Structure of the human volume regulated anion channel
Figures
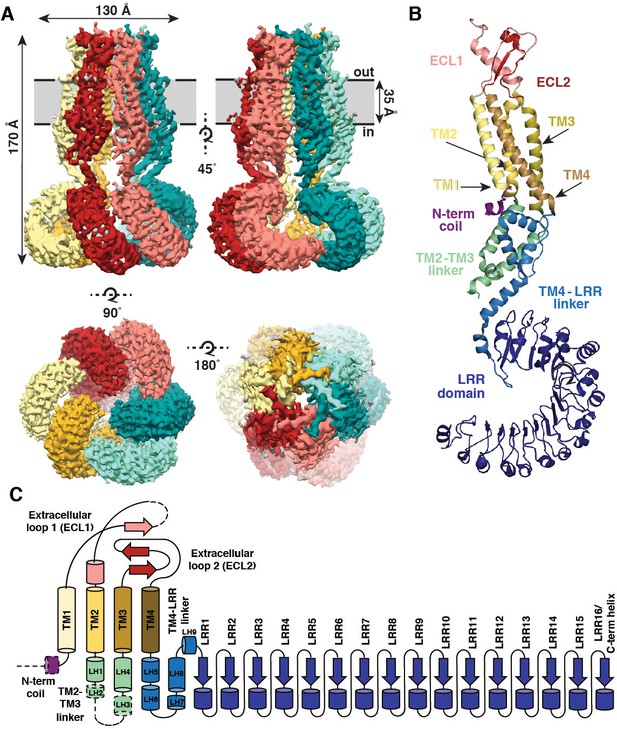
Overall architecture of homomeric SWELL1.
(A) Cryo-EM reconstruction of SWELL1 homohexamer viewed from the membrane plane highlighting a dimer pair (top left, red and pink subunits) and an interface between dimers (top right, pink and green subunits), from the extracellular side (bottom left), and from the cytosolic side (bottom right). (B) Detailed view of SWELL1 ‘inner’ protomer. (C) Topology diagram denoting secondary structural elements. Dashed lines indicate unresolved regions on both protomers in a dimer pair, while dashed shape borders indicate regions that are only resolved on one protomer.
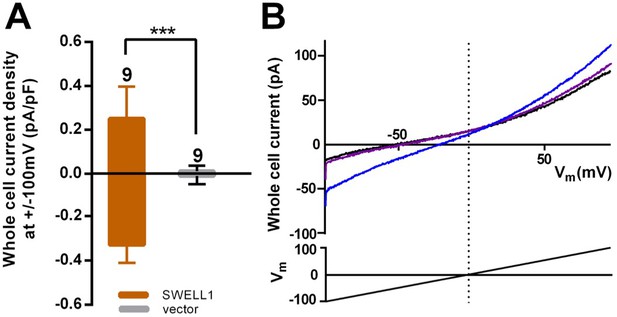
SWELL1 overexpression in cells lacking other LRRC8 subunits produces small DCPIB-sensitive swelling-induced whole cell currents.
(A) HeLa LRRC8-/- cells were transfected with SWELL1-pIres-GFP or empty pIres-GFP vector and tested for hypotonic (230 mOsm/kg) solution-induced currents using voltage ramp protocols (Qiu et al., 2014). Whole cell currents were measured at −100 and +100 mV before and 5–7 min after hypotonic challenge and normalized to cell capacitance. Shown are means ± s.e.m. for the number of cells indicated from 3 separate experiments. (B) Hypotonic solution-induced currents (blue) were blocked by DCPIB (20 μM) (purple); ramp-induced current in isotonic solution prior to challenge with hypotonic solution is shown in black. Traces were filtered at 1900 Hz and averaged (ten sweeps each). Extracellular solution (in mM): 90 NaCl, 2 KCl, 1 MgCl2, 1 CaCl2, 10 HEPES, 110 mannitol (isotonic, 300 mOsm/kg) or 30 mannitol (hypotonic, 230 mOsm/kg), pH 7.4 with NaOH. Recording pipettes were filled with intracellular solution containing (in mM): 133 CsCl, 5 EGTA, 2 CaCl2, 1 MgCl2, 10 HEPES, 4 Mg-ATP, 0.5 Na-GTP (pH 7.3 with CsOH; 106 nM free Ca2+) and had resistances of 2–3 MΩ.
