Orphan receptor GPR158 controls stress-induced depression
Figures
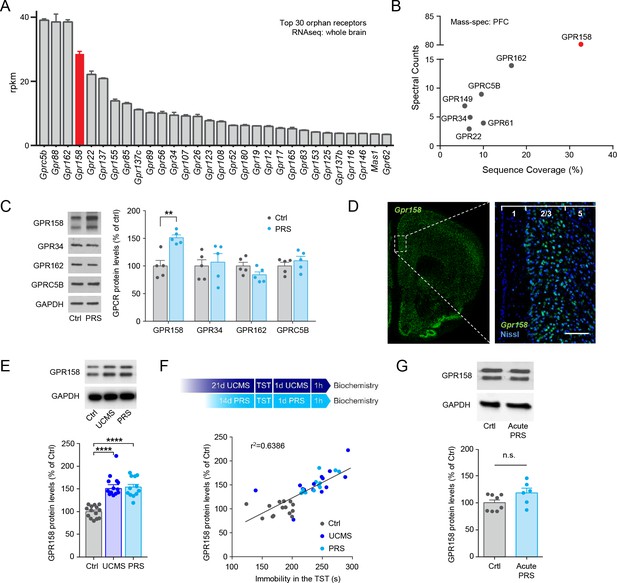
Stress upregulates GPR158 expression in the mPFC.
(A) RNA sequencing results from the whole mouse brain showing the top 30 orphan receptors. (B) Mass spectrometry analysis of the orphan receptors expressed in the PFC of mice. (C) Representative western blots and protein level quantification of several orphan receptors identified in the mPFC of control non-stressed (Crtl) and mice subjected to chronic Physical Restraint Stress (PRS) (n = 4–5 mice/group; Student’s t test; *p<0.05). (D) In situ hybridization showing the expression of GPR158 in the mPFC of mouse brain. (E) Mice subjected to PRS and unpredictable chronic mild stress (UCMS) show elevated GPR158 protein levels in the mPFC (n = 12–14 mice/group, one-way ANOVA with Bonferroni post hoc test). (F) Top panel shows scheme of stress paradigms. Graph showing a significant correlation between GPR158 protein levels and immobility in the TST of mice subjected to chronic stress (Pearson R2 = 0.6386, p<0.0001). (G) Mice subjected to 1 day of physical resistant stress (Acute PRS, 3 hr, n = 6) show no difference in GPR158 protein levels compared to control non-stressed mice (Crtl, n = 8) in the mPFC.
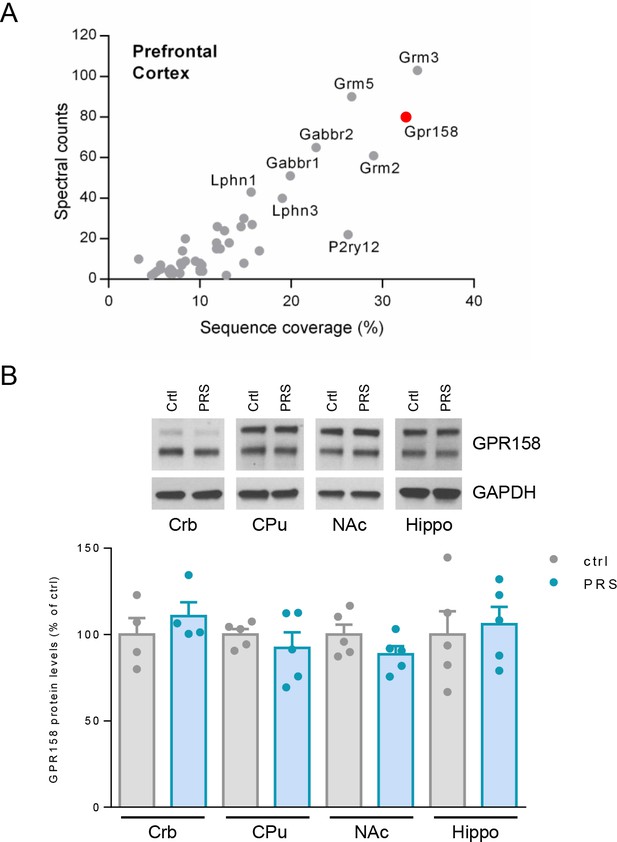
GPR158 is among the most expressed GPCR in the PFC.
(A) A proteomic analysis of the GPCR expression pattern in a membrane-enriched fraction of mouse mPFC identified 48 GPCRs with GPR158 being one of the most expressed. (B) Representative western blots and quantification of GPR158 protein levels in the indicated brain regions in control non-stressed (Crtl) and PRS mice (n = 4–5 mice/group; Student’s t test; *p<0.05; Crb = Cerebellum; CPu = Caudate Putamen; NAc = Nucleus Accumbens; Hippo = Hippocampus).
