Discovery and characterization of a prevalent human gut bacterial enzyme sufficient for the inactivation of a family of plant toxins
Figures
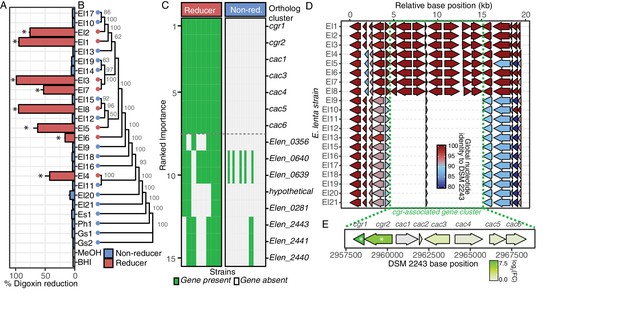
Comparative genomics expands the boundaries of the cgr operon.
(A) Survey of digoxin reduction in 21 strains of E. lenta (El#), 2 strains of Gordonibacter spp. (Gs#), E. sinensis (Es1), and Paraeggerthella hongkongesis (Ph1) (Figure 1—source data 1 and 2) revealed eight strains capable of reducing digoxin to dihydrodigoxin (*p<0.05, ANOVA with Dunnett’s test vs. vehicle controls). Data represents mean ± standard error of the mean (SEM) over three biological replicates. (B) Digoxin reduction did not correlate with phylogeny in E. lenta species (cladogram displayed with bootstrap values indicated at nodes; p=0.275, K = 0.049, Blomberg’s K). (C) Comparative genomics using a random forest classifier (see Materials and methods) revealed seven genes with perfect predictive accuracy for digoxin reduction. The orthologous cluster identified as hypothetical corresponds to an open reading frame present at position 299442..2995131 in the DSM 2243 reference genome. (D) Analysis of genomic context revealed a highly conserved 10.4 kb locus of 7 genes that flank a short, conserved hypothetical gene, herein termed the cgr-associated gene cluster (cac). (E) Analysis of gene expression in the cgr-associated gene cluster revealed only the cgr-locus was significantly upregulated by exposure to digoxin. * FDR < 0.1 (Figure 1—source data 3).
-
Figure 1—source data 1
Bacterial strains used in study.
Acronyms: ND (Not Detected), NA (Not Available), BHI (Brain Heart Infusion), BHIA (Brain Hearth Infusion with supplemented arginine), FAA (Fastidious Anaerobe Agar), GAM (Gifu Anaerobic Medium), DDMM (Dopamine Dehydroxylation Minimal Medium), Nag (Nutrient Agar).
- https://doi.org/10.7554/eLife.33953.004
-
Figure 1—source data 2
Digoxin Reduction by E.lenta and related bacterial isolates.
- https://doi.org/10.7554/eLife.33953.005
-
Figure 1—source data 3
Differential gene expression analysis of cgr-associated genes in E.lenta DSM 2243.
- https://doi.org/10.7554/eLife.33953.006
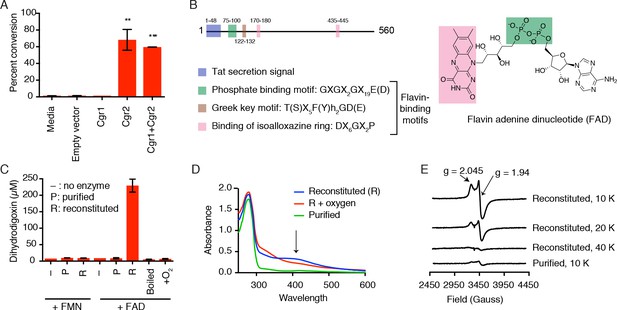
Cgr2 is sufficient for digoxin reduction and requires FAD and [4Fe-4S] cluster(s) for activity.
(A) Whole cell assays using R. erythropolis expressing Cgr1 and Cgr2 constructs demonstrated that Cgr2 is sufficient for reducing digoxin. Data represents the mean ± SEM (n = 3 biological replicates). Asterisks indicate statistical significance of each variant as compared to empty vector by Student’s t test (**p<0.01, ***p<0.001) (Figure 2—source data 1). (B) Annotation and amino acid numbering of Cgr2, including the predicted Tat secretion signal and three conserved flavin-binding motifs from the glutathione reductase family (X = any amino acid; h = hydrophobic residue). (C) In vitro activity of Cgr2 for digoxin reduction using reduced methyl viologen as an electron donor, analyzed by liquid chromatography-tandem mass spectrometry (LC-MS/MS). [Fe-S] cluster reconstitution, FAD, and anaerobic conditions are required for Cgr2 activity. Data represents the mean ± SEM (n = 3 independent experiments) (Figure 2—source data 2). FAD = flavin adenine dinucleotide; FMN = flavin mononucleotide. (D) Ultraviolet-visible (UV-Vis) absorption spectra of Cgr2 revealed an oxygen-sensitive peak centered around 400 nm that increased upon [Fe-S] cluster reconstitution, supporting the presence of [4Fe-4S] clusters in Cgr2. (E) Electron paramagnetic resonance (EPR) spectra of sodium dithionite-reduced Cgr2 reconstituted with iron ammonium sulfate hexahydrate ((NH4)2Fe(SO4)2·6H20) and sodium sulfide (Na2S·9H20). G-values and decreased EPR signal intensity at higher temperatures (10 – 40 K) indicated the presence of low potential [4Fe-4S]1+ clusters. Experimental conditions were microwave frequency 9.38 GHz, microwave power 0.2 mW, modulation amplitude 0.6 mT, and receiver gain 40 dB.
-
Figure 2—source data 1
Digoxin metabolism by R. erythropolis overexpressing Cgr proteins.
- https://doi.org/10.7554/eLife.33953.011
-
Figure 2—source data 2
Digoxin metabolism by Cgr2 in vitro.
- https://doi.org/10.7554/eLife.33953.012
-
Figure 2—source data 3
Examples of [2Fe-2S], [3Fe-4S], and [4Fe-4S] cluster binding motifs that are not found in Cgr2 (Zhang et al., 2010; Nakamaru-Ogiso et al., 2002; Lee et al., 2004; Pandelia et al., 2011; Schnackerz et al., 2004; Leech et al., 2003; Gorodetsky et al., 2008; Lee et al., 2010; Weiner et al., 2007; Klinge et al., 2007; Dickert et al., 2002; Conover et al., 1990; Schneider and Schmidt, 2005; Iwasaki et al., 2000; Banci et al., 2013; Dailey and Dailey, 2002; Jung et al., 2000).
