Structures of two aptamers with differing ligand specificity reveal ruggedness in the functional landscape of RNA
Figures
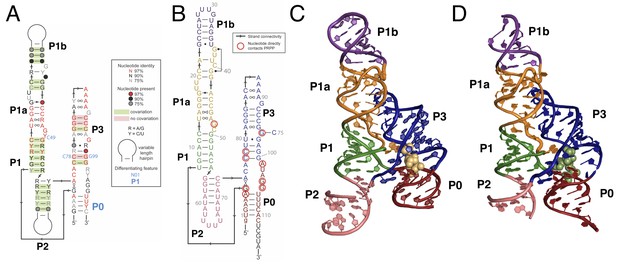
Overall structure of the PRPP riboswitch and its G96A mutant, which is a ppGpp aptamer.
(A) Consensus sequence of the PRPP riboswitch, adapted from Sherlock et al. (Sherlock et al., 2018b). The secondary structure has been updated to show structural information gained from the present study. The sequence is depicted as in Sherlock et al. (see key). Nucleotides noted in blue are important bioinformatic differences between PRPP riboswitches and guanidine riboswitches. Base pair notation is as published previously (Leontis and Westhof, 2001). (B) Secondary structure of the PRPP riboswitch aptamer from T. mathranii. Nucleotides are colored by paired region. Paired regions are indicated in bold. Sequence numbering is indicated in gray. Nucleotides that directly contact PRPP are circled in red, and arrows indicate strand connectivity. (C) Crystal structure of the PRPP riboswitch. Chain A is shown. The RNA is depicted as a cartoon and PRPP is depicted as yellow spheres. Nucleotides are colored by paired region as in B. (D) Crystal structure of the G96A mutant. Chain A is shown. The RNA is depicted as a cartoon and ppGpp is depicted as green spheres. Nucleotides are colored by paired region as in B.
-
Figure 1—source data 1
Summary of fitted binding data without Bmax constraints.
- https://doi.org/10.7554/eLife.36381.006
-
Figure 1—source data 2
Raw binding data.
- https://doi.org/10.7554/eLife.36381.007
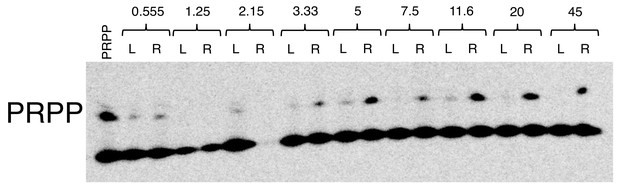
Autoradiograph of a representative PAGE gel from dissociation constant determination for the PRPP aptamer.
The higher of the two bands represents intact PRPP, while the lower band represents degradation products. The leftmost lane is purified PRPP. This leftmost lane represents a sample of PRPP that was purified and eluted but not subjected to the reaction conditions. Each pair of lanes thereafter represents one dialysis cassette. The concentration of aptamer added to the cassette in micromolar is depicted above each pair. The concentration of aptamer in crystallization experiments was 150 μM, three-fold greater than the highest concentration used in determining the dissociation constant for wild type RNA. ‘L’ is the side of the cassette to which PRPP ligand was initially added. ‘R’ is the side of the cassette to which RNA aptamer was initially added. After overnight incubation at room temperature, material recovered from dialysis cassettes was electrophoresed to separate intact PRPP from degradation products. PRPP incubated in the presence of aptamer experienced significant protection relative to unbound PRPP, permitting the use of this unstable metabolite in the present structural and biophysical studies.
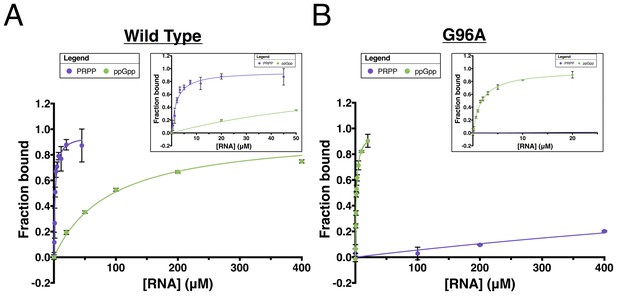
Data from equilibrium dialysis experiments and fits used to calculate dissociation constants.
