C/EBPδ drives interactions between human MAIT cells and endothelial cells that are important for extravasation
Figures
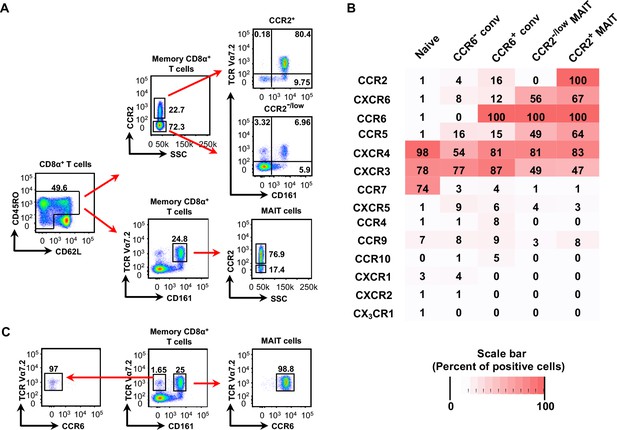
Most CD8α+CCR2+ cells are MAIT cells and all CD8α+ MAIT cells express CCR6.
(A) Expression of CCR2 on CD8α+ memory-phenotype cells and frequencies of TCR Vα7.2+ CD161+ (MAIT) cells within the CCR2+ and CCR2- subsets (top), and expression of CCR2 within the MAIT cell population (bottom). Cells are shown from one representative of more than 30 donors. (B) Expression of chemokine receptors on naive CD8α+ T cells and memory-phenotype CD8α+ T cells divided into the following subsets: TCR Vα7.2− CCR6− (CCR6- conventional), TCR Vα7.2− CCR6+ (CCR6+ conventional), TCR Vα7.2+ CCR6+ CCR2-/low (CCR2-/low MAIT), and TCR Vα7.2+ CCR6+ CCR2+ (CCR2+ MAIT); shades of red reflect percentages of cells positive for each chemokine receptor as determined by flow cytometry and as displayed in each box. Data are averaged from cells from nine donors. (C) Expression of CCR6 on MAIT cells (right) and TCR Vα7.2+ CD161- cells (left). Numbers are percentages of cells within a quadrant or demarcated region. Cells are shown from one representative of six donors.
-
Figure 1—source data 1
Data for Figure 1B, flow cytometry results for cells from individual donors.
- https://doi.org/10.7554/eLife.32532.004
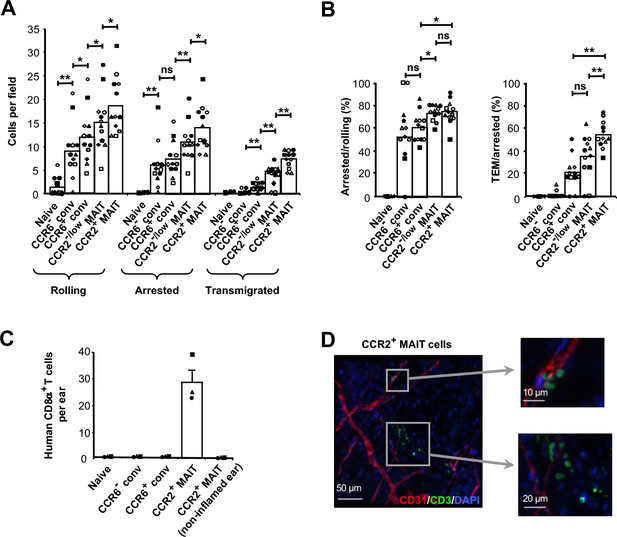
CCR2+ MAIT cells are highly efficient at TEM.
(A) Numbers of cells rolling, arrested, and transmigrated on TNFα-activated HUVECs for CD8α+ T cells divided into subsets as in Figure 1B and shown in Figure 2—figure supplement 1. (B) Percentages of rolling and arrested cells that arrested and transmigrated, respectively, calculated from the data in (A). Bars in (A) and (B) show means, and data are from cells from 12 donors, each identified by a unique symbol. The p values were calculated using the Wilcoxon signed rank test. (C) Numbers of CFSE-labeled human CD8α+ T cells recovered from TNFα/IL-1β-injected mouse ears after intra-cardiac injections of T cell subsets. Bars show means and SEMs from three experiments, each with one mouse injected with 1 × 106 cells from each subset. (D) Confocal microscopy of ear sheets prepared from mice after injection of purified CCR2+ MAIT cells as in (C) and stained for human T cells (anti-human CD3, green), mouse endothelial cells (anti-murine CD31, red) and nuclei (DAPI, blue). Squares indicate areas magnified at right. Images are representative of four experiments. (A and B) ns, not significant; *, p<0.05; **, p<0.01.
-
Figure 2—source data 1
Data for Figure 2A, B, flow chamber results for cells from individual experiments.
- https://doi.org/10.7554/eLife.32532.008
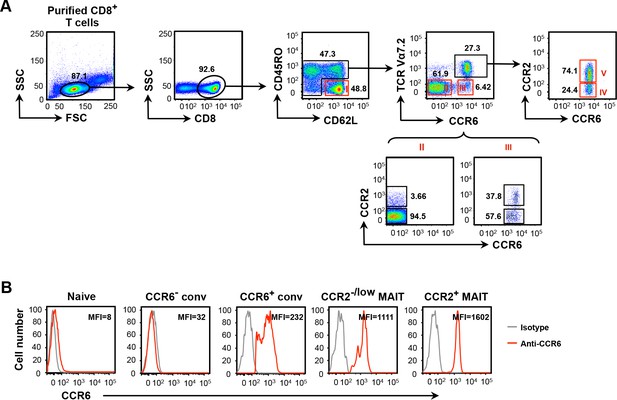
CCR2+ MAIT cells are CCR6high.