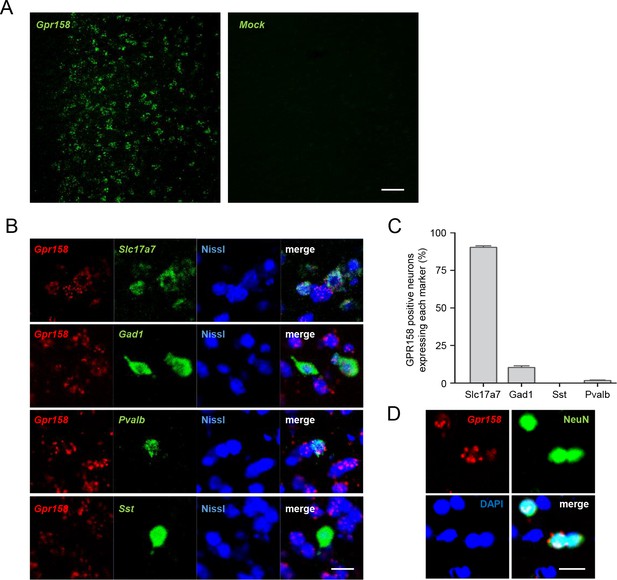
GPR158 expression in several neuronal populations.
(A) In situ hybridization specificity control. In situ hybridization of mouse GPR158 (left) and of the bacterial gene DapB (right) in the mPFC (scale bar = 50 µm). (B) Double in situ hybridization of GPR158 with the indicated neuronal markers. (C) Quantification of the percentage of cells in each population expressing GPR158 (92.0 ± 0.9% of vGlut1 positive neurons, 91.2 ± 0.8% of GAD1 positive neurons, 93.3 ± 6.7% of parvalbumin (Pvalb) positive neurons and 0.0% of somatostatin (Sst) positive neurons) (Nissl staining in blue, scale bar = 20 µm). (D) GPR158 lack of expression in glia cells is visualized by in situ hybridization of GPR158 mRNA (red), immunostaining with the neuronal marker NeuN (green) and nuclear staining of all the cells with DAPI (blue) (scale bar = 20 µm).
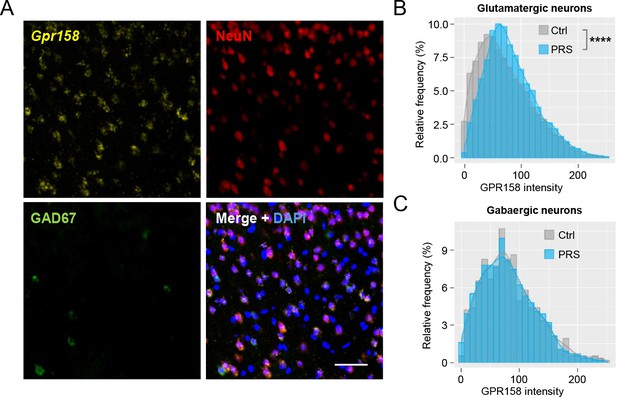
Chronic stress regulates GPR158 levels in glutamatergic neurons of the layer 2/3 of the mPFC.
(A) Representative image of in situ hybridization using a probe against GPR158 (yellow) and co-immunostaining with antibodies against NeuN (red) and GAD67 (green) in the layer 2/3 of mPFC (nuclei stained with DAPI in blue, scale bar = 50 µm). (B) Frequency distribution of GPR158 mRNA levels in glutamatergic neurons in the mPFC layer 2/3 (Ctrl n = 5 mice/6486 neurons; PRS n = 5 mice/6400 neurons; bin width = 10; p<0.0001 Kolmogorov–Smirnov test). (C) Frequency distribution of GPR158 expression in GAD67- positive neurons (Ctrl n = 5 mice/569 cells; PRS n = 5 mice/564 cells; bin width = 10; p=0.9599 Kolmogorov-Smirnov test).
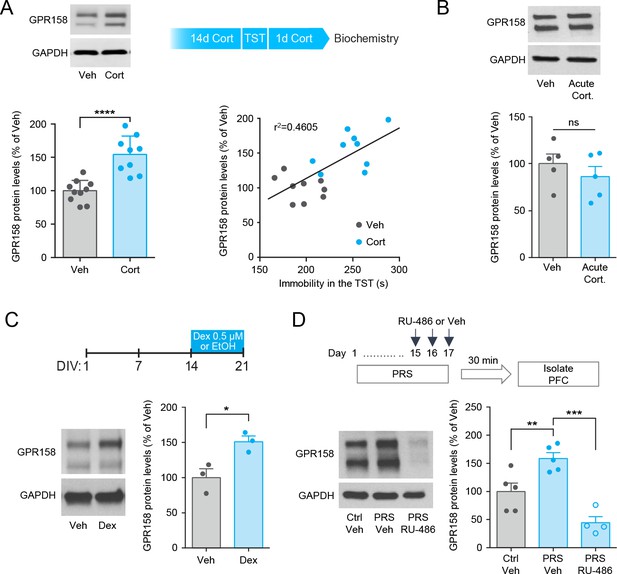
GPR158 is increased in the mouse mPFC by a corticosterone-mediated mechanism.
