Intracellular antibody signalling is regulated by phosphorylation of the Fc receptor TRIM21
Figures
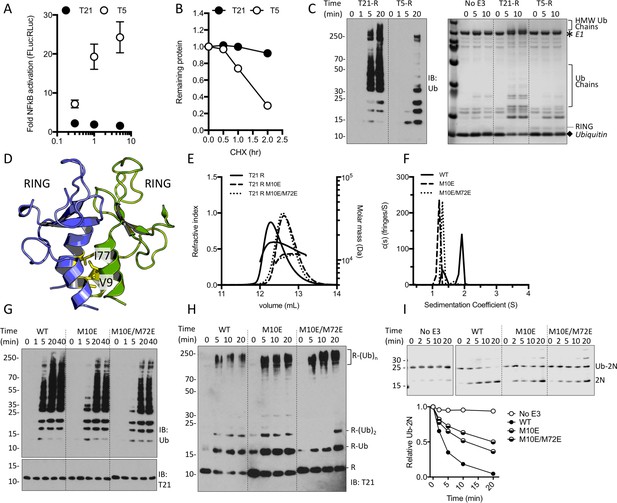
TRIM21 NFκB signalling is constitutively silent despite monomeric RING K63 chain synthesis.
(A) TRIM5α (T5) but not TRIM21 (T21) overexpression in 293Ts activates an NFκB-luciferase reporter. (B) Reduction in endogenous T5 and T21 protein levels after cycloheximide treatment. (C) (left) Immunoblot for in vitro polyubiquitin synthesis catalysed by T5 or T21 RING (R) domains in the presence of Ube2N/Ube2V2; (right) SDS-PAGE showing T21 R autoubiquitination in the presence of Ube2W and Ube2N/Ube2V2. (D) RING dimerization interface in T5 structure (4TKP). (E–F) Oligomerisation of T21 R and mutants M10E and M10E/M72E as assessed by (E) SEC MALS and (F) AUC. (G–H) Wild-type (WT) T21 R or dimerization mutants catalyzing unanchored polyubiquitin synthesis using Ube2N (G) or anchored chains using Ube2N/Ube2V2 and Ube2W (H). (I) T21 (WT or mutants) catalysed discharge of ubiquitin from conjugated Ube2N/Ube2V2.
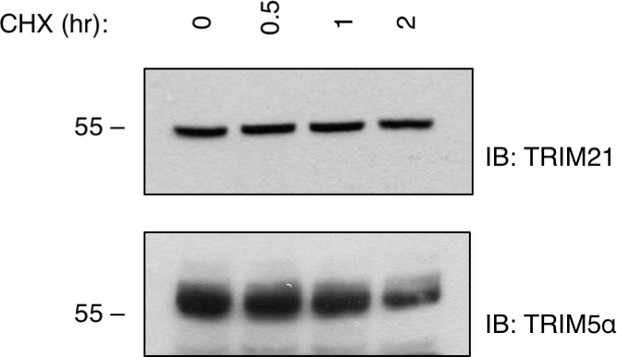
Cellular Turnover of endogenous TRIM5α and TRIM21.
https://doi.org/10.7554/eLife.32660.004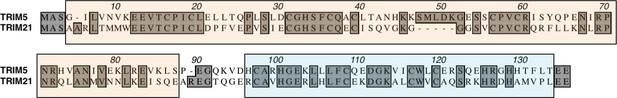
Sequence alignment of TRIM5 and TRIM21.
Boxed regions show the RING (orange) and B-Box (blue). Identical residues are highlighted.
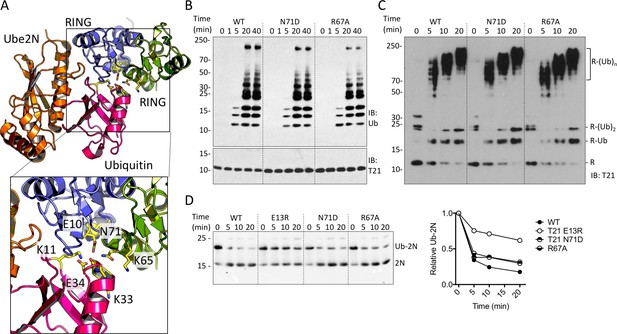
TRIM21 RING dimerization is not required to catalyse ubiquitin release from E2:Ub.