All experiments were performed in triplicate and the standard deviation for each data point is shown using black error bars. (A) wild type RNA binding to PRPP (purple) and ppGpp (green). The ppGpp complex did not quite reach saturation, so the Bmax was constrained to equal the Bmax value from the fit for ppGpp binding to the G96A mutant. (B) G96A mutant binding to PRPP (purple) and ppGpp (green). PRPP reached just 20% bound at 400 μM RNA, so the Bmax was constrained to equal the Bmax value from the fit for PRPP binding to wild type RNA.
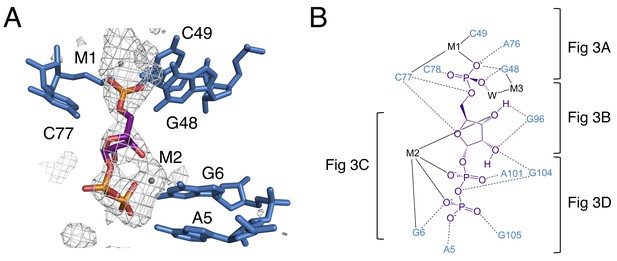
The binding pocket of the PRPP riboswitch.
(A) Crystal structure of the ligand-binding site in chain A. Relative to Figure 1, the structure is rotated 180° about the y axis. PRPP is depicted as sticks and colored by element with purple carbons. Nucleotides are depicted as blue sticks. Metal ions are depicted as gray spheres. Individual nucleotides and metals are labeled. An FO–FC map contoured at 2.5 σ is shown as a gray mesh. The map was calculated using an otherwise complete model lacking PRPP, M1, and M2. (B) Ligand interaction map. The map is colored essentially as in A. All RNA and metal contacts to PRPP are shown. Dashed black lines indicate hydrogen bonds. Solid black lines indicate coordination to a metal ion. Brackets indicate interactions shown in individual panels of Figure 3.
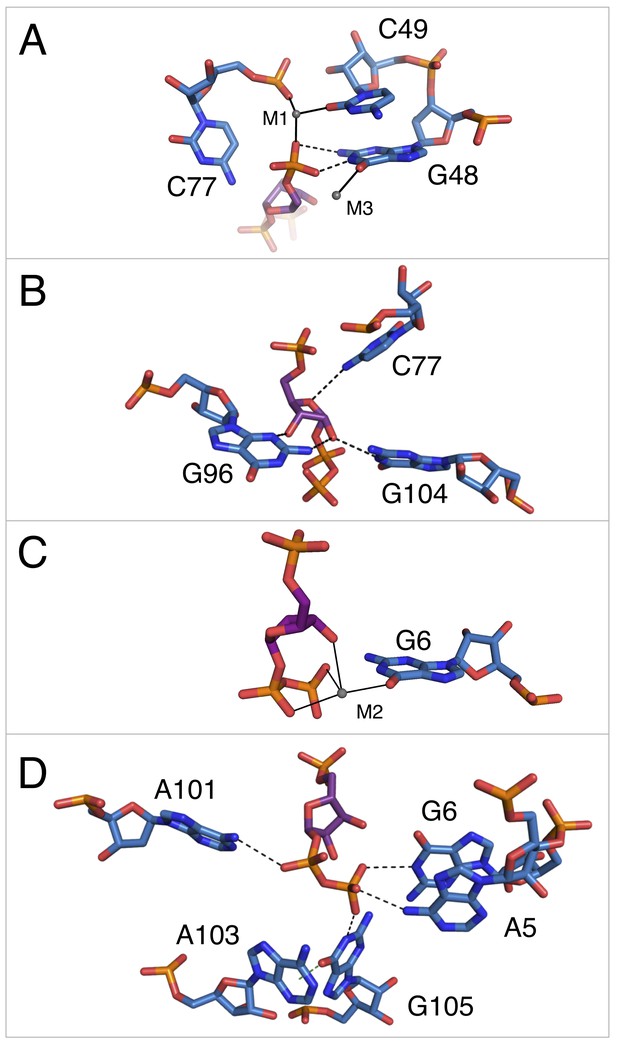
Notable contacts to the PRPP ligand in chain A. PRPP is depicted as sticks and colored by element with purple carbons.
Nucleotides are depicted as sticks and colored by element with blue carbons. Individual nucleotides and metals are labeled. Dashed black lines indicate hydrogen bonds. A dashed green line shows the lone pair-π interaction between A103 and G105. Solid black lines indicate coordination to a metal ion. Relative to Figure 1, the structure is rotated 180° about the y axis. Panel A is additionally rotated nearly 90° about the x axis. Panel D is rotated approximately 45° about the x axis in the opposite direction. (A) Contacts among the 5-phosphate of PRPP, residues G48, C49, C77, and metal ions M1 and M3. (B) Hydrogen bonds between the ribose of PRPP and residues C77, G96, and G104. (C) Coordination of metal M2 by PRPP and residue G6. (D) Recognition of the pyrophosphate group of PRPP by residues A5, G6, A101, and G105.
