Multifactorial processes underlie parallel opsin loss in neotropical bats
Figures

Distribution of an intact open reading frame (ORF), mRNA transcript, and protein for the OPN1SW, OPN1LW, and RHO photopigments in ecologically diverse noctilionoid bats.
The composition of species diet follows Rojas et al. (2018), dietary types are indicated with the following symbols: invertebrates – moth, vertebrates – frog, fruit – fruit and nectar/pollen – flower. The species phylogeny follows Rojas et al. (2016) and Shi and Rabosky (2015). Vertical black bars, from left to right, indicate: (1) Noctilionoidea, (2) Phyllostomidae, (3) and Stenodermatinae. RNA-Seq data was generated to both infer the presence of an intact ORF (in combination with genomic and PCR sequence data) and to determine the presence of an expressed mRNA transcript. The presence of an intact ORF and mRNA transcript for RHO was verified across all transcriptomes. The presence/absence of a protein product for S- and L-opsins was assayed by IHC on flat mounted retinas. The presence of an intact ORF, mRNA, and protein are indicated by a filled color marker (OPN1SW – purple, OPN1LW – green and RHO – blue), and its absence by a white marker. Missing data (i.e. species for which we were unable to obtain tissue) are indicated with a grey marker with grey outline. Mismatches between intact ORFs and transcripts, or between transcripts and protein data are indicated by an inequality symbol. Note: OPN1SW protein assays for P. quadridens revealed polymorphisms within the sample, and we recorded positive OPN1SW assays in some P. poeyi individuals despite an apparent disrupted ORF. Finally, a grey marker with no outline indicates the failure of protein assay for some species represented by museum specimens (Tadarida brasiliensis, Phyllostomus hastatus, Sturnira tildae, Sturnira ludovici, Platyrrhinus dorsalis and Chiroderma villosum).
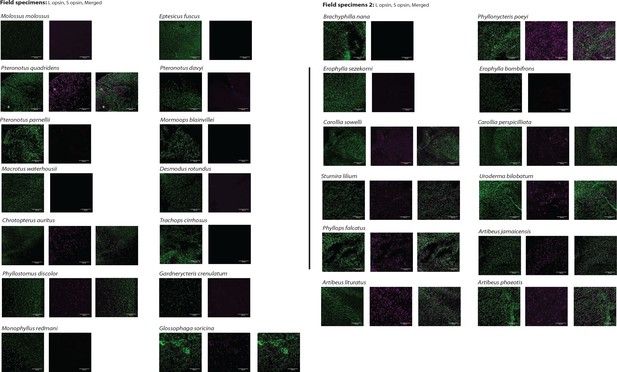
L- and S-opsin protein expression in the L- and S-cones of field samples
L- and S-opsin presence was assayed by IHC on field samples for 26 species with antibodies recognizing each of L- and S-opsin. For each species, L- and S-cone labeling are shown. For species with both L- and S-cones, the merged column is the result of the merged images of L- and S-cones from the same individual. No evidence of dual cones was found. Scale bar: 100 μm.
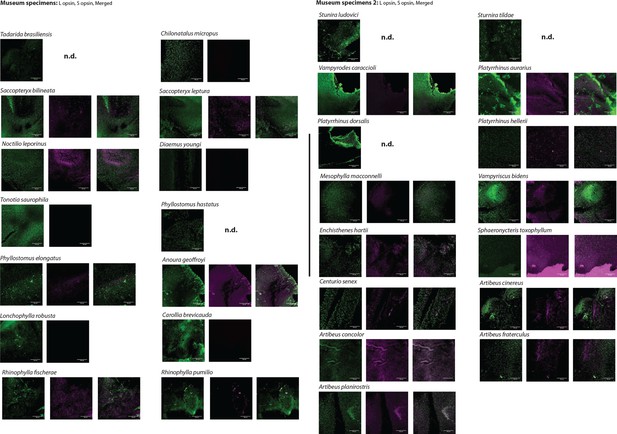
L- and S-opsin protein expression in the L- and S-cones of museum samples.