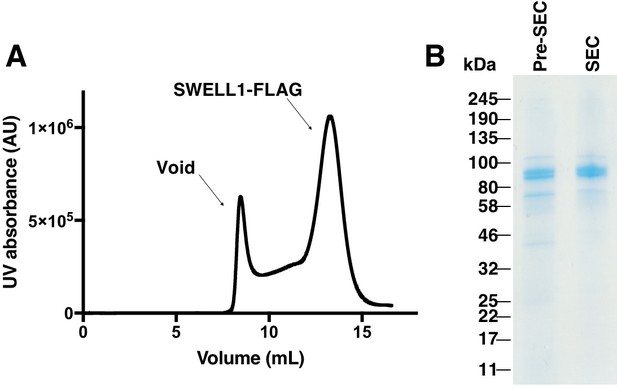
Purification of SWELL1-FLAG.
(A) Gel filtration trace of SWELL1-FLAG after affinity purification. (B) SDS-PAGE gel of SWELL1-FLAG. Lane 1 is eluate after affinity purification (Pre-SEC) and lane 2 is combined fractions from SWELL1-FLAG peak post size exclusion. Expected molecular weight is 94 kDa.
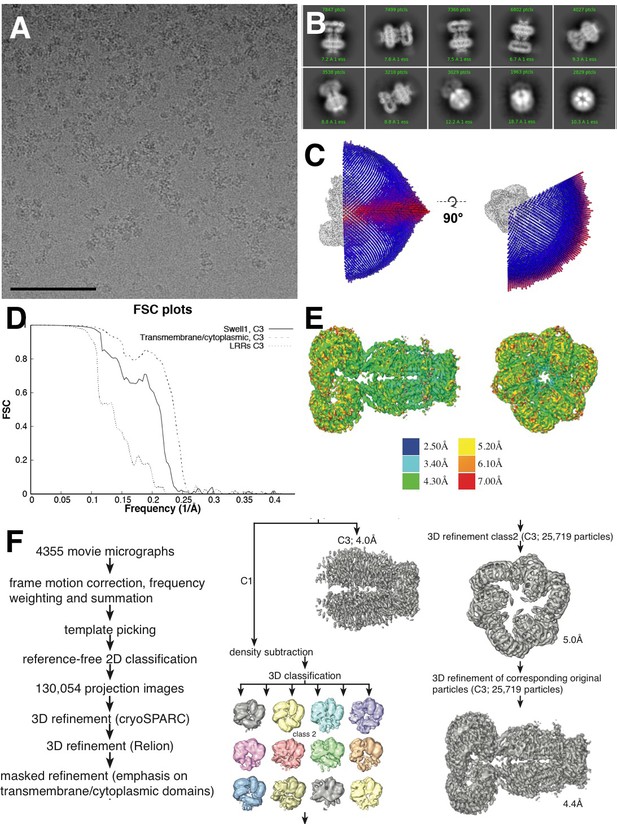
Cryo-EM data collection.
(A) Representative aligned micrograph (scale bar, 100 µm). 4355 movies were collected of SWELL1-FLAG in vitreous ice. (B) Representative 2D classes showing range of orientations of SWELL1-FLAG particles. (C) Euler distribution for final map. Applied C3 symmetry was used, thus only one third of the sphere is shown. (D) Fourier shell correlation between two independently refined cryo-data half sets. (E) Local resolution estimates of the final reconstruction. (F) Data-processing flow chart.

Model-to-map fit of electron density.
Selected regions of the model are shown with superimposed electron density. Density is derived from the final C3 symmetry-imposed map sharpened with a b-factor of −110 Å2. The cytosolic loop helices (LH1-4) and LRR 15–16 are from the outer subunit.