(A) GPR158 protein levels are increased in the mPFC of mice chronically treated with corticosterone (n = 9–10 mice/group, Student’s t test). Top panel depicts corticosterone paradigm. Mice treated with corticosterone exhibited a significant correlation between GPR158 protein levels and immobility in the TST (Pearson R2 = 0.4605, p=0.0014). (B) Mice injected acutely with corticosteroid (Acute Cort., 20 mg/kg) show no difference in GPR158 levels compared to vehicle (Veh) treatment (n = 5, Student’s t test; ns, not significant). (C) Western blot quantification of GPR158 in primary cortical neurons cultured for 21 days in vitro and treated with 0.5 µM dexamethasone (Dex) from DIV 14 to DIV21 (n = 3, Student’s t test, *p<0.05). (D) GPR158 protein levels in the mPFC of mice subjected to physical restraint stress and treated with RU-486. To induce chronic stress mice underwent a 17 day PRS period. On the last three days of the stress paradigm, mice were injected with RU-486 (10 mg/kg) or vehicle prior to being subjected to PRS. The mPFC was isolated 30 min following last episode of PRS (day 17). Stressed mice treated with RU-486 have a decrease in GPR158 protein levels compared to saline treatment (n = 4–5 mice/group, Two-way ANOVA with Tukey post hoc test, **p<0.01, ***p<0.001). Data shown as means ± SEM (Veh, vehicle; Ctrl, control; PRS, physical restraint stress; Cort., corticosterone).
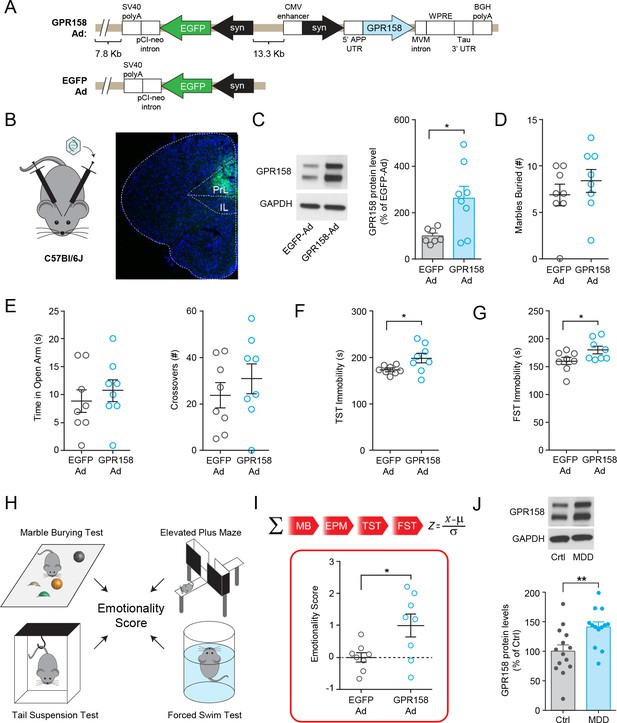
GPR158 overexpression in mouse mPFC and in dlPFC of MDD patient.
(A) Scheme of Punisher-Adenovirus expressing GPR158 (GPR158-Ad) and EGFP (EGFP-Ad) injected bilaterally into the mPFC of C57Bl/6J mice. (B) Representative viral expression in the mPFC (right panel). (C) Representative western blots and quantification of GPR158 protein levels of mice injected with GPR158-Ad and EGFP-Ad. Injected mice underwent marble burying test (D), Elevated Plus Maze (E), Tail Suspension Test (F), and Forced Swim Test (G) (n = 8 mice/group). (H) Emotionality scores were integrated from four behavioral tests (marble burying, elevated plus maze, tail suspension test and force swim test) and normalized to mice injected with EGFP-Ad. (I) Mice injected with GPR158-Ad showed a higher emotionality score reflective of a depressive-like phenotype. (J) Representative western blot and quantification of GPR158 protein levels in the dlPFC show an increase in subjects with MDD compared to healthy unaffected controls (Ctrl) (Ctrl n = 14; MDD n = 13). The different groups were matched as closely as possible for sex, age, and postmortem interval (see Materials and methods for further information). Data are mean ± SEM (Student’s t test, *p<0.05, **p<0.01).
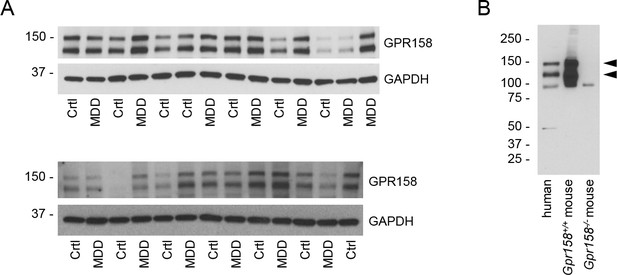
GPR158 expression levels in patients affected by MDD.