- https://doi.org/10.7554/eLife.33953.013
-
Figure 2—source data 4
Digoxin metabolism by R. erythropolis overexpressing Cgr2 cysteine to alanine point mutants.
- https://doi.org/10.7554/eLife.33953.014
-
Figure 2—source data 5
Digoxin metabolism by Cgr2 cysteine to alanine point mutants in vitro.
- https://doi.org/10.7554/eLife.33953.015
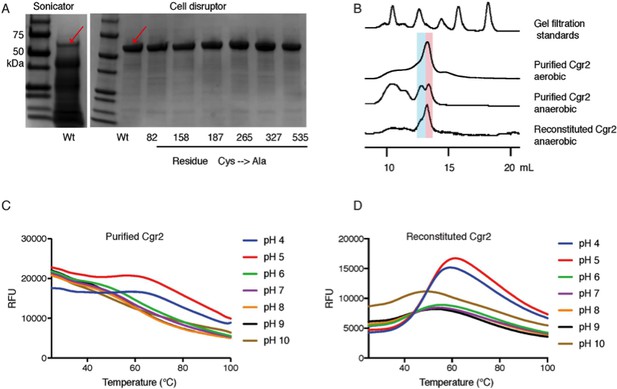
[Fe-S] cluster(s) affect Cgr2 stability and oligomerization.
(A) SDS-PAGE analysis of heterologously expressed Cgr2(–48aa)-NHis6 constructs (expected mass = 55 kDa) purified on HisPur Ni-NTA resin. Heat-generating lysis methods (e.g. sonication) led to substantial protein degradation as compared to cell disruption. (B) Analytical fast protein liquid chromatography (FPLC) performed under aerobic and anaerobic conditions. Colored bars highlight molecular weights corresponding to dimeric (blue) or monomeric (pink) Cgr2. (C) Thermal melt curves displaying relative fluorescence of Sypro Orange bound to purified and (D) reconstituted Cgr2 in various pH buffers. Protein melting temperature (Tm) of purified protein was < 37°C before reconstitution and increased to 40 – 50°C after reconstitution.
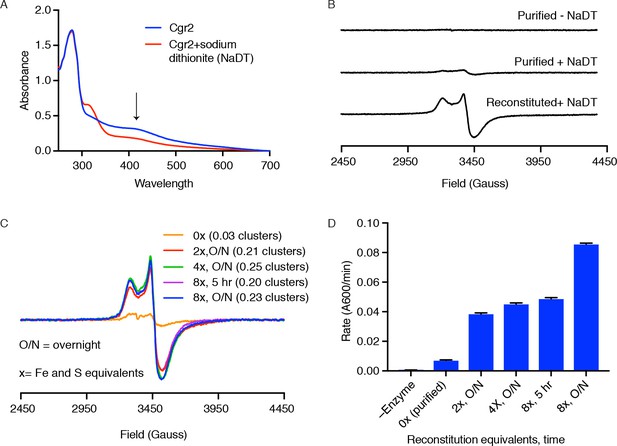
Cgr2 activity, but not EPR-active [4Fe-4S] clusters, increase with higher Fe and S equivalents.
(A) Ultraviolet-visible (UV-Vis) spectra of reconstituted Cgr2 in the absence or presence of reducing agent sodium dithionite (NaDT) revealed that the [Fe-S] clusters in Cgr2 are redox active. (B) Purified Cgr2 did not exhibit a detectable EPR signal. Upon reduction with NaDT, an EPR signal corresponding to [4Fe-4S]1+ clusters was detected in purified Cgr2. This signal was amplified in reconstituted Cgr2, showing incorporation of additional [4Fe-4S] cluster(s). Samples contained 200 µM protein, 0.2 mM sodium dithionite, and EPR measurements were conducted at 10 K. (C) EPR spectra of dithionite-treated Cgr2 samples that had been reconstituted with 0 (purified), 2, 4, or 8 equivalents of iron and sulfide for 5 hours or overnight (O/N). Samples contained 150 µM protein, 0.3 mM sodium dithionite, and measurements were conducted at 10 K. Number of EPR-active clusters per Cgr2 monomer under each reconstitution condition is shown in parentheses. Spin quantitation was determined against a 150 µM Cu2+-EDTA standard measured under non-saturating conditions. (D) In vitro reaction rates of Cgr2 reconstituted under different conditions revealed increasing activity with higher reconstitution equivalents. Data represents mean ± SEM (n = 3 independent experiments).
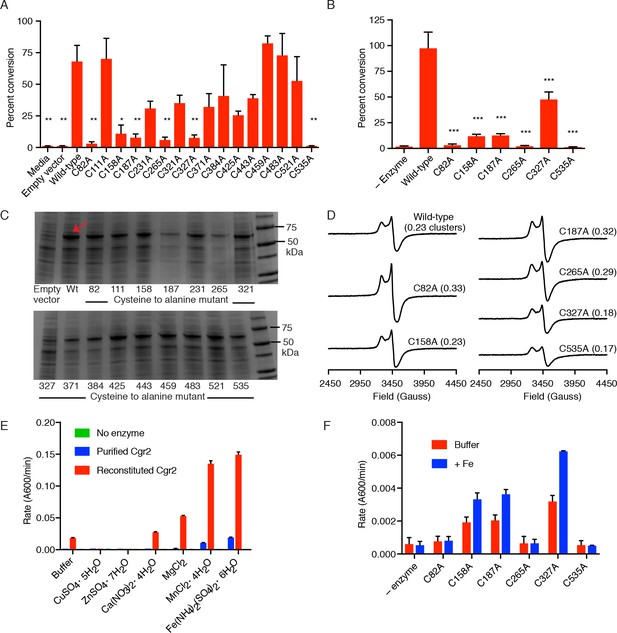
Identification of 6 cysteine residues important for Cgr2 activity.