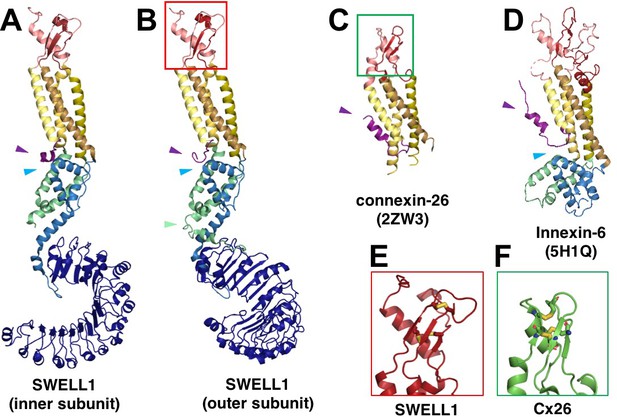
SWELL1 subunit pair and structural homology of SWELL1 structure to connexin-26 and innexin-6 structures.
(A–B) Comparison of the SWELL1 ‘inner subunit’ and ‘outer subunit’ in a dimer pair. In the inner subunit, the TM2-TM3 linker contributes two long parallel helices (LH1 and LH4) with a flexible loop of res175-231 between them, while the outer subunit has an additional pair of parallel helices (LH2 and LH3) kinked to place them between the two subunits, though still with an unresolved loop of residues 176–213 between them. LH2, corresponding to residues R214-E236, sits atop of the LRR in the outer subunit (green arrow). Additionally, there is a rotation of the LRR domains relative to the TM domains that allows them to dimerize. (A–D) The LRRC8 family has been shown to be related by weak sequence homology to pannexins (Abascal and Zardoya, 2012), which are in turn related by structural homology to connexins and innexins. The transmembrane helices (TM1-4) of SWELL1 share the same order and arrangement as those of connexin-26 (Cx26; PDB: 2ZW3) (Maeda et al., 2009) and innexin-6 (PDB: 5H1Q) (Oshima et al., 2016) with TM1 closest to the central axis and TM3 and TM4 facing the membrane environment. An N-terminal helix in both Cx26 and innexin-6 creates a pore funnel that forms the narrowest constriction of the channel (Maeda et al., 2009; Oshima et al., 2016), which may be recapitulated in SWELL1 (purple arrows) (E–F) The extracellular loops of SWELL1 (red box) share 3 structurally conserved disulfide bonds with connexin-26 (green box), as well as a three-strand antiparallel beta sheet composed of the antiparallel beta hairpin of ECL2, and a beta strand from ECL1 (Maeda et al., 2009). (A–B, D) In SWELL1, cytoplasmic core formed by the intracellular linker domains share the same topology with portions of the C-terminal domains of distantly related innexin-6 (blue arrows) (Oshima et al., 2016).

Alignment of LRRC8 subunits (res 1–435).
Human LRRC8 family aligned using ClustalW. Secondary structure and domain assignments are annotated above. Residues mentioned in text are boxed red.

Alignment of LRRC8 subunits (res 436–810).
Human LRRC8 family aligned using ClustalW. Secondary structure and domain assignments are annotated above. Residues mentioned in text are boxed red.
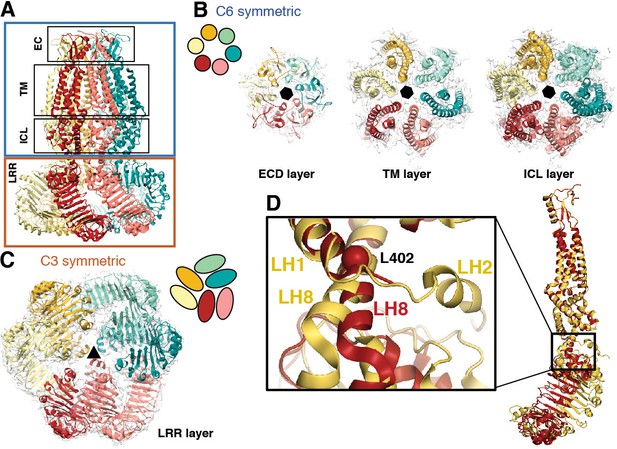
Subunit arrangement exhibits symmetry mismatch.