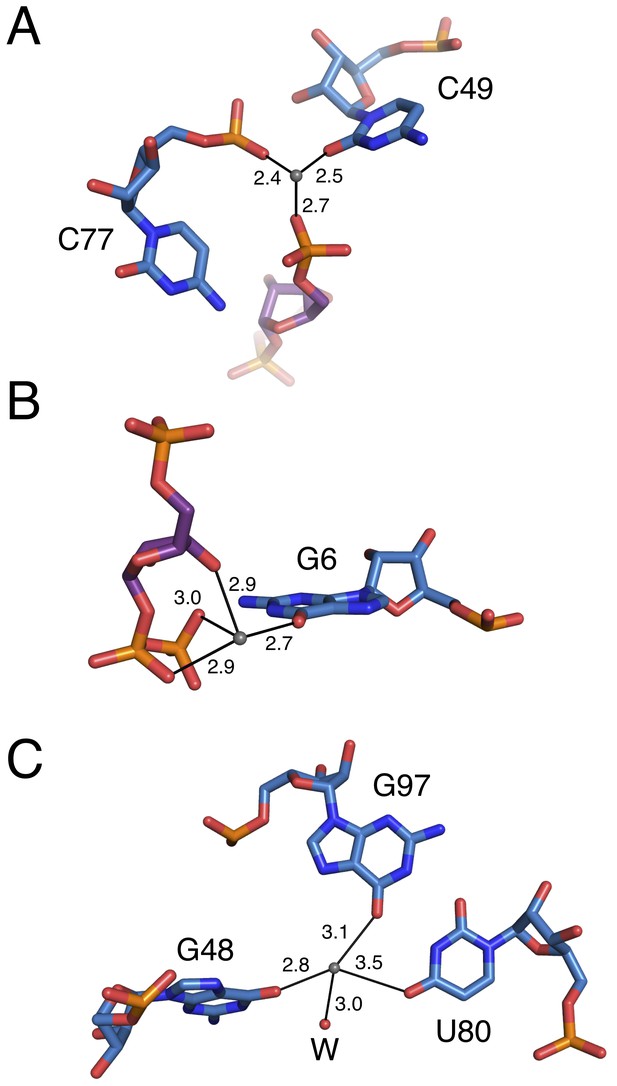
Coordination of metals by chain A of the PRPP aptamer.
PRPP is depicted as sticks and colored by element with purple carbons. Nucleotides are depicted as sticks and colored by element with blue carbons. Individual nucleotides and metals are labeled. Solid black lines indicate coordination to a metal ion, and coordination distances in Å are indicated. (A) Coordination of M1. (B) Coordination of M2. (C) Coordination of M3.
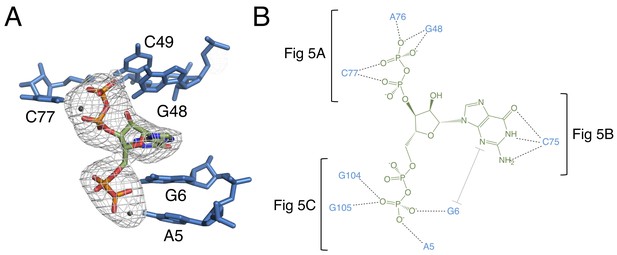
The binding pocket of the G96A mutant in complex with ppGpp.
(A) Crystal structure of the ligand-binding site in chain A. ppGpp is depicted as sticks and colored by element with green carbons. Nucleotides are depicted as blue sticks. Metal ions are depicted as gray spheres. Individual nucleotides are labeled. An FO–FC map contoured at 3.0 σ is shown as a gray mesh. The map was calculated using an otherwise complete model lacking ppGpp and nearby metals. Relative to Figure 1, the structure is rotated 180° about the y axis. (B) Ligand interaction map. The map is colored essentially as in A. All RNA contacts to ppGpp are shown. Dashed black lines indicate hydrogen bonds. The grey bracket indicates base stacking. Black brackets indicate interactions shown in individual panels of Figure 5.
