Selective increases in inter-individual variability in response to environmental enrichment in female mice
Figures
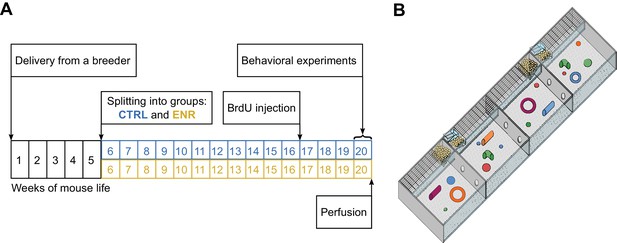
Experimental setup.
(A) Experimental outline. At an age of 5 weeks, 80 female mice were split equally into two groups: one group lived in an enriched environment (ENR) for 15 weeks and one group lived in standard mouse cages in groups of five mice per cage (CTRL) for the same period of time. To analyze neurogenesis in the hippocampus, mice received intraperitoneal BrdU injections three weeks before perfusion. Behavioral phenotyping was performed in the last eight days before perfusion. (B) The enriched environment enclosure covered a total area of 2.2 m2 and consisted of four sub-compartments, which were connected via tunnels. Food, toys and nesting material were provided in every compartment.
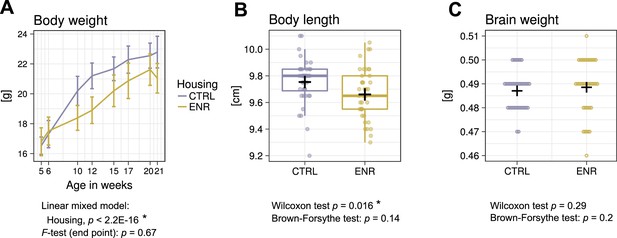
Environmental enrichment does not increase variance in gross body morphology.
(A) Longitudinal measurement of body weight. Presented are means ± standard deviations. (B) Body length and (C) brain weight were assessed at the end of the experiment. Box and whisker plot: center line, median; plus sign, mean; upper and lower hinges, first and third quartiles; whiskers, highest and lowest values within 1.5 times the interquartile range outside hinges; dots, individual data points. Asterisks indicate significant effects at 5% threshold in the indicated statistical tests. Full information on the statistical tests is available in Supplementary file 2. CTRL, control group; ENR, enriched group.
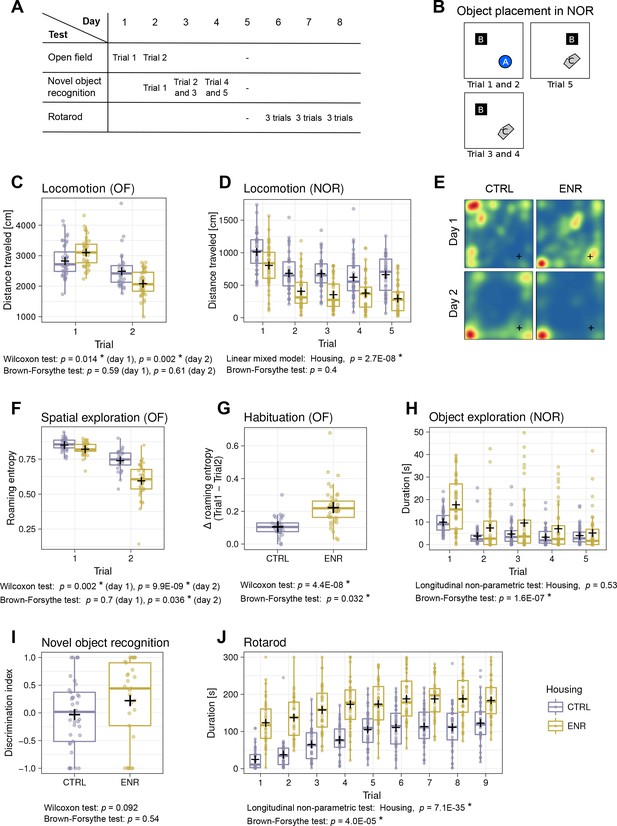
Mice living in an enriched environment exhibit inter-individual differences in motor abilities, spatial exploration and object exploration.