(A) Western blots of GPR158 expression in the dorsolateral prefrontal cortex (dlPFC) of patients with major depressive disorder (MDD) or health unaffected controls (Crtl). (B) Western blot of human (dlPFC) and mouse (mPFC) samples to show GPR158CT antibody specificity for human samples.
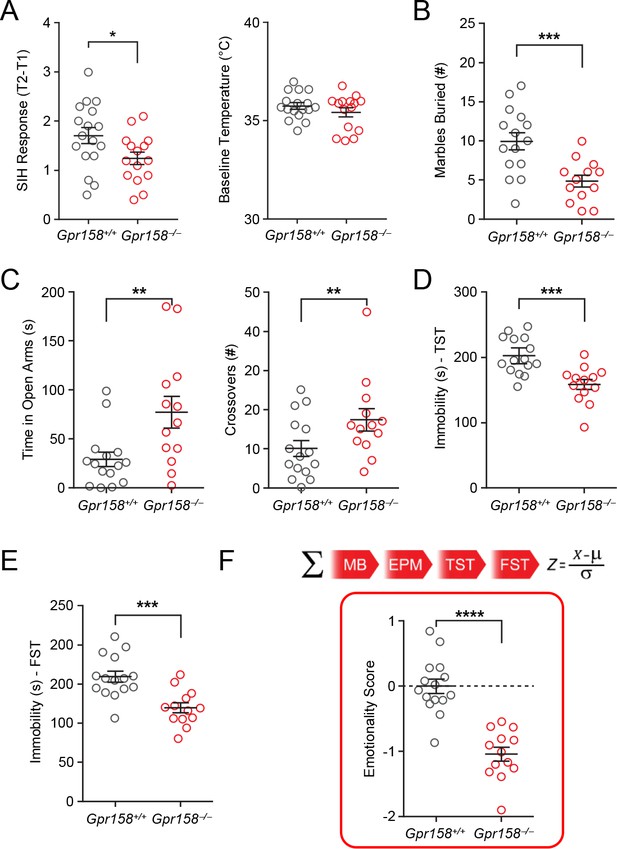
Ablation of GPR158 in mice results in an antidepressant-like phenotype and resiliency to chronic stress.
(A) Gpr158−/− mice display a lower SIH response (T2–T1) compared to Gpr158+/+ mice without affecting their baseline temperature (right panel) (n = 15–17/genotype, Student’s t test, *p<0.05). (B) Gpr158−/− mice buried fewer marbles compared to Gpr158+/+ mice in the marble burying test. (C) Gpr158−/− mice spent significantly more time in the open arm and entered more frequently in the open arm during the EPM test. Gpr158−/− mice displayed decreased immobility time in the (D) TST and (E) FST. (F) Emotionality scores were integrated from four behavioral tests (marble burying, elevated plus maze, tail suspension test and force swim test) and normalized to Gpr158+/+ mice. (G) Gpr158−/− mice had a lower emotionality score reflective of an antidepressant-like phenotype (right panel; Student’s t test). Data shown as means ± SEM (*p<0.05, **p<0.01, ***p<0.001, ****p<0.0001; ns = not significant).
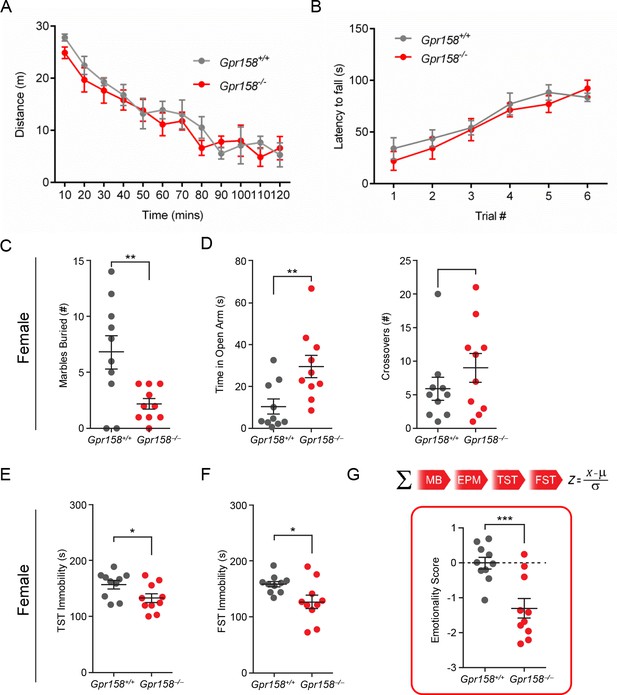
Depressive- and anxiety-related behaviors are altered in both male and female Gpr158−/− mice.