(A) Whole cell assays of R. erythropolis expressing individual cysteine to alanine point mutants and incubated with digoxin demonstrated that six cysteine residues are important for activity. Data represents mean ± SEM (n = 3 biological replicates). Asterisks indicate statistical significance of each variant compared to wild-type Cgr2 by Student’s t test (*p<0.05, **p<0.01) (Figure 2—source data 4). (B) Mass spectrometry analysis of in vitro assays (quenched at 15 min) containing reconstituted wild-type Cgr2 or single cysteine to alanine point mutants. Data represents mean ± SEM (n = 3 independent experiments). Asterisks indicate statistical significance of each variant compared to wild-type Cgr2 by Student’s t test (***p<0.001) (Figure 2—source data 5). (C) SDS-PAGE analysis of clarified lysate from R. erythropolis cells transformed with empty pTipQC vector or expressing cytoplasmic wild-type Cgr2 (wt) or individual cysteine to alanine point mutants (~55 kDa). All point mutants were soluble. (D) [4Fe-4S]1+ clusters were detected by EPR in all Cgr2 point mutants treated with sodium dithionite. Spin quantitation against a Cu2+-EDTA standard revealed similar levels of [4Fe-4S]1+ clusters per Cgr2 monomer for all variants. Number of clusters shown in parentheses. (E) Divalent metal cations (Fe2+, Mg2+, Mn2+) stimulated the activity of Cgr2 in vitro. Data represents mean ± SEM (n = 3 independent experiments). (F) Addition of Fe2+ stimulated the activity of three cysteine residues, potentially implicating C92, C265, and C535 in metal binding. Data represents mean ± SEM (n = 3 independent experiments).
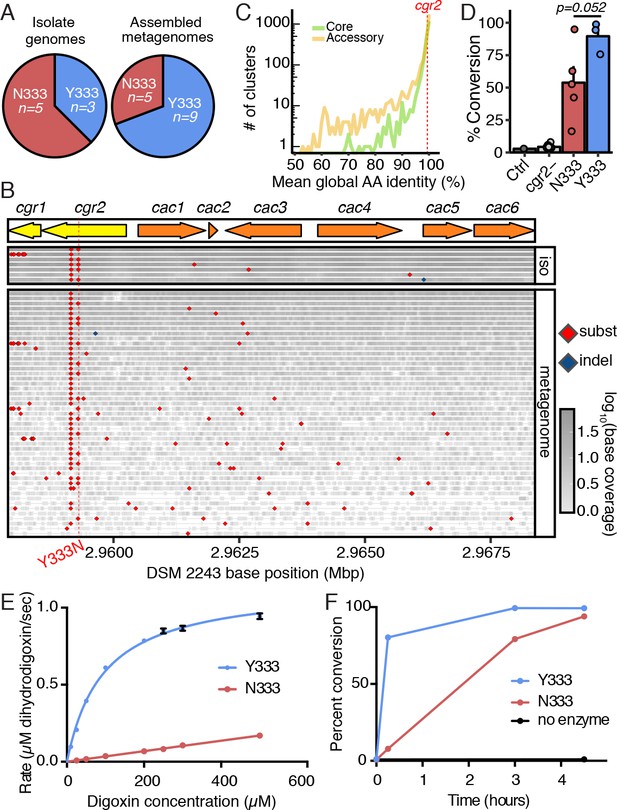
A single polymorphism in Cgr2 at position 333 (Y/N) leads to altered metabolism of digoxin.
(A) Analysis of Cgr2 amino acid sequence composition of isolate genomes (n = 8) and reconstructed sequences from gut microbiome datasets (n = 14) revealed a single non-conservative Y333N variant in isolate strains supported by metagenomes (12Y/10N). (B) Nucleotide variation in the cgr-associated gene cluster. Reads aligned from both isolate genomes (iso) and metagenomes were aligned to the DSM 2243 reference assembly and plotted if there was coverage of the Y333N variant position (CP001726.1: 2959294 bp). Variants were called when at least one read was mapped to the position and > 50% of reads supported an alternative base. Read depth at any given position is indicated by shading. Confirming assembly-based methods, cgr2 amino acid position 333 was bi-allelic with 5 of 8 isolate genomes and 15 of 49 metagenomes showing the N333 variant (four metagenomes have evidence of both alleles) and minimal variation in other regions of the cluster. (C) Average amino acid conservation in the E. lenta core (n = 1832) and non-singleton accessory genome (n = 2557) demonstrates that cgr2 is in the 67th percentile for conservation in the pan-genome (78.8th in the core genome, and 58.5th in the non-singleton accessory genome) with higher average conservation observed in the core genome (98.6 ± 2.41% core, 97.4 ± 5.6% accessory, mean ± SD). (D) Comparison of digoxin metabolism in culture by E. lenta cgr2- (n = 13 strains), Cgr2Y333 (n = 3 strains), and Cgr2N333 (n = 5 strains). Control refers to digoxin in BHI media. Each point represents the mean percent conversion to dihydrodigoxin of each individual strain cultured in biological triplicate. Bars represent the mean ± SEM percent conversion per E. lenta group. Statistical significance between Y333 and N333 groups was calculated using two-tailed Welch’s t test (p=0.052) (Figure 3—source data 1). (E) Michaelis–Menten kinetics of Cgr2 towards digoxin revealed that the Y333 variant is significantly more active than the N333 variant. Data represents mean ± SEM (n = 3 independent experiments) (Figure 3—source data 2; Figure 3—source data 4). (F) In vitro time course (0 – 4.5 hr) of the conversion of digoxin to dihydrodigoxin by Cgr2 Y333 and N333 variants. Reaction aliquots were quenched in methanol and analyzed by liquid chromatography-tandem mass spectrometry. Values represent mean ± SEM (n = 3 independent experiments). Asterisks indicate statistical significance at each timepoint of Y333 vs. N333 percent conversion, by Student’s t test (*p<0.05, ***p<0.001) (Figure 3—source data 3).
-
Figure 3—source data 1
Whole cell activity of E. lenta strains with Y333 vs N333 Cgr2 variants.
- https://doi.org/10.7554/eLife.33953.017
-
Figure 3—source data 2
Kinetics of Y333 and N333 Cgr2 variants towards digoxin.