(A) Upper panels show gating for separation of CD8+ T cells, purified by negative selection, into CD8α+ naive cells (I), CCR6- conventional memory-phenotype cells (II), CCR6+ conventional memory-phenotype cells (III), CCR2-/low MAIT cells (IV), and CCR2+ MAIT cells (V), for purification by cell sorting. Sequential gating is shown for cells demarcated by black boxes. Lower panels show CCR2-expressing cells in the CCR6- conventional memory-phenotype cells (II) and the CCR6+ conventional memory-phenotype cells (III). Data are from one representative of more than 20 experiments. (B) Expression of CCR6 on CD8α + T cell subsets. Staining with an isotype-matched antibody served as a negative control. MFI is mean fluorescent intensity for CCR6 staining. Data are from one representative of more than 20 experiments.
-
Figure 2—figure supplement 2—Source data 1
Data for Figure 2—figure supplement 2C, flow chamber results for individual cells.
- https://doi.org/10.7554/eLife.32532.009
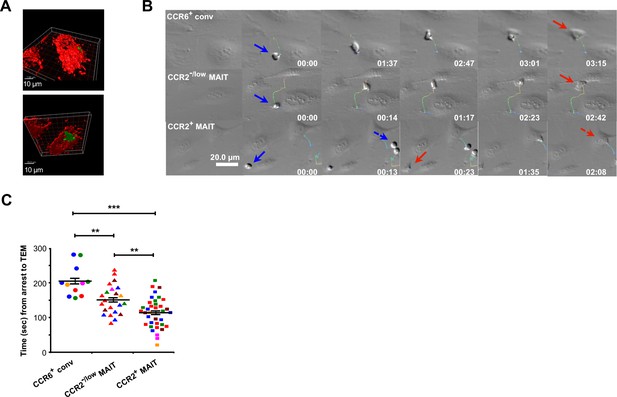
CCR2+ MAIT cells undergo rapid TEM.
(A) View from the top (above) and bottom (below) using three-dimensional volume renderings of CFSE-stained CCR2+ MAIT cells migrating across a monolayer of Red CMTPX-stained TNFα-activated HUVECs under flow. Data are from one representative of four experiments. (B) Time-lapse images of CD8α+ CCR6+ conventional T cells, CCR2−/low MAIT cells, and CCR2+ MAIT cells on TNFα-activated HUVEC under flow. Time 0 is at the arrest of the first cell within each subset (row), and times elapsed are shown in minutes:seconds. For crawling cells, tracks are color-coded for time from early (blue-green) to late (red). Blue arrows indicate cells at the time of arrest and red arrows at the time of completing TEM. Two transmigrating CCR2+ MAIT cells are shown, marked separately with solid and dashed arrows. (C) Times between arrest and initiating TEM for cells within the CD8α + T cell subsets on TNFα-activated HUVECs. Each symbol represents an individual cell and colors identify data from individual donors. Means and SEMs are indicated for each subset. Data in (B) are of cells from one representative of more than ten donors. Cells from five of these donors were used for the analysis in (C), for which p values were calculated using the unpaired t test. **, p<0.01; ***, p<0.001.
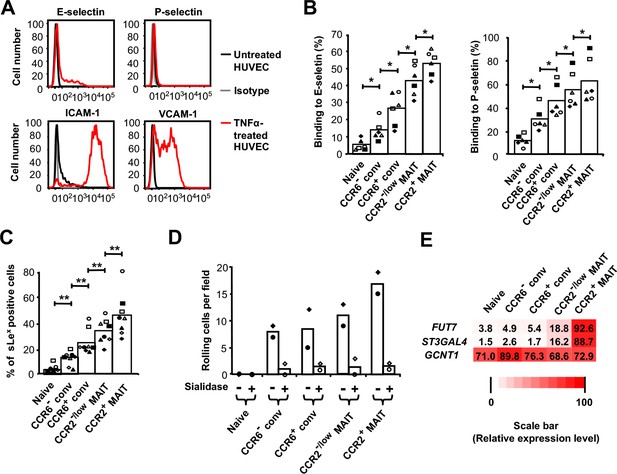
Rolling correlates with expression of selectin ligands and FUT7 and ST3GAL4.
(A) Expression of adhesion molecules on untreated HUVECs and TNFα-treated HUVECs. Staining of untreated HUVECs with the four different antibodies and TNFα-treated HUVECs with isotype-matched antibodies are shown as negative controls. Data are from one representative of three experiments. (B) Percentages of cells binding to the E-selectin-Fc and P-selectin-Fc chimeric proteins. CD8α+ T cells were divided into subsets as in Figure 1B and shown in Figure 2—figure supplement 1. (C) Percentages of sLeX-positive cells within each CD8α+ T cell subset. (B and C) Bars show means, and data are from cells from six (B) or those six plus two additional (C) donors, each identified by a unique symbol within each panel. The p values were calculated using the Wilcoxon signed rank test. (D) Numbers of cells rolling per field on TNFα-activated HUVECs for CD8α+ T cell subsets, either untreated or pre-treated with sialidase. Bars show means from cells from two donors as represented by the two symbols. (E) Expression of FUT7, ST3GAL4 and GCNT1 in CD8α+ T cell subsets; shades of red and numbers displayed in each box represent relative levels of expression based on values for 2−ΔCT obtained by real-time RT-PCR. Data are averaged from cells from three donors. (B and C) *, p<0.05; **, p<0.01.