(A) Model of the TRIM25 RING:Ube2N:Ub complex (5EYA), showing interactions between ubiquitin and both RING monomers. (B–D) Ubiquitination activity of WT T21R and mutants of putative ubiquitin-contacting residues in the second monomer. (B) Catalysis of unanchored ubiquitin chains using Ube2N. (C) Synthesis of anchored K63-chains on TRIM21 in presence of both Ube2N/Ube2V2 and Ube2W. (D) Catalysis of ubiquitin discharge from conjugated Ube2N/Ube2V2.
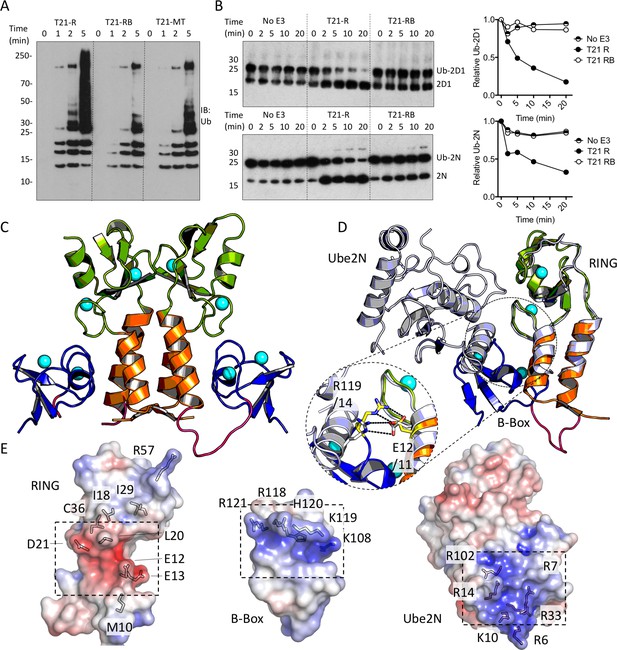
The Box is an autoinhibitory domain which supresses E3 ligase activity.
(A) Catalysis of Ube2N/Ube2V2 driven polyubiquitin-chain synthesis by T21 RING (R), RING-Box (RB) and MiniTRIM21 (MT). (B) T21 R or RB catalysed ubiquitin discharge from Ube2D1 or Ube2N. (C) 2 Å X-ray structure of T21 RB coloured by domain with zinc ions in cyan. (D) Superposition of T21 with T5 model 4TKP (grey), showing Ube2N and the Box are structurally competitive. (E) Surface representation of (left-to-right) T21 R, Box and Ube2N binding sites coloured by electrostatic potential from −5 kT/e (red) to + 5 kT/e (blue).
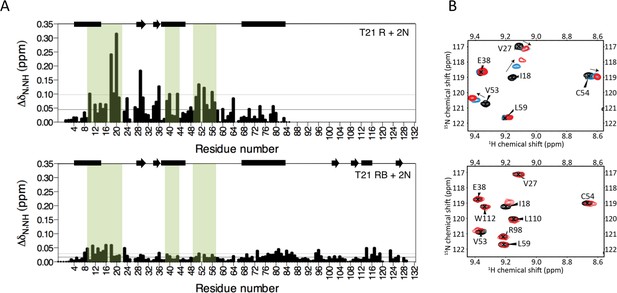
The Box prevents binding of E2 enzyme to the RING E3.
(A) Chemical shift perturbations recorded upon addition of unlabelled Ube2N into 15N labelled TRIM21 RING (R) (top) or RING-Box (RB) (bottom). E2 interacting regions are highlighted in green and secondary structure elements comprising either helices (rectangles) or b-strands (arrows) are indicated at the top of each plot. (B) Examples of the 15N HSQC spectra; residues in the RING (top) are shifted upon adding E2 (RING:E2 ratio: black 1:0, blue, 1:0.5, red, 1:1), while the same residues in the RING-Box (bottom) show little or no movement at a 1:1 titration point (red). Figure 4—figure supplement 1 shows the complete spectra.
