A multicellular rosette-mediated collective dendrite extension
Figures
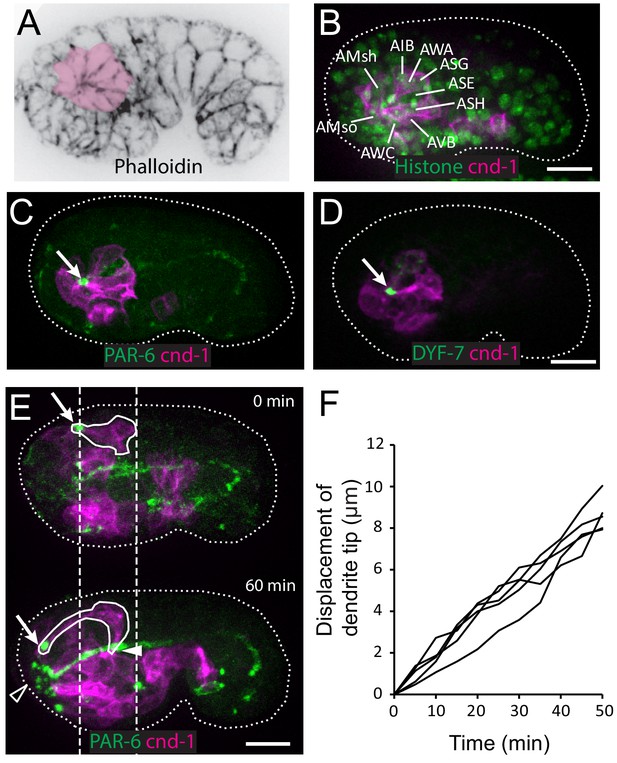
Multicellular rosette precedes collective dendrite extension.
(A, B) Amphid neurons form a multicellular rosette. (A) Image of embryo prior to amphid dendrite extension stained with phalloidin. A multicellular rosette is shaded in pink. In this and all subsequent figures, embryos are oriented with anterior to the left. (B) cnd-1p::PH::GFP labels amphid neurons in the rosette. Lineage-derived cell identities are shown. (C, D) PAR-6 and DYF-7 are localized to the rosette vertex. cnd-1p::PH::mCherry labels neurons. (E) Amphid dendrite tip migrates anteriorly. PAR-6 accumulates in the dendrite tip. cnd-1p::PH::mCherry labels neurons. Vertical dashed lines show initial position of amphid dendrite tip and posterior extent of amphid cell bodies. Arrows indicate dendrite tip. Closed arrowhead indicates amphid axon commissure. Open arrowhead indicates sensory depression. (F) Measured displacement of amphid dendrite tips from five embryos during dendrite extension. Scale bars in A-E, 10 µm.
-
Figure 1—source data 1
Displacement of dendrite tip from five embryos.
- https://doi.org/10.7554/eLife.38065.004
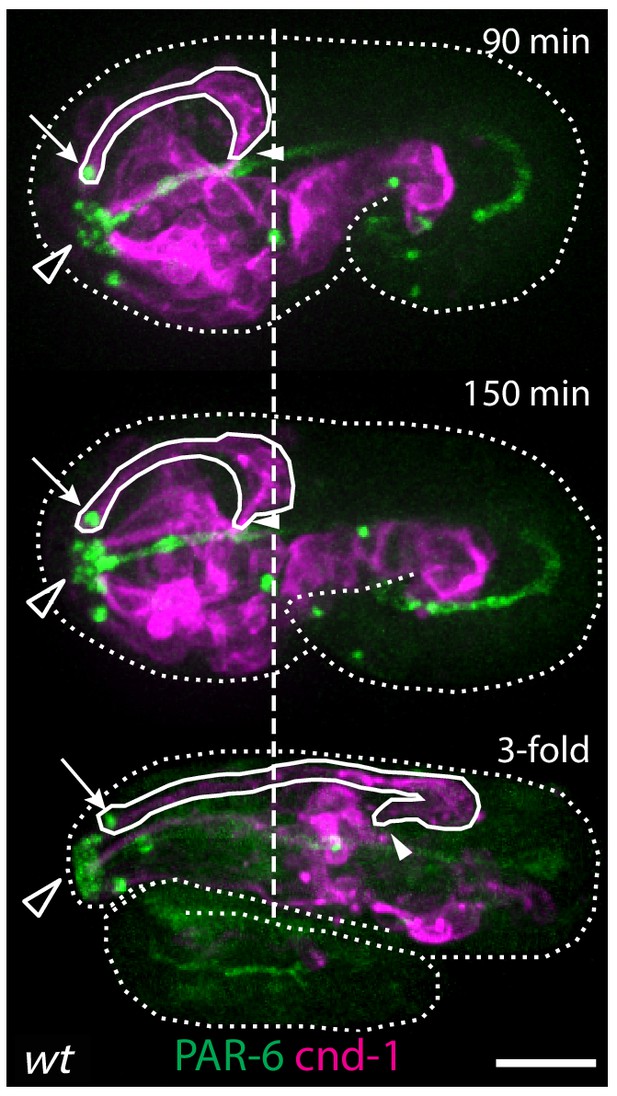
Retrograde extension in wild-type embryos.
