The HIV-1 Tat protein recruits a ubiquitin ligase to reorganize the 7SK snRNP for transcriptional activation
Figures
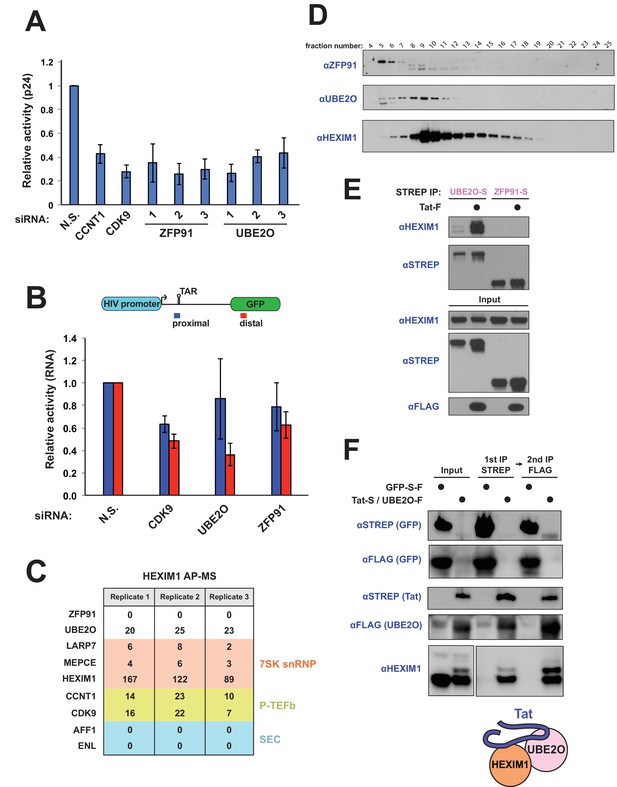
UBE2O is a critical regulator of Tat-dependent transcription and binds HEXIM1 of the 7SK snRNP.
(A) UBE2O and ZFP91 RNAi knockdown inhibit Tat-dependent transcription. Normalized, relative Tat activities ([p24host RNAi/FFLhost RNAi] / [p24N.S. RNAi/FFLN.S. RNAi]) for CCNT1, CDK9, UBE2O, and ZFP91 are shown, and data are represented as the mean ± SEM of at least three biological replicates (defined as independent transfections. Cells were transfected independently on at least 2 different days) for independent siRNA transfections in the HeLaprovirusΔtat cell line. (B) HIV RNA elongation assay with ZFP91 (si2), UBE2O (si1), or CDK9 RNAi. Knockdowns were performed in the same cells as (A), which contain GFP cloned into the Nef locus. qPCR was used to monitor the Tat-dependent production of proximal (blue) and distal (red) RNA transcripts, with the relative positions of the primer sets shown above. Data are presented as the mean ± SEM of relative Tat activity of at least three biological replicates (same definition as above). (C) AP-MS results using HEXIM1-FLAG as the bait protein. Peptide counts are shown for selected prey proteins from triplicate biological purifications (as defined above). See also Supplementary file 1 for a complete list of interacting proteins. (D) Gel filtration of HEK 293T whole cell lysate. Fractions were collected after the void volume and Western blotted for the indicated proteins. ZFP91 is shown as a specificity control. (E) STREP purification of transfected proteins ±Tat co-transfection. The endogenous HEXIM1 interaction was monitored by western blot using an anti-HEXIM1 antibody. (F) Tandem affinity purification of Tat-STREP and UBE2O-FLAG from transfected cells. GFP-S-F (which is STREP and FLAG tagged) was used as a negative control in the experiment. The endogenous HEXIM1 interaction was detected by antibody. Different exposure times from the same blot for the HEXIM1 input and elutions are shown for ease of comparison. Schematic illustrates the Tat-HEXIM1-UBE2O complex.
-
Figure 1—source data 1
Source data for luciferase-normalized p24 values of siRNA knockdown coupled to Tat activity of Figure 1A.
- https://doi.org/10.7554/eLife.31879.004
-
Figure 1—source data 2
Source data for proximal and distal Tat-dependent RNA species of Figure 1B.
- https://doi.org/10.7554/eLife.31879.005
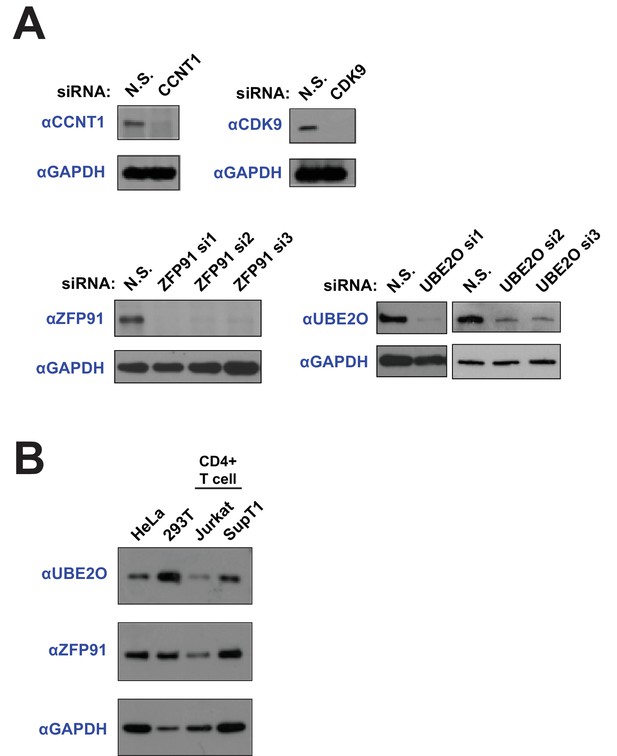
RNAi-mediated protein knockdown and endogenous expression.
