Emergence of trait variability through the lens of nitrogen assimilation in Prochlorococcus
Figures
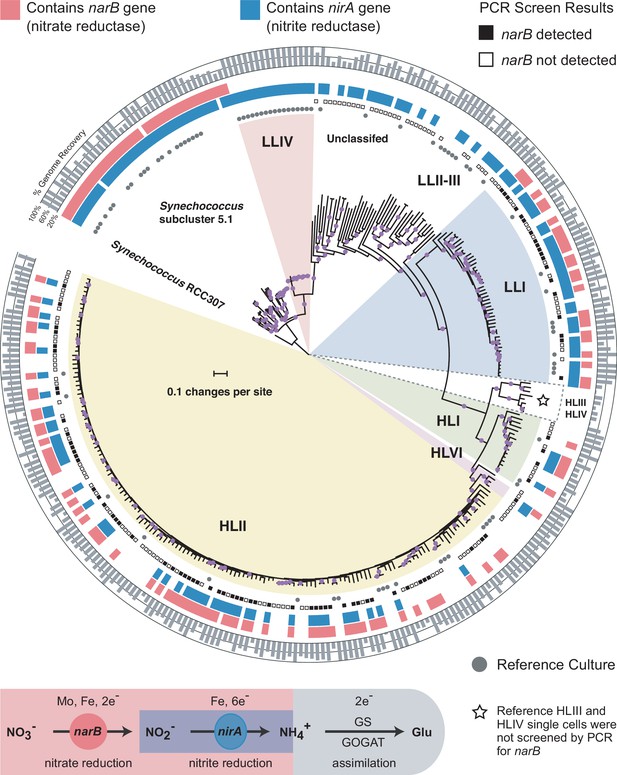
Distributions of the nitrate reductase (narB) and nitrite reductase (nirA) genes across a core marker gene phylogeny of 329 Prochlorococcus and Synechococcus genomes.
Selected Prochlorococcus clades are highlighted as pie slices. Red bars indicate genomes with the potential to use nitrate based on the presence/absence of a narB gene. Blue bars indicate genomes with an annotated nirA gene in the genome assembly. The outer ring indicates the estimated percentage of the genomes recovered as a gray bar chart. Reference culture genomes are indicated by gray circles and the results of the PCR screen for narB are indicated by filled (present) and open (absent) squares. Single cells belonging to the HLIII and HLIV clades of Prochlorococcus were not screened by PCR and are only included as additional reference genomes. The nucleotide phylogeny is based on a concatenated alignment of 37 marker genes in the PhyloSift software package and inferred using maximum likelihood in RAxML (GTRCAT model) with automatic bootstopping criteria (250 replicate trees). Filled purple circles on branches indicate that the associated genomes clustered together in at least 75% of trees.
-
Figure 1—source data 1
Compressed tar archive (zip format) containing the concatenated codon alignment (fasta format) and tree file (newick format) used to generate Figure 1.
- https://doi.org/10.7554/eLife.41043.003
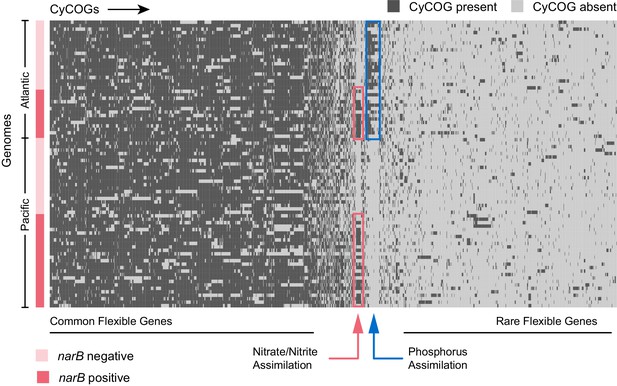
Hierarchical clustering of presence and absence distributions for flexible CyCOGs found in 83 Prochlorococcus HLII single cell genomes with genome recoveries of at least 75% (median 90%).
Genomes are sorted by Atlantic and Pacific Oceans and by the presence/absence of the narB gene – a marker for the capacity to assimilate nitrate. Other than genes in the nitrate assimilation gene cluster (green box), no CyCOGs were over- or under-represented among the flexible genes in genomes containing narB. Genomes from the Atlantic, regardless of whether or not they contained the narB marker gene, were enriched in phosphorus assimilation genes (blue box).
-
Figure 2—source data 1
Binary matrix containing the raw presence and absence data for each CyCOG in each genome analyzed for Figure 2 and Figure 2—figure supplement 1.
- https://doi.org/10.7554/eLife.41043.006
-
Figure 2—source data 2
Compressed tar archive (zip format) containing input and output files for gene enrichment analysis using BiNGO 3.0.3 (Maere et al., 2005) in Cytoscape 3.4 (Shannon et al., 2003).