L- and S-opsin presence was assayed by IHC on museum samples for 29 species with antibodies recognizing each of L- and S-opsin. For each species, L- and S-cone labeling are shown. For species with both L- and S-cones, the merged column is the result of the merged images of L- and S-cones from the same individual. No evidence of dual cones was found. Scale bar: 100 μm.

Partial amino acid alignments for OPN1SW across the bat species studied.
For OPN1SW, the putative alternative start codon is indicated by the square box. Missing data is indicated by ‘X’. conserved amino acids by ‘.’, and the terminal STOP codon by ‘*’.
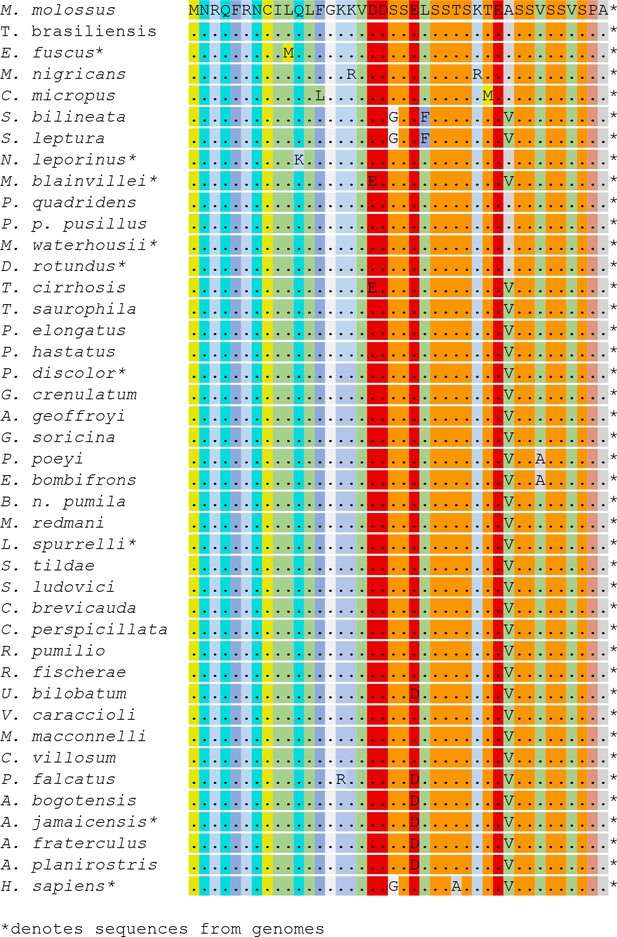
Partial amino acid alignments for OPN1LW across the bat species studied.
Missing data is indicated by ‘X’. conserved amino acids by ‘.’, and the terminal STOP codon by ‘*’.
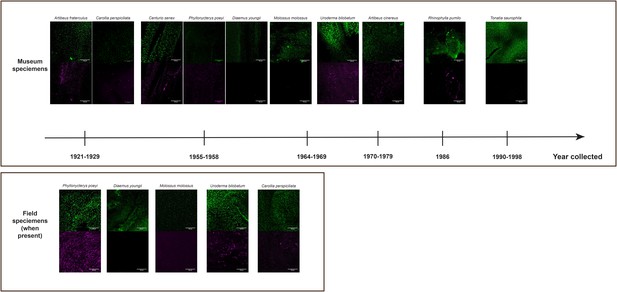
L- and S-opsin protein expression in L- and S-cones visualized in museum specimen of various ages (from 1921 to 1998) showing the consistency of the staining in old specimens.
For some of these samples, some fresh specimens that have been assayed are shown, demonstrating the consistency of the staining between field and museum specimens. For each species, L- and S-cone labeling are represented by the following colors: L-opsins – green and S-opsins – purple. Scale bar: 100 μm.