(A) Western blots demonstrate specific protein knockdown for the indicated siRNAs used in Figure 1A. (B) Protein expression comparison across multiple cell lines. 100,000 cells of HeLa, HEK 293T, Jurkat (T cell line), or SupT1 (T cell line) were collected and lysed in SDS loading buffer. Western blots of whole cell lysate with specific antibodies demonstrate endogenous protein expression.
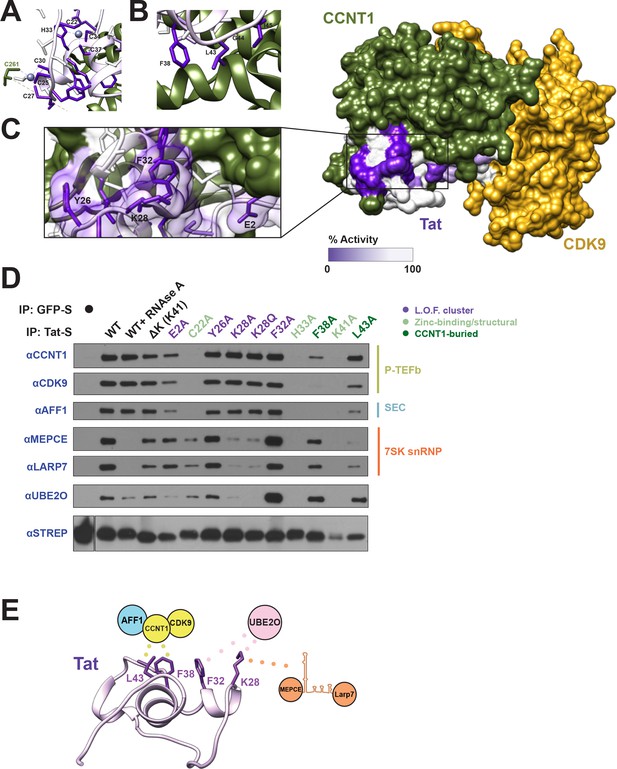
Tat uses a novel interaction surface to interact with the 7SK snRNP and UBE2O.
(A) Tat transcriptional activity data collected from our systematic set of alanine point mutants (D'Orso et al., 2012; Fernandes et al., 2016) was mapped onto the crystal structure of the Tat-P-TEFb complex (PDB i.d. 3MI9). The most severe loss-of-function (L.O.F.) mutations are shaded in deep purple while those with no effect on activity are shaded in white. The cysteine-rich zinc-binding clusters (A) and a hydrophobic stretch buried in CCNT1 (B) are critically important for Tat function. (C) A surface-exposed patch of severe loss-of-function mutants (Glu2, Tyr26, Lys28, and Phe32) does not contact CCNT1 or CDK9. (D) IP of transiently transfected wild type or mutant STREP-tagged Tat proteins. GFP-S was used as a negative control. Western blots were performed on STREP elutions using antibodies to the endogenous proteins indicated. Important residues are color coded as indicated. L.O.F. cluster is the four surface-exposed mutants from (C). See also Figure 2—figure supplement 1 for inputs. (E) Model of Tat interaction surfaces. Tat uses a hydrophobic patch including Leu43 and Phe38 buried at the CCNT1 interface, which also results in the binding of CDK9 and AFF1. Mutation of Lys28 reduces the interactions with both the 7SK snRNP proteins and UBE2O, whereas mutation of Phe32 dramatically increases the UBE2O interaction, associating the ligase with this residue.
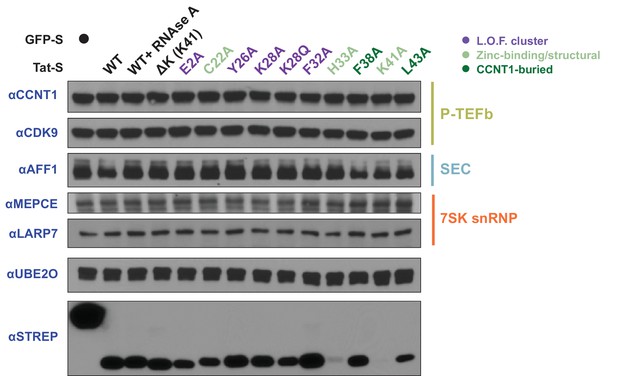
Protein input blots for Tat mutant IPs.
Input western blots for STREP IPs from Figure 2D using endogenous antibodies.