- https://doi.org/10.7554/eLife.41043.007
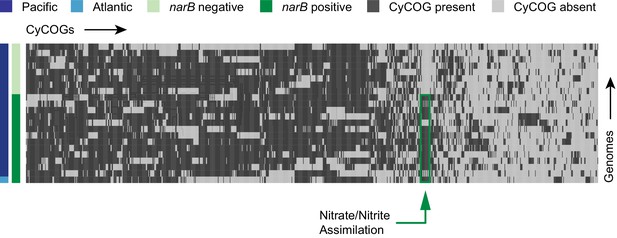
Hierarchical clustering of presence and absence distributions for flexible CyCOGs found in 22 Prochlorococcus single cell genomes belonging to the LLI clade with genome recoveries of at least 75% (median 87%).
Genomes are sorted by Atlantic and Pacific Oceans and by the presence/absence of the narB gene – a marker for the capacity to assimilate nitrate. Other than genes in the nitrate assimilation gene cluster (green box), no CyCOGs were over- or under-represented among the flexible genes in genomes containing narB.
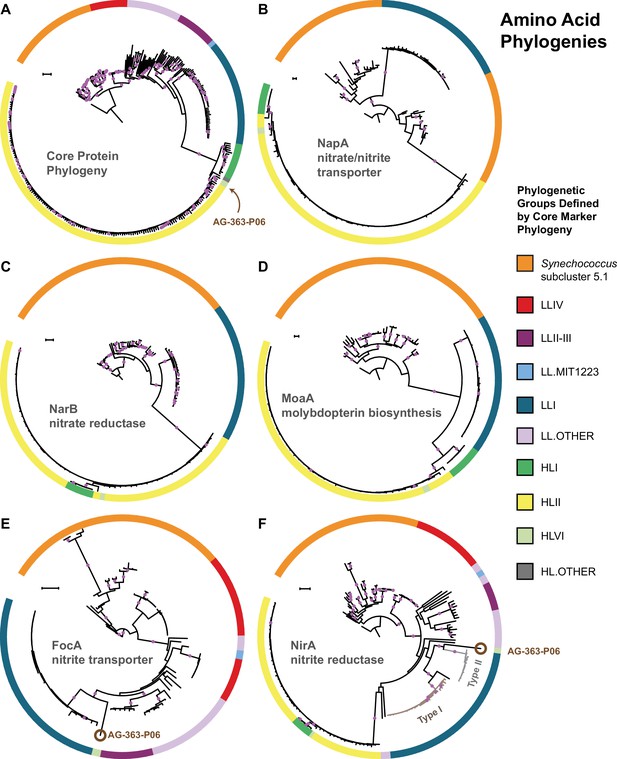
The core marker protein phylogeny of Prochlorococcus and Synechococcus.
(A) in comparison to phylogenies for the nitrate/nitrite transporter, NapA (B), the nitrate reductase, NarB (C), the molybdopterin biosynthesis protein, MoaA (D), the nitrite transporter, FocA (E), and the nitrite reductase, NirA (F). Interclade horizontal gene transfer is minimal for genes encoding proteins in the upstream half of the nitrate assimilation pathway (B–D) since clades defined by the core phylogeny (A) remain separate. Horizontal gene transfer is observed in a few instances for genes encoding proteins in the downstream half of the nitrate assimilation pathway (E–F). One single cell (AG-363-P06; brown circle) from the high-light adapted HLVI clade possesses FocA and NirA proteins most similar to those from low-light adapted Prochlorococcus. Prochlorococcus belonging to the LLI clade possess two types of NirA as indicated by well supported phylogenetic divergence of NirA among LLI cells. Filled purple circles on branches indicate that the associated taxa clustered together in at least 75% of trees. Scale bars are 0.1 changes per site.
-
Figure 3—source data 1
Compressed tar archive (zip format) containing codon alignments (fasta format) and tree files (newick format) used to generate Figure 3 and Figure 3—figure supplement 1.
- https://doi.org/10.7554/eLife.41043.010
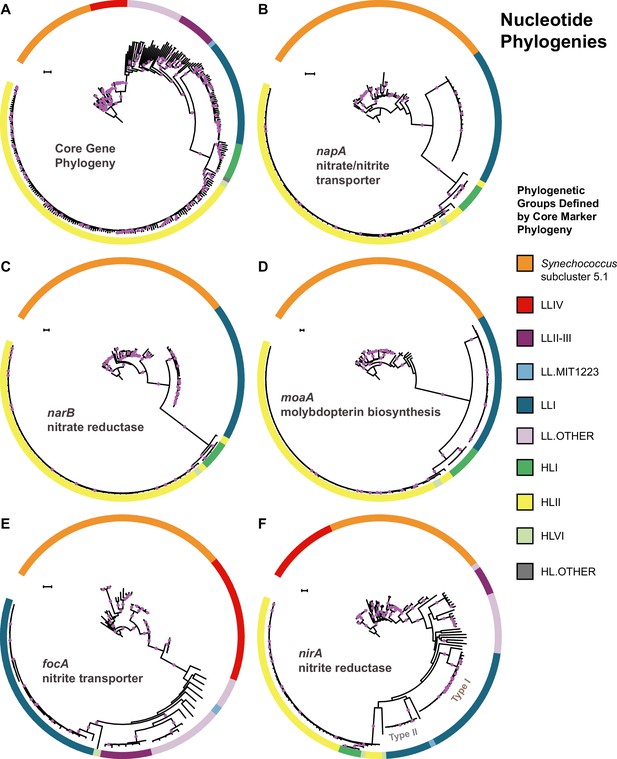
The core marker gene phylogeny of Prochlorococcus and Synechococcus.