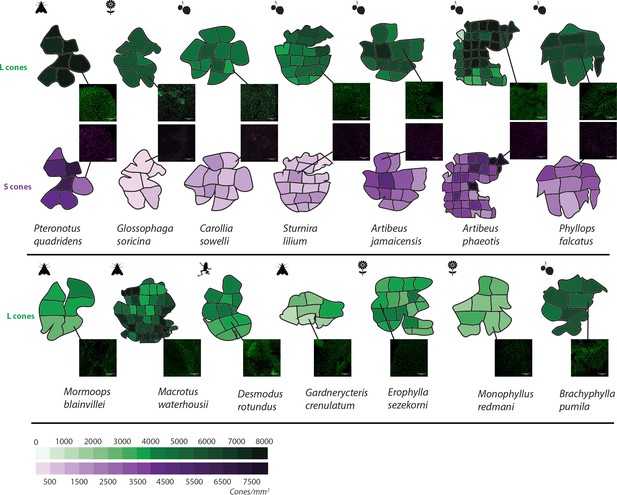
L and S opsin cone distribution in 14 representative noctilionoid bat species.
Density maps of L and S opsin cone topography in 14 noctilionoid bat species. For each species, a representative dissected flat-mounted retina is shown. Insets are representative IHC magnifications of flat mounted retinas immune-stained for either L- or S-opsins in the highlighted region. Dietary types are indicated with the following symbols: invertebrates – moth, vertebrates – frog, fruit – fruit and nectar/pollen – flower. Measured opsin densities (0–8000 cones/mm2) are represented by the following color scales: L-opsins – green and S-opsins – purple.
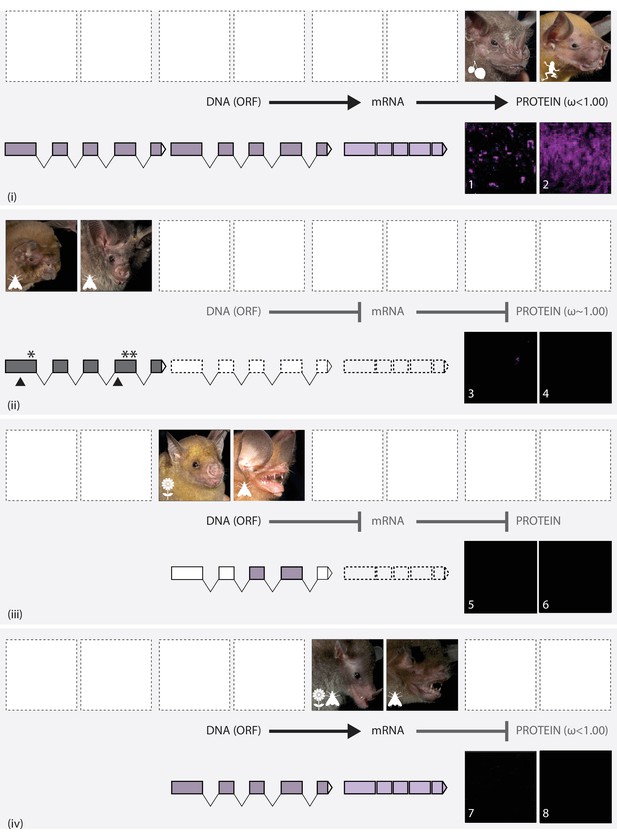
The putative routes explaining variation in S-cone presence in noctilionoid bats.