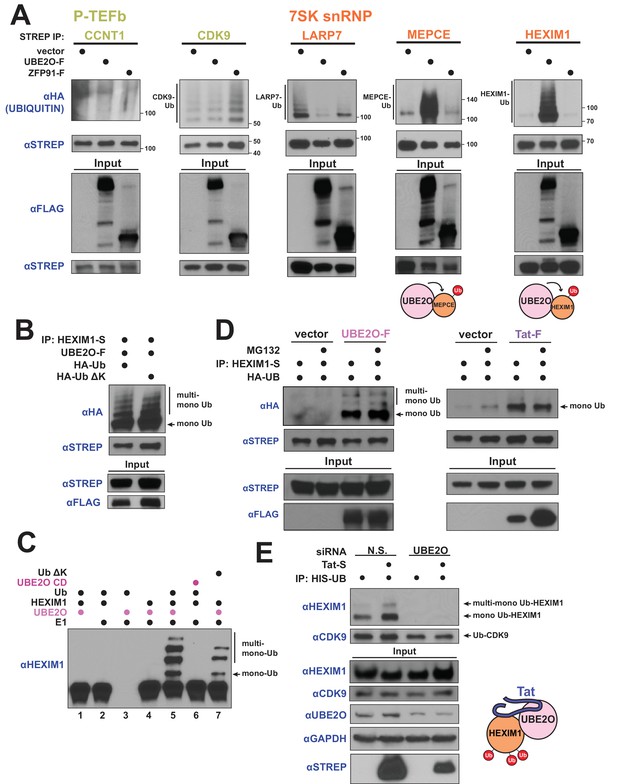
HIV-1 Tat hijacks UBE2O for the non-degradative monoubiquitination of HEXIM1.
(A) In vivo ubiquitination of transfected, STREP-tagged P-TEFb and 7SK snRNP proteins co-transfected with HA-ubiquitin ±co transfection of UBE2O-F or ZFP91-F (-F designates FLAG tagged protein). ZFP91 is used as a specificity control. Denaturing lysis and IPs were performed to detect ubiquitination by HA western blot. Ubiquitinated species are indicated. Schematics illustrate that UBE2O increases ubiquitination of the 7SK snRNP proteins MEPCE and HEXIM1. (B) In vivo ubiquitination of HEXIM1-S in HEK 293 T cells co-transfected with UBE2O-F and either wild type or lysine-free (ΔK) HA-ubiquitin. Monoubiquitinated HEXIM1 and additional ubiquitinated species are indicated. (C) In vitro ubiquitination of HEXIM1 by UBE2O. HEXIM1-S purified from HEK 293T cells was incubated with recombinant E1 and ubiquitin, UBE2O-F purified from HEK 293T cells, and ATP. Control reactions lacked individual components as indicated. Additionally, one reaction was incubated with UBE2O CD (catalytically dead) purified from HEK 293T cells in place of wild type UBE2O (lane 6) or recombinant ubiquitin ΔK in place of wild type ubiquitin (lane 7). See also Figure 3—figure supplement 2B. (D) In vivo ubiquitination of HEXIM1-S co-transfected with UBE2O-F or Tat-F and subsequently treated with DMSO control or MG132 to block proteasomal degradation. See also Figure 3—figure supplement 2C. (E) HEK 293 T cells were transfected with non-silencing or UBE2O-directed siRNA (si1). Knockdown cells were then transfected with HIS-Ubiquitin L73P followed by a denaturing nickel purification. The L73P ubiquitin mutant was used to enrich for ubiquitinated species, which were detected by western blot using specific antibodies (anti-CDK9, anti-HEXIM1) on the nickel-purified elution and are indicated. Schematic illustrates that Tat enhances HEXIM1 ubiquitination through UBE2O.
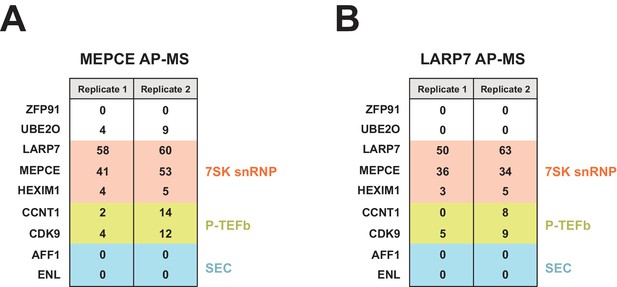
FLAG-tagged MEPCE and LARP7 AP-MS.
Peptide counts are shown for selected prey proteins from MEPCE (A) or LARP7 (B) duplicate (independent transfections on different days) FLAG purifications.
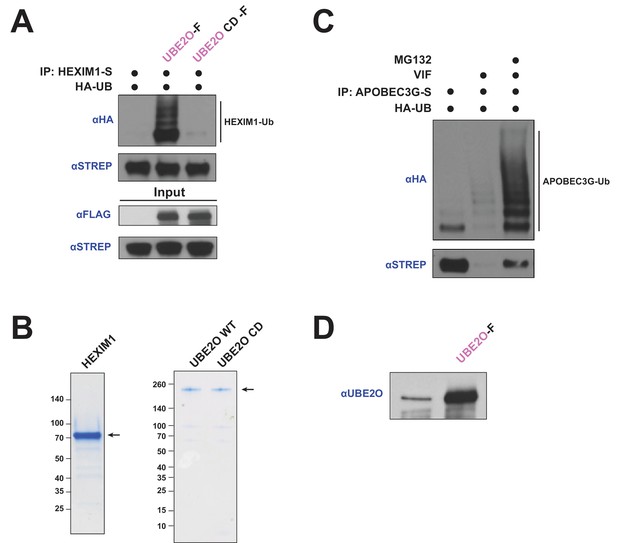
A catalytically inactive UBE2O does not induce HEXIM1 ubiquitination.