(A) in comparison to gene phylogenies for the nitrate/nitrite transporter, napA (B), the nitrate reductase, narB (C), the molybdopterin biosynthesis protein, moaA (D), the nitrite transporter, focA (E), and the nitrite reductase, nirA (F). Filled purple circles on branches indicate that the associated taxa clustered together in at least 75% of trees. Scale bars are 0.1 changes per site.
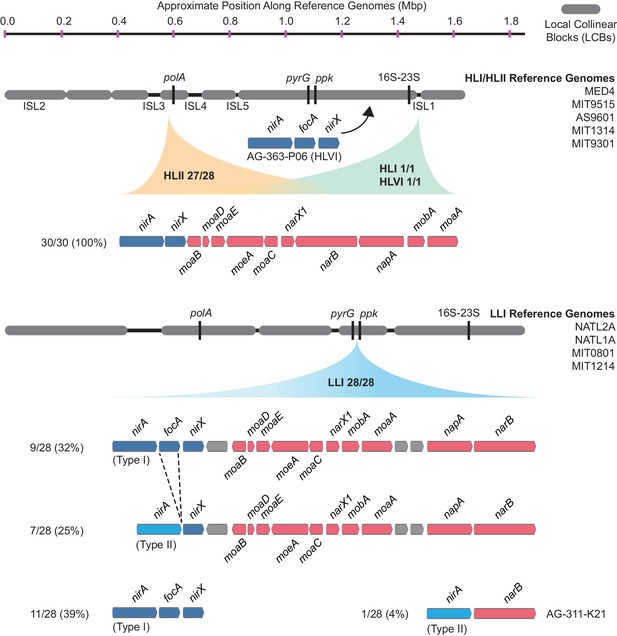
The gene order and genomic location of the nitrate assimilation gene cluster in high-light and low-light adapted Prochlorococcus genomes.
The location of the nitrate and nitrite assimilation genes is shown relative to conserved core marker genes (black bars). The proportion of genomes in each clade with the indicated location is shown next to the clade names. The percentage of genomes in each group with a specific gene content and order is shown to the left of each gene order plot.

Mauve alignments visualized for representative contigs from HLII Prochlorococcus single cell genome assemblies in comparison to the reference genome Prochlorococcus AS9601 (HLII; non-nitrate assimilating).
The nitrate assimilation gene cluster is found in a local collinear block shared with AS9601. CyCOG_60001297 and DNA polymerase I genes are marked as reference points.
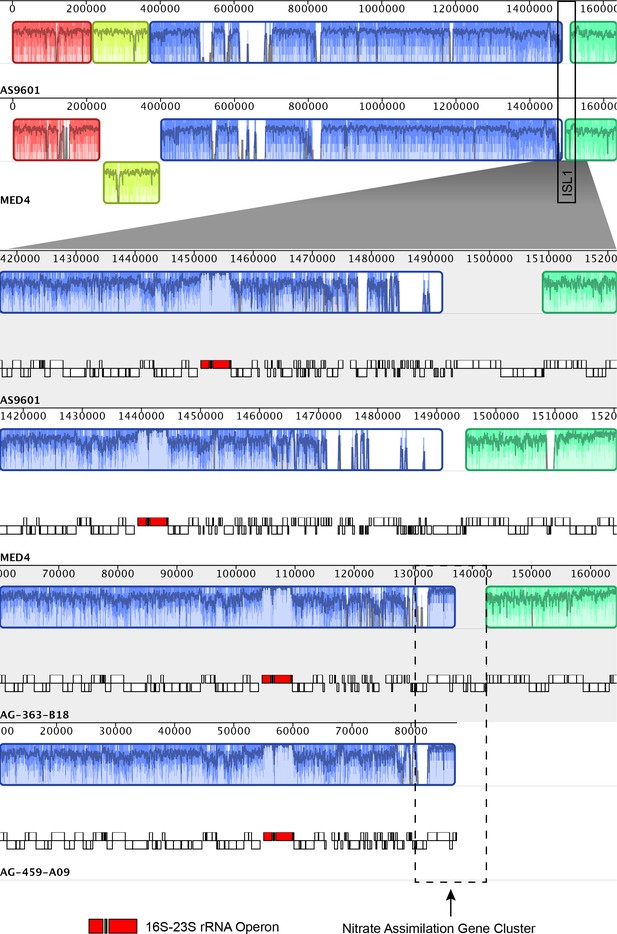
Mauve alignments visualized for representative contigs from HLI and HLVI Prochlorococcus single cell genome assemblies in comparison to the reference genomes Prochlorococcus MED4 (HLI) and Prochlorococcus AS9601 (HLII).