In each panel, upper images (left and right) show the gross phenotype of the eye in representative bat species, and lower images (numbered) show IHC magnifications of their respective flat mounted retinas immune-stained for S-opsin. Diets are depicted with the following symbols: invertebrates – moth, vertebrates – frog, fruit – fruit and nectar/pollen – flower. (i): Information in the intact DNA Open Reading Frame (dark purple) is transcribed to form mRNA (light purple), which is then translated into OPN1SW. Codon analyses reveal purifying selection. Example species: (L + 1) Artibeus jamaicensis; (R + 2) Noctilio leporinus. (ii): The DNA ORF is disrupted (grey) by the presence of STOP codons (*) and indels (black triangles). Neither OPN1SW mRNA (dashed boxes) nor OPN1SW are detected. Codon analyses reveal relaxed selection. Example species: (L + 3) Mormoops blainvillei; (R + 4) Macrotus waterhousii. (iii): Although the DNA ORF (dark purple) appears to be intact, information is not transcribed to mRNA (dashed boxes), and no OPN1SW is detected. Example species: (L + 5) Brachyphylla pumila (R + 6) Chilonatalus micropus. (iv): Information in the intact DNA ORF (dark purple) is transcribed to form mRNA (light purple); however, the OPN1SW is not detected. Codon analyses reveal purifying selection. Example species: (L + 7) Monophyllus redmani (R + 8) Pteronotus parnellii..
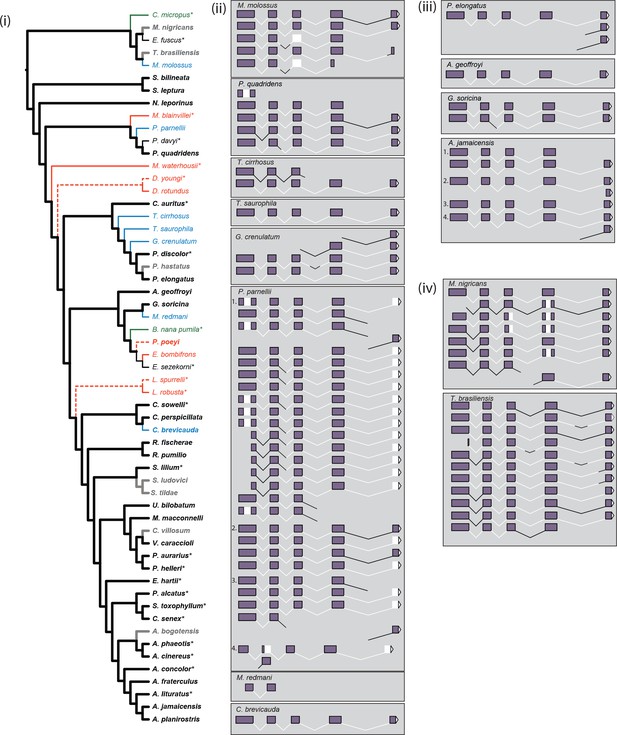
Inferred parallel losses of S-opsins mapped on to the species phylogeny and exonic content of reconstructed mRNA.
(i) Taxa and branches are colored as follows: presence of protein – black; presence of intact ORF but no protein – green; presence of mRNA but absence of protein – blue; evidence of pseudogenization (disrupted ORF) – red; protein status not determined – grey. Weight of branches indicates: inferred presence of protein – heavy; inferred absence of protein – light; protein absence based on evidence of gene loss but not confirmed by IHC – dashed light. We were either not able to recover mRNA, or preserved material was not available, for species marked with ‘*’, evidence for ORF status for Diaemus youngi taken from Kries et al., 2018. The species phylogeny follows Rojas et al., 2016 and Shi and Rabosky, 2015. (ii) Reconstructed mRNA transcript variants of seven species (M. molossus, T. cirrhosus, T. saurophila, G. crenulatum, P. parnellii, M. redmani, and C. brevicauda) with OPN1SW mRNA present but no detected protein, and P. quadridens for which presence of detected protein varied across individuals. The four biological replicates of P. parnellii are numbered 1–4. Sections of the mRNA are indicated as follows: exons 1–5 – purple filled boxes; introns 1–4 – black lines; the 3’UTR – white filled triangle; and missing regions – white regions. (iii) Reconstructed mRNA transcript variants of four species (P. elongatus, A. geoffroyi, G. soricina and A. jamaicensis) with OPN1SW mRNA present and detected protein. The four biological replicates of A. jamaicensis are numbered 1–4. Sections of the mRNA are indicated as above. (iv) Reconstructed mRNA transcript variants of two species (M. nigricans and T. brasiliensis) with OPN1SW mRNA present but protein status not determined. Sections of the mRNA are indicated as above.
