Release and spread of Wingless is required to pattern the proximo-distal axis of Drosophila renal tubules
Figures
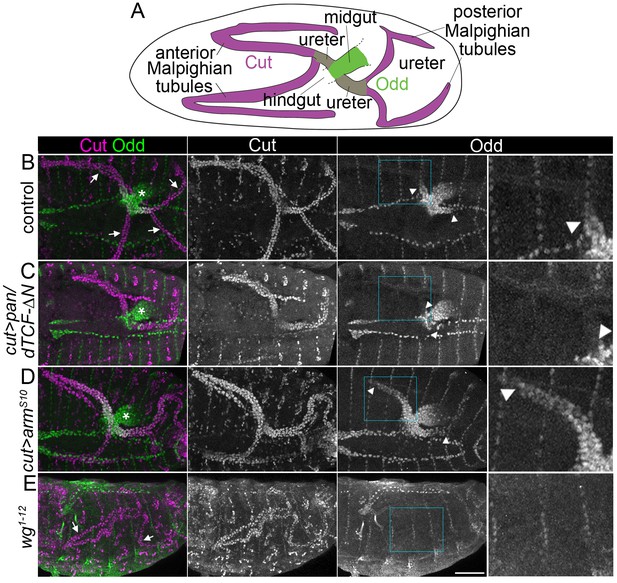
Wingless signalling activates expression of Odd skipped in the proximal tubule.
(A) Diagram to indicate organisation of the anterior and posterior Malpighian tubule pairs in the embryo, which emanate from the gut at the midgut-hindgut junction. Wild-type expression domains of Cut (magenta) and Odd (green) are indicated. B-E show maximum intensity projections of approximately stage 16 Drosophila embryos, oriented anterior left, stained with anti-Cut to mark the entire tubule and anti-Odd to highlight the proximal tubule. Arrowheads mark the distal extent of the Odd-expressing domain in the tubule. Asterisks indicating Odd expression in posterior midgut. An enlargement of the Odd-expression domain in the anterior tubule corresponding to boxed region is provided (right panel). Note that not all tubules may be visible due to orientation. (B) Control (w1118) embryo showing Odd staining in the nuclei of both ureters at the proximal end of the Malpighian tubules (arrows indicate tubules). (C) Embryo in which dominant negative pangolin/dTCF-∆N (pan/dTCF-∆N) is expressed in tubules under control of the cut promoter, to inhibit Wg signalling. (D) Embryo in which constitutively active armadilloS10 (armS10) is expressed in tubules under control of the cut promoter, to activate Wg signalling. (E) A wg temperature sensitive mutant (homozygous wg1-12), switched to the restrictive temperature during later stages of tubule development (see Materials and methods for timings). Arrows indicate the Malpighian tubules (displaying somewhat abnormal morphology) which show no clear anti-Odd staining. Scale bar = 50 μm.
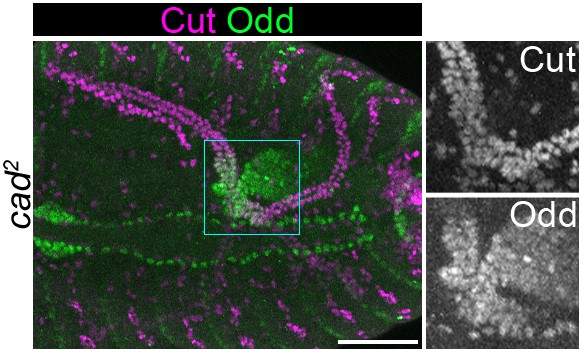
Odd expression in proximal tubule is normal upon loss of zygotic Caudal.
Maximum intensity projection of approximately stage 16 Drosophila embryo, oriented anterior left. Embryo is homozygous mutant for caudal2 (cad2). Embryos is stained with anti-Cut and anti-Odd. Panels to right show enlargement of boxed area. Scale bar = 50 μm.