(A) SWELL1 model viewed from the membrane plane with domain layers viewed perpendicular to the symmetry axis. (B–C) Domain layers viewed from the top of the channel grouped according to shared symmetry with simple schematic to demonstrate subunit arrangement. (B) From left to right: extracellular domain layer (EC), transmembrane domain layer (TM), and intracellular linker domain layer (ICL) all share the same 6-fold rotation symmetry axis (black hexagon). (C) The LRR domain layer has 3-fold rotational symmetry (black triangle), resulting from parallel pairing of three sets of LRR domains. (D) Asymmetry in LRR pairing arises from a hinge at L402 on LH8 that allows rotation of the LRR domain as a rigid body in a dimer pair. The first two TM domains of the inner (red) and outer (yellow) subunits are aligned to one another using the PyMOL align function.
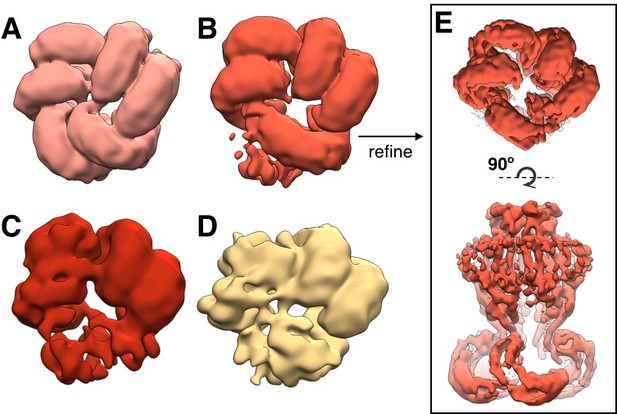
Flexibility in LRR domains observed during 3D classification.
Bottom view of arrangement of LRR domains in representative subclasses of density-subtracted 3D classification (see Materials and methods). Pre-cleaned particle picks were distributed into classes with clear organization of three (A; 25,607 particles), two and one half (B; 29,638 particles), two (C; 7197 particles), and one (D; 8071 particles) pairs of LRRs are observed. The highest resolution map was produced with the particles in the classes in which all three pairs are resolved, while the classes in which one or more LRR pairs are flexible produce lower resolution refinements. (E) Example refinement of LRR class (B) with two and one half LRR pairs resolved and one LRR flexible (~5.7 Å resolution).
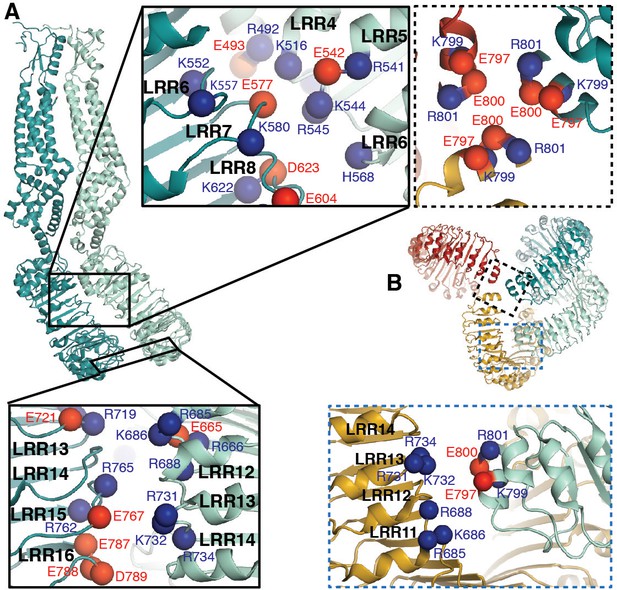
LRRs interact via charged residues at dimer interfaces and C-termini.
(A) One dimer of SWELL1 subunits. Charged residues both of opposite and similar charges face each other in the interface between the two LRR domains (insets, top middle and bottom left; blue spheres are positively charged residues (Arg, Lys, and His), red spheres are negatively charged residues (Asp and Glu)). (B) C-termini of the two protomers in a dimer interact with regions of the neighboring LRR domain. Two of three ‘outer’ subunits are removed for clarity. ‘Inner’ subunits may be able to coordinate with one another via a triad of charged residues (E800) at their C-termini (inset, dashed border, top right), while the C-termini of the ‘outer subunit’ may interact with the edge of the neighboring outer subunit via charged residues R688 on LRR12 and/or K732 on LRR13 (inset, dashed blue border, bottom right).