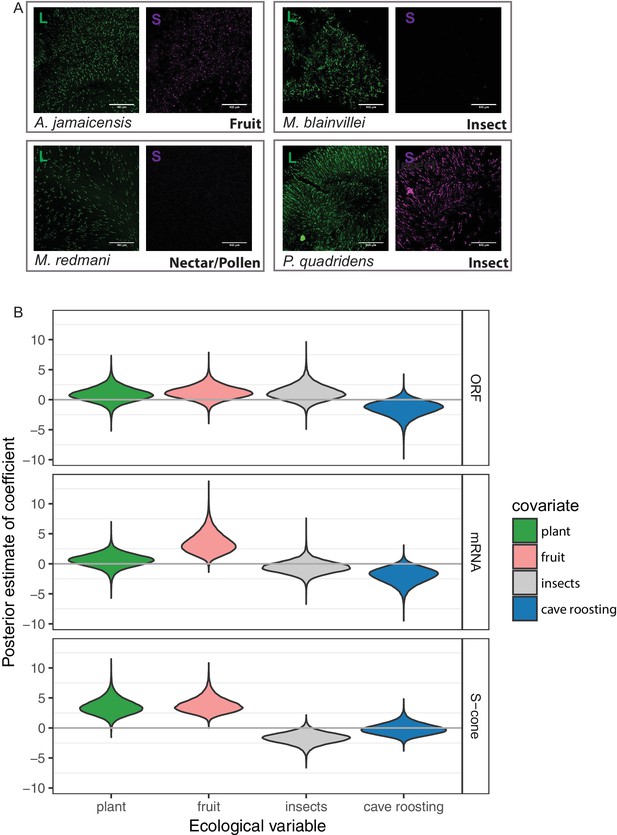
S-opsin presence is correlated with diet.
(A) Representative IHC magnifications of flat mounted retinas immune-stained for both L and S opsin for four species representatives of the diversity of phenotypes observed. Fruit-based diet: Artibeus jamaicensis, pollen/nectar-based diet: Monophyllus redmani, and insect-based diet: Mormoops blainvillei and Pteronotus quadridens. (B) Violin-plots of the posterior estimates of the coefficients for the presence of an OPN1SW ORF, mRNA, or cone as a function of ecological covariates. The gray horizontal lines indicate a coefficient of 0, or no effect of the covariate on the response. The high-probability density estimates for all coefficients are given in Tables S5-S7, and for fruit in Table 1.
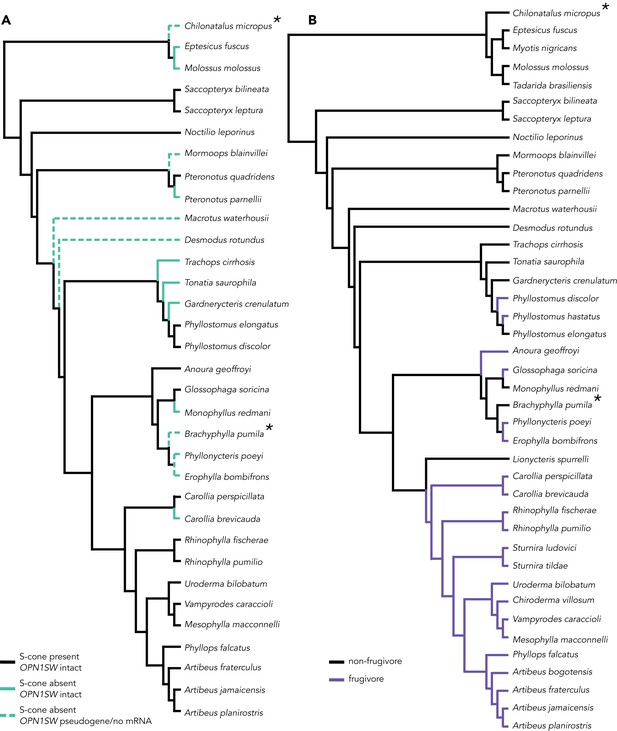
Branch class coding for molecular evolution analyses in PAML.