The reference genomes do not contain the nitrate assimilation gene cluster. In single cells, this cluster is found in the genomic island ISL1 (sensu Kettler et al., 2007).
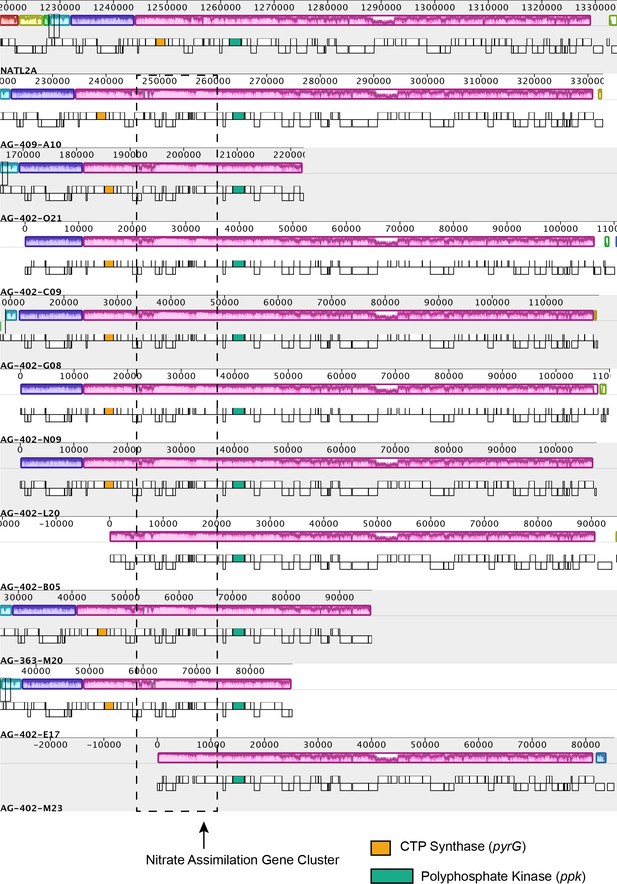
Mauve alignments visualized for representative contigs from LLI Prochlorococcus single cell genome assemblies in comparison to the reference genome Prochlorococcus NATL2A (LLI; nitrite assimilation only).
The nitrate assimilation gene cluster is found in a local collinear block shared with NATL2A. The pyrG and ppk genes are marked as reference points.
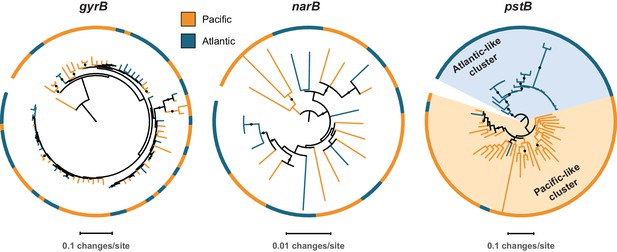
Representative phylogenetic patterns for DNA gyrase subunit B (gyrB) in comparison to nitrate reductase (narB) and the phosphate assimilation gene pstB (encoding the ABC transporter ATP binding subunit) from single cells in surface populations at HOT (Hawai’i Ocean Time-series) and BATS (Bermuda Atlantic Timeseries Study).
The gyrB and narB genes do not exhibit significant phylogenetic divergence between the two sites. The pstB gene, in contrast, has significantly diverged into Atlantic-like and Pacific-like clusters of sequences due to frequent recombination, gene transfer, and/or selection (sensu Coleman and Chisholm, 2010).
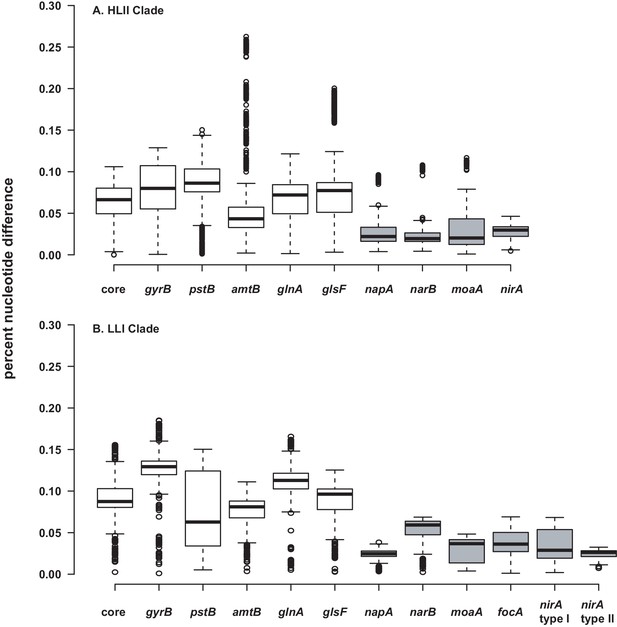
Percent nucleotide difference of selected genes for the HLII.