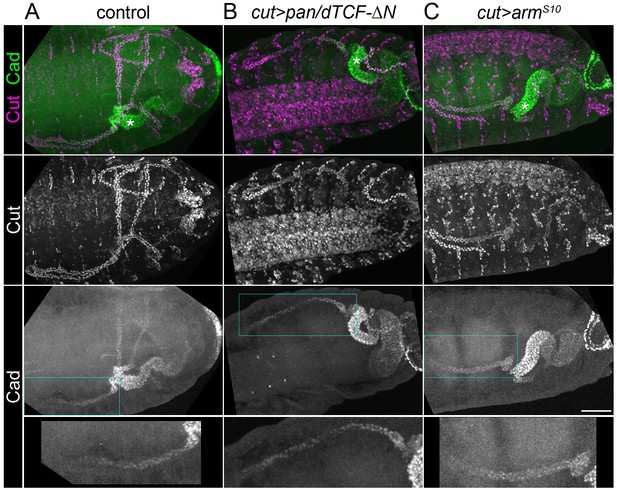
Caudal activation in the proximal tubule is independent of Wingless signalling.
Maximum intensity projections of approximately stage 16 Drosophila embryos, oriented anterior left, stained with anti-Cut to mark the entire tubule and anti-Caudal (Cad). (A) Control (w1118) embryo in which Caudal can be observed staining the nuclei of the posterior midgut and the proximal halves of the tubules Note Cad expression extends more distally than Odd (compare with Figure 1B). (B) Embryo expressing pan/dTCF-∆N using the cut promoter. (C) Embryo expressing armS10 using the cut promoter. An enlargement of the Caudal-expressing domain from an anterior tubule corresponding to boxed region is provided (bottom panel). Scale bar = 50 μm.
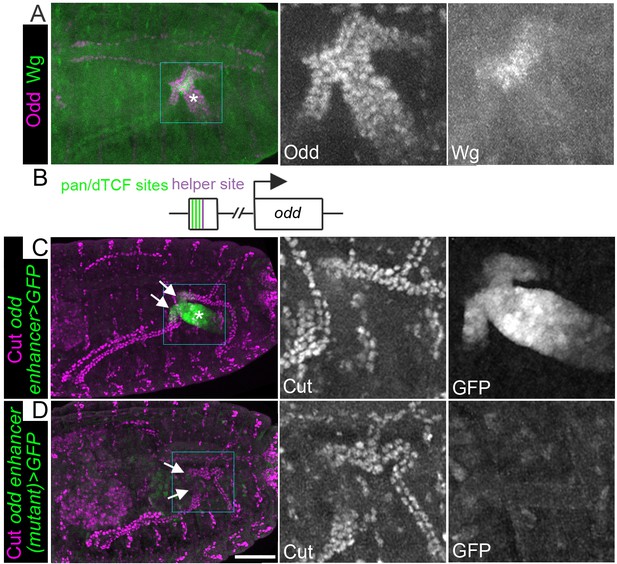
Wingless activity extends from the midgut into the proximal tubule to directly activate Odd skipped expression.
Maximum intensity projections of approximately stage 16 embryos oriented anterior left. Asterisks indicate midgut and arrows indicate proximal ends of tubules. In each case the two right panels show enlargement of areas marked with squares. (A) Staining for anti-Wg and anti-Odd revealing Wg staining in a small patch within the midgut and Odd staining in the posterior midgut and both tubule ureters. (B) Cartoon to indicate features of odd enhancer region containing pangolin/dTCF binding sites and helper site. (C) Transgenic embryo containing the putative odd enhancer upstream of a GFP reporter gene (odd enhancer > GFP) stained for anti-Cut and anti-GFP. (D) Transgenic embryo containing the putative odd enhancer bearing mutated pangolin/dTCF binding sites upstream of a GFP reporter gene (odd enhancer (mutant) > GFP) stained for anti-Cut and anti-GFP. Scale bar = 50 μm.