(A) Locomotor activity in 10 min bins (n = 5–6 mice/genotype). No significant difference between genotypes were found. (B) Effects of Gpr158−/− and Grp158+/+ mice on the accelerated rotarod test show no difference between genotype over six trials (n = 7 mice/genotype). (C) Female Gpr158−/− mice buried fewer marbles compared to Gpr158+/+ mice in the marble burying test. (D) Female Gpr158−/− mice spent significantly more time in the open arm and entered more frequently in the open arm during the elevated plus maze (EPM) test. Female Gpr158−/− mice displayed decreased immobility time in the (E) tail suspension test (TST) and (F) forced swim test (FST). (G) Emotionality score for Gpr158+/+ and Gpr158−/− female mice. Individual behavioral measures were integrated to calculate the emotionality score for each animal (top panel) (n = 10 mice/genotype). Data shown as means ± SEM (Student’s t test, *p<0.05, **p<0.01, ***p<0.001).
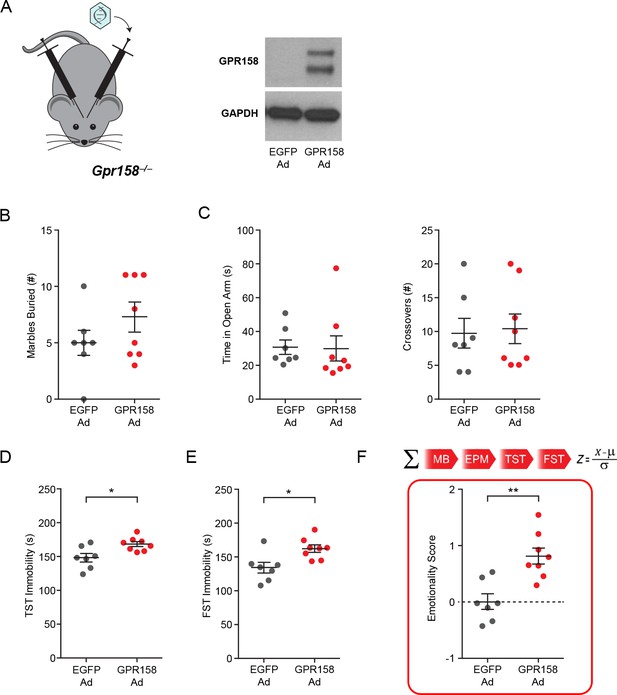
Viral-mediated expression of GPR158 in mPFC of Gpr158−/− mice rescues the antidepressant-like phenotype.
(A) mPFC of Gpr158−/− mice was injected bilaterally with Adenovirus expressing either EGFP or GPR158. Western blot showing the expression of GPR158 only in mice injected with GPR158-Ad (right panel). Gpr158−/− mice injected with EGFP-Ad or GPR158-Ad into the mPFC underwent the (B) marble burying test, (C) EPM, (D) TST and (E) FST. (F) Emotionality score was calculated using the individual behavioral measures from the MB, EPM, TST and FST. Data shown as means ± SEM (n = 7–8 mice/group, Student’s t test,*p<0.05, **p<0.01).
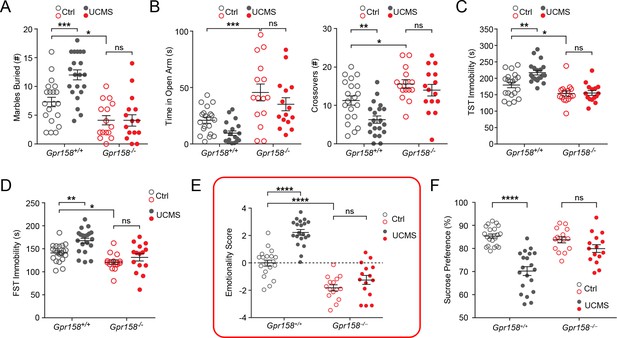
Gpr158−/− mice are resilient to unpredictable chronic mild stress (UCMS).
Individual behavioral measures were integrated to calculate the emotionality score for each animal. Behavioral analysis of Gpr158+/+ and Gpr158−/− mice subjected to UCMS in the (A) marble burying test, (B) elevated plus maze (EPM), (C) tail suspension test (TST) and (D) forced swim test (FST). (E) Emotionality score and (F) sucrose preference test for Gpr158+/+ and Gpr158−/− mice subjected to UCMS (n = 15–20 mice/group, Two-way ANOVA with Bonferroni post hoc test). Data shown as means ± SEM (n = 15–20 mice/group; Two-way ANOVA with Bonferroni post hoc; *p<0.05, **p<0.01, ***p<0.001, ****p<0.0001; ns, not significant).