(A) and LLI (B) clades of Prochlorococcus. The 'core' genes are a concatenated alignment of up to 37 PhyloSift marker genes. Genes in the nitrate assimilation cluster are in gray. Center lines show the medians, box limits indicate the 25th and 75th percentiles, whiskers extend 1.5 times the interquartile range from the 25th and 75th percentiles, and outliers are represented by dots. For HLII genes (A), n = 5151, 4186, 3321, 3570, 4371, 3240, 378, 351, 351, 351 sample points. For LLI genes (B), n = 496, 406, 435, 435, 406, 406, 153, 136, 136, 171, 210, 28 sample points.
-
Figure 6—source data 1
Compressed tar archive (zip format) containing the alignments (fasta format) and column distance matrices used in the preparation of Figure 6.
- https://doi.org/10.7554/eLife.41043.021
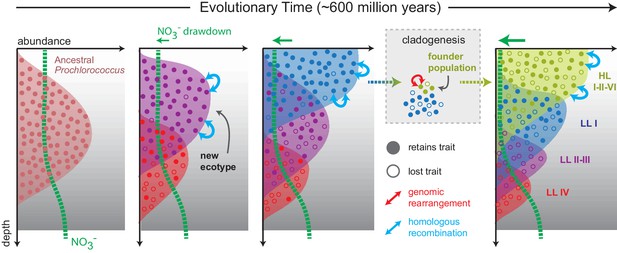
Proposed model mapping the vertical inheritance of the nitrate assimilation pathway onto a general model of speciation in Prochlorococcus (Braakman et al., 2017).
We assume different cost:benefit ratios at top and bottom of the water column based on the energetic requirements of the nitrate assimilation pathway and the selective advantage of carrying this trait in environments with low nitrogen availability (see main text for further discussion). We further assume that ancestral Prochlorococcus, similar to Synechococcus, were capable of nitrate assimilation. As new Prochlorococcus clades/ecotypes emerged to more efficiently harness light energy and facilitate the draw-down of nitrogen at the surface, basal lineages were partitioned to the deeper regions of the euphotic zone (Braakman et al., 2017). In these basal lineages, higher relative costs of the pathway combined with access to other nitrogen sources (e.g. amino acids) hastened the loss of this trait through stochastic gene loss. In more recently emerging lineages (LLI, HLI, HLII, HLVI), the trait has been retained with intra-clade frequencies influenced by the specific chemical and physical characteristics of the environment in which they are found (Berube et al., 2016). The founder effect has driven punctuated changes (e.g. genome-wide rearrangements) during speciation, while homologous recombination has acted to constrain the divergence of gene sequence and order within clades.
-
Figure 7—source data 1
Source data used to create Figure 7—figure supplement 1.
This file is provided in nexus format suitable for analysis in Mesquite. Contents include the taxa, phylogeny, and character matrix required for reconstructing the ancestral state of Prochlorococcus with respect to the nitrate assimilation trait.
- https://doi.org/10.7554/eLife.41043.030
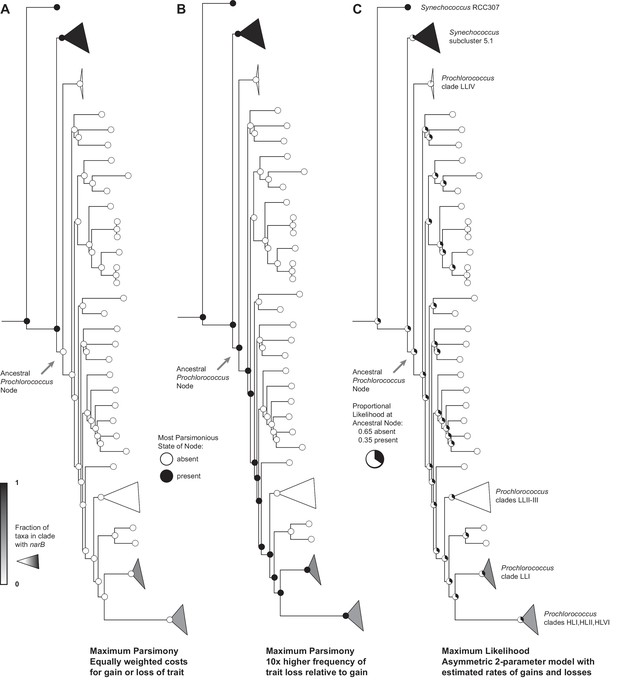
Reconstruction of the state of the ancestral Prochlorococcus node with regards to the presence or absence of narB.