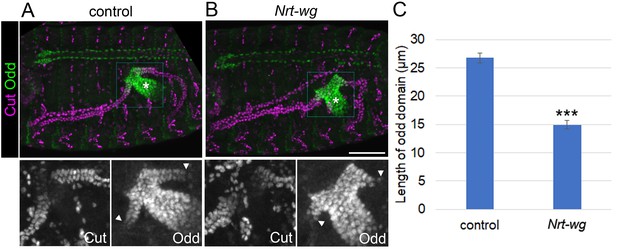
Membrane-tethered Wingless is unable to correctly pattern the proximal tubule.
Maximum intensity projections of approximately stage 16 embryos stained with anti-Cut and anti-Odd, oriented anterior left. Asterisks indicate midgut and arrowheads mark distal extent of Odd expression in the tubule. Lower panels show enlargement of boxed areas. (A) Sibling control embryo (Nrt-wg/+ or +/+). (B) Embryo which is homozygous mutant for wg, and expressing membrane-tethered Nrt-wg. Scale bar = 50 μm. (C) Graph showing mean length of Odd expressing region along tubule in control (mixture of Nrt-wg/+ and +/+) and Nrt-wg embryos. Error bars indicate SEM. Control n = 29, Nrt-wg n = 37. P (Mann-Whitney) <0.00001***.
-
Figure 4—source data 1
Quantifications of Odd skipped domain length in Nrt-wg embryos.
- https://doi.org/10.7554/eLife.35373.011
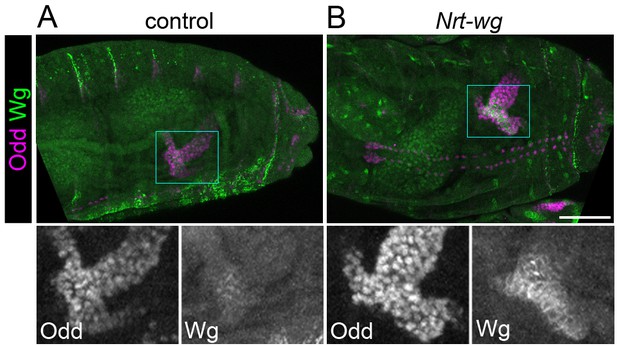
Expression of Nrt-wg.
Maximum intensity projections of approximately stage 16 Drosophila embryos, oriented anterior left, stained with anti-Odd and anti-Wg. Bottom panels show enlargement of areas marked with squares. (A) Control (w1118) embryo. (B) Embryo expressing Nrt-wg in homozygous wg mutant background (Nrt-wg). Images were taken using identical microscope settings. Scale bar = 50 μm.
-
Figure 4—figure supplement 2—source data 1
Quantifications of Odd skipped domain length upon overexpression of Wg or Nrt-Wg.
- https://doi.org/10.7554/eLife.35373.009
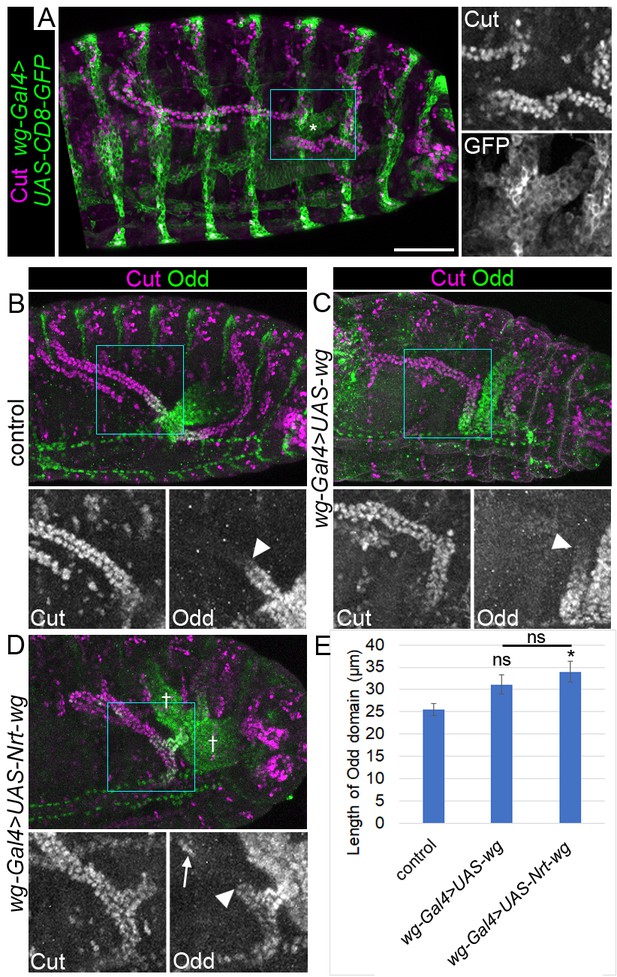
Incorrect patterning of proximal tubule is not result of dominant effect of Nrt-wg.
