three independent experiments is shown. *p-value<0.05. The number above each of the three mutants is the fold change of RNA pulldown relative to wild type with background reduction by subtracting the negative control value.
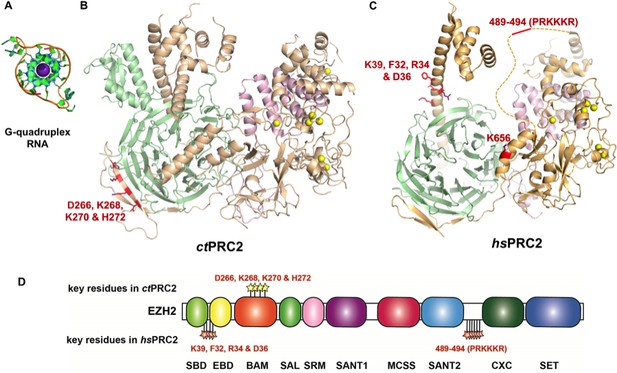
Summary of residues on EZH2 of ctPRC2 and hsPRC2 that are important for binding G-quadruplex RNA.
(A) Crystal structure of a telomeric RNA quadruplex (PDB: 3IBK, sequence: UAGGGUUAGGGU) allows comparison of its size with that of PRC2 structures. Blue spheres in the center are potassium ions. (B and C) Identified key residues for RNA binding are shown in red. Subunits of PRC2 are colored differently. (C) Locations of the key residues for RNA binding are annotated along the polypeptide chain. Key residues in ctPRC2 are listed above the chain, while those in hsPRC2 are below the chain. Individual domains are color-coded as in Figure 2A. K656 mutation increases catalytic activity and is therefore omitted from this annotation map.
Additional files
-
Supplementary file 1
List of all proteins used in this study
- https://doi.org/10.7554/eLife.31558.016
-
Supplementary file 2
Mutations in ctPRC2 that had <3 fold effect on affinity for (G4A4)4 RNA
- https://doi.org/10.7554/eLife.31558.017
-
Transparent reporting form
- https://doi.org/10.7554/eLife.31558.018