Tests for S-cone variation and diet were run for all three genes. The * indicates lineages absent from the OPNSW1 alignment. (A) Branch coding for species with variations S-cone presence. In the two-branch class test, black branches (S-cone present) were compared against teal branches (S-cone absent). In the three-branch class test, teal branches were subdivided between lineages in which OPN1SW is intact (solid teal branch) and lineages in which OPN1SW is a pseudogene or the mRNA was absent (dashed teal branch). (B) Branch coding for frugivorous (purple) v. non-frugivorous lineages and background branches (black).
Tables
Summary of Bayesian regression models of the presence of S-cones or the ln-transformed density of L-cones as a function of predictor variables.
R2, variance explained by sample-wide factors; Σ, species-specific phylogenetic effect for species.
| Formula | R2 | Parameter | Median | 2.5% | 97.5% |
|---|---|---|---|---|---|
| Presencei ~ a + b. fruit_prevalenti + Σ | 0.503 | intercept (a) | −1.10 | −3.19 | 0.52 |
| b | 3.66 | 1.85 | 6.76 | ||
| Σ | 1.94 | 0.02 | 8.64 | ||
| ln(density)i ~ a + b. fruit_prevalentj + Σj | 0.000 | intercept (a) | 8.42 | 8.00 | 8.84 |
| b | 0.22 | −0.21 | 0.66 | ||
| Σj | 0.14 | 0.04 | 0.46 | ||
| ln(density)i ~ a + b. S-cones_presentj + Σj | 0.240 | intercept (a) | 8.28 | 7.95 | 8.63 |
| b | 0.43 | 0.09 | 0.78 | ||
| Σj | 0.07 | 0.000 | 0.51 |
| Reagent type (species) or resource | Designation | Source or reference | Identifiers | Additional information |
|---|---|---|---|---|
| Antibody | sc-14363, goat anti-OPN1SW | Santa-Cruz Biotechnologies | RRID: AB_2158332 | (1∶1000) |
| Antibody | ab5405, rabbit anti-opsin red/green | Millipore Ibérica, Madrid | RRID: AB_177456 | (1∶750) |
| Antibody | donkey anti-goat Alexa Fluor 568 | Thermofisher | RRID:AB_2534104 | (1∶500) |
| Antibody | donkey anti-rabbit Alexa Fluor 647 | Thermofisher | RRID: AB_10891079 | (1∶500) |
| Software, algorithm | FIJI | https://fiji.sc/ | ||
| Commercial assay or kit | RNeasy Mini kits | Qiagen | ||
| Commercial assay or kit | Qiagen DNeasy Blood and Tissue Kit | Qiagen |
Additional files
-
Supplementary file 1
Specimen and sampling information for the tissues used by this study.
Unknown collection month indicated by ‘unk’, and (?) indicates uncertainty in the museum specimen collection date.
- https://doi.org/10.7554/eLife.37412.015
-
Supplementary file 2
Results of molecular evolution branch analyses for each of the three opsin genes tested for differences in rates of nonsynonymous to synonymous substitutions (ω) for lineages that lack the S-cone and lineages that have retained the S-cone.
Grey boxes indicate the preferred model inferred from a likelihood ratio test. lnL: log-likelihood; np: number of parameters; TL: tree length; k: kappa (transition/transversion ratios); LR: likelihood ratio; p: p-value of likelihood ratio of alternative relative to null for each test
- https://doi.org/10.7554/eLife.37412.016
-
Transparent reporting form
- https://doi.org/10.7554/eLife.37412.017