(A) In vivo ubiquitination of transfected HEXIM1 with the co-transfection of wild type or catalytically dead (CD) UBE2O in HEK 293T cells. HA western blot indicates the ubiquitinated species. (B) Protein gels demonstrating purity of HEXIM1, UBE2O WT, and UBE2O CD purified from HEK 293 T cells and used in the in vitro ubiquitination assay in Figure 3C. Arrows indicate HEXIM1 or UBE2O species. (C) In vivo ubiquitination of APOBEC3G-S ±HIV-1 Vif co-transfection and ±MG132 treatment. This control indicates that MG132 treatment restores APOBEC3G protein levels from Vif-mediated degradation (Yu et al., 2003). (D) Extent of UBE2O overexpression compared to endogenous levels. Shown are the input UBE2O levels from Figure 3D, +MG132 with an anti-UBE2O antibody.
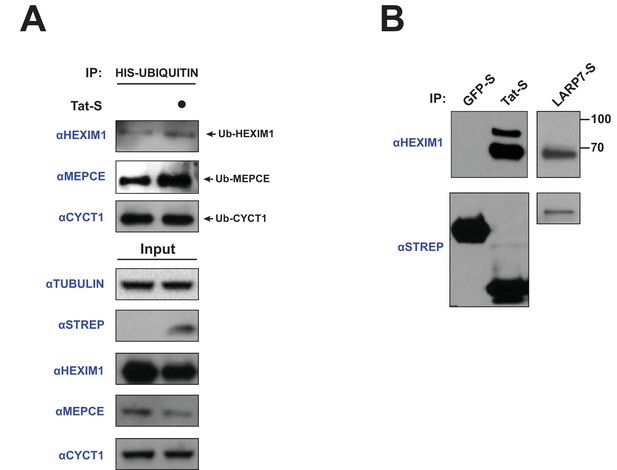
Tat induces the ubiquitination of endogenous HEXIM1 and MEPCE.
(A) HeLa cells were transfected with HIS-Ubiquitin and increasing amounts of Tat-STREP plasmid (0.33, 1, and 3 μg). A denaturing nickel pull-down followed by western blotting with specific antibodies was used to assess the ubiquitination state of each protein. (B) HEK 293 T cells were transfected with GFP-STREP or Tat-STREP plasmids, followed by a STREP IP and western blotting for endogenous HEXIM1. Unmodified HEXIM1 runs slightly under 70 kDa. A LARP7-STREP IP was used as a comparison for the state of bound HEXIM1.
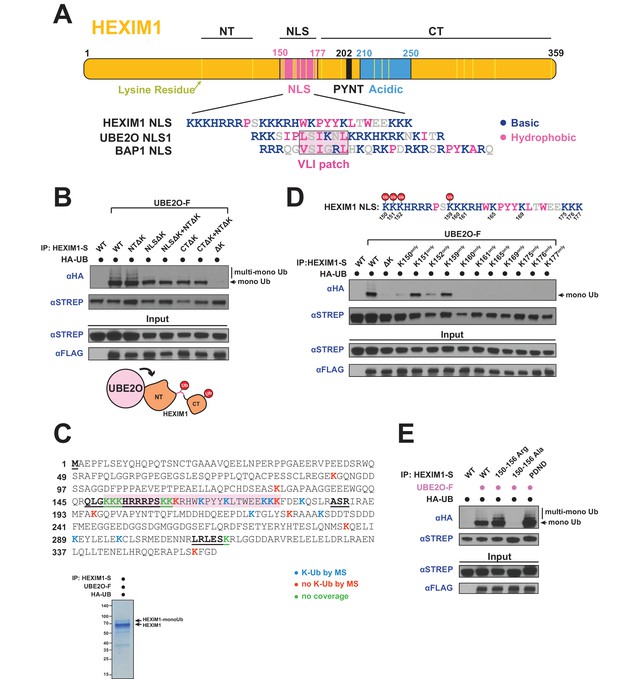
UBE2O ubiquitinates the NLS of HEXIM1.
(A) Domain organization of HEXIM1. The positions of the 23 lysines in HEXIM1 are indicated by yellow bars in the C-terminal (CT) and N-terminal (NT) regions and the NLS. (Below) Alignment of the NLS sequences from HEXIM1, UBE2O, and a known UBE2O substrate, BAP1 (Mashtalir et al., 2014). UBE2O and several of its target proteins contain a consensus bipartite NLS architecture that shares similarities with the HEXIM1 NLS. Basic residues are labeled in blue while hydrophobic residues are labeled in pink. (B) In vivo ubiquitination of HEXIM1 lysine mutants. Combinations of STREP-tagged lysine-to-arginine mutants were generated in the HEXIM1 N-terminal region (NTΔK), NLS (NLSΔK), and C-terminal region (CTΔK). A complete lysine-free HEXIM1 is designated as ΔK. UBE2O-F was co-transfected where indicated, and ubiquitination was detected by anti-HA western after denaturing lysis and IP. Monoubiquitin and multi-monoubiquitin species are indicated. The NTΔK mutant had no effect on ubiquitination, suggesting that there are no acceptor sites in the N-terminal region. However, both the NLSΔK and CTΔK mutants displayed decreased ubiquitination, demonstrating the presence of acceptor lysines in these regions. Schematic illustrates that lysines in the NLS (pink connection between domains) and in the C-terminal region function as acceptor sites. See also Supplementary file 2. (C) Coverage of the HEXIM1 sequence by MS. Sequences not identified by MS are bolded and underlined. Shaded in red is the HEXIM1 bipartite NLS. Color coding reflects whether a lysine was identified with a ubiquitin di-glycine tryptic remnant fragment. Shown below is a protein gel of HEXIM1-S purified under denaturing conditions from HEK 293 T cells with the co-expression of UBE2O-F. Arrows indicate unmodified and monoubiquitinated HEXIM1 species. (D) In vivo ubiquitination of individual lysine residues in the HEXIM1 NLS re-introduced in the context of the lysine-free (ΔK) HEXIM1 background. Ubiquitination of single-lysine HEXIM1 proteins was detected by HA Western blot. (top) All lysines are numbered in the HEXIM1 NLS and those that restore ubiquitination are indicated. (E) In vivo ubiquitination of wild-type STREP-tagged HEXIM1 or mutants deficient in binding P-TEFb (202PYNT205 -> 202PDND205) or the 7SK snRNA (150–156 Ala). HEXIM1-S was co-transfected with UBE2O-F as indicated.