-
Figure 3—source data 1
Data for Figure 3B, C (flow cytometry results for cells from individual donors), Figure 3D (flow chamber results for cells from individual experiments), and Figure 3E, (normalized mRNA expression in cells from individual experiments).
- https://doi.org/10.7554/eLife.32532.014
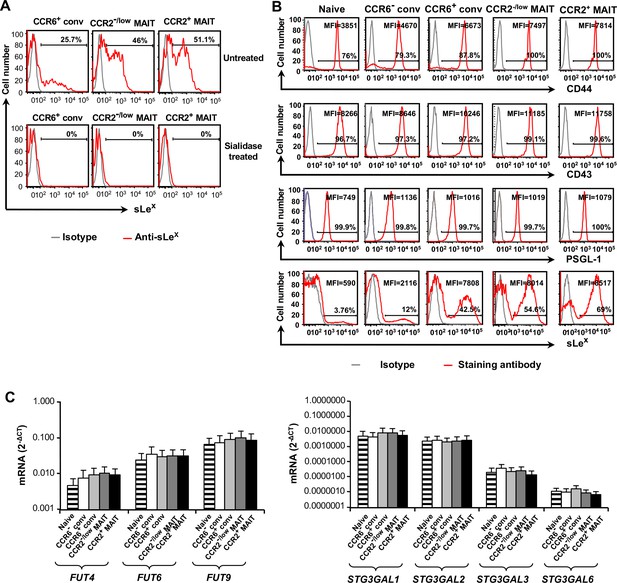
Selectin ligands and glycosyltransferases in CD8α+ T cells.
(A) Staining of CD8α+ T cell subsets either untreated (top) or after treatment with sialidase (bottom) for sLeX. Staining with an isotype-matched antibody served as a negative control. Percentages of cells staining positive are indicated by the horizontal lines. Data are from one of two experiments. (B) Staining of CD8α+ T cell subsets is shown for sLeX and proteins that can serve as selectin ligands. Percentages of cells staining positive are indicated by the horizontal lines. MFI numbers are mean fluorescent intensities. Staining with isotype-matched antibodies served as negative controls. For each surface molecule data are from cells from one representative of four donors. (C) Expression of mRNAs encoded by the glycosyltransferase genes normalized to GAPDH expression in the CD8α+ T cell subsets. Bars show means and SEMs for cells from three donors.
-
Figure 3—figure supplement 1—source data 1
Data for Figure 3—figure supplement 1B, mRNA expression in cells from individual experiments.
- https://doi.org/10.7554/eLife.32532.015
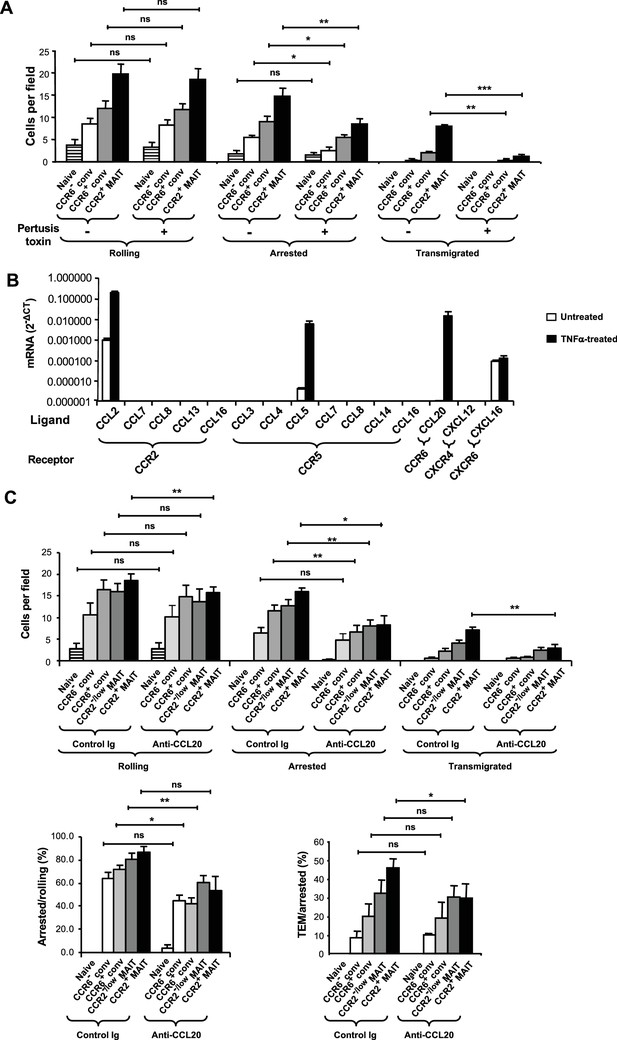
CCR6 mediates arrest.
(A) Numbers of cells rolling, arrested, and transmigrated per field on TNFα-activated HUVECs for CD8α+ T cell subsets, either untreated or treated with pertussis toxin. CD8α+ T cells were divided into subsets as in Figure 1B and shown in Figure 2—figure supplement 1, except that the CCR2-/low MAIT cells were not studied. (B) Expression of mRNAs, normalized to GAPDH expression, encoding chemokine ligands for the listed receptors in HUVECs either untreated or treated with TNFα. Bars show means and SEMs from two experiments. (C) Numbers of T cells rolling and arrested per field on TNFα-activated HUVEC that had been pre-treated with either control IgG or anti-human CCL20 antibody (top); percentages of rolling and arrested cells that arrested and transmigrated, respectively, calculated from the data shown above. CD8α+ T cells were divided into subsets as in Figure 1B and shown in Figure 2—figure supplement 1. (A and C) Bars show means and SEMs. Data are from cells from four (A) and five (C) donors. The p values were calculated using the paired t test. (A and C) ns, not significant; *, p<0.05; **, p<0.01; ***, p<0.001.