- https://doi.org/10.7554/eLife.33953.019
-
Figure 3—source data 3
In vitro time course of digoxin reduction by Y333 and N333 Cgr2 variants.
- https://doi.org/10.7554/eLife.33953.020
-
Figure 3—source data 4
Comparison of Cgr2 kinetics with related enzymes towards their respective substrates (Kemp et al., 2010; Rohman et al., 2013; Morris et al., 1994; Bogachev et al., 2012).
- https://doi.org/10.7554/eLife.33953.021
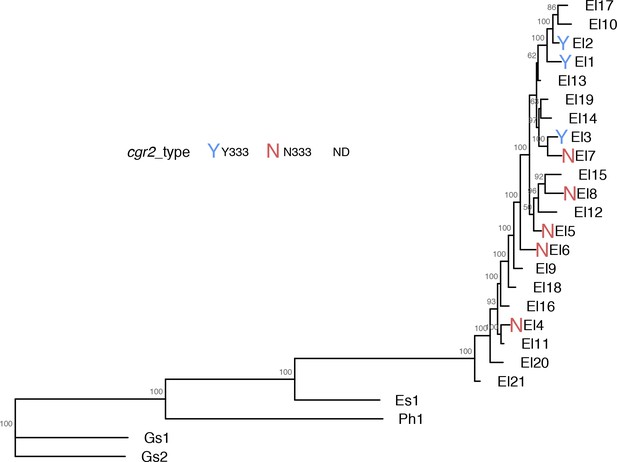
Phylogenetic tree of assayed E. lenta strains showing cgr2 Y333/N333 variants.
https://doi.org/10.7554/eLife.33953.018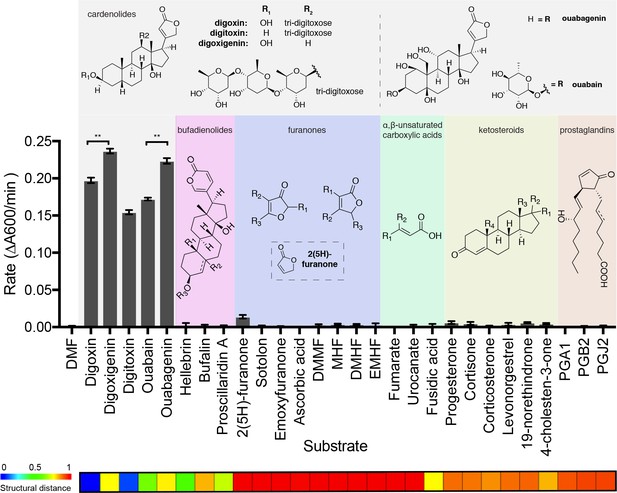
The substrate scope of Cgr2 is restricted to cardenolides.
Rate of methyl viologen oxidation coupled to substrate reduction by Cgr2. Colors denote different substrate classes. With the exception of the cardenolides, a representative substrate structure is shown. Values represent mean ± SEM (n = 3 independent experiments). **p<0.01, Student’s t test (Figure 4—source data 1). The heatmap generated in ChemMine (Backman et al., 2011) represents the structural similarity of each compound relative to digoxin. Structural distance matrix is calculated as (1- Tanimoto coefficient), where lower values represent more structurally similar compounds.
-
Figure 4—source data 1
Rate of methyl viologen oxidation coupled to substrate reduction by Cgr2.
- https://doi.org/10.7554/eLife.33953.025
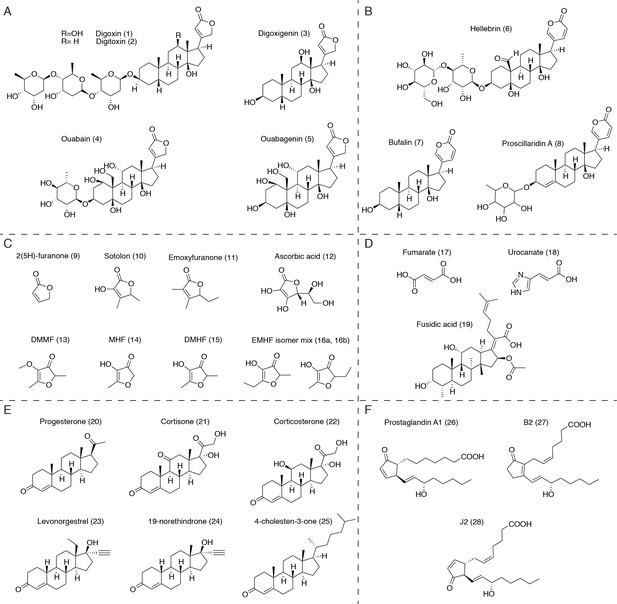
Putative substrates for Cgr2 in the context of the human gut.
(A) Plant-derived cardenolides. (B) Bufadienolides. (C) Dietary furanones, including sotolon (4,5-dimethyl-3-hydroxy-2,5-dihydrofuran-2-one), emoxyfuranone (5-ethyl-3-hydroxy-4-methyl-2(5 hr)-furanone), DMMF (2,5-dimethyl-4-methoxy-3(2 hr)-furanone), MHF (4-hydroxy-5-methyl-3-furanone), DMHF (4-Hydroxy-2,5-dimethyl-3(2 hr)-furanone), and EMHF (5-ethyl-4-hydroxy-2-methyl-3(2 hr)-furanone). (D) α,β-unsaturated carboxylic acids, including the antibiotic fusidic acid and substrates of similar bacterial reductases. (E) Ketosteroids, including naturally occurring hormones, synthetic steroid drugs, and putative cholesterol metabolites. (F) Unsaturated prostaglandins involved in host inflammation.
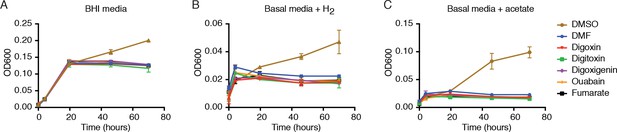
Digoxin and related cardenolides do not affect E. lenta growth in rich or minimal media.
E. lenta DSM 2243 growth in (A) rich media or in basal media lacking terminal electron acceptors and including (B) 5% H2 or (C) 10 mM sodium acetate as electron donors. Cells were grown in 10 mL of media supplemented with 10 µM of each substrate or an equivalent volume (0.1% v/v) of solvent (DMSO and DMF) at 37°C. DMSO serves as a positive control to show that E. lenta is capable of anaerobic respiration. Data represents mean ± SEM (n = 3).