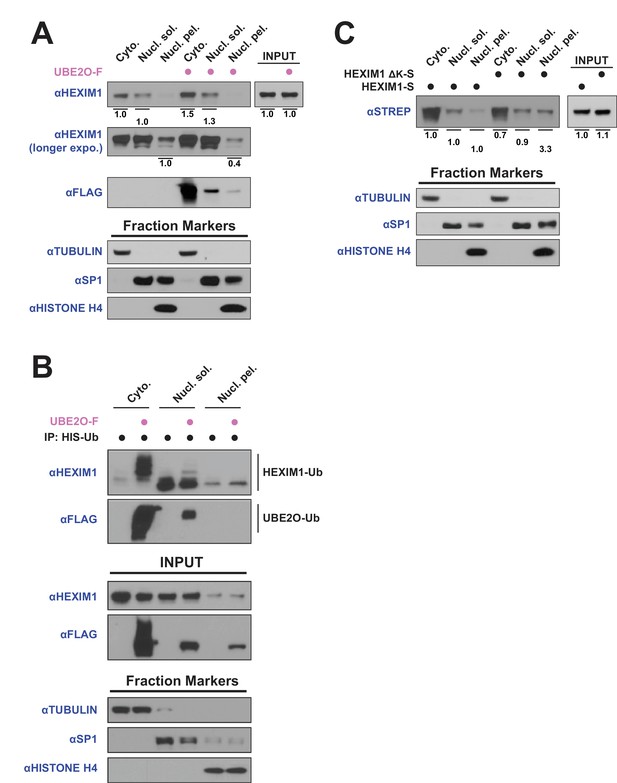
UBE2O influences HEXIM1 subcellular localization.
(A) Subcellular localization of endogenous HEXIM1. HEK 293T were transfected ±3 μg UBE20-F plasmid and fractionated into cytoplasmic (Cyto.), soluble nuclear (Nucl. sol.), and nuclear pellet (Nucl. pel.) fractions. Numbers represent quantified pixel intensity for HEXIM1 bands relative to un-transfected cells for the same fraction. Tubulin, SP1, and histone H4 were used as markers for each fraction and show the quality of the separations. HEXIM1 bands were first normalized to the pixel intensity of appropriate fraction controls (i.e. Cyto. normalized to tubulin, Nucl. sol. normalized to SP1, Nucl. pel. normalized to histone H4) prior to comparison between treatments. Two exposures of HEXIM1 from the same blot are shown. (B) Subcellular fractionation of HEK 293 T cells transfected with HIS-Ubiquitin. UBE2O-F was co-transfected as indicated and denaturing nickel pull-downs were used to detect HEXIM1 ubiquitination by anti-HEXIM1 western in the various subcellular fractions. Ubiquitinated species are indicated. (C) Subcellular localization of 0.25 μg of transfected HEXIM1-S or HEXIM1 ΔK-S plasmid in HEK 293 T cells. Numbers represent quantified pixel intensity for HEXIM1 (STREP) bands relative to wild-type expression for the same fraction. Bands were normalized as in (A).
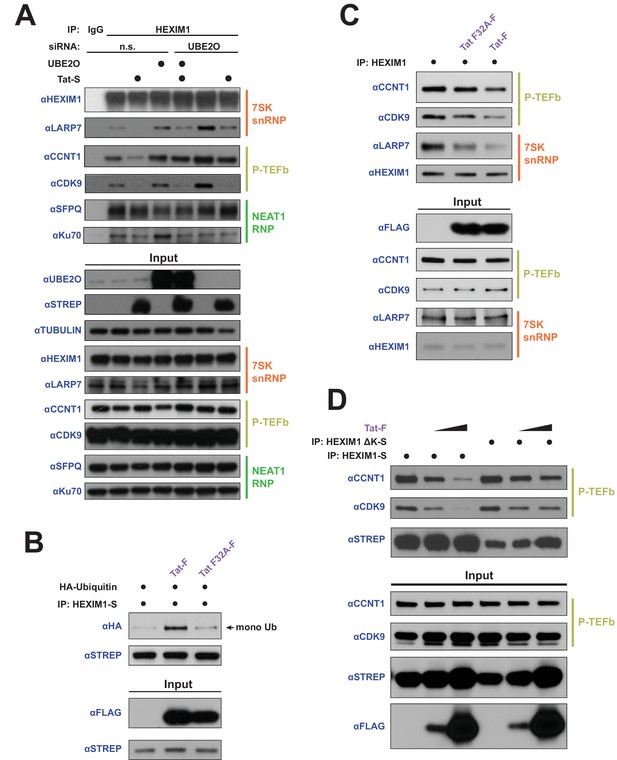
HEXIM1 ubiquitination is required for Tat-dependent release of P-TEFb from the inhibitory 7SK snRNP complex.