-
Figure 4—source data 1
Data for Figure 4A, C (flow chamber results for cells from individual experiments) and Figure 4B (mRNA expression in individual experiments).
- https://doi.org/10.7554/eLife.32532.018
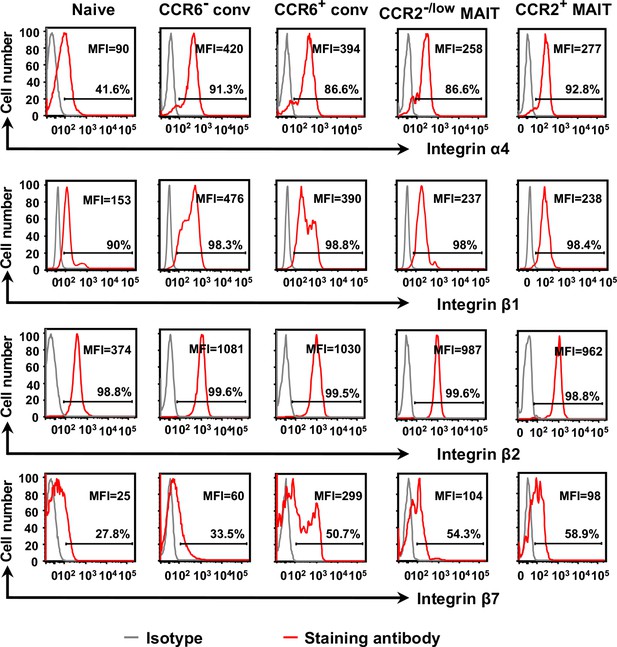
Expression of integrins does not account for MAIT cells’ enhanced arrest on activated endothelial cells.
Staining of CD8α+ T cell subsets is shown for integrin subunits. Percentages of cells staining positive are indicated by the horizontal lines. MFI numbers are mean fluorescent intensities. Staining with isotype-matched antibodies served as negative controls. For each surface molecule, data are from cells from one representative of four donors.
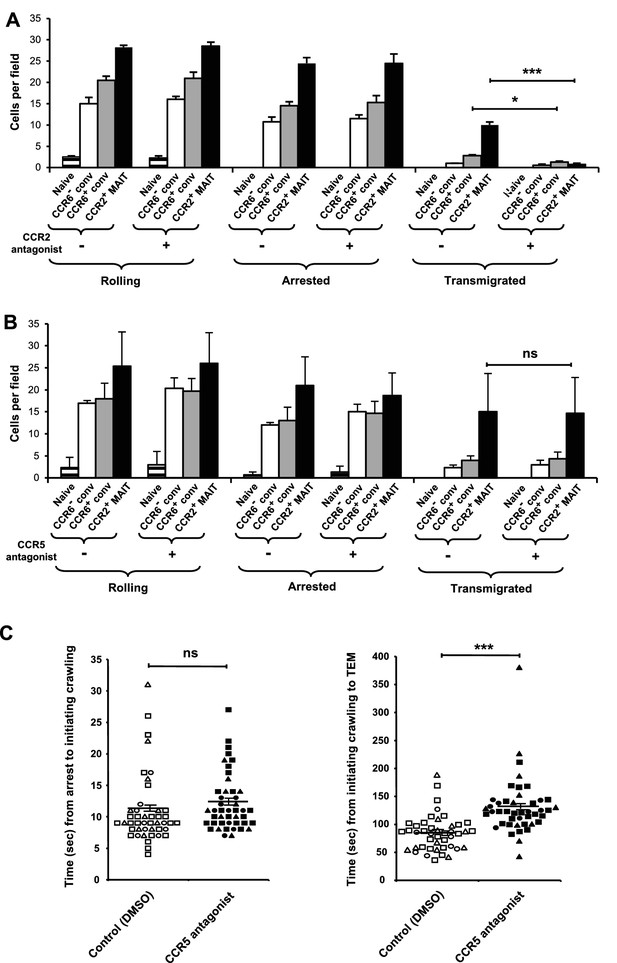
CCR2 and CCR5 contribute to TEM.
(A) Numbers of cells rolling, arrested, and transmigrated per field on TNFα-activated HUVECs for CD8α+ T cells, either untreated or treated with a CCR2 antagonist. CD8α+ T cells were divided into subsets as in Figure 1B and shown in Figure 2—figure supplement 1, except that the CCR2-/low MAIT cells were not studied. (B) Numbers of cells rolling, arrested, and transmigrated as described in (A) for cells either untreated or treated with a CCR5 antagonist. (A and B) Bars show means and SEMs. Data are from cells from four (A) and three (B) donors. The p values were calculated using the paired t test. (C) Times between arrest and initiating crawling and between initiating crawling and initiating TEM for CCR2+ MAIT cells on TNFα-activated HUVECs, either control-treated (open symbols) or treated with a CCR5 antagonist (closed symbols). Each symbol represents an individual cell; open-ended horizontal lines show means and SEMs. Data are from cells from three donors as represented by the three types of symbols. The p value was calculated using the unpaired t test. ns, not significant; *, p<0.05; ***, p<0.001.
-
Figure 5—source data 1
Data for Figure 5A, B (flow chamber results for cells from individual experiments) and Figure 5C (flow chamber results for individual cells).