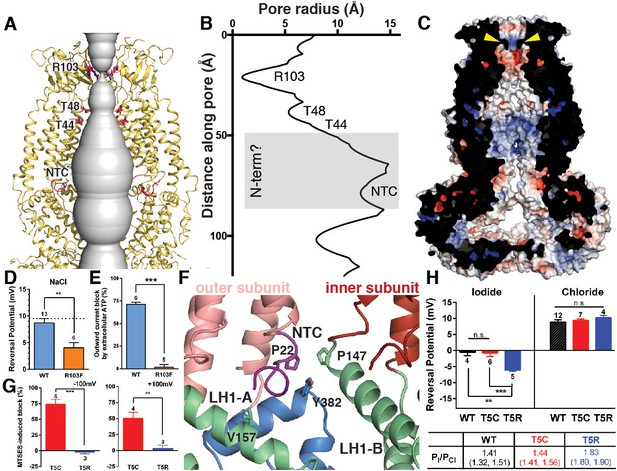
Ion pore structure and characterization of channels with mutations at pore-contributing residues R103 and T5.
(A) Cartoon model of the SWELL1 pore, with two subunits removed for clarity. A surface representation of the radial distance between the protein surface and the pore axis is shown in grey. Pore-facing residues R103, T48 and T44, and N-terminal coil (NTC) are labeled in pink. (B) Graph of van der Waals radii of the pore, plotted against distance along the pore axis. Locations of residues R103, T48, T44, and NTC are labeled along 2D plot. Grey box covers potential area the N-terminus might occupy. (C) Electrostatic surface potential of channel pore, viewed by vertical cross-section. Narrow constriction on the extracellular side of the channel is formed by a ring of R103 residues (yellow arrows). Calculated using APBS implemented by Pymol2.0 with potentials ranging from −10 kT (red) to +10 kT (blue). (D–E) Cells expressing heteromeric VRACs composed of mutant SWELL1-R103F + LRRC8C show reduced chloride selectivity and insensitivity to external ATP block. (D) For highly Cl- selective channels, the voltage at which there is no net current (Vrev) is close to the equilibrium potential for Cl- (in these experiments ECl = +9.75 mV; indicated by the dotted line). Vrev of currents mediated by SWELL1-R103F-containing channels (orange bar; +4.6 ± 1.0 mV (mean ± s.e.m., n = 6 cells from 3 separate transfections)) is significantly reduced compared to WT (blue bar; +8.8 ± 0.8 mV (n = 13 from 6 separate transfections); p = 0.003, Student’s t-test. (E) The percent block of whole cell leak subtracted hypotonic-induced currents by extracellular applied Na2ATP (2 mM) was determined at +100 mV. Outward WT SWELL1 + LRRC8C-mediated currents are blocked 72 ± 2% (mean ± s.e.m., n = 7 from 4 separate transfections; blue bar). Outward currents mediated by SWELL1-R103 + LRRC8C are not blocked by extracellular ATP (2 ± 3% (mean ± s.e.m., n = 5 from 3 separate transfections; orange bar); this difference is highly significant (p = 5.4e^-8, Student’s t-test). (F) Detailed view of coordination of NTC (purple). The NTC makes intrasubunit contacts with V157 on LH1 and a conserved Y382 at the kink between LH6 and LH7 of the TM4-LRR linker. Additionally, P22 of the NTC makes an intersubunit contact with a conserved P147 at the kink between TM2 and the TM2-TM3 linker of the neighboring subunit. (G–H) SWELL1-T5 is close to or part of the pore. (G) A cysteine mutation at SWELL1-T5 confers sensitivity to the polar MTS reagent MTSES applied extracellularly; maximum percent block of T5C-containing channels (red bars) by 3.33 mM MTSES was 74.2 ± 7.7% at −100 mV (left) and 50.4 ± 9.5% at +100 mV (right) (n = 5; mean ± s.e.m.; four separate transfections). The unmodifiable T5R-containing heteromeric channels (blue bars) are unaffected (n = 3 from 3 separate transfections; p=0.0009 at −100 mV and p=0.010 at +100 mV, Student’s t-test). (H) Relative permeability PI/PCl is enhanced by the T5R mutation. Reversal potentials in iodide (left) and chloride (right) 230 mOsm/kg solutions are shown for the number of cells from 3 to 7 separate transfections (WT-, T5C- and T5R-expressing cells were from 6, 4, and 3 transfections, respectively, in the Cl- condition, and 3, 3, and 3 transfections, respectively, in the I- condition). The Vrev of currents mediated by SWELL1-T5R + LRRC8C (blue) in I- solution was significantly more negative than either WT- (black) or T5C (red)-containing channels (p=0.0088 (**) and 0.0047 (***), respectively. Table: PI/PCl is shown as means with lower and upper 95% confidence intervals.