(A) Endogenous IP of HEXIM1 in non-silencing (n.s.) and UBE2O RNAi cells, ±Tat expression. Rabbit IgG was used as a negative IP control. Western blots were performed with the specific endogenous antibodies indicated to detect HEXIM1 protein-protein interactions. (B) In vivo ubiquitination assay of HEXIM-S. Wild type Tat or the F32A mutant was co-transfected as indicated. HEXIM1 ubiquitination was monitored by anti-HA western. The monoubiquitinated HEXIM1 species is indicated. (C) Endogenous IP of HEXIM1. Prior to lysis, cells were transfected with vector, wild type Tat, or Tat F32A. HEXIM1 interactions were detected by western blot with the indicated antibodies. (D) IP of HEXIM1-S wild type or ΔK mutant co-transfected with increasing Tat plasmid.
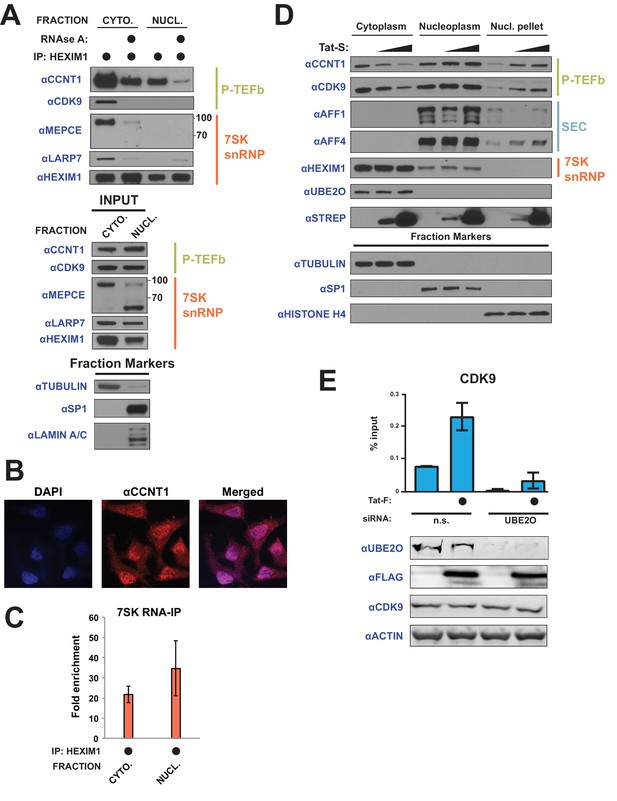
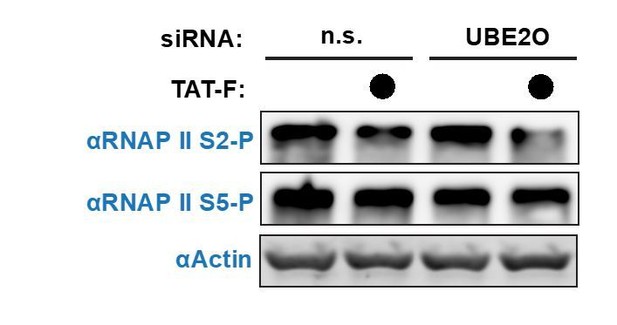
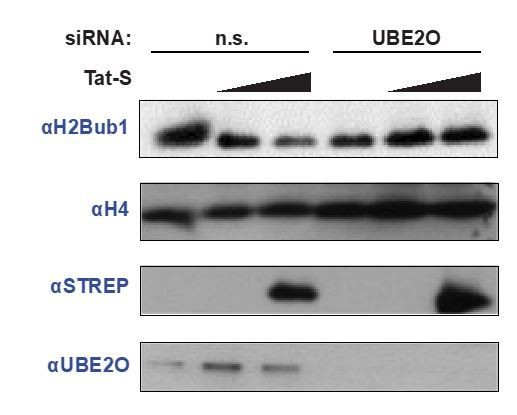
Tat utilizes UBE2O for the chromatin recruitment of P-TEFb released from a cytoplasmic pool of 7SK snRNP.
(A) Endogenous HEXIM1 IP using cytoplasmic and soluble nuclear fractions. One sample from each fraction was treated with RNAse A prior to the IP to evaluate RNA-dependence of interactions. (B) Immunofluorescence analysis of endogenous CCNT1 in HeLaprovirusΔtat cells, demonstrating nuclear and cytoplasmic staining. Nuclei were stained with DAPI. (C) RNA-IP assay of the 7SK snRNA. An endogenous HEXIM1 IP was performed using cytoplasmic and soluble nuclear fractions. RNA was then purified from the IPs and 7SK snRNA was detected by RT-qPCR. Fold enrichments were calculated as relative amounts of 7SK snRNA [HEXIM1 IP/rabbit IgG IP], with the average fold enrichments shown ± SEM for biological triplicates (purifications performed on different days from independently seeded cells). (D) HeLa cells were transfected with increasing amounts of Tat-S plasmid (0, 0.5, or 5 μg) followed by subcellular fractionation, and western blots of endogenous proteins were performed using the indicated antibodies. Fraction markers are shown to demonstrate the quality of the fractions. (E) Chromatin immunoprecipitation (ChIP) of CDK9 at the HIV-1 promoter in the HeLaprovirusΔtat cell line, ±UBE2O RNAi, ±Tat expression. Values represent mean % input (IP/input) minus a non-specific IgG control from biological duplicate experiments (defined as independent transfections on different days). Error bars represent standard error. Western blots below demonstrate efficient UBE2O knockdown and Tat expression in the cells used for ChIP.