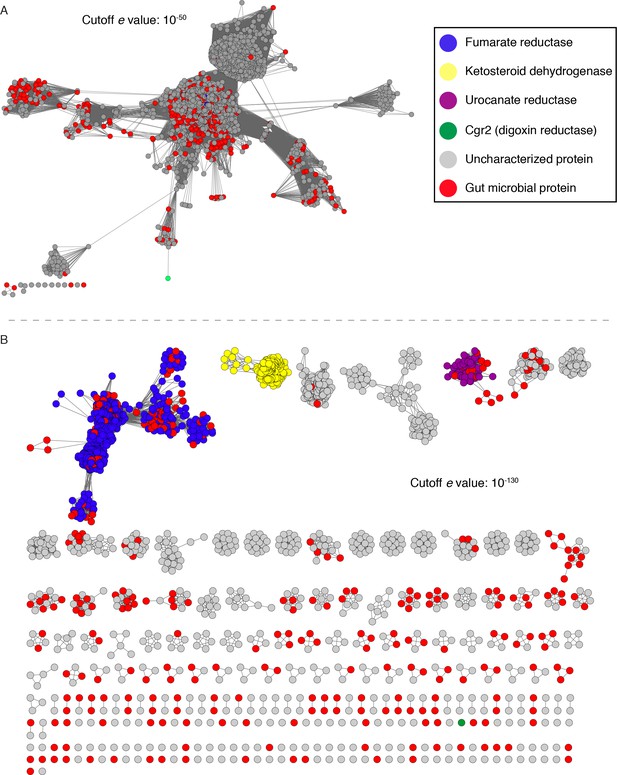
Sequence similarity network (SSN) analysis reveals that the gut bacterial enzyme Cgr2 is a highly distinct member of a large enzyme family that is widespread in gut microbes.
The SSN was constructed using the top 5000 most similar proteins to Cgr2 from the UniprotKB database. Nodes represent proteins with 100% sequence identity. (A) SSN displayed with an e-value threshold of 10−50. The seven previously characterized enzymes (PDB ID: 1D4D, 1E39; UniProtKB ID: Q07WU7, Q9Z4P0, 8CVD0, P71864, Q7D5C1) and Cgr2 are colored according to biochemical function. (B) SSN displayed with an e-value threshold of 10−130. All nodes that co-clustered with characterized enzymes are shown in the same color, denoting putative isofunctional activity. With the exception of Cgr2, if a node comes from a gut bacterium, it is colored red rather than the color of the corresponding cluster.
-
Figure 5—source data 1
Characterized enzymes within the Cgr2 sequence similarity network.
- https://doi.org/10.7554/eLife.33953.031

Multiple sequence alignment of fumarate reductases.
UniProtKB ID numbers are shown in parentheses. Active site residues (marked with an asterisk) were conserved in characterized fumarate reductases and clustered proteins from the sequence similarity network. These residues were not conserved in Cgr2 and another predicted fumarate reductase (Cac4) associated with the cgr gene cluster.
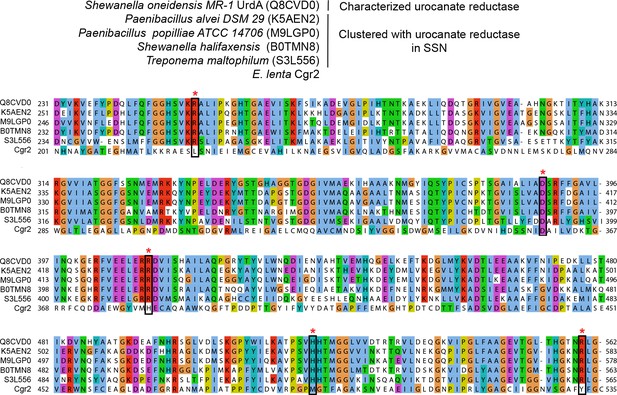
Multiple sequence alignment of urocanate reductases.
UniProtKB ID numbers are shown in parentheses. Active site residues (marked with an asterisk) were conserved in characterized urocanate reductases and clustered proteins from the sequence similarity network and were not conserved in Cgr2.
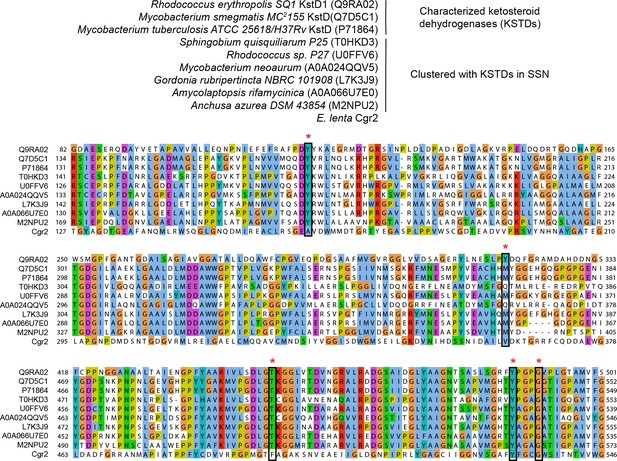
Multiple sequence alignment of ketosteroid dehydrogenases.
UniProtKB ID numbers are shown in parentheses. Active site residues (marked with an asterisk) were conserved in characterized ketosteroid dehydrogenases and clustered proteins from the sequence similarity network. Two residues involved in substrate binding and activation were conserved in Cgr2 (Y532, G536).