(A) Timeline of behavioral testing. (B) Object placement in trials of the novel object recognition (NOR) task. (C) Total distance that control (CTRL, blue) and enriched animals (ENR, yellow) moved in the arena on the two days of open field (OF) testing. (D) Total distance that mice moved during each trial in the NOR test. (E) Representative heatmaps for two mice depicting the probabilities that each mouse was to be found at a specific location in the OF arena. Blue indicates lowest and red highest probabilities, respectively. The corner in which the light source was located is marked with a cross (+). (F) Roaming entropy in the OF arena describes spatial exploration. (G) Habituation to the OF expressed as a difference in roaming entropy between trials. (H) Object exploration in the NOR test. (I) Discrimination index indicating preference for the novel (+1) or the old (−1) object. (J) Duration that mice spent on the rotating rod during the individual trials of the rotarod task. Box and whisker plots, see Figure 2. Asterisks, significant effects at 5% threshold.
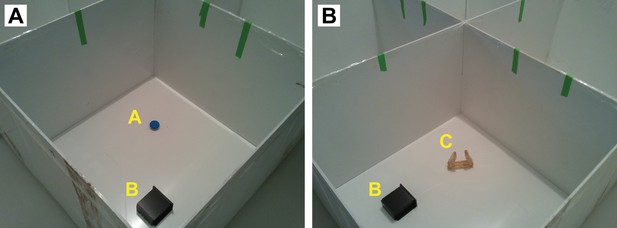

Photographs showing examples of the OF arena with objects (labeled in yellow) during NOR testing.
(A) Trial 1 and 2; (B) trial 5. Markings with green tape provide additional spacial clues.
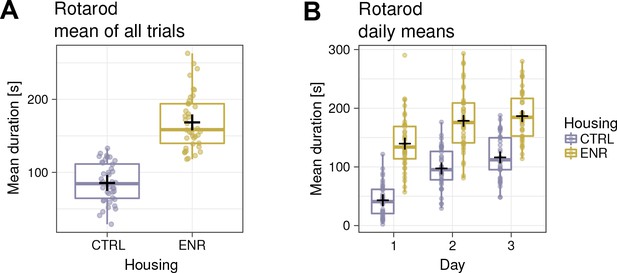
Rotarod mean duration from all trials (A) and daily sessions (B).
https://doi.org/10.7554/eLife.35690.007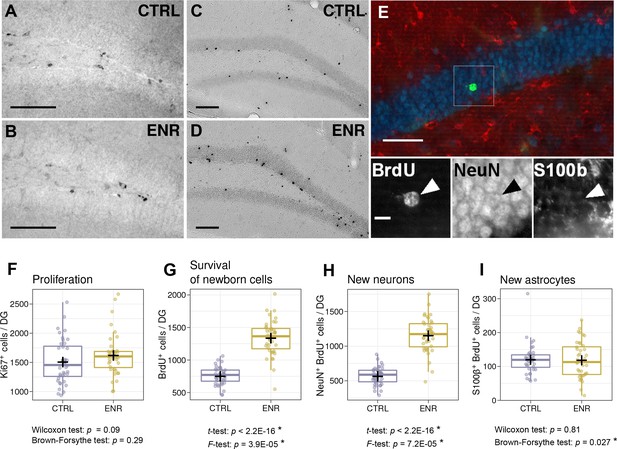
Environmental enrichment leads to the development of individual levels of adult hippocampal neurogenesis.
(A, B) Representative images of Ki67 immunostaining, which marks proliferating cells in control (CTRL) and enriched (ENR) mice. (C, D) New-born cells were identified by BrdU immunoreactivity three weeks after the injection of BrdU. (E) The proportions of new-born neurons and astrocytes were determined by co-localization of BrdU (green) with NeuN (blue) and S100β (red), respectively. The image shows a single optical section. The arrowhead highlights a new-born neuron. (F) No difference in the number of proliferating cells can be observed between mice housed under CTRL and ENR conditions. (G–I) ENR mice have significantly higher means and variances in the numbers of new-born BrdU-positive cells (G) and new neurons (H), whereas only the variance of the number of new astrocytes was increased (I). Scale bars are as follows: (A–D) 100 µm; (E), 50 µm; (E inset), 10 µm. Box and whisker plots, see Figure 2. Asterisks indicate significant effects at a 5% threshold.