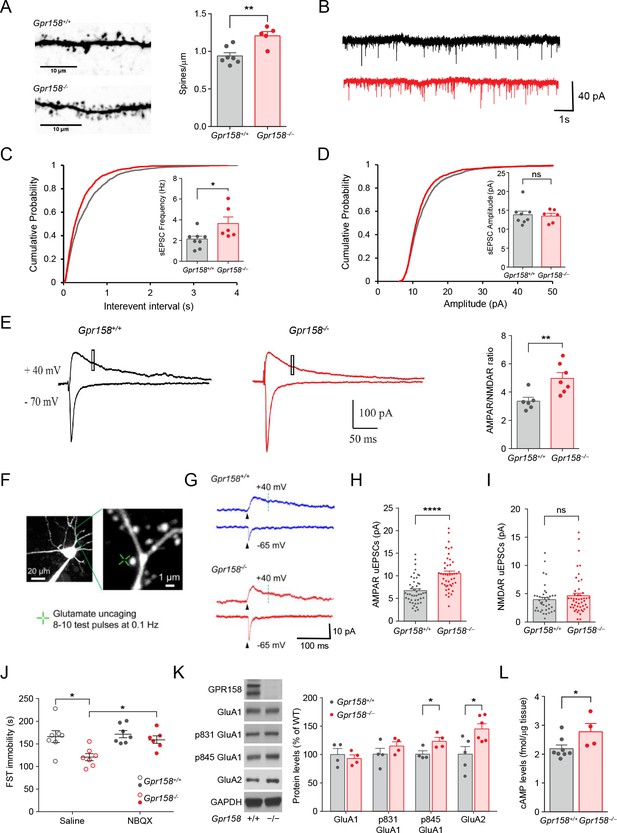
Synaptic adaptations in layer 2/3 neurons of mPFC induced by GPR158 loss.
(A) Representative images and summarized data showing that the spine density was increased in Gpr158−/− neurons (n = 7–5). (B) Representative traces and summarized data showing that the frequency (C) (n = 8–6 mice/genotype; Student’s t test), but not the amplitude (D) (n = 8–6 mice/genotype; Student’s t test), of sEPSC was increased in Gpr158−/− neurons. (E) Representative traces and summarized data showing that AMPAR/NMDAR current ratio was increased in Gpr158−/− neurons (n = 6–7 mice/genotype). The AMPAR component was measured as the maximal response while neurons were held at −70 mV. The NMDAR component was measured as the average current between 50–55 ms (square) following the stimulation while the neurons were held at +40 mV. (F) Representative image of a spine and dendrite showing location of glutamate uncaging. (G) AMPAR- and NMDAR-mediated uEPSCs at −65 and +40 mV, respectively. The green dotted line (70 ms after uncaging pulse) indicates the time at which the NMDAR-uEPSCs were measured. Summary of (H) AMPAR uEPSC (n = 47–48 neurons/genotype) and (I) NMDAR uEPSC (n = 40–47 neurons/genotype) amplitude are shown for each genotype. (J) FST for Gpr158−/− and Gpr158+/+ mice injected with NBQX (K) Representative western blots and quantification of total and phosphorylated levels of AMPAR subunits, GluA1 and GluA2 (n = 4–6). (L) Basal levels of cAMP in the mPFC of Gpr158+/+ and Gpr158−/− mice (n = 4–8 mice/genotype). Data are mean ± SEM (Student’s t test, *p<0.05, **p<0.01, ****p<0.0001, ns = not significant).
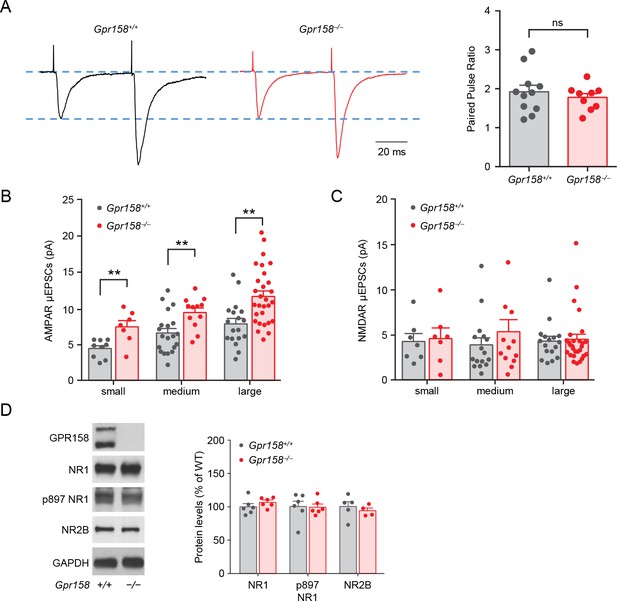
AMPA receptors are involved in the synaptic adaptations of Gpr158−/− mice.