- https://doi.org/10.7554/eLife.32532.021
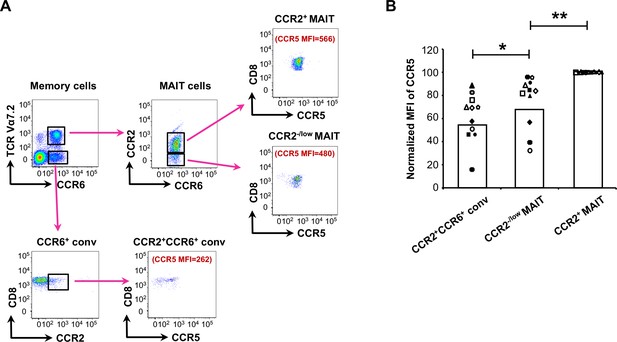
Levels of CCR5 expression differ among CD8α+ subsets.
(A) Gating for identifying CCR2+ MAIT cells, CCR2-/low MAIT cells, and CCR2+CCR6+ conventional memory-phenotype cells for analyzing CCR5 mean fluorescent intensities (MFI). Sequential gating is shown for cells demarcated by black boxes. Data are from one representative of 11 donors. (B) MFI’s for CCR5 were determined for the three subsets shown in A from 11 donors, with each donor identified by a unique symbol. For each donor, the subset with the highest MFI for CCR5 was used to establish the value of 100, and the CCR5 MFI’s for the remaining two subsets were normalized accordingly. Bars show means. The p values were calculated using the Wilcoxon signed rank test. *, p<0.05; **, p<0.01.
-
Figure 5—figure supplement 1—source data 1
Data for Figure 5—figure supplement 1B, flow cytometry results for cells from individual donors.
- https://doi.org/10.7554/eLife.32532.022
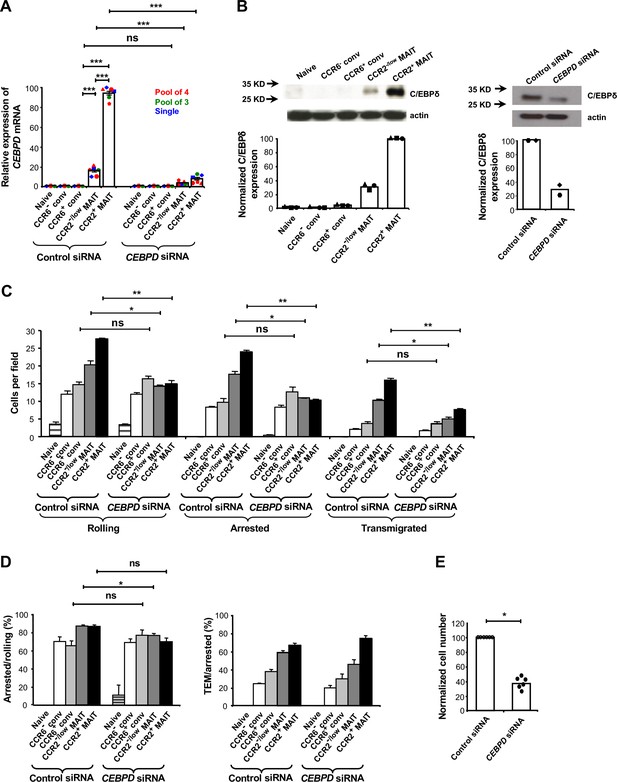
C/EBPδ supports rolling and arrest of MAIT cells.
(A) Relative expression of CEBPD mRNA in CD8α+ T cells after transfections with control or CEBPD siRNA. CD8α+ T cells were divided into subsets as in Figure 1B and shown in Figure 2—figure supplement 1. Each symbol type shows results from an individual donor. Red and green symbols are from experiments using non-overlapping pools of four or three CEBPD siRNAs, respectively, and blue symbols are from experiments using one of the siRNAs from the pool of three. Other experiments in this figure used the pool of four siRNAs and in all experiments using siRNAs, cells were harvested 3–4 days after transfections. Bars show means and SEMs. Data are from cells from six donors. The p values were calculated using the paired t test. (B) Expression of C/EBPδ in CD8α+ T cell subsets, either untreated (left) or after transfections with control or CEBPD siRNA (right). Actin bands demonstrate equal loading and arrows indicate positions of molecular weight markers. Blots are from cells from one representative of three (left) and two (right) donors, with quantification of the blots shown below. (C) Numbers of cells rolling, arrested, and transmigrated per field on TNFα-activated HUVECs for CD8α+ T cell subsets after transfections with control or CEBPD siRNA. (D) Percentages of rolling and arrested cells that arrested and transmigrated, respectively, calculated from the data in (C). (C and D) Bars show means and SEMs. Data are from cells from three donors. The p values were calculated using the paired t test. (E) Relative numbers of CCR2+ MAIT cells transfected with CEBPD versus control siRNAs recovered from TNFα/IL-1β-injected mouse ears eight minutes after intra-cardiac injection of a 1:1 mixture of differentially labeled cells. Values were normalized to numbers of the cells transfected with control siRNA. Bars show means. Data are from four experiments with a total of six mice. The p value was calculated using the Wilcoxon signed rank test. (A, C, D, and E) ns, not significant; *, p<0.05; **, p<0.01; ***, p<0.001.
-
Figure 6—source data 1
Data for Figure 6A (normalized mRNA expression in cells from individual experiments), Figure 6E, (quantification of Western blots from individual experiments), Figure 6C, D, (flow chamber results for cells from individual experiments), and Figure 6E, (normalized cell numbers from ears from individual experiments).