After the initial anterior extension is complete and the dendrite tips reach the sensory depression (top panel), cell bodies start to move posteriorly as the embryo elongates, further extending the dendrites through retrograde extension. Arrows indicate amphid dendrite tips. Closed arrowheads indicate amphid commissure. Open arrowheads indicate the sensory depression. Dashed line marks the posterior end of the amphid neurons at the time when anterior dendrite extension is complete. Scale bar, 10 µm.
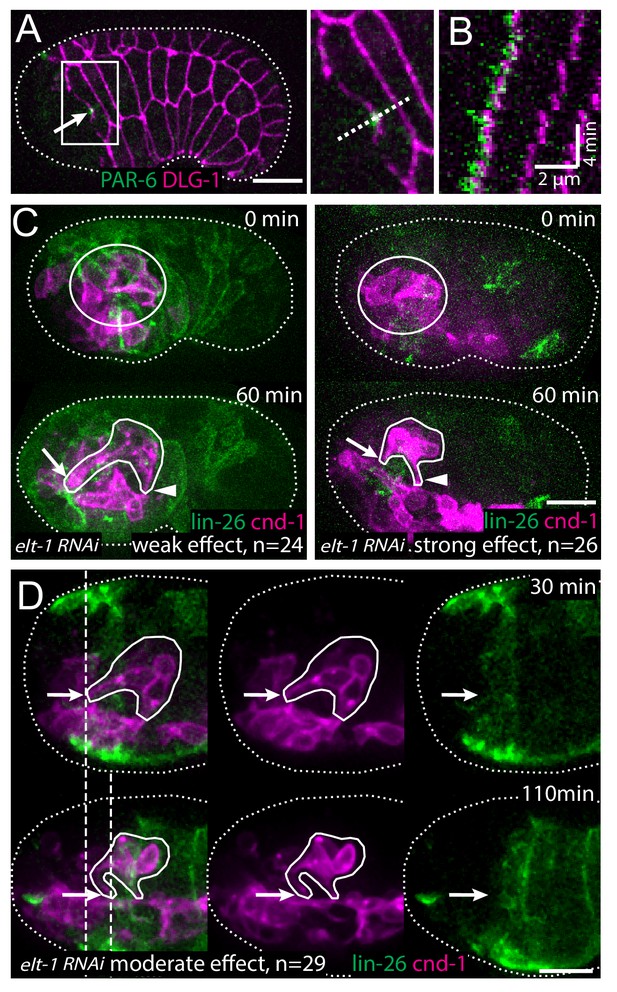
The amphid rosette vertex is attached to the epidermis.
(A) PAR-6-labeled amphid tip (arrows) is localized at the hyp5 leading edge. The apical junction of epidermal cells was labeled by DLG-1::GFP with xnIs17 transgene. The inset is shown magnified on the right. (B) A kymograph generated along the dashed line in the inset in A, showing the correlated anterior movement of the PAR-6 signal and the leading edge of the epidermal cell. (C) Example of an elt-1(RNAi) embryo with weak RNAi effect which is similar to the wild type (left) and an elt-1(RNAi) embryo with strong RNAi effect and minimal lin-26::GFP expression (right). In the latter group dendrites failed to extend (arrow), even though the amphid rosette (circle in the upper panel) and the amphid axon commissure both formed normally (arrowheads). (D) A representative elt-1(RNAi) embryo with moderate lin-26::GFP expression shows a partial anterior migration of the epidermis (upper panel) followed by posterior retraction. Dashed lines mark the leading edge of the epidermis before and after the retraction. The amphid dendrite tips follow the leading edge of the epidermis. Arrows mark the position of the dendrite tip. Scale bars in A-D, 10 µm.