(A) Representative current traces are shown and Paired pulse ratio comparison showing no significant differences in Gpr158+/+ and Gpr158−/− mice (n = 11–9 neurons/3 mice). (B) AMPAR and (C) NMDAR µESPCs amplitudes for each spine size in layer 2/3 of the prelimbic cortex for Gpr158+/+ and Gpr158−/− mice (n = 3 mice/group). (D) Representative western blot and quantification of total and phosphorylated levels of NMDAR subunits, NR1 and NR2B (n = 4–6). Data shown as means ± SEM (Student’s t test, **p<0.01, ns = not significant).
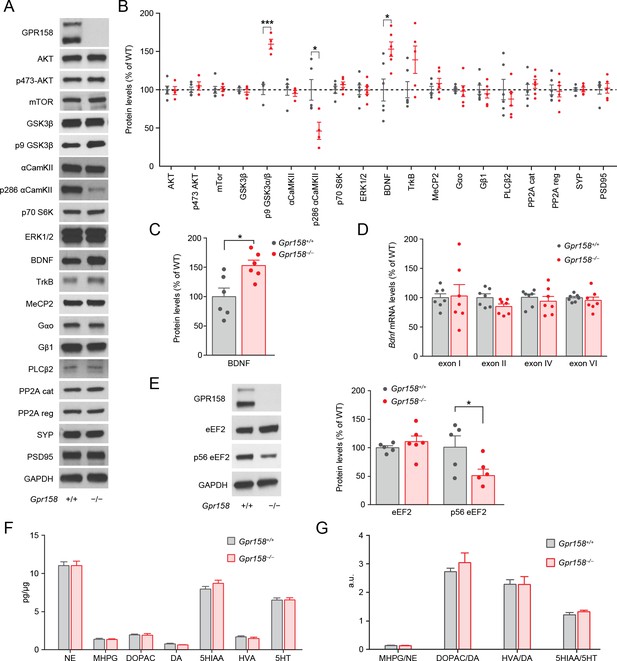
GPR158 modulates BDNF protein levels in the mPFC.
Representative western blots (A) and quantification (B) of the protein levels and phosphorylation state of the indicated proteins in the mPFC of Gpr158+/+ and Gpr158−/− mice (n = 4–6). (C) Western blot quantification of BDNF levels in the mPFC of Gpr158+/+ and Gpr158−/− mice (n = 6). (D) Comparison of mRNA levels of 4 BDNF splicing isoforms in the mPFC of Gpr158+/+ and Gpr158−/− do not show any significant difference. GPR158 levels were normalized against the ribosomal subunit 18S RNA (n = 7). (E) Western blot and quantification of total and phosphorylated levels of eEF2 (n = 5–6). (F) Quantitative analysis of the levels of norepinephrine (NE), 3-methoxy-4-hydroxyphenylglycol (MHPG), 3,4-dihydroxyphenylacetic acid (DOPAC), dopamine (DA), 5-hydroxyindoleacetic acid (5HIAA), homovanillic acid (HVA) and serotonin (5HT) in the mPFC of Gpr158+/+ and Gpr158−/− (G) Neurotransmitter turnover calculated as metabolite/neurotransmitter ratio. Data are mean ± SEM (Student’s t test, *p<0.05, ***p<0.001).
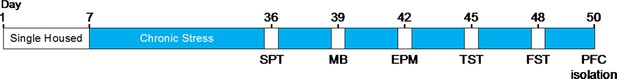
Timeline for behavioral evaluation in stress-induced depression models.
https://doi.org/10.7554/eLife.33273.016Tables
| Reagent type (species) or resource | Designation | Source or reference | Identifiers | Additional information |
|---|---|---|---|---|
| Antibody | GAPDH | Millipore | Cat# AB2302 | (1:30000) |
| Antibody | GluA1 | Abcam | Cat# ab76321 | (1:1000) |
| Antibody | p831 GluA1 | Millipore | Cat# 04–823 | (1:1000) |
| Antibody | p845 GluA1 | Cell Signaling | Cat# 8084 | (1:1000) |
| Antibody | GluA2 | Millipore | Cat# MAB N71 | (1:1000) |
| Antibody | NR1 | Zymed | Cat# 32–500 | (1:1000) |
| Antibody | p897 NR1 | Millipore | Cat# 06–641 | (1:1000) |
| Antibody | NR2B | Millipore | Cat# AB1557P | (1:1000) |
| Antibody | Rabbit anti-GPR158CT | PMID:25792749 | N/A | (1:1000) |
| Antibody | Rabbit anti-Gβ5 | PMID:15632198 | N/A | (1:1000) |
| Antibody | Rabbit anti-Gβ1 | gift from Dr. Willardson PMID:15485848 | N/A | 1:6000 |
| Antibody | GPR34 | ThermoFisher Scientific | Cat# PA5-45717 | (1:1000) |
| Antibody | GPR162 | Aviva Systems Biology | Cat# ARP68400 | (1:1000) |
| Antibody | GPRC5B | Aviva Systems Biology | Cat# ARP59799 | (1:1000) |
| Antibody | AKT | Cell Signaling | Cat# 9272 | (1:1000) |
| Antibody | p473 AKT | Cell Signaling | Cat# 4060 | (1:1000) |
| Antibody | mTOR | Cell Signaling | Cat# 2983 | (1:1000) |
| Antibody | S6 Kinase (P70) | Cell Signaling | Cat# 9202 | (1:1000) |
| Antibody | ERK1/2 | Cell Signaling | Cat# 9102 | (1:1000) |