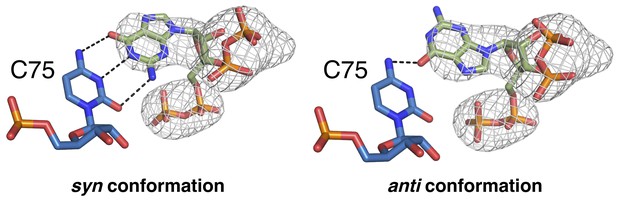
A comparison of ppGpp modeled in the syn conformation and the anti conformation.
Chain A is shown. ppGpp has green carbons and is colored by element. The Fo-Fc map was generated using an otherwise complete model lacking ppGpp and is shown contoured at 3.0 σ as a grey mesh.
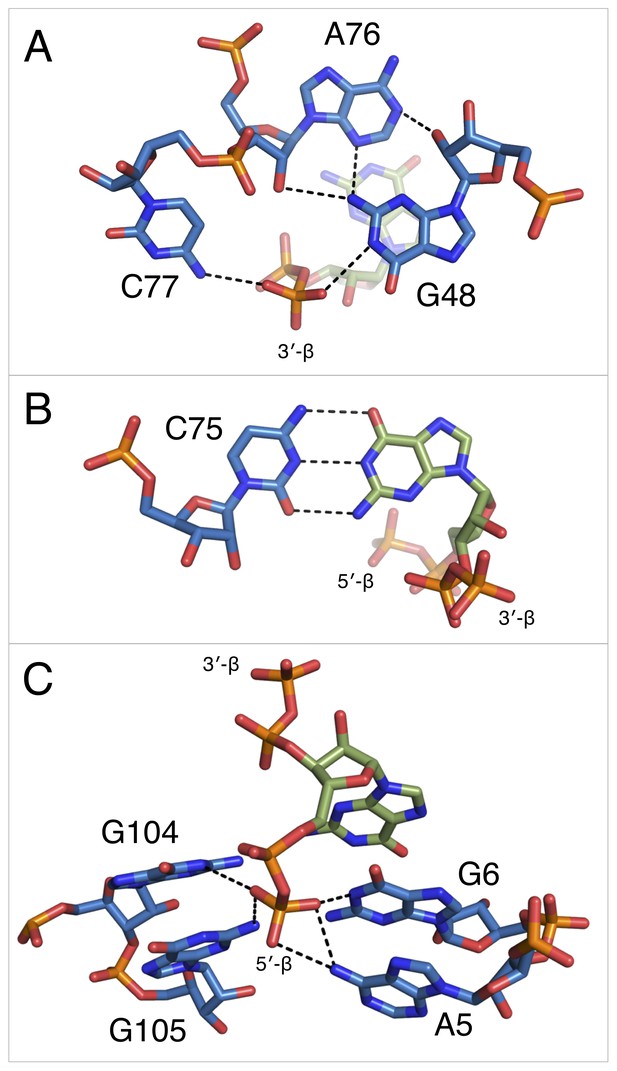
Notable contacts to the ppGpp ligand in chain A. ppGpp is depicted as sticks with green carbons and is colored by element.
RNA is depicted as sticks with blue carbons and is colored by element. Dashed black lines indicate hydrogen bonds. (A) hydrogen bonds donated from amino groups in C77 and G48 to the 3′-β-phosphate of ppGpp, as well as the hydrogen bond network that constructs this part of the binding pocket. The 2′OH of A76 is close enough in chains B and C to form an additional hydrogen bond to the 3′-β-phosphate. (B) The Watson-Crick base pair between C75 in the RNA and the ppGpp ligand. (C) hydrogen bonds donated from amino groups in A5, G6, G104, and G105 to the 5′-β-phosphate of ppGpp.
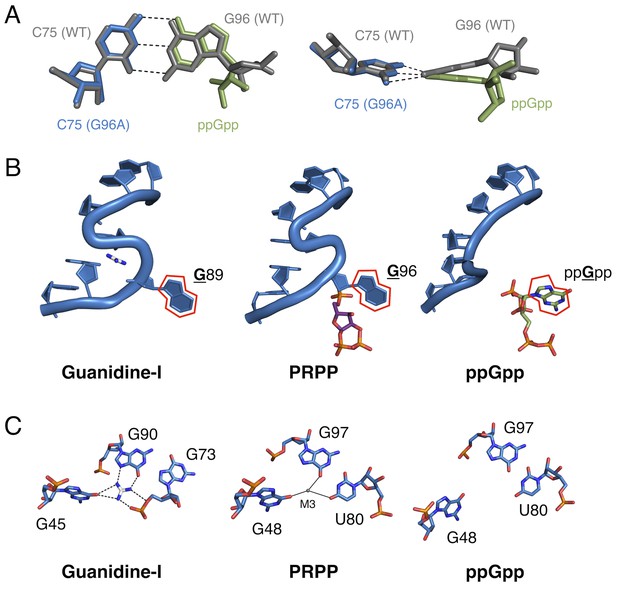
Comparison of the guanidine binding site in the guanidine aptamer, the M3 binding site in the PRPP aptamer, and the corresponding site in the G96A mutant.