Maximum intensity projections of approximately stage 16 Drosophila embryos, oriented anterior left. Scale bar = 50 μm. (A) An embryo in which the expression pattern of wg-Gal4 is determined using a UAS-CD8-GFP reporter (marking cell membranes) and staining with anti-GFP. Embryo is co-stained with anti-Cut to allow visualisation of tubules. Asterisk marks expression in presumptive midgut. Right panel shows enlargement of area marked with square. B-D show embryos stained with anti-Cut and anti-Odd, with lower panels showing enlargements of boxed areas, to highlight the region where the proximal tubule of the anterior tubule pair meets the gut. Arrowheads show the distal extent of the Odd expressing tubule domain. (B) Control embryo (wg-Gal4). (C) An embryo in which wg-Gal4 is used to drive expression of UAS-wg, in a wild-type wg background. (D) An embryo in which wg-Gal4 is used to drive expression of UAS-Nrt-wg in a wild-type wg background. In this context Odd is expressed in the gut beyond both sides of the junction with the Malpighian tubules (†) that is in both the midgut and hindgut. Arrow marks patch of cells in tubule expressing Odd. (E) Graph showing mean length of Odd expressing region along tubule in control embryos (UAS-wg), and embryos in with wg-Gal4 is driving the expression of UAS-wg or UAS-Nrt-wg. Error bars represent SEM. One-way ANOVA indicated a significant statistical difference between the three genotypes (p=0.04*). Control n = 14, wg-Gal4 >UAS wg n = 20, p=0.052 (Mann-Whitney, compared to control), wg-Gal4 >UAS Nrt-wg n = 25, p=0.012* (Mann-Whitney, compared to control) and p=0.35 (Mann-Whitney, compared to wg-Gal4 >UAS wg.
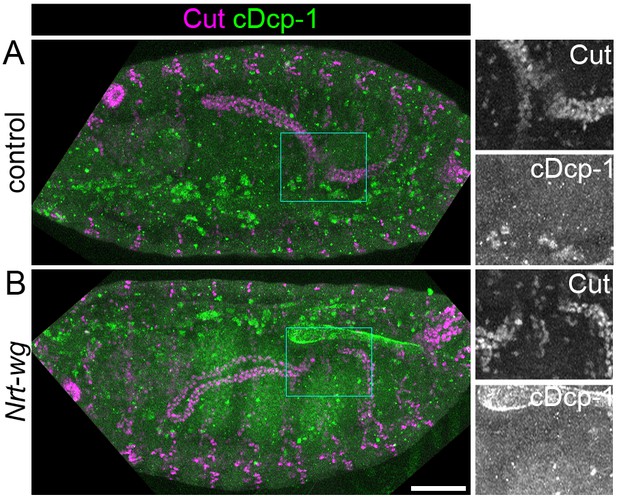
Apoptosis is not induced in tubules of Nrt-wg embryos.