-
Figure 7—source data 1
Source data for RIP assay of Figure 7C.
- https://doi.org/10.7554/eLife.31879.016
-
Figure 7—source data 2
Source data for ChIP experiment in Figure 7E.
- https://doi.org/10.7554/eLife.31879.017
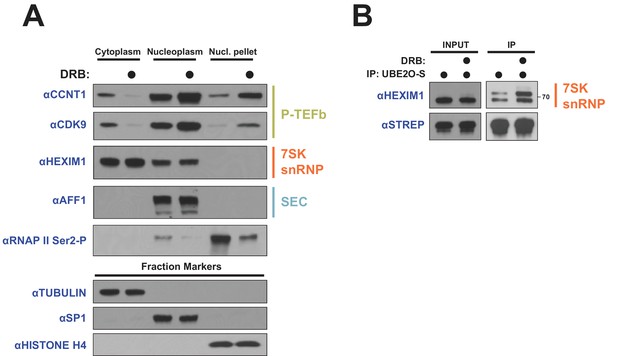
DRB releases P-TEFb from cytoplasmic 7SK snRNP.
(A) HeLa cells were treated with 100 μM DRB or an equal volume of DMSO for 1 hr prior to subcellular fractionation and Western blotting. (B) HEK 293 T cells were transfected with UBE2O-S. Cells were then treated with 100 μM DRB or DMSO 1 hr prior to lysis and STREP IP. The endogenous HEXIM1 interaction was detected by anti-HEXIM1 western.
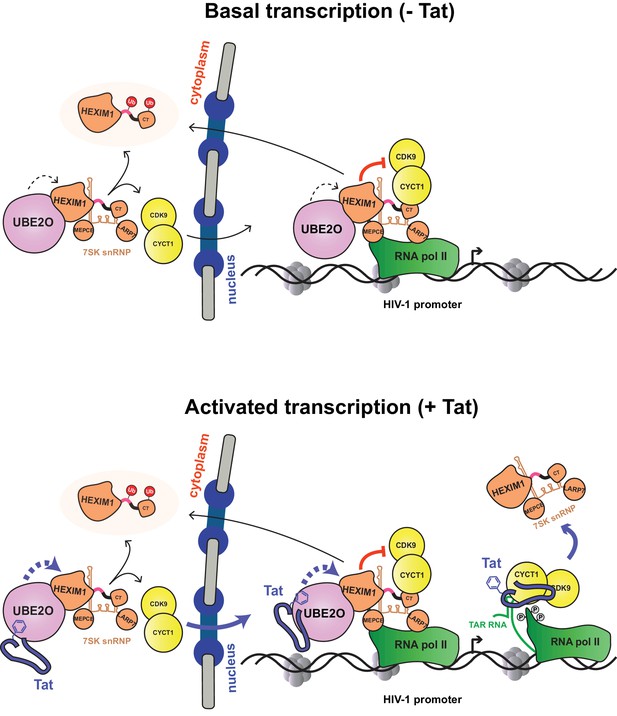
Model of basal and Tat-activated HIV-1 transcription.
(Top) Basal transcription. The inhibited P-TEFb-7SK snRNP complex resides in the cytoplasm and bound to RNAP II at the viral promoter. Inhibition of P-TEFb kinase activity pauses the RNAP II-7SK snRNP complex at the promoter. UBE2O, also residing in the nucleus and cytoplasm, ubiquitinates HEXIM1, resulting in the sequestration of HEXIM1 in the cytoplasm and the nuclear import/retention of P-TEFb. This freed P-TEFb is then able to phosphorylate RNAP II, leading to low-level activation. (Bottom) Activated transcription. Once Tat is expressed, it can assemble with the inhibited P-TEFb complex at the HIV-1 promoter. The synthesis of TAR RNA results in the displacement of the inhibitory snRNP to activate the kinase activity of CDK9 (D'Orso and Frankel, 2010). Further, Tat enhances the association of UBE2O with HEXIM1, leading to increased HEXIM1 ubiquitination and P-TEFb release and nuclear import. Residue F32 in Tat is critically important for the UBE2O-dependent activation mechanism. Tat combines multiple P-TEFb targeting mechanisms to supply activated kinase to RNAP II at the HIV-1 promoter.