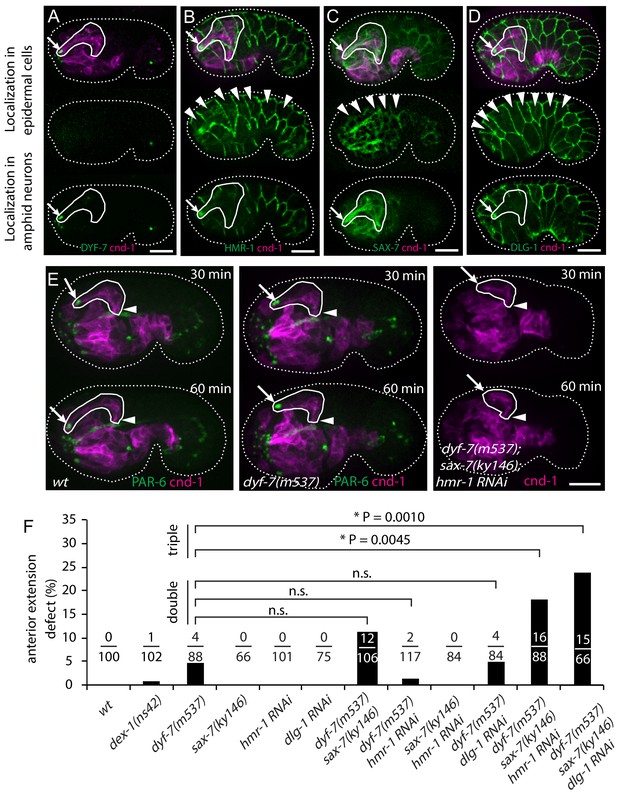
Multiple adhesion molecules function redundantly in amphid dendrite anterior extension.
(A–D) Localization of DYF-7, HMR-1, SAX-7 and DLG-1 (dlg-1(cp301)) in amphid neurons and epidermal cells. Middle panels, superficial focal plane showing localization in epidermal cells. Lower panels, deeper focal plane showing localization in amphid neurons. Arrows indicate amphid tips and arrowheads indicate epidermal cells. (E) Dynamics of dendrite anterior extension in wt, dyf-7(m537) and dyf-7;sax-7;hmr-1(RNAi) triple loss of function. (F) Frequency of anterior extension defects in single, double and triple loss of function embryos. Number of amphids scored is indicated. P-values were calculated with Fisher’s exact test (two-tailed). Threshold for significance was adjusted by Bonferroni correction to 0.01 for five comparisons. Scale bars in A-E, 10 µm.
-
Figure 3—source data 1
P values in multiple comparasions.
- https://doi.org/10.7554/eLife.38065.005
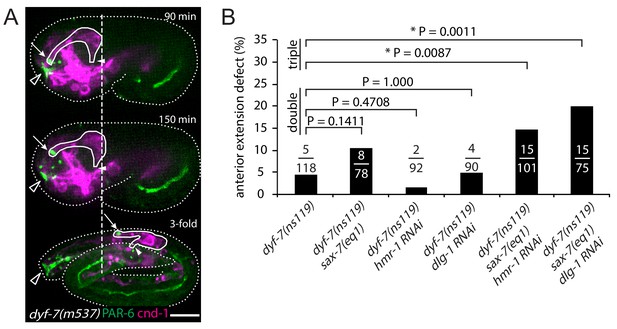
dyf-7 phenotype and genetic redundancy.
(A) Retrograde extension defects in dyf-7(m537) embryos.The dendrite tips detach from the sensory depression during embryo elongation, after successful anterior extension. See Figure 1—figure supplement 1 for the wild-type control. Arrows indicate amphid dendrite tips. Closed arrowheads indicate amphid commissure. Open arrowheads indicate the sensory depression. Dashed line marks the posterior end of the amphid neurons at the time when anterior dendrite extension is complete. Scale bar, 10 µm. (B) Frequency of anterior extension defects in single, double and triple loss of function embryos with dyf-7(ns119), sax-7(eq1), hmr-1(RNAi) or dlg-1(RNAi). Number of amphids scored is indicated. P-values were calculated with Fisher’s exact test (two-tailed). Threshold for significance was adjusted by Bonferroni correction to 0.01 for five comparisons.