- https://doi.org/10.7554/eLife.32532.025
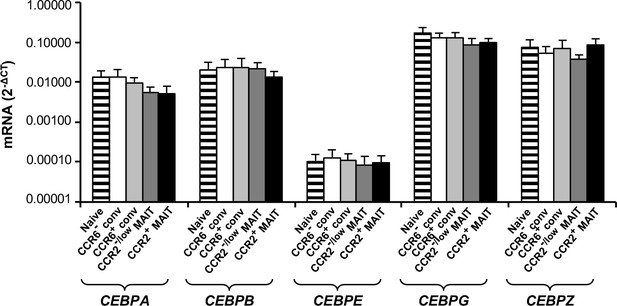
Multiple CEBP genes are similarly expressed among CD8α+ T cell subsets.
Expression of mRNAs encoded by the CEBP genes, normalized to GAPDH expression, as indicated in the CD8α+ T cell subsets. Bars show means and SEMs for cells from four donors.
-
Figure 6—figure supplement 1—source data 1
Data for Figure 6—figure supplement 1, mRNA expression from individual experiments.
- https://doi.org/10.7554/eLife.32532.026
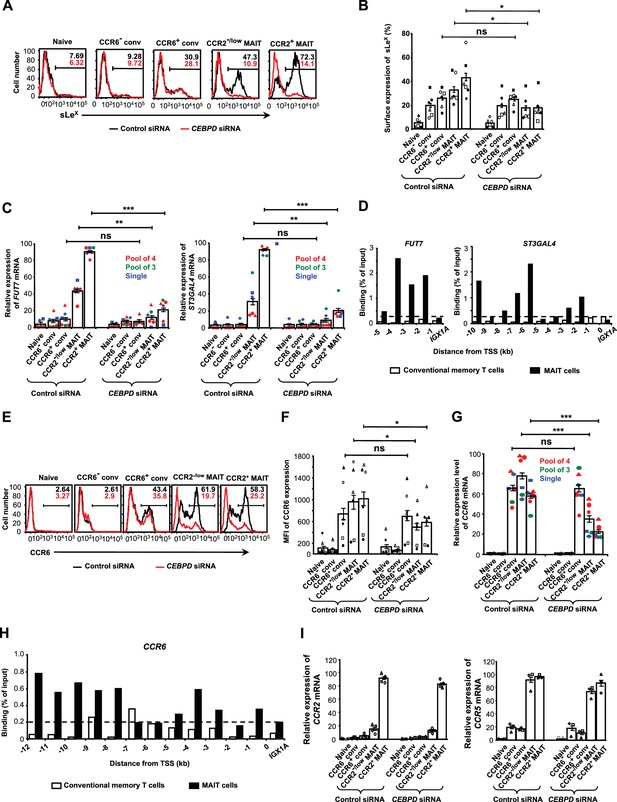
C/EBPδ regulates sLeX, FUT7, ST3GAL4 and CCR6/CCR6 in MAIT cells.
(A) Expression of sLeX on CD8α+ T cells after transfections with control or CEBPD siRNA. CD8α+ T cells were divided into subsets as in Figure 1B and shown in Figure 2—figure supplement 1. Percentages of cells staining positive are indicated by the horizontal lines. Unless otherwise noted, experiments in this figure used the pool of four siRNAs, and in all experiments using siRNAs cells were harvested 3–4 days after transfections. (B) Expression of sLeX on T cells from multiple donors treated as in (A). (C) Relative expression of FUT7 and ST3GAL4 mRNAs in CD8α+ T cell subsets after transfections with control or CEBPD siRNAs. Each symbol type shows results from an individual donor. Red and green symbols are from experiments using non-overlapping pools of four or three CEBPD siRNAs, respectively, and blue symbols are from experiments using one of the siRNAs from the pool of three. (D) ChIP analysis of conventional memory-phenotype T cells and MAIT cells using anti-C/EBPδ antibodies and primers for amplifying sequences 5’ to the transcription start sites (TSS) for FUT7 and ST3GAL4 at 1 kb intervals. Data are expressed as percent of input DNA, and the dashed line indicates ‘background’ signal based on results for the intergenic region, IGX1A. (E) Expression of CCR6 on T cells after transfections with control or CEBPD siRNAs. Percentages of cells staining positive are indicated by the horizontal lines. (F) MFIs for CCR6 on T cells treated as in (E). (G) Relative expression of CCR6 mRNA in CD8α+ T cell subsets after transfections with control or CEBPD siRNAs as in (C). (H) ChIP analysis as in (D), except using primers for CCR6. (I) Relative expression of CCR2 and CCR5 mRNAs in CD8α+ T cell subsets after transfections with control or CEBPD siRNAs. (A, D, E and H) Data are from one representative of six (A), five (D), left panel), six (D), right panel), six (E), and two (H) donors. (B, C, F, G and I) Bars show means and SEMs, and data are from cells from a total of six (B, C, F, and G), four (I, left panel), or three (I, right panel) donors, each represented by a unique symbol. The p values were calculated using the ratio paired t test. (B, C, F, and G) ns, not significant; *, p<0.05; **, p<0.01; ***, p<0.001.
-
Figure 7—source data 1
Data for Figure 7B, F (flow cytometry results from individual experiments), Figure 7C, G, I (normalized mRNA expression in cells from individual experiments), Figure 7D, H (normalized ChIP-PCR results from individual experiments).