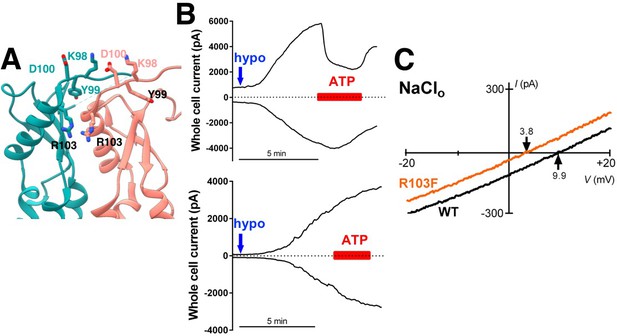
Additional characterization of selectivity at extracellular residues.
(A) Detailed view of neighboring extracellular loops with KYD motif and pore-constricting residue R103 labeled. (B) Representative data from HeLa LRRC8-/- cells transfected 1–3 days earlier with WT SWELL1 (top) and SWELL1-R103F (bottom) together with LRRC8C in a 2:1 ratio. Whole cell currents at +100 mV (upper trace) and −100 mV (lower trace) were measured from voltage ramp-induced currents and plotted. Blue arrow indicates addition of hypotonic solution; red bar, Na2ATP. Chloride solutions described in bianionic experiments (Qiu et al., 2014; Syeda et al., 2016) were used (Cl- in: 130 mM; Cl- out: 88 mM). For cells expressing wildtype SWELL1 +LRRC8C-containing channels outward currents were rapidly reduced by ATP block, while essentially no ATP block was observed in cells expressing R103F-containing channels. (C) Representative leak subtracted whole cell ramp-induced currents elicited by hypotonic solutions (230 mOsm/kg) containing 88 mM NaCl/10 mM HEPES from HeLa LRRC8-/- cells overexpressing WT (black trace) or the R103F mutant (orange trace) together with LRRC8C at a 2:1 ratio. The calculated Pother/PCl was 0.05 + 0.04 (n = 12; mean + s.e.m.) and 0.27 + 0.05 (n = 5) for WT and R103F, respectively (p=0.00036 using Student’s t-test). The relative permeability of other ions to Cl- was determined from the Erev observed in individual cells using the equation (Yang et al., 2012). Pother/PCl = ([Cl-]o*e(F(Erev)/RT) – [Cl-]i)/[Na+]o where [Cl-]o = 88mM, [Cl-]i = 130mM, [Na+]o = 88mM.
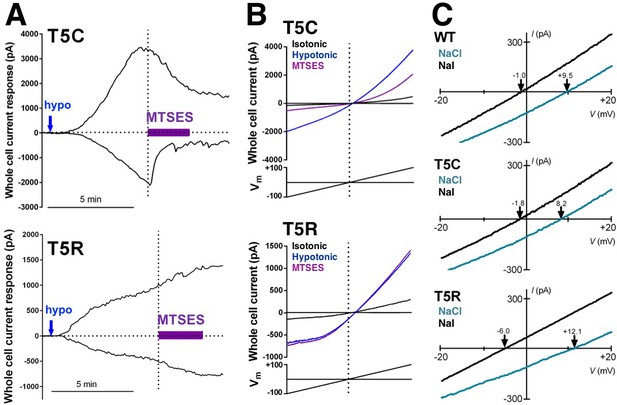
Representative data from cells expressing mutations in SWELL1 at T5 in the unresolved N-terminal region in heteromeric VRAC.