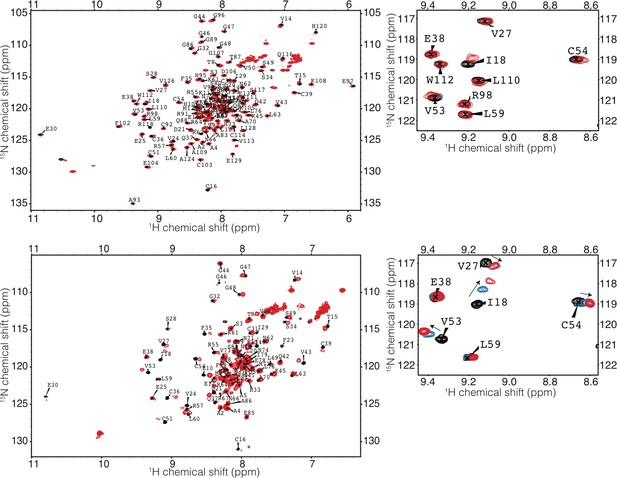
HSQC of TRIM21 RING and RING-Box with Ube2N.
(top) RING-Box (bottom) RING). Spectra of TRIM21 alone is in black, a 1:1 complex of TRIM21:Ube2N is coloured red. A 1:0.5 complex of TRIM21 RING:Ube2N is shown in blue.
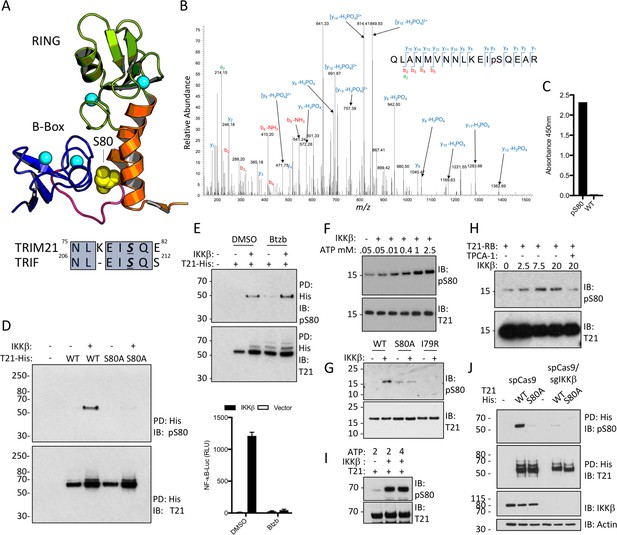
TRIM21 possesses a pLxxIS motif in its RING domain and is phosphorylated in vitro and in cells.
(A) Residue S80 (yellow spheres) is located at the centre of the Box:RING interface and is part of an epitope related to the IKKβ phosphorylation motif found in TRIF. (B) MS/MS spectra of TRIM21-His purified from 293 T cells indicating the presence of phosphorylated S80 (pS80) within a tryptic peptide precursor/m/z/680.0^3 ± 0.39 ppm. (C) ELISA of pS80 antisera at 1:50000 against recombinant WT or pS80 TRIM21. (D) IKKβ phosphorylates C-terminally His-tagged WT but not S80A T21-His, as detected by specific pS80 polyclonal sera. (E) As D except in presence of 30 nM bortezomib (Btzb) or DMSO and, below, NFκB stimulation by IKKβ in parallel. (F–H) In vitro phosphorylation of T21 RB by IKKβ with increasing ATP (F), impaired phosphorylation of S80A and I79R RB (G), in presence of 20 µM IKKβ inhibitor TPCA-1 (H). (I) In vitro IKKβ phosphorylation of full-length lipoyl-T21. (J) Phosphorylation of WT or S80A T21-His from control 293T (spCas9) or IKKβ-deleted 293T (spCas9/sgIKKβ) cells.
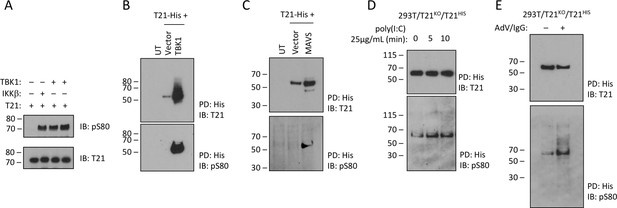
TRIM21 is phosphorylated upon immune stimulation.