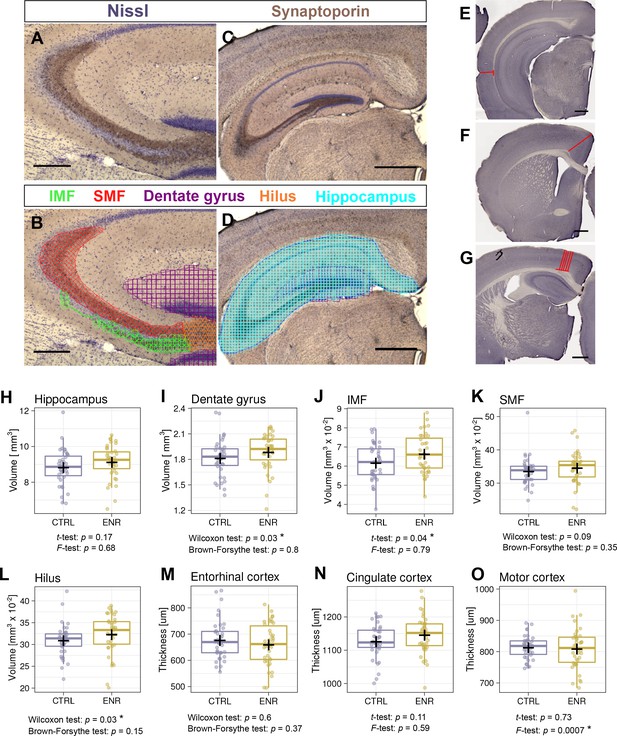
Environmental enrichment does not increase variability in gross brain morphology.
(A–D) Representative images of sections immunostained with a synaptoporin antibody (brown) and counterstained with Nissl (purple). (B, D) Examples of contour tracing with overlaid Cavalieri probe estimator markers for the indicated brain structures. (E–G) Thickness measurement on Ki67-DAB stained sections of entorhinal (E), cingulate (F), and motor cortex (G). (H–L) Results from volumetric analyses of the hippocampus (H), dentate gyrus (I), infrapyramidal mossy fiber tract (IMF; J), the suprapyramidal mossy fiber tract (SMF; K), and the hilus (L). (M–O) Thickness of three cortical areas: the entorhinal cortex (M), the cingulate cortex (N) and the motor cortex (O). Scale bars are 200 µm in (A-B), and 500 µm in (C-G). CTRL, control; ENR, enriched mice. Box and whisker plot, see Figure 2. Asterisks indicate significant effects at 5% threshold.
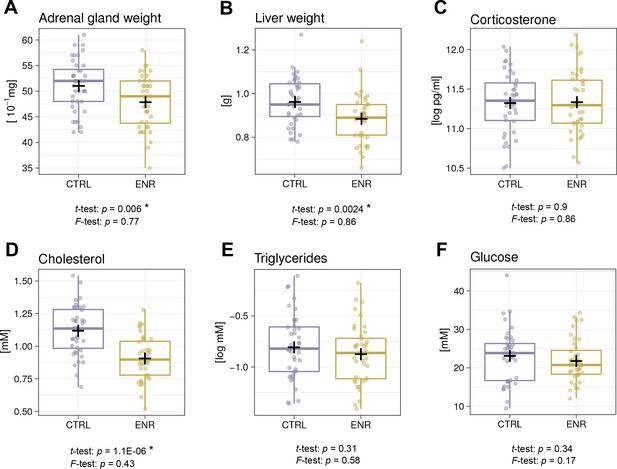
Environmental enrichment does not induce metabolic variability.
(A, B) Environmental enrichment (ENR) mice have adrenal gland (A), and liver (B) weights that are lower than those of control (CTRL) animals. (C) Housing does not affect acute corticosterone levels. (D–F) Effects of ENR on plasma biomarkers: cholesterol (D), triglycerides (E) and glucose (F). Box and whisker plots, see Figure 2. Asterisks indicate significant effects at a 5% threshold.
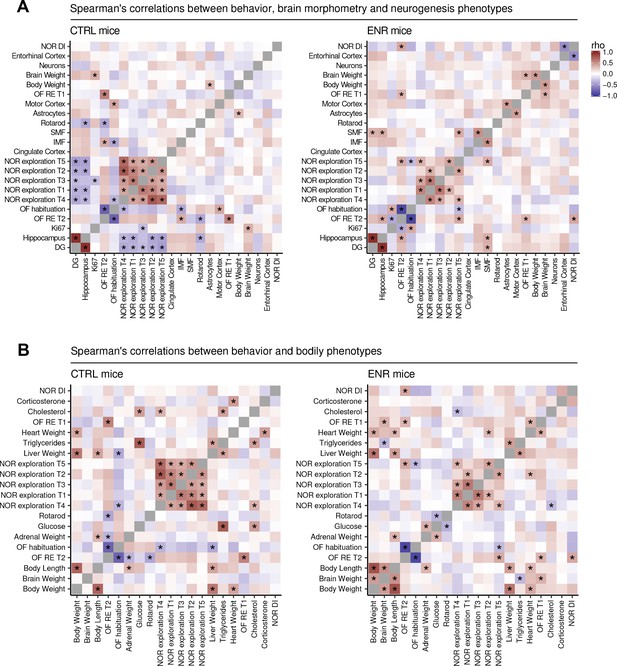
Environmental enrichment leads to partial restructuring of relationships between phenotypes.