When parsimony is used to infer whether or not ancestral Prochlorococcus possessed the nitrate assimilation trait (A, B), the results are highly dependent on the relative costs associated with the gain or loss of the trait. If these costs are equally weighted (A), all changes are estimated to occur at the leaves of the tree and consist primarily of acquisition events – this explanation seems unlikely given the conservation of gene order and location within clades (Figure 4). But, if a bias towards loss of the nitrate assimilation trait is imposed, the more parsimonious explanation is that ancestral Prochlorococcus possessed this trait (B). We further note that maximum likelihood estimates a reasonable probability that ancestral Prochlorococcus were capable of nitrate assimilation (C). Further, maximum likelihood generally supports a higher rate of loss relative to gain of the nitrate assimilation trait in Prochlorococcus given that the estimated forward (trait gain) rate was 14.5 and the estimated backward (trait loss) rate was 27.4 in this asymmetrical model.
Tables
Rates of recombination relative to mutation for representative genomic regions in high-light and low-light adapted Prochlorococcus.
https://doi.org/10.7554/eLife.41043.015| Region | Functional group | Sequences in alignment | Alignment length (bp) | κ | δ | ν | R/θ | ρ/θ | R/m |
|---|---|---|---|---|---|---|---|---|---|
| High-light adapted Prochlorococcus (HLII and HLVI clades) | |||||||||
| nirA-moaA | nitrate | 33 | 11428 | 3.32 | 4202 | 0.012 | 0.68 | 1.36 | 34.0 |
| polA region* | core flanking | 19 | 11554 | 3.72 | 8035 | 0.020 | 0.20 | 0.41 | 33.2 |
| Low-light adapted Prochlorococcus (LLI clade) | |||||||||
| moaB-narB | nitrate | 17 | 11985 | 2.91 | 435 | 0.021 | 1.78 | 3.57 | 16.5 |
| pyrG region† | core flanking | 15 | 10080 | 2.81 | 964 | 0.029 | 2.17 | 4.34 | 59.7 |
| ppk region‡ | core flanking | 15 | 11918 | 3.19 | 1056 | 0.031 | 0.93 | 1.85 | 30.3 |
-
* 3’ flanking region, containing polA, adjacent to the nitrate assimilation gene cluster in the HLII clade.
† 5’ flanking region, containing pyrG, adjacent to the nitrate assimilation gene cluster in the LLI clade.
-
‡ 3’ flanking region, containing ppk, adjacent to the nitrate assimilation gene cluster in the LLI clade.
κ, Transition/transversion rate ratio as estimated by PhyML under the HKY85 model.
-
δ, Mean length of DNA imported by homologous recombination.
ν, Divergence rate, per site, of DNA imported by homologous recombination.
-
R/θ, Per-site rate of initiation of recombination relative to the population mutation rate.
ρ/θ, Population recombination rate relative to the population mutation rate (ρ = 2R).
-
r/m, Relative impact of recombination versus mutation on the per-site substitution rate. Equal to (R/θ) × δ × ν.
-
Table 1—source data 1
Compressed tar archive (zip format) containing alignments used in CLONALFRAMEML analyses.
- https://doi.org/10.7554/eLife.41043.016
Significance (p value) for divergent phylogenetic gene clusters separating populations in the North Pacific Subtropical Gyre (HOT) and in the North Atlantic Subtropical Gyre (BATS).
https://doi.org/10.7554/eLife.41043.018| Gene | Product | Unifrac | P-test |
|---|---|---|---|
| amtB | Ammonium transporter | 0.651, 0.135, 0.199 | 0.576, 0.006, 0.554 |
| glnA | Glutamine synthetase | 0.058, 0.129, 0.136 | 0.045, 0.238, 0.214 |
| glsF | Glutamate synthase | 0.277, 0.066, 0.428 | 0.547, 0.235, 0.249 |
| gyrB | DNA gyrase subunit B | 0.425, 0.110, 0.477 | 0.867, 0.058, 0.006 |
| moaA | Molybdopterin biosynthesis | 0.040, 0.138, 0.090 | 0.067, 0.577, 0.555 |
| napA | Nitrate/nitrite transporter | 0.427, 0.082, 0.600 | 0.229, 0.051, 0.225 |
| narB | Nitrate reductase | 0.326, 0.547, 0.355 | 0.558, 0.060, 0.221 |
| nirA | Nitrite reductase | 0.004, 0.039, 0.072 | 0.010, 0.066, 0.056 |
| pstB | Phosphate transporter ATP-binding | 0.001,<0.001,<0.001 | 0.001,<0.001,<0.001 |
| pstS | Phosphate transporter substrate binding | <0.001,<0.001,<0.001 | <0.001,<0.001,<0.001 |
-
P values are reported for three replicate analyses using 18 single cells subsampled from HOT (AG-347; n = 9) and BATS (AG-355; n = 9). Trees exhibiting significant phylogenetic divergence between the two populations (p<0.05) are in bold face type.