(A) Watson-Crick base pairs with C75 in the wild-type and G96A aptamers. Wild-type and G96A aptamer structures are overlaid. Wild-type RNA is shown as gray sticks. G96A RNA is shown as blue sticks. The base and ribose of ppGpp are shown as green sticks. Hydrogen bonds are shown as dashed lines. Left: face-on view of the preserved base pair. Right: edge-on view of the same interaction. (B) The S-turn motif in the guanidine and PRPP aptamers, and the equivalent position in the ppGpp aptamer. The RNA is depicted as a blue cartoon. Guanidine is colored by element with white carbons. PRPP is colored by element with purple carbons. ppGpp is colored by element with green carbons. A red outline showcases the position of a conserved guanine base in all three RNA elements. Relative to Figure 1, the structure is rotated 180° about the y axis. (C) Nucleotides in the guanidine or M3 binding site, or the equivalent site in the ppGpp aptamer. The RNA is colored by element with blue carbons. Guanidine is colored by element with white carbons. Black dashed lines indicate hydrogen bonds. Solid black lines indicate coordination to a metal ion. Individual nucleotides, guanidine and M3 are labeled. Chain A of the PRPP and ppGpp aptamers is shown.
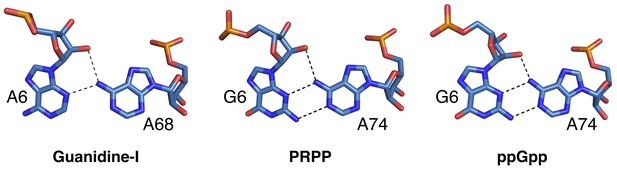
Conserved interactions between P1 and P3.
Nucleotides are labeled, depicted as sticks and colored by element with blue carbons. Dashed lines indicate hydrogen bonds. (A) Nucleotides A6 and A68 of the guanidine aptamer. (B) Nucleotides G6 and A74 of the PRPP aptamer. (C) Nucleotides G6 and A74 of the G96A mutant, a ppGpp aptamer. Chain A of the PRPP and ppGpp aptamers is shown.
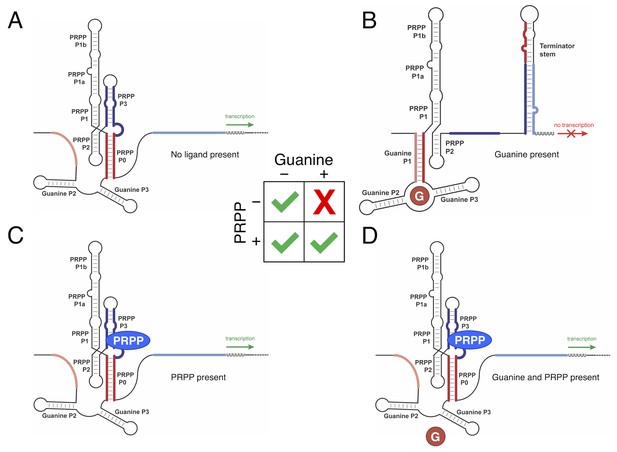
Predicted model of switching in the tandem guanine-PRPP riboswitch from T. mathranii.
The secondary structure of the RNA is represented as black lines. P0 nucleotides are colored dark red. Guanine aptamer nucleotides that may pair with P0 nucleotides are colored light red. P3 nucleotides are colored dark blue. Nucleotides in the terminator stem that may pair with P3 nucleotides are colored light blue. PRPP is represented as a blue ellipse, and guanine is represented as a red circle. The behavior of this IMPLY Boolean logic gate is shown in the center. (A) Predicted structure in the absence of PRPP and guanine. (B) Predicted structure in the presence of guanine. (C) Predicted structure in the presence of PRPP. (D) Predicted structure in the presence of PRPP and guanine. In this model, the presence of PRPP stabilizes P0 and destabilizes the guanine aptamer domain.