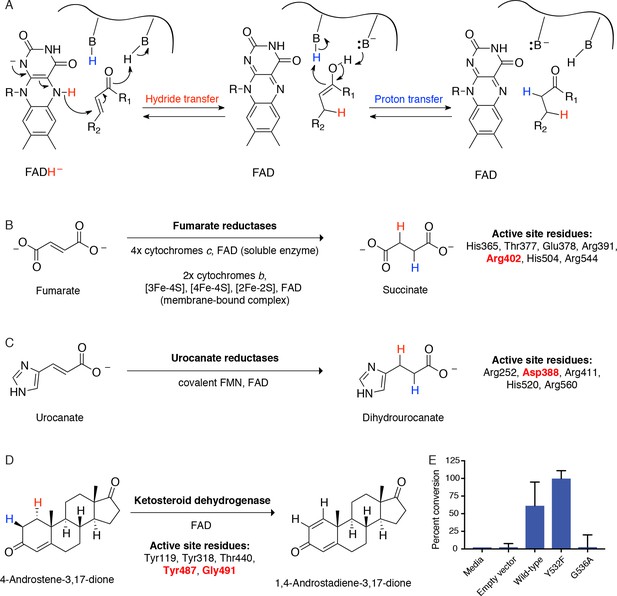
Cgr2 is a distinct flavoprotein reductase.
(A) General mechanism of catalysis by Cgr2 homologs. Cgr2 appeared to lack most of the conserved active site residues found in the most similar related enzymes, including (B) 6/7 residues utilized by fumarate reductases, (C) 4/5 residues utilized by urocanate reductases, and (D) 3/5 residues utilized by ketosteroid dehydrogenases. Active site residues are shown with numbering based on S. putrefaciens fumarate reductase, S. oneidensis MR-1 urocanate reductase, and R. erythropolis SQ1 ketosteroid dehydrogenase. Residues shown in red were conserved in Cgr2. (E) Two residues involved in substrate binding and activation in ketosteroid dehydrogenases were conserved in Cgr2 (Y532, G536). Whole cell assays in R. erythropolis overexpressing putative active site mutants in Cgr2 showed that Y532 was not essential for Cgr2 activity towards digoxin. Data represents mean ± SEM (n = 3 biological replicates).
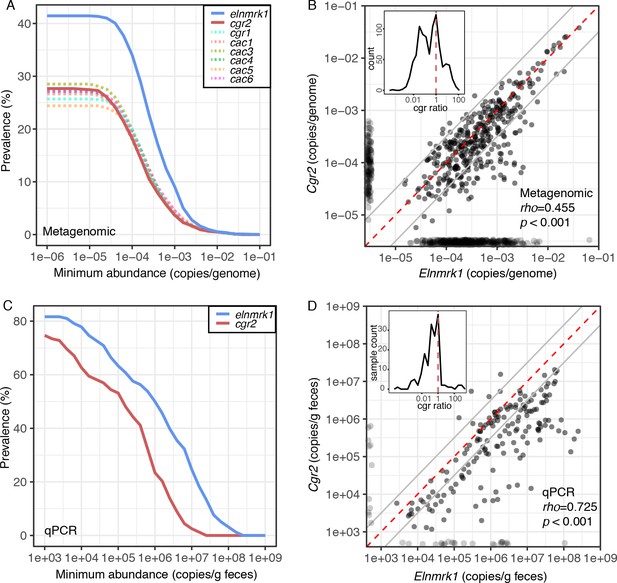
Cgr2 is widespread in the human gut microbiome.
(A) Analysis of the cgr-associated gene cluster and E. lenta (via elenmrk1) prevalence in the gut metagenomes of 1872 individuals (see Materials and methods) revealed that both E. lenta and cgr2 are highly prevalent (41.5% and 27.7% respectively) but frequently low in abundance. (B) Quantification of E. lenta and cgr2 abundances in individual gut metagenomes revealed a tight correlation between the two, providing evidence that cgr2 is restricted to E. lenta and that individuals may harbor sub-populations of both cgr2+ and cgr2- strains. Red line denotes the expected linear relationship and dashed lines represent a ± half log deviation. (Inset) Histogram of cgr-ratio (cgr/elnmrk1) demonstrates a significant skew away from communities that would have more cgr2 than expected by E. lenta abundance (p<0.001, D’Agostino skewness test). (C) Replication in an additional 158 individuals located in the USA (n = 85) and Germany (n = 73) via duplexed qPCR increased prevalence estimates to 74.7% and 81.6% at the extremes of detection limit (1e3 copies/g). qPCR samples were run in technical triplicate (Figure 6—source datas 1 and 2). (D) Similarly, qPCR-derived abundances of E. lenta and cgr2 were correlated, corroborating metagenome-based analysis. (Inset) Histogram of cgr-ratio demonstrating significant skew (p<0.001).
-
Figure 6—source data 1
qPCR based E. lenta and cgr2 prevalence.
Prevalence based on median of abundance in individuals with repeated sampling.
- https://doi.org/10.7554/eLife.33953.033
-
Figure 6—source data 2
Replicate qPCR data of for cgr2 and elnmrk1 in human fecal samples.
- https://doi.org/10.7554/eLife.33953.034
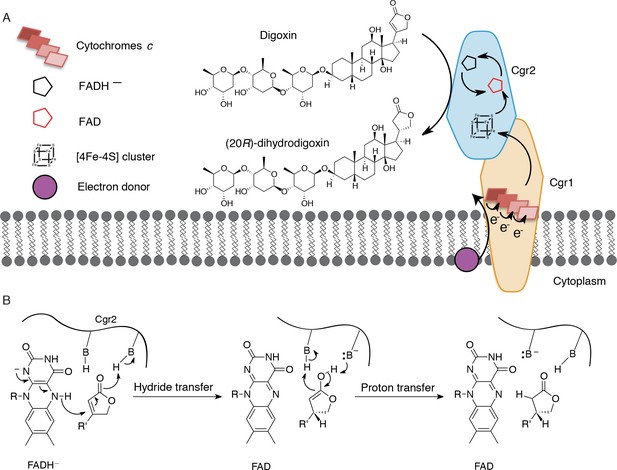
Preliminary model for digoxin metabolism by Cgr1 and Cgr2.
(A) Proposed biochemical model and (B) mechanism of digoxin reduction by Cgr proteins. Cgr1 is predicted to transfer electrons from a membrane-associated electron donor to the [4Fe-4S]2+ cluster of Cgr2 via covalently bound heme groups. The reduced [4Fe-4S]1+ cluster of Cgr2 could sequentially transfer two electrons to FAD, generating FADH–, which could mediate hydride transfer to the β-position of the digoxin lactone ring. Protonation of the resulting intermediate would yield (20R)-dihydrodigoxin.