Tables
| Reagent type (species) or resource | Designation | Source or reference | Identifiers | Additional information |
|---|---|---|---|---|
| Recombinant DNA reagent | UBE2O-FLAG (plasmid) | this paper | vector:pcDNA4/TO, insert: full-length Homo sapiens UBE2O | |
| Recombinant DNA reagent | HEXIM1-STREP (plasmid) | this paper | vector:pcDNA4/TO, insert: full-length Homo sapiens HEXIM1 | |
| Sequence-based reagent | ZFP91 siRNA #1 | Qiagen | SI05109223 | |
| Sequence-based reagent | ZFP91 siRNA #2 | Qiagen | SI05109230 | |
| Sequence-based reagent | ZFP91 siRNA #3 | Qiagen | SI05150985 | |
| Sequence-based reagent | UBE2O siRNA #1 | Qiagen | SI04153863 | |
| Sequence-based reagent | UBE2O siRNA #2 | Dharmacon | J-008979–05 | |
| Sequence-based reagent | UBE2O siRNA #3 | Dharmacon | J-008979–06 | |
| Sequence-based reagent | CDK9 siRNA | Qiagen | SI00605066 | |
| Sequence-based reagent | CCNT1 siRNA | Qiagen | SI02625707 | |
| Sequence-based reagent | Non-silencing siRNA | Qiagen | SI03650318 | |
| Cell line (Homo sapiens) | HeLa (provirusΔtat) | PMID: 28345603 | ||
| Peptide, recombinant protein | HA-ubiquitin | this paper | purified from E. coli BL21 | |
| Peptide, recombinant protein | HA-ΔK ubiquitin | this paper | purified from E. coli BL21 | |
| Peptide, recombinant protein | UBE2O | this paper | purified from HEK 293 T cells with high salt | |
| Peptide, recombinant protein | UBE2O, catalytically inactive | this paper | purified from HEK 293 T cells with high salt | |
| Peptide, recombinant protein | HEXIM1 | this paper | purified from HEK 293 T cells with high salt | |
| Commercial assay or kit | HiPerFect transfection reagent | Qiagen | used for transfecting siRNA | |
| Commercial assay or kit | PolyJet Transfection reagent | Signa Gen | used for transfecting DNA | |
| Commercial assay or kit | STREP-tactin superflow resin | IBA Lifesciences | ||
| Antibody | anti-HA | Covance | MMS-101P RRID:AB_2314672 | 1:2000 dilution |
| Antibody | anti-HEXIM1 | Abcam | ab25388 RRID:AB_2233058 | 1:2000 dilution, also used for IP |
| Antibody | anti-CDK9 | Santa Cruz Biotechnology | sc-13130 RRID:AB_627245 | used for ChIP |
| Antibody | anti-IgG | Millipore | 12–371 RRID:AB_145840 | used for ChIP |
| Antibody | anti-IgG | Santa Cruz Biotechnology | sc-2027 RRID:AB_737197 | For IP |
| Antibody | anti- RNA polymerase II | Santa Cruz Biotechnology | sc-899x RRID:AB_632359 | 1:2000 |
| Antibody | anti- CCNT1 | Santa Cruz Biotechnology | sc-8127 RRID:AB_2073892 | 1:2000 |
| Antibody | anti- CCNT1 | Santa Cruz Biotechnology | sc-10750 RRID:AB_2073888 | 1:2000 |
| Antibody | anti- MEPCE | Santa Cruz Biotechnology | sc-82542 RRID:AB_2266399 | 1:1000 |
| Antibody | anti- SP1 | Santa Cruz Biotechnology | sc-14027 RRID:AB_2171049 | 1:5000 |
| Antibody | anti-AFF1 | Bethyl | A302-344A RRID:AB_1850255 | 1:1000 |
| Antibody | anti- AFF4 | Bethyl | A302-538A RRID:AB_1998985 | 1:1000 |
| Antibody | anti- UBE2O | Bethyl | A301-873A RRID:AB_1309799 | 1:1000 |
| Antibody | anti-Ku70 | Bethyl | A302-624A-T RRID:AB_10554672 | 1:1000 |
| Antibody | anti- SFPQ | Bethyl | A301-322A-T RRID:AB_937995 | 1:1000 |
| Antibody | anti-LARP7 | Aviva | ARP40847_P050 RRID:AB_1294403 | 1:2000 |
| Antibody | anti-LARP7 | Santa Cruz Biotechnology | sc-515209 RRID:AB_2728652 | 1:2000 |
| Antibody | anti-HEXIM1 | Everest | EB06964 RRID:AB_2118260 | 1:5000 |
| Antibody | anti-CDK9 | Santa Cruz Biotechnology | sc-8338 RRID:AB_2260303 | 1:2000 |
| Antibody | anti-CDK9 | ProMab | 30212 RRID:AB_2728653 | 1:2000 |
| Antibody | anti-TUBULIN | Sigma-Aldrich | T6199 RRID:AB_477583 | 1:10,000 |
| Antibody | anti-FLAG | Sigma-Aldrich | F1804 RRID:AB_262044 | 1:5000 |
| Antibody | anti-RNA pol II phosphor-Ser2 | Abcam | ab5095 RRID:AB_304749 | 1:2000 |
| Antibody | anti-STREP-HRP | Millipore | 71591 RRID:AB_10806716 | 1:5000 |
| Antibody | anti-histone H4 | Active Motif | 39269 RRID:AB_2636967 | 1:5000 |
Additional files
-
Supplementary file 1
Full HEXIM1, MEPCE, and LARP7 AP-MS lists.
Each tab is a biological replicate FLAG affinity purification for the indicated protein.
- https://doi.org/10.7554/eLife.31879.020
-
Supplementary file 2
HEXIM1-Ub diGly data, related to Figure 4.
Tabs show raw data for all detected unmodified and modified HEXIM1 peptides (HEXIM1 tab) or the sum of MS/MS counts for peptides generated from trypsin or Arg-C treatment (HEXIM1 summ. tab).
- https://doi.org/10.7554/eLife.31879.021
-
Transparent reporting form
- https://doi.org/10.7554/eLife.31879.022