-
Table 2—source data 1
Compressed tar archive (zip format) containing the alignments and group files used in beta diversity analyses.
- https://doi.org/10.7554/eLife.41043.019
Estimates of dN/dS and results of site model tests for adaptive evolution among codon sites for Prochlorococcus and Synechococcus.
Bolded LRT statistic values are chi-square critical values that meet a significance level of <0.001. For all genes, the inclusion of a class of neutral sites (M1) fits the data better than one dN/dS value for all sites (M0). While the inclusion of a class of sites under positive selection may be statistically justified under the M2 and M8 models, all dN/dS values are well below one suggesting that most sites are under purifying or neutral selection.
| log-likelihood of site models for adaptive evolution | likelihood ratio test (LRT) statistic for model pairs (degrees of freedom) | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Gene | Taxa | dN/dS | M0 | M1 | M2 | M7 | M8 | M0 vs. M1 (1) | M1 vs. M2 (2) | M7 vs. M8 (2) |
| gyrB | 226 | 0.036 | −105998 | −101367 | −101367 | −102434 | −100472 | 9263 | 0 | 3924 |
| pstB | 200 | 0.045 | −39945 | −39466 | −39466 | −38543 | −38484 | 957 | 0 | 118 |
| amtB | 200 | 0.066 | −70319 | −68351 | −68234 | −68226 | −67620 | 3935 | 235 | 1212 |
| glnA | 229 | 0.031 | −66948 | −65905 | −65905 | −65462 | −65118 | 2086 | 0 | 688 |
| glsF | 215 | 0.103 | −267696 | −249591 | −249591 | −253085 | −246724 | 36210 | 0 | 12722 |
| napA | 76 | 0.054 | −23709 | −23028 | −23028 | −23201 | −22961 | 1363 | 0 | 480 |
| narB | 78 | 0.119 | −40791 | −38790 | −38790 | −39102 | −38498 | 4001 | 0 | 1209 |
| moaA | 67 | 0.206 | −24794 | −22973 | −22931 | −23268 | −22770 | 3643 | 84 | 996 |
| focA | 60 | 0.078 | −18477 | −17701 | −17701 | −17895 | −17615 | 1553 | 0 | 562 |
| nirA | 115 | 0.109 | −54829 | −52361 | −52361 | −52063 | −51470 | 4937 | 0 | 1187 |
-
Table 3—source data 1
Compressed tar archive (zip format) containing example codeml control files, codon alignments (phylip format), and tree files (newick format) used for site model tests of adaptive evolution.
- https://doi.org/10.7554/eLife.41043.023
Branch-site model tests for adaptive evolution among codon sites for the foreground HLII branch.
Background lineages include Synechococcus and all other Prochlorococcus. Bolded LRT statistic values are chi-square critical values that meet a significance level of <0.001 with 1 degree of freedom.
| Gene | Hypothesis log-likelihood | Site class 0 | Site class 1 | Site class 2a | Site class 2b | LRT statistic | |
|---|---|---|---|---|---|---|---|
| gyrB | H0 −104105 | proportion of sites | 0.98759 | 0.00138 | 0.01102 | 0.00002 | 6074 |
| background dN/dS | 0 | 1 | 0 | 1 | |||
| foreground dN/dS | 0 | 1 | 1 | 1 | |||
| H1 −101068 | proportion of sites | 0.99666 | 0.00177 | 0.00157 | 0 | ||
| background dN/dS | 0.00908 | 1 | 0.00908 | 1 | |||
| foreground dN/dS | 0.00908 | 1 | 1 | 1 | |||
| pstB | H0 −39463 | proportion of sites | 0.99801 | 0.00161 | 0.00038 | 0 | 0 |
| background dN/dS | 0.03255 | 1 | 0.03255 | 1 | |||
| foreground dN/dS | 0.03255 | 1 | 1 | 1 | |||
| H1 −39463 | proportion of sites | 0.99801 | 0.00161 | 0.00038 | 0 | ||
| background dN/dS | 0.03255 | 1 | 0.03255 | 1 | |||
| foreground dN/dS | 0.03255 | 1 | 1 | 1 | |||
| amtB | H0 −68231 | proportion of sites | 0.9926 | 0.00663 | 0.00076 | 0.00001 | -2 |
| background dN/dS | 0.02052 | 1 | 0.02052 | 1 | |||
| foreground dN/dS | 0.02052 | 1 | 1 | 1 | |||