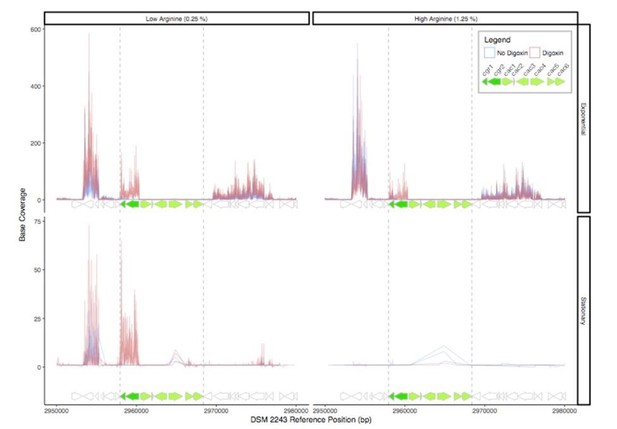
RNA-Seq read depth across the cgr-associated gene cluster and neighboring regions.
While the cgr-operon is induced in the presence of digoxin in exponential phase and stationary phase under low arginine conditions, the remainder of the cluster is relatively transcriptionally dormant, with the exception of a small degree of transcription of the cac4 reductase independent of digoxin in stationary phase.

Clustal Omega alignment of Sanger-sequenced cgr2 confirms high degree of conservation.
https://doi.org/10.7554/eLife.33953.041
Variant calling within the cgr-associated gene cluster.
Mapping of reads to the reference assembly and calling of variants confirms assembly-based analysis wherein an average of 4.14 variants are called in the cluster (median = 3, range 2-14).
Tables
| Reagent type (species) or resource | Designation | Source or reference | Identifiers | Additional information |
|---|---|---|---|---|
| Gene (Eggerthella lenta) | cgr1 | PMCID: PMC3035228 | cgr1 | |
| Gene (Eggerthella lenta) | cgr2 | PMCID: PMC3035228 | cgr2 | |
| Gene (Eggerthella lenta) | 16S rRNA | Ref 11 | ||
| Gene (Eggerthella lenta) | E. lenta marker gene | Ref 11 | elnmrk1 | |
| Strain, strain background (Escherichia coli) | One Shot Top10 | Thermo Fisher Scientific | ||
| Strain, strain background (Rhodococcus erythropolis) | L88 | doi: 10.1128/JB.187.8.2582–2591.2005 | ||
| Strain, strain background (Eggerthella lenta) | Eggerthella lenta strains | Ref 11 | El1 - El21 | see Figure 1—source data 1 for full descriptions |
| Strain, strain background (Eggerthella sinensis) | Eggerthella sinensis DSM16107 | Ref 11 | Es1 | |
| Strain, strain background (Gordonibacter) | Gordonibacter strains | Ref 11 | Gs1, Gs2 | |
| Strain, strain background (Paraeggerthella hongkongensis) | Paraeggerthella hongkongensis | Ref 11 | Ph1 | |
| Recombinant DNA reagent | pTip expression vectors | doi: 10.1128/AEM.70.9.5557–5568.2004 | ||
| Sequence-based reagent | Amplification cgr1, cgr2 | Integrated DNA technologies | see Table 1 for primers and constructs | |
| Sequence-based reagent | Point mutants of Cgr2 | Integrated DNA technologies | see Table 2 for primers and mutants | |
| Sequence-based reagent | cgr2 sequencing primer | This work | Confirmed sequence of E. lenta isolates with primers: cgr2_fwd (TGCAATCAAGACAACCACGA), cgr2_internal (TCGGTGTACAACCACAATGC), and cgr2_rev (GTTGCGCTGTGATTAGACTG) | |
| Sequence-based reagent | qPCR primers cgr2 | This work | cgr2_F (GAGGCCGTCGATTGGATGAT), cgr2_R (ACCGTAGGCATTGTGGTTGT), and cgr2_probe ([HEX]CGACACGGAGGCCGATGTCG[BHQ1]) | |
| Sequence-based reagent | qPCR primers elnmrk | This work | ElentaUni_F (GTACAACATGCTCCTTGCGG), ElentaUni_R (CGAACAGAGGATCGGGATGG), ElentaUni_Probe ([6FAM]TTCTGGCTGCACCGTTCGCGGTCCA[BHQ1]), | |
| Chemical compound, drug | BBL Brain Heart Infusion (BHI) media | Becton Dickinson | BD:L007440 | |
| Chemical compound, drug | L-arginine | Sigma Aldrich | SA:A5006 | |
| Chemical compound, drug | Digoxin | Sigma Aldrich | SA:D6003 | |
| Chemical compound, drug | Dihydrodigoxin | doi: 10.1126/science.1235872 | ||
| Chemical compound, drug | Digitoxin | Sigma Aldrich | SA:D5878 | |
| Chemical compound, drug | Digoxigenin | Sigma Aldrich | SA:D9026 | |
| Chemical compound, drug | Ouabain | Sigma Aldrich | SA:O3125 | |
| Chemical compound, drug | Ouabagenin | Sigma Aldrich | SA:O2627 | |
| Chemical compound, drug | Sypro Orange protein gel stain | Thermo Fisher Scientific | SA:S6650 | |
| Chemical compound, drug | Thiostrepton | Sigma Aldrich | SA:T8902 | |
| Chemical compound, drug | Methyl viologen | Sigma Aldrich | SA:856177 | |
| Chemical compound, drug | Sodium dithionite | Sigma Aldrich | SA:157953 | |
| Chemical compound, drug | FAD | Sigma Aldrich | SA:F6625 | |
| Chemical compound, drug | Iron (II) ammonium sulfate hexahydrate | Sigma Aldrich | SA:F1543 | |
| Chemical compound, drug | Sodium sulfide nonahydrate | Sigma Aldrich | SA:208043 | |
| Chemical compound, drug | Dithothreitol | Sigma Aldrich | SA:D0632 | |
| Chemical compound, drug | HEPES | Sigma Aldrich | SA:RDD002 | |
| Software, algorithm | Cytoscape | doi:10.1101/gr.1239303 | ||
| Software, algorithm | Prism software | Prism software | Graphpad Software v 7 | |
| Other | Anaerobic chambers | Coy Laboratory products; Mbraun | ||
| Other | LC-MS/MS | Agilent | Agilent:6410 Triple Quad LC/MS | |
| Other | Electron paramagentic resonance (EPR) spectrometer | Bruker | ||
| Other | CFX96 Touch Real-Time PCR machine | Bio-Rad | ||
| Other | PowerWave HT Microplate Spectrophotometer | BioTek |
Primers and constructs for heterologous expression of Cgr proteins in R. erythropolis.