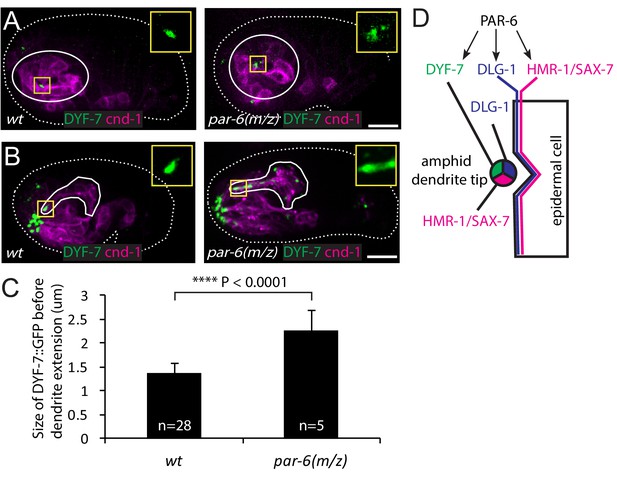
PAR-6 regulates localization of DYF-7.
(A, B) DYF-7 shows more dispersed localization in par-6(M/Z). (C) Quantification of the size of DYF-7::GFP in WT and par-6(M/Z) embryos before the dendrite tips migrate anteriorly. Data was shown as mean ±SD. P value was calculated with student t-test. (D) Schematic model of molecular localization between the amphid dendrite tips and epidermal cell. Scale bars in A-B, 10 µm.
-
Figure 4—source data 1
Size of DYF-7::GFP in wt and par-6(M/Z) embryos.
- https://doi.org/10.7554/eLife.38065.006
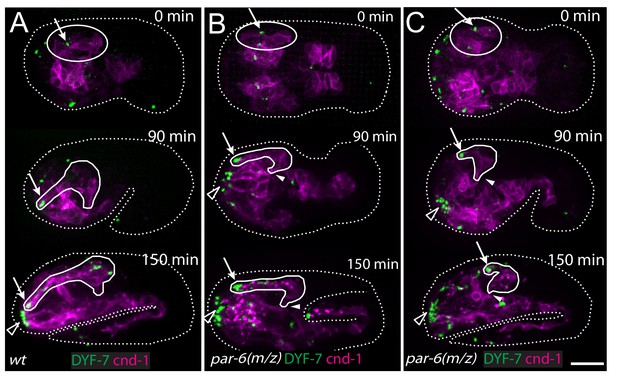
PAR-6 is required for amphid dendrite extension.
(A) The wild type. (B) Partial extension of amphid dendrites in par-6(M/Z) embryos that arrested at the 1.5-fold stage, indicating abnormal epidermal morphogenesis. (C) Partial extension of amphid dendrites in a par-6(M/Z) embryo that developed beyond the 1.5-fold stage, indicating more or less normal epidermal morphogenesis during dendrite extension. Arrows indicate amphid dendrite tips. Closed arrowheads indicate amphid commissure. Open arrowheads indicate the sensory depression. Scale bar, 10 µm.
Videos
Amphid dendrite anterior extension.
Time-lapse imaging of a wild-type embryo expressing cnd-1p::PH::mCherry to label sensory neurons and PAR-6::GFP to label dendrite tips. Dashed lines mark the initial position of the amphid neuron cell bodies at 0 min. The dendrite tip is tracked with an arrow. Scale bar, 10 µm.