(A) Full-length recombinant lipoyl-T21 protein phosphorylation in vitro by IKKβ and TBK1. (B–C) Expression of TBK1 (B) or upstream adaptor MAVS (C) in 293 T cells phosphorylates and stabilises C-terminally His-tagged T21. (D) 293 T cells lacking T21 reconstituted with His-tagged T21 and challenged with poly(I:C). Increased phosphorylation of T21 detected by immunoblot with pS80 antisera. (E) As in (D), except cells infected with adenovirus 5 (AdV) in the presence of IgG and T21 immunoprecipitated 7 hr post-infection.
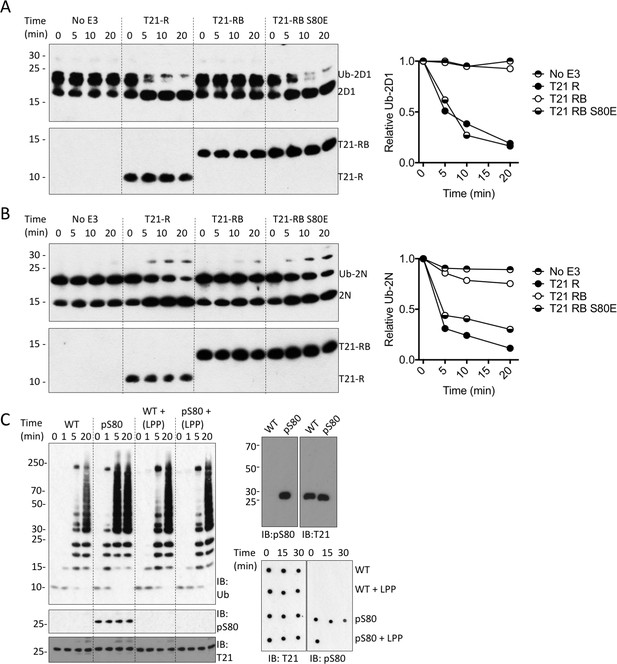
Phosphorylation of S80 or phosphomimetic mutation S80E releases Box inhibition and promotes TRIM21 ubiquitination activity.
(A–B) Catalysis of ubiquitin discharge from Ube2D1 (A) or Ube2N (B) by TRIM21 RING (R), RING-Box (RB) or RING-Box containing S80E mutation. The phosphomimetic mutation is sufficient to confer RING-like discharge kinetics on the RING-Box (offset graphs). (C) (top) Ubiquitin-chain catalysis by MiniTRIM21 either WT or with a phosphoserine incorporated co-translationally at position 80 by amber suppression (pS80). WT and pS80 MiniTRIM21 blotted with pS80 or T21 sera (top-right) or after treatment with lambda protein phosphatase (LPP) (bottom-right). LPP dephosphorylates S80 and restores Box autoinhibition.
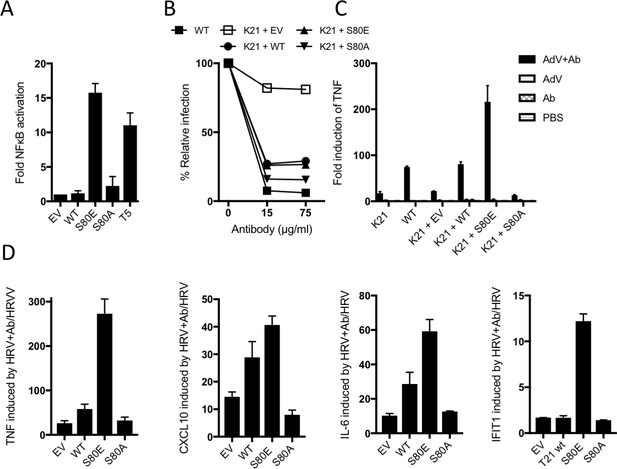
B-Box autoinhibition regulates antibody-dependent immune signalling by TRIM21.