Maximum intensity projections of approximately stage 16 Drosophila embryos, oriented anterior left. Embryos are stained with anti-cleaved Dcp-1 (cDcp-1) to mark apoptotic cells, and anti-Cut to mark the Malpighian tubules. (A) Control (Nrt-wg/+ or +/+) embryo. (B) Embryo expressing Nrt-wg in a wg homozygous mutant background. Panels to right show enlargements of boxed areas. Scale bar = 50 μm.
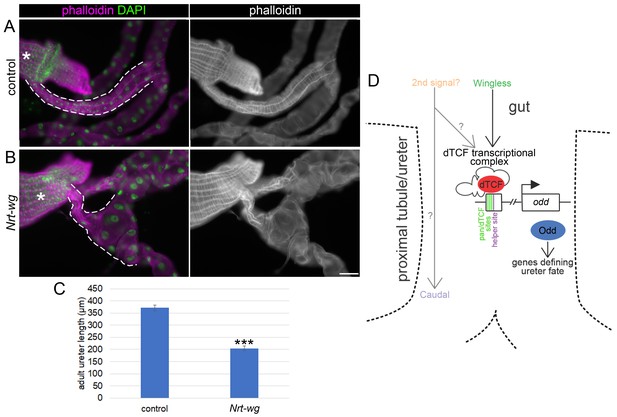
Disruption of the Wingless pathway leads to abnormal morphology of the adult ureters.
Single focal plane epifluorescence images of dissected guts and Malpighian tubules of adult flies, stained to show actin (phalloidin) and nuclei (DAPI). White dashed line indicates most proximal tubule/ureter and asterisks indicate midgut. (A) Centred on ureter from sibling control (Nrt-wg/+) adult. (B) Centred on ureter from fly which is homozygous mutant for wg, and expressing membrane-tethered Nrt-wg. Scale bar = 50 μm. (C) Graph showing mean length of adult ureters in control (Nrt-wg/+) and Nrt-wg flies. Error bars indicate SEM. Control n = 47, Nrt-wg n = 30. P (Mann-Whitney) <0.00001***. (D) Model of signals from the gut patterning the proximal Malpighian tubule, necessary for correct ureter development. Enhancer region upstream of odd coding region as in Figure 3B.
-
Figure 5—source data 1
Quantifications of adult tubule ureter length in Nrt-wg flies.
- https://doi.org/10.7554/eLife.35373.014
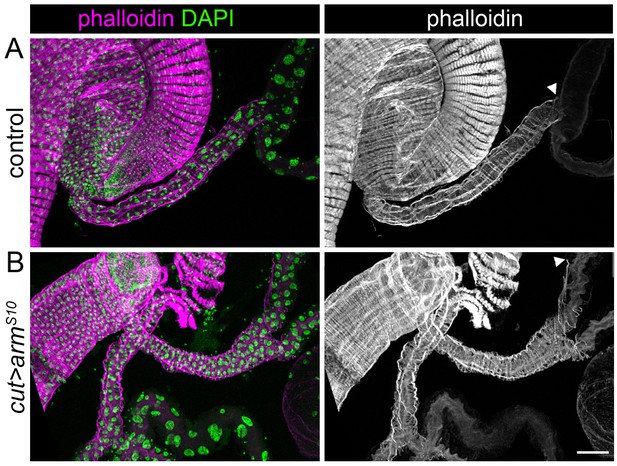
Changes to morphology of adult tubules is not the result of a dominant effect of Nrt-wg.
maximum intensity projections of dissected guts and Malpighian tubules of adult flies, stained to show actin (phalloidin) and nuclei (DAPI). (A) Control (w1118) adult. (B) Adult expressing armadilloS10 using the cut promoter. Arrowheads mark the distal extent of the ensheathing visceral muscle along the tubule. Scale bar = 50 μm.
Additional files
-
Transparent reporting form
- https://doi.org/10.7554/eLife.35373.015