Tables
| Reagent type (species) or resource | Designation | Source or reference | Identifiers | Additional information |
|---|---|---|---|---|
| Genetic reagent (E. coli) | OP50 | Caenorhabditis Genetics Center | OP50 | |
| Genetic reagent (C. elegans) | dex-1(ns42) III | PMID: 19344940 | ||
| Genetic reagent (C. elegans) | dyf-7 (ns119) X | PMID: 19344940 | ||
| Genetic reagent (C. elegans) | dyf-7 (m537) X | PMID: 19344940 | ||
| Genetic reagent (C. elegans) | sax-7(ky146) IV | Caenorhabditis Genetics Center | CX2993 | |
| Genetic reagent (C. elegans) | sax-7(eq1) IV | Caenorhabditis Genetics Center | LH81 | |
| Genetic reagent (C. elegans) | par-6(zu170) I | Caenorhabditis Genetics Center | FT36 | |
| Genetic reagent (C. elegans) | par-6(tm1425)/hIn1 [unc-54(h1040)] I | Caenorhabditis Genetics Center | JJ1743 | |
| Genetic reagent (C. elegans) | unc-101(m1) I | Caenorhabditis Genetics Center | DR1 | |
| Genetic reagent (C. elegans) | zbIs3[cnd-1p::PH::GFP] | PMID: 28441532 | ||
| Genetic reagent (C. elegans) | ujIs113[pie-1p::mCherry::H2B; nhr-2p::mCherry::HIS-24 -let-858UTR; unc-119(+)] II | Caenorhabditis Genetics Center | JIM113 | |
| Genetic reagent (C. elegans) | itIs1024[par-6::PAR-6::GFP] | Dr. Kenneth Kemphues | KK1024 | |
| Genetic reagent (C. elegans) | zyIs36[cnd-1p::PH::mCherry; myo-2p::mCherry]X | PMID: 28441532 | ||
| Genetic reagent (C. elegans) | mcIs40 [lin-26p::ABDvab- 10::mCherry; myo-2p::GFP] | Caenorhabditis Genetics Center | ML916 | |
| Genetic reagent (C. elegans) | xnIs17[dlg-1::GFP; rol-6(su1006)] | Caenorhabditis Genetics Center | FT63 | |
| Genetic reagent (C. elegans) | axIs1928[mCherry::PAR-6] | Caenorhabditis Genetics Center | JH2648 | |
| Genetic reagent (C. elegans) | dlg-1(cp301[dlg-1:: mNG-C1^3xFlag])X | Caenorhabditis Genetics Center | LP598 | |
| Genetic reagent (C. elegans) | ntIs1[gcy-5::GFP] | Caenorhabditis Genetics Center | OH3192 | |
| Genetic reagent (C. elegans) | oyIs44[odr-1::RFP + lin-15(+)]V | Caenorhabditis Genetics Center | PY2417 | |
| Genetic reagent (C. elegans) | zuIs43[pie-1::GFP::PAR-6::ZF1; unc-119(+)] | Caenorhabditis Genetics Center | JJ1743 | |
| Genetic reagent (C. elegans) | ddIs290[sax-7::TY1::EGFP:: 3xFLAG(92C12); unc-119(+)] | Caenorhabditis Genetics Center | TH502 | |
| Genetic reagent (C. elegans) | xnIs96 [hmr-1p::HMR-1::GFP:: unc-54 3'UTR; unc-119(+)] | Caenorhabditis Genetics Center | FT250 | |
| Genetic reagent (C. elegans) | kyIs4[ceh-23-unc-76-gfp::lin-15]X | Caenorhabditis Genetics Center | CX2565 | |
| Genetic reagent (C. elegans) | hmnEx149[dyf-7p::DYF-7 (ZP-sfGFP)-mCherry; rol-6(su1006)] | DOI: 10.1101/393850 | ||
| Genetic reagent (E. coli) | elt-1(RNAi) expressed in HT115 (DE3) | Source BioScience | C. elegans RNAi collection (Vadel) | RNAi Bacteria, Clone ID: 10019-B-8 |
| Genetic reagent (E. coli) | hmr-1(RNAi) expressed in HT115 (DE3) | Source BioScience | C. elegans RNAi collection (Ahringer) | RNAi Bacteria, Clone ID: I-5F23 |
| Genetic reagent (E. coli) | dlg-1(RNAi) expressed in HT115 (DE3) | Source BioScience | C. elegans RNAi collection (Ahringer) | RNAi Bacteria, Clone ID: X-8A08 |
| Software, algorithm | Fiji | https://fiji.sc/ |
Additional files
-
Transparent reporting form
- https://doi.org/10.7554/eLife.38065.013