| Antibody | GSK-3β | Cell Signaling | Cat# 9315 | (1:1000) |
| Antibody | p9 GSK3α/β | Cell Signaling | Cat# 9331 | (1:1000) |
| Antibody | BDNF | Abcam | Cat# ab108319 | (1:1000) |
| Antibody | TrkB | Cell Signaling | Cat# 4603 | (1:1000) |
| Antibody | MeCP2 | Millipore | Cat# 07–013 | (1:1000) |
| Antibody | Gαo | Cell Signaling | Cat# 3975 | (1:1000) |
| Antibody | PLCβ2 | Santa Cruz | Cat# sc206 | (1:200) |
| Antibody | PP2Acat | Assay Biotech | Cat# B0555 | (1:1000) |
| Antibody | PP2Areg | Cell Signaling | Cat# 2041 | (1:1000) |
| Antibody | SYP | Assay Biotech | Cat# C0333 | (1:1000) |
| Antibody | PSD95 | Cell Signaling | Cat# 3450 | (1:1000) |
| Antibody | eEF2 | Cell Signaling | Cat# 2332 | (1:1000) |
| Antibody | p56 eEF2 | Cell Signaling | Cat# 2331 | (1:1000) |
| Antibody | NeuN | Abcam | Cat# 104225 | (1:1000) |
| Antibody | GAD67 | Millipore | Cat# MAB5406 | (1:1000) |
| Genetic reagent (adenovirus) | HdAd-23E4-Pun-GPR158-syn-EGFP | This Paper | N/A | |
| Genetic reagent (adenovirus) | HdAd-23E4-Pun-Syn-EGFP | This Paper | N/A | |
| Biological sample (brain tissue) | Adult human brain tissue (healthy and MDD samples) | NIH NeuroBioBank (University of Pittsburg, University of Miami Brain Endowment Bank and the Human Brain and Spinal Fluid Resource Center) | 13138, 13054, 13082, 5289, 002, 006, 5293, AN04642, AN04917, AN07465, AN14368, AN15392, AN01077, AN05364, 13006, 13011, 13022, 13047, 13075, 13086, 13145, 13057, 13051, 13181, 535, 556, 711 | |
| Chemical compound, drug | Corticosterone | Sigma-Aldridge | Cat# C2505 | |
| Chemical compound, drug | RU-486 | Sigma-Aldridge | Cat# M8046 | |
| Chemical compound, drug | Dexamethasone | Tocris | Cat# D4902 | |
| Chemical compound, drug | NBQX | Tocris | Cat# 10441R | |
| commercial assay or kit | QuantiGene ViewRNA ISH Tissue 2-Plex Assay | Affymetrix | Cat# QVT0012 | |
| commercial assay or kit | Gpr158 NM_001004761; ViewRNA TYPE 1 Probe | Affymetrix | Cat# VB1-11518 | |
| commercial assay or kit | Slc17a7 NM _182993; ViewRNA TYPE 6 Probe | Affymetrix | Cat# VB6-14162 | |
| commercial assay or kit | Gad1 NM_008077; ViewRNA TYPE 6 Probe | Affymetrix | Cat# VB6-12632 | |
| commercial assay or kit | Pvalb NM_013645; ViewRNA TYPE 6 Probe | Affymetrix | Cat# VB6-13220 | |
| commercial assay or kit | Sst NM_009215; ViewRNA TYPE 6 Probe | Affymetrix | Cat# VB6-13754 | |
| commercial assay or kit | DapB NC_000913; ViewRNA TYPE 1 Probe | Affymetrix | Cat# VF1-10272 | |
| commercial assay or kit | Direct cAMP ELISA kit | ENZO Life Sciences | Cat# ADI-900–066 | |
| strain, strain background (mus musculus) | GPR158 KO | PMID:25792749 | N/A | |
| software, algorithm | ImageJ | Wayne Rasband, NIH, USA | SCR_003070 | |
| software, algorithm | Zen2.1 SP1 (Black) | Carl Zeiss | SCR_013672 | |
| software, algorithm | Prism6 | GraphPad Software | SCR_002798 | |
| software, algorithm | Neurolucida | MBF Bioscience |
Additional files
-
Supplementary file 1
G protein Coupled Receptors identified by mass-spectrometry in the PFC.
- https://doi.org/10.7554/eLife.33273.017
-
Transparent reporting form
- https://doi.org/10.7554/eLife.33273.018