(A,B) Representative examples of whole cell currents tested for sensitivity to the extracellularly applied polar MTS reagent MTSES. (A) MTSES blocks SWELL1-T5C-containing channels (top) but has no effect on T5R-containing channels that are not modifiable by MTS reagents. (B) Whole cell currents induced by hypotonic solution (230 mOsm/kg; blue arrows) in HeLa LRRC8-/- cells heterologously expressing SWELL1-T5C (top) or SWELL1-T5R (bottom) with LRRC8C in a 2:1 ratio (0.8 and 0.4 γ/ml). MTSES (purple bar, 3.33 mM) strongly reduced T5C-but not T5R-mediated currents in a manner consistent with covalent modification since reversibility of the T5C block was not observed during washout. Representative currents elicited by voltage ramps from −100 mV to +100 mV are shown (black, before; blue, in hypotonic solution before MTSES; purple, during MTSES). (C) Representative leak subtracted whole cell ramp-induced currents elicited by hypotonic solutions (230mOsm/kg) containing either 88 mM NaCl/10 mM HEPES (green traces) or 88 mM Na-Iodide/10 mM HEPES (black traces) from HeLa LRRC8-/- cells overexpressing WT, T5C or T5R mutants together with LRRC8C at 2:1 ratio. Bianionic solutions described in Qiu et al. (2014) were used to determine relative permeability (PI/PCl).
Tables
| Reagent type (species) or resource | Designation | Source or reference | Identifiers | Additional information | |||
|---|---|---|---|---|---|---|---|
| Gene (Homo sapiens) | LRRC8A | Origene | Uniprot: Q8IWT6 | ||||
| Cell line (Homo sapiens) | Freestyle 293-F | ThermoFisher | RRID: CVCL_D603 | ||||
| Cell line (Homo sapiens) | HeLa | ATCC | RRID: CVCL_0030 | ||||
| Recombinant DNA reagent | pcDNA3.1/Zeo(-) | ThermoFisher | cat no: V86520 | ||||
| Chemical compound | digitonin | Sigma-Aldrich | CAS Number: 11024-24-1 | ||||
| Software, algorithm | MotionCor2 | doi:10.1038/nmeth.4193 | |||||
| Software, algorithm | EMHP | doi:10.1093/bioinformatics/btx500 | |||||
| Software, algorithm | Gctf | doi:10.1016/j.jsb.2015.11.003 | |||||
| Software, algorithm | FindEM template correlator | doi:10.1016/j.jsb.2003.11.007 | |||||
| Software, algorithm | cryoSPARC | doi: 10.1038/nmeth.4169 | https://cryosparc.com/ | ||||
| Software, algorithm | RELION | RRID:SCR_016274 | |||||
| Software, algorithm | Rosetta | RRID:SCR_015701 | https://www.rosettacommons.org/software | ||||
| Software, algorithm | Robetta | http://robetta.bakerlab.org/ | |||||
| Software, algorithm | COOT | RRID:SCR_014222 | http://www2.mrc-lmb.cam.ac.uk/personal/pemsley/coot/ | ||||
| Software, algorithm | Phenix | RRID:SCR_014224 | https://www.phenix-online.org/ | ||||
| Software, algorithm | PyMOL | PyMOL Molecular Graphics System, Schrodinger LLC | RRID:SCR_000305 | http://www.pymol.org/ | |||
| Software, algorithm | UCSF Chimera | UCSF | RRID:SCR_004097 | http://plato.cgl.ucsf.edu/chimera/ | |||
Additional files
-
Supplementary file 1
Cryo-EM data collection, refinement and validation statistics.
- https://doi.org/10.7554/eLife.38461.017
-
Transparent reporting form
- https://doi.org/10.7554/eLife.38461.018
-
Reporting standard 1
- https://doi.org/10.7554/eLife.38461.019