- https://doi.org/10.7554/eLife.32532.030
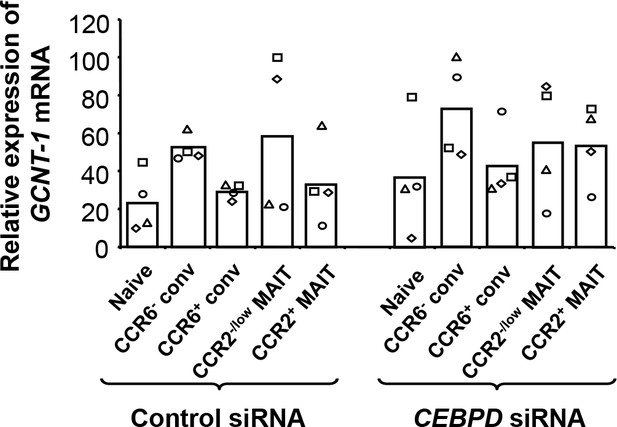
No evidence that C/EBPδ regulates GCNT1 in MAIT Cells.
Relative expression of GCNT1 mRNA in CD8α+ T cell subsets after transfections with control or CEBPD siRNA. Bars show means and data are from cells from four donors where each symbol type shows results from an individual donor.
-
Figure 7—figure supplement 1—source data 1
Data for Figure 7—figure supplement 2, predicted C/EBPd binding sequences.
- https://doi.org/10.7554/eLife.32532.031
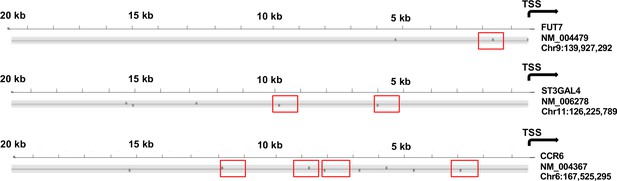
The 5’ flanking regions of FUT7, ST3GAL4, and CCR6 contain sequences predicted to bind C/EBPδ.
Schematic presentation of computationally predicted C/EBPδ-binding sites. Bindings sites for each gene were predicted in a 20,000 bp region immediately upstream from the start of transcription (TSS, indicated by the arrow). For each gene, the corresponding NCBI RefSeq number and the transcription start site genomic coordinate are provided below the gene name. Each black vertical bar indicates a predicted binding site on the sense strand (pointing up) or anti-sense strand (pointing down). Red boxes indicate C/EBPδ binding regions revealed by ChIP assays in Figure 7 that contain predicted C/EBPδ binding sites. * Because FUT7 is annotated on the anti-sense strand, the reverse-complement of the region was queried and displayed.
-
Figure 7—figure supplement 2—source data 2
Data for Figure 7—figure supplement 1, normalized mRNA expression in cells from individual experiments.
- https://doi.org/10.7554/eLife.32532.032
Videos
CCR2+ MAIT cells transmigrate soon after arresting on TNFα-activated HUVECs.
https://doi.org/10.7554/eLife.32532.010CCR2+ MAIT cells extravasate into the inflamed mouse ear.
https://doi.org/10.7554/eLife.32532.011Summary cartoon, C/EBPδ blocks early steps in MAIT cell trafficking.
https://doi.org/10.7554/eLife.32532.033Tables
| Reagent type (species) or resource | Designation | Source or reference | Identifiers |
|---|---|---|---|
| Biological sample (human) | Primary human umbilical vein endothelial cell (HUVEC) | ATCC, Manassas, VA | Cat#: PCS-100–013 |
| Biological sample (human) | Human whole blood and elutriated lymphocytes | Department of Transfusion Medicine, Clinical Center, National Institutes of Health | |
| Antibody | Purified-anti-CCL20/MIP-3α (67310) | Minneapolis, MN | Cat#: MAB360 |
| Antibody | Biotin-anti-CCR2 (48607) | R and D Systems | Cat#: FAB151B |
| Antibody | Allophycocyanin-anti-PSGL-1 (688101) | R and D Systems | Cat#: FAB9961R |
| Antibody | FITC-anti-CCR5 (2D7/CCR5) | Franklin Lakes, NJ | Cat#: 561747 |
| Antibody | PE-Cy5-anti-CCR5 (2D7/CCR5) | BD Biosciences | Cat#: 556889 |
| Antibody | Alexa Fluor 647-anti-CCR4 (1G1) | BD Biosciences | Cat#: 557863 |
| Antibody | Alexa Fluor 488-anti-CCR9 (112509) | BD Biosciences | Cat#: 112509 |
| Antibody | PE-anti-CCR10 (1B5) | BD Biosciences | Cat#: 563656 |
| Antibody | FITC-anti-CXCR1 (5A12) | BD Biosciences | Cat#: 555939 |
| Antibody | Allophycocyanin-anti-CXCR2 (6C6) | BD Biosciences | Cat#: 551127 |
| Antibody | Allophycocyanin-anti-CXCR3 (1C6/CXCR3) | BD Biosciences | Cat#: 561324 |
| Antibody | Allophycocyanin-anti-CXCR4 (12G5) | BD Biosciences | Cat#: 560936 |
| Antibody | Alexa Fluor 647-anti-CXCR5 (RF8B2) | BD Biosciences | Cat#: 558113 |
| Antibody | APC-Cy7-anti-CD8 (SK1) | BD Biosciences | Cat#: 557834 |
| Antibody | Alexa Fluor 700-anti-CD8 (RPA-T8) | BD Biosciences | Cat#:565165 |
| Antibody | PE-Cy5-anti-CD62L (DREG-56) | BD Biosciences | Cat#: 561915 |
| Antibody | FITC-anti-CD62L (DREG-56) | BD Biosciences | Cat#: 555543 |
| Antibody | PE-Cy5-anti-CD45RO (UCHL1) | BD Biosciences | Cat#: 561888 |
| Antibody | Brilliant Violet 605-anti-CD45RO (UCHL1) | BD Biosciences | Cat#: 562641 |