Restriction sites are bolded.
| Construct | For/ | Sequence | Restriction sites | Vector | Anneal temp (°C) | Extend time (s) |
|---|---|---|---|---|---|---|
| Rev | ||||||
| cgr operon | For | ACTGACCCATGGATGGAATACGGAAAGTGCC | n/a | n/a | 71 | 75 |
| Rev | GTTTTACTGCAGTTACGCCGCCGTCGAA | |||||
| Cgr1 + Cgr2 | For | TGACGAATTCTAATGGAATACGGAAAGTGCCG | EcoRI, | pTipQT2 | 70 | 90 |
| Rev | TTATAAGATCTCGCCGCCGTCGAAAG | BglII | ||||
| Cgr1 | For | TCGAACATATGATGGCTGAGGAACCTGTGG | NdeI, | pTipQT1 | 65 | 60 |
| Rev | ATAACTCGAGTCACGCCGCCGTCGAAA | XhoI | ||||
| Cgr2 (native) | For | ACTGACCCATGGGCATGGAATACGGAAAGTGCC | NcoI, HindIII | pTipQC2 | 65 | 60 |
| Rev | ATTAGAAGCTTTCACTCCCACGGCTCGAG | |||||
| Cgr2-CHis6 (native) | For | ACTGACCCATGGGCATGGAATACGGAAAGTGCC | NcoI, HindIII | pTipQC1 | 65 | 60 |
| Rev | GTTAGAAGCTTCTCCCACGGCTCGAG | |||||
| Cgr2(−48aa)-NHis6 | For | TATTACCATGGATCAGACCGCGCCTGC | NcoI, HindIII | pTipQC2 | 65 | 60 |
| Rev | ATACTAAGCTTCTCCCACGGCTCGA | |||||
| Cgr2(−48aa)-CHis6 | For | TATTACCATGGATCAGACCGCGCCTGC | NcoI, HindIII | pTipQC1 | 65 | 60 |
| Rev | ATACTAAGCTTTTACTCCCACGGCTCGA | |||||
| Sequencing primers | For | CGTGGCACGCGGAAC | n/a | All pTip vectors | n/a | n/a |
| Rev | GTGCAGGTTTCGCGTG |
Primers for site-directed mutagenesis of Cgr2.
Amino acid numbering is based on full length Cgr2 sequence. Introduced mutations are bolded.
| Mutant | F/R | Sequence |
| C82A | For | CAGCGGCGGCACGGCCGCGGCCATCG |
| Rev | CCTCGATGGCCGCGGCCGTGCCGCCG | |
| C111A | For | GCGGCAACTCGGCACTAGCCGGTGGATACAT |
| Rev | CCAGCATGTATCCACCGGCTAGTGCCGAGTTG | |
| C158A | For | ATATGATCCGCGAGGCGGCCTTGCGCTCCGGC |
| Rev | GCCTCGCCGGAGCGCAAGGCCGCCTCGCGGAT | |
| C187A | For | GCCCCCGGTCTGGTCAGCCGGCGACACGG |
| Rev | GGCCTCCGTGTCGCCGGCTGACCAGACCGG | |
| C231A | For | CGAAATCGAGATGGGCGCCGAGGTGGCGCAC |
| Rev | GATGTGCGCCACCTCGGCGCCCATCTCGAT | |
| C265A | For | GGCGTGGTCATGGCGGCCGCTTCGGTGGA |
| Rev | GTTGTCCACCGAAGCGGCCGCCATGACCA | |
| C321A | For | GATCGGTGCTGAGCTTGCCATGCAGCAGGC |
| Rev | CACGGCCTGCTGCATGGCAAGCTCAGCACC | |
| C327A | For | CATGCAGCAGGCCGTGGCCATGAACGATTCT |
| Rev | GATAGAATCGTTCATGGCCACGGCCTGCTG | |
| C371A | For | GACCGGCAGACGGTTTGCCCAGGACGATGCCG |
| Rev | CTCGGCATCGTCCTGGGCAAACCGTCTGCC | |
| C384A | For | CTATGTCATGCACGAGGCCGCGCAAGCTGCA |
| Rev | CCATGCAGCTTGCGCGGCCTCGTGCATGAC | |
| C425A | For | CATACGCCCGACACGGCCGATACTACGTTC |
| Rev | CGAGAACGTAGTATCGGCCGTGTCGGGCGT | |
| C443A | For | GCCGAGTTTATCGGCGCCGATCCGACCGC |
| Rev | GAGGGCGGTCGGATCGGCGCCGATAAACTC | |
| C459A | For | GAGGTGGAACTCTTTCGCCGAGGCCGGTTTG |
| Rev | CATCCAAACCGGCCTCGGCGAAAGAGTTCCA | |
| C483A | For | GACGCCGCCGTTCTACGCCGATGTCGTGCGC |
| Rev | GGGGCGCACGACATCGGCGTAGAACGGCGG | |
| C521A | For | CTGTACGGCGCCGGGGCCATCATCGGGGGT |
| Rev | GTTACCCCCGATGATGGCCCCGGCGCCGTA | |
| C535A | For | GCCTTCTACTTCGGCGCCGGCTGGTCCATC |
| Rev | CGTGATGGACCAGCCGGCGCCGAAGTAGAA | |
| Y333N | For | GCATGAACGATTCTATCAACGTAGGCGGCATCA |
| Rev | TCGCTGATGCCGCCTACGTTGATAGAATCGTTCA | |
| Y532F | For | GATGCCGAGTGGGGCTTTGTCATGCACG |
| Rev | GCACTCGTGCATGACAAAGCCCCACTCG | |
| G536A | For | TTCTACTTCGGCTGCGCCTGGTCCATCA |
| Rev | GTTCGTGATGGACCAGGCGCAGCCGAAG |
Additional files
-
Transparent reporting form
- https://doi.org/10.7554/eLife.33953.038