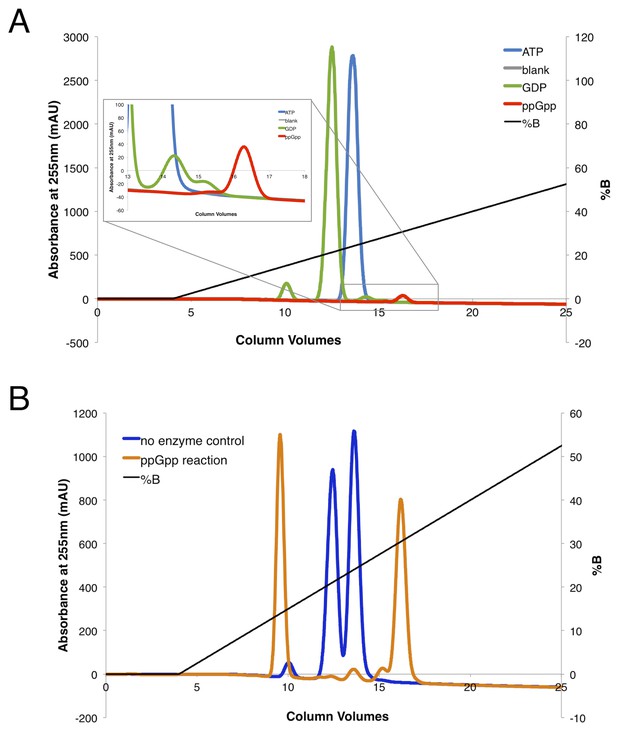
ppGpp purification and validation.
(A) Anion-exchange chromatography of substrate and product standards. The ppGpp sample standard was purchased from TriLink BioTechnologies. (B) Anion-exchange chromatography of reactions with substrates GDP and ATP in the presence or absence of SAS1. To purify ppGpp for use in crystal trays, only fractions from the peak at ~16 column volumes were collected and pooled.
Tables
Dissociation constants for PRPP and ppGpp binding to the wild type and G96A T. mathranii aptamers with calculated fold specificity changes.
https://doi.org/10.7554/eLife.36381.011| Dissociation constants for WT and G96A binding to PRPP and ppGpp | ||||
|---|---|---|---|---|
| Construct | Kd for PRPP | Kd for ppGpp | Fold specificity for PRPP over ppGpp | Estimated magnitude of overall specificity switch |
| Wild type | 2.0 ± 0.3 μM | 91 ± 3 μM | 46 | ~40,000 |
| G96A | 1600 ± 200 μM | 1.8 ± 0.1 μM | ~0.001 | |
| Reagent type (species) or resource | Designation | Source or reference | Identifiers |
|---|---|---|---|
| Chemical compound, drug | ppGpp standard | TriLink BioTechnologies | TriLink Biotechnologies: N-6001 |
| Chemical compound, drug | PRPP | Millipore Sigma | Millipore Sigma:P8296- 25MG |
| Commercial assay or kit | Hampton Research Natrix HT Screen | Hampton Research | Hampton Research:HR2- 131 |
| Commercial assay or kit | Harvard Apparatus cassettes | Harvard Apparatus | Harvard Apparatus:742203 |
| Peptide, recombinant protein | Ribose phosphate pyrophosphokinase | Abbexa | Abbexa:abx072019 |
| Sequence-based reagent | T. mathranii genome | NCBI | NCBI:NC_014209.1 |
| Software, algorithm | CCP4 | https://doi.org/10.1107/S0907444910045749 | CCP4:7.0.042; RRID:SCR_007255 |
| Software, algorithm | Coot | https://doi.org/10.1107/S0907444904019158 | Coot:0.8.6.1; RRID:SCR_014222 |
| Software, algorithm | GraphPad | GraphPad | GraphPad:7.0a; RRID:SCR_002798 |
| Software, algorithm | HKL2000 | https://doi.org/10.1016/S0076-6879(97)76066-X | HKL2000:v0.98.714; RRID:SCR_015547 |
| Software, algorithm | Open Source PyMol | SourceForge https://sourceforge.net/projects/pymol/ | PyMol:v1.8.x; RRID:SCR_000305 |
Additional files
-
Supplementary file 1
X-ray crystallography statistics from data collection and refinement.
- https://doi.org/10.7554/eLife.36381.019
-
Transparent reporting form
- https://doi.org/10.7554/eLife.36381.020
-
Reporting file 1
- https://doi.org/10.7554/eLife.36381.021