(A) NFκB activation in 293Ts expressing WT, S80A or S80E T21 or WT T5. Phosphomimetic mutation S80E is sufficient to render TRIM21 constitutively active for NFκB activity. (B) Adenoviral neutralization is reconstituted in T21 knockout MEFs (K21) equally upon overexpression with WT or S80 mutants. (C) T21-mediated TNFA induction upon adenoviral (AdV) infection in the presence of antibody (Ab) is restored in K21 by overexpression of WT TRIM21 but not S80A, while S80E is hyperactive. Fold change in TNFA transcripts as determined by qPCR. (D) Immune gene transcription induced by antibody-coated human rhinovirus (HRV) in reconstituted MEFs. Phosphomimetic mutation S80E potentiates transcriptional activation by TRIM21, while S80A inhibits activity.
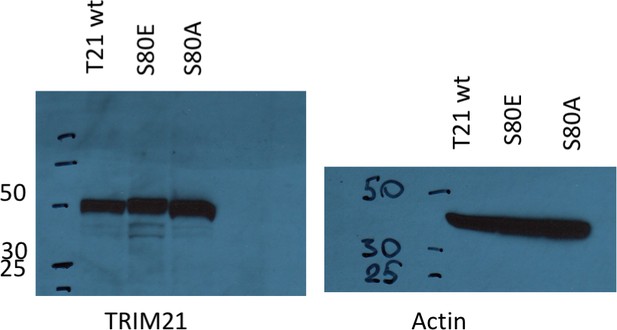
Ectopic expression of TRIM21 in knockout MEFs.
https://doi.org/10.7554/eLife.32660.014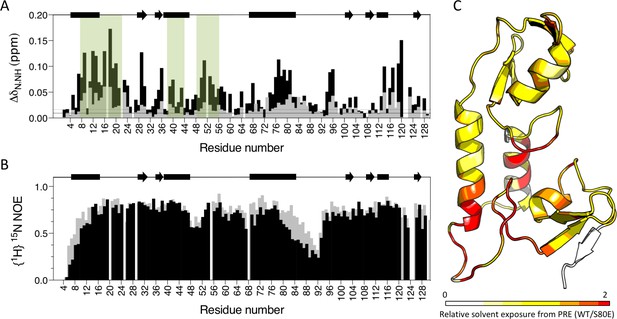
Phosphomimetic mutation S80E displaces the Box from the RING and allows E2 enzyme binding.
(A) Comparison of chemical shift perturbations upon E2 enzyme Ube2N titration into T21 RB (grey) or T21 RB S80E (black). E2 interacting regions are highlighted in green and secondary structure is indicated by rectangles (helices) and arrows (β-strands). (B) Comparison of heteronuclear NOEs for T21 RB (grey) or T21 RB S80E (black) in absence of E2. The data is consistent with increased dynamics in the linker region between the RING and the Box upon introduction of S80E. (C) Relative solvent exposure in S80E vs. WT, as detected by solvent PREs (shown as intensity ratio WT/S80E mapped onto the RB structure; primary data in Supplementary file 2). White areas indicate regions more solvent-exposed in WT while red areas indicate parts of the surface more solvent-exposed in S80E.
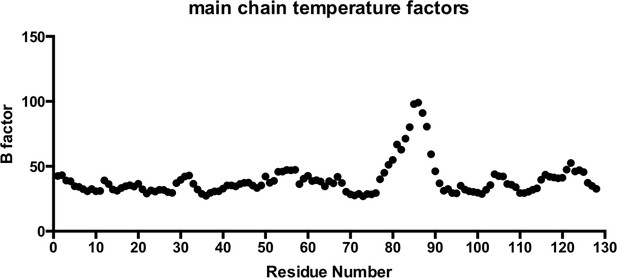
Main chain temperature factors for T21 RING-Box structure.
https://doi.org/10.7554/eLife.32660.016Additional files
-
Supplementary file 1
Data collection and refinement statistics
- https://doi.org/10.7554/eLife.32660.017
-
Supplementary file 2
Solvent PRE raw data
- https://doi.org/10.7554/eLife.32660.018
-
Transparent reporting form
- https://doi.org/10.7554/eLife.32660.019