| Antibody | PE-Cy7-anti-CD45RO (UCHL1) | BD Biosciences | Cat#: 337168 |
| Antibody | PE-Cy7-anti-CCR6 (11A9) | BD Biosciences | Cat#: 560620 |
| Antibody | Allophycocyanin-anti-CCR6 (11A9) | BD Biosciences | Cat#: 560619 |
| Antibody | Allophycocyanin-anti-CD161(DX12) | BD Biosciences | Cat#: 550968 |
| Antibody | Non-conjugated anti-sLeX (CSLEX1) | BD Biosciences | Cat#: 551344 |
| Antibody | PE-conjugated streptavidin | BD Biosciences | Cat#: 349023 |
| Antibody | Alexa Fluor 647-anti-CD31 (390) | San Diego, CA | Cat#: 102416 |
| Antibody | Brilliant Violet 605-anti-CD3 (17A2) | BioLegend | Cat#: 100237 |
| Antibody | FITC-anti-integrin α4 (9F10) | BioLegend | Cat#: 304316 |
| Antibody | Alexa Fluor 647-anti-integrinβ1 (TS2/16) | BioLegend | Cat#: 303017 |
| Antibody | Allophycocyanin-anti-integrin β2 (m24) | BioLegend | Cat#: 363410 |
| Antibody | Allophycocyanin-anti-integrin β7 (FIB504) | BioLegend | Cat#: 321208 |
| Antibody | FITC-anti-TCR Vα7.2 (3C10) | BioLegend | Cat#: 351704 |
| Antibody | PE-anti-CXCR6 (K041E5) | BioLegend | Cat#: 356004 |
| Antibody | FITC-anti-CX3CR1 (2A9-1) | BioLegend | Cat#: 341606 |
| Antibody | Allophycocyanin-anti-CD43 (CD43-10G7) | BioLegend | Cat#: 343206 |
| Antibody | Allophycocyanin-anti-CD44 (BJ18) | BioLegend | Cat#: 338806 |
| Antibody | Biotin-anti-IgG Fc (HP6017) | BioLegend | Cat#:409308 |
| Antibody | Anti-CEBPδ (mouse monoclonal) | Dallas, TX | Cat#: sc-135733 |
| Antibody | Anti-CEBPδ | Other | BD 19 |
| Peptide, recombinant protein | Human recombinant TNFα | R and D Systems | Cat#: 210-TA/CF |
| Peptide, recombinant protein | Murine recombinant TNFα | R and D Systems | Cat#: 410-MT/CF |
| Peptide, recombinant protein | Murine recombinant IL-1β | R and D Systems | Cat#: 401 ML-025/CF |
| Peptide, recombinant protein | Human E-selectin Fc chimera | R and D Systems | Cat#: 724-ES |
| Peptide, recombinant protein | Human P-selectin Fc chimera | R and D Systems | Cat#: 137-PS |
| Chemical compound, drug | BMS CCR2 22 | R and D Systems/Tocris | Cat#: 3129 |
| Chemical compound, drug | Maraviroc | R and D Systems/Tocris | Cat#: 3756 |
| Chemical compound, drug | Sialidase (Vibrio cholera) | St. Louis, MO | Cat#: N7885-2UN |
| Chemical compound, drug | Pertussis toxin | R and D Systems | Cat#: 3097 |
| Chemical compound, drug | CFSE | Waltham, MA | Cat#: C34554 |
| Chemical compound, drug | CMTPX | Life Technologies | Cat#: C34572 |
| Chemical compound, drug | DAPI | Life Technologies | Cat#: D13076 |
| Sequence-based reagent | CEBPD SMARTpool siRNA | Lafayette, CO | Cat#: L-010453; D-010453–01; D-010453–02; D-01-453-03 |
| Sequence-based reagent | SAMRTpool siRNA control | Dharmacon | Cat#: D-001810-01-05; D-001210–01; |
| Commercial assay or kit | RosetteSep for human CD8 + T cell enrichment | Vancouver, Canada | Cat#: 15063 |
| Commercial assay or kit | qScript cDNA SuperMix | Quanta Biosciences | Cat#: 95048–500 |
| Commercial assay or kit | Perfecta qPCR FastMix UNG ROX | Beverly, MA | Cat#: 95077–012 |
| Commercial assay or kit | RT2 SYBR Green/ROX qPCR Master Mix | Frederick, MD | Cat#: 330522 |
| Commercial assay or kit | Human T Cell Nucleofector Kit | Walkersville, MD | Cat#: VPA-1002 |
| Commercial assay or kit | Magna ChIP A/G kit | Burlington, MA | Cat#: MAGNA0017 |
| Commercial assay or kit | SuperSignal West Pico Chemiluminescent Substrate | Rockford, lL | Cat#: 34080 |
| Software, algorithm | ImageJ | https://imagej.nih.gov/ij/ | |
| Software, algorithm | LAM510 version 4.2 | Wetzlar, Germany | |
| Software, algorithm | Flowjo | Ashland, OR | |
| Software, algorithm | Imaris (Bitplane) | Leica Microsystems | |
| Software, algorithm | Genome Analyzer's Common TF | Ann Arbor, MI | |
| Software, algorithm | Prism | La Jolla, CA |
Additional files
-
Transparent reporting form
- https://doi.org/10.7554/eLife.32532.034