Heat maps show Spearman’s rank correlation coefficient between selected behavioral, morphometric and neurogenic (A) or metabolic traits (B) within the control (CTRL, left panels) and the enriched (ENR, right panels) groups. Rotarod performance was summarized as a mean of data from all individual trials for each animal. Phenotypes were ordered on the basis of hierarchical clustering in the ENR group. Significant correlations (p<0.05) are marked with asterisks. NOR, novel object recognition test; OF, open field test; T1–5, trials 1–5 of the NOR and OF tests; RE, roaming entropy; DG, dentate gyrus; DI, discrimination index. Source files listing the rho values are available in Figure 7—source data 1 (CTRL) and in Figure 7—source data 2 (ENR).
-
Figure 7—source data 1
Correlation matrix, CTRL mice.
Spearman’s correlation coefficients for correlations between phenotypes in CTRL mice. A tab-delimited text file.
- https://doi.org/10.7554/eLife.35690.012
-
Figure 7—source data 2
Correlation matrix, ENR mice.
Spearman’s correlation coefficients for correlations between phenotypes in ENR mice. A tab-delimited text file.
- https://doi.org/10.7554/eLife.35690.013
Tables
| Reagent type (species) or resource | Designation | Source or reference | Identifiers | Additional information |
|---|---|---|---|---|
| Strain, strain background (M. musculus) | C57BL/6JRj | Janvier Labs | ||
| Antibody | Anti-Ki67 (rabbit polyclonal) | Novocastra | Novocastra: NCL-Ki67p; RRID:AB_442102 | (1:500) |
| Antibody | Anti-BrdU (rat monoclonal) | AbD Serotec | AbD Serotec: OBT0030; RRID:AB_609568 | (1:500) |
| Antibody | Anti-synaptoporin (rabbit polyclonal) | Synaptic Systems | Synaptic Systems:102002; RRID:AB_887841 | (1:500) |
| Antibody | Anti-NeuN (mouse monoclonal) | Merck Millipore | Merck: MAB377; RRID:AB_2298772 | (1:100) |
| Antibody | Anti-S100beta (rabbit monoclonal) | Abcam | Abcam: ab52642; RRID:AB_882426 | (1:200) |
| Antibody | Biotin-conjugated secondary (donkey polyclonal) | Jackson ImmunoResearch | (1:500) | |
| Antibody | Alexa 488-, Cy5-, Cy3- secondaries (donkey polyclonal) | Jackson ImmunoResearch | (1:500) | |
| Commercial assay or kit | Vectastain ABC Elite kit | Vector Laboratories | Vector: PK-6100; RRID:AB_2336819 | |
| Commercial assay or kit | Amplex Red Glucose/Glucose Oxidase Assay | Invitrogen | Invitrogen: A22189 | |
| Commercial assay or kit | Amplex Red Cholesterol Assay | Invitrogen | Invitrogen: A12216 | |
| Commercial assay or kit | Triglyceride Assay | Abcam | Abcam: ab65336 | |
| Commercial assay or kit | Corticosterone ELISA kit | Enzo | Enzo: ADI-901–097 | |
| Chemical compound, drug | 5-Bromo-2'-deoxyuridine | Sigma Aldrich | Sigma: B5002 | |
| Software, algorithm | Stereoinvestigator7 software | MBF Bioscience | ||
| Software, algorithm | Ethovision | Noldus | ||
| Software, algorithm | ZEN blue edition | Zeiss |
Additional files
-
Supplementary file 1
Phenotype data table.
Abbreviations: NOR, novel object recognition test; OF, open field test; IMF, infrapyramidal mossy fibers; SMF, suprapyramidal mossy fibers; DG, dentate gyrus.
- https://doi.org/10.7554/eLife.35690.014
-
Supplementary file 2
Summary data, test statistics and p-values from statistical analyses of differences between CTRL and ENR mice.
The p-values were not corrected for multiplicity of tests. For longitudinal data, individual comparisons were made only when the omnibus test indicated significant difference between groups. In this case, both uncorrected and adjusted p-values are reported. Correction was performed using the Holm method (Walsh and Cummins, 1979) implemented using the p.adjust function in R. Abbreviations: LRT, likelihood ratio test; WTS, Wald-type statistic; NOR, novel object recognition test; OF, open field test; IMF, infrapyramidal mossy fibers; SMF, suprapyramidal mossy fibers; DG, dentate gyrus; st. dev., standard deviation.
- https://doi.org/10.7554/eLife.35690.015
-
Transparent reporting form
- https://doi.org/10.7554/eLife.35690.016