| H1 −68232 | proportion of sites | 0.99288 | 0.00657 | 0.00054 | 0 | ||
| background dN/dS | 0.0206 | 1 | 0.0206 | 1 | |||
| foreground dN/dS | 0.0206 | 1 | 2.17216 | 2.17216 | |||
| glnA | H0 −65896 | proportion of sites | 0.99839 | 0.00144 | 0.00018 | 0 | 0 |
| background dN/dS | 0.01722 | 1 | 0.01722 | 1 | |||
| foreground dN/dS | 0.01722 | 1 | 1 | 1 | |||
| H1 −65896 | proportion of sites | 0.99839 | 0.00144 | 0.00018 | 0 | ||
| background dN/dS | 0.01722 | 1 | 0.01722 | 1 | |||
| foreground dN/dS | 0.01722 | 1 | 1 | 1 | |||
| glsF | H0 −249282 | proportion of sites | 0.99092 | 0.00679 | 0.00227 | 0.00002 | 0 |
| background dN/dS | 0.0227 | 1 | 0.0227 | 1 | |||
| foreground dN/dS | 0.0227 | 1 | 1 | 1 | |||
| H1 −249282 | proportion of sites | 0.99092 | 0.00679 | 0.00227 | 0.00002 | ||
| background dN/dS | 0.0227 | 1 | 0.0227 | 1 | |||
| foreground dN/dS | 0.0227 | 1 | 1 | 1 | |||
| napA | H0 −23009 | proportion of sites | 0.97966 | 0.01219 | 0.00805 | 0.0001 | 0 |
| background dN/dS | 0.00928 | 1 | 0.00928 | 1 | |||
| foreground dN/dS | 0.00928 | 1 | 1 | 1 | |||
| H1 −23009 | proportion of sites | 0.97966 | 0.01219 | 0.00805 | 0.0001 | ||
| background dN/dS | 0.00928 | 1 | 0.00928 | 1 | |||
| foreground dN/dS | 0.00928 | 1 | 1 | 1 | |||
| narB | H0 −38711 | proportion of sites | 0.94311 | 0.02164 | 0.03446 | 0.00079 | 0 |
| background dN/dS | 0.02489 | 1 | 0.02489 | 1 | |||
| foreground dN/dS | 0.02489 | 1 | 1 | 1 | |||
| H1 −38711 | proportion of sites | 0.94311 | 0.02164 | 0.03447 | 0.00079 | ||
| background dN/dS | 0.02489 | 1 | 0.02489 | 1 | |||
| foreground dN/dS | 0.02489 | 1 | 1 | 1 | |||
| moaA | H0 −22958 | proportion of sites | 0.94068 | 0.03375 | 0.02468 | 0.00089 | 0 |
| background dN/dS | 0.03882 | 1 | 0.03882 | 1 | |||
| foreground dN/dS | 0.03882 | 1 | 1 | 1 | |||
| H1 −22958 | proportion of sites | 0.94068 | 0.03375 | 0.02468 | 0.00089 | ||
| background dN/dS | 0.03882 | 1 | 0.03882 | 1 | |||
| foreground dN/dS | 0.03882 | 1 | 1 | 1 | |||
| nirA | H0 −52348 | proportion | 0.97996 | 0.01302 | 0.00693 | 0.00009 | 14 |
| background dN/dS | 0.04327 | 1 | 0.04327 | 1 | |||
| foreground dN/dS | 0.04327 | 1 | 1 | 1 | |||
| H1 −52341 | proportion | 0.97254 | 0.0124 | 0.01487 | 0.00019 | ||
| background dN/dS | 0.04293 | 1 | 0.04293 | 1 | |||
| foreground dN/dS | 0.04293 | 1 | 1 | 1 |
-
Table 4—source data 1
Compressed tar archive (zip format) containing example codeml control files, codon alignments (phylip format), and tree files (newick format) used for branch-site model tests of adaptive evolution in the HLII clade.
- https://doi.org/10.7554/eLife.41043.025
Branch-site model tests for adaptive evolution among codon sites for the foreground LLI branch.
Background lineages include Synechococcus and all other Prochlorococcus. Bolded LRT statistic values are chi-square critical values that meet a significance level of <0.001 with 1 degree of freedom.
| Gene | Hypothesis log-likelihood | Site class 0 | Site class 1 | Site class 2a | Site class 2b | LRT statistic | |
|---|---|---|---|---|---|---|---|
| gyrB | H0 −101367 | proportion of sites | 0.99813 | 0.00187 | 0 | 0 | 22 |
| background dN/dS | 0.01062 | 1 | 0.01062 | 1 | |||
| foreground dN/dS | 0.01062 | 1 | 1 | 1 | |||
| H1 −101356 | proportion of sites | 0.99768 | 0.00186 | 0.00046 | 0 | ||
| background dN/dS | 0.01053 | 1 | 0.01053 | 1 | |||
| foreground dN/dS | 0.01053 | 1 | 1 | 1 | |||
| pstB | H0 −39407 | proportion of sites | 0.98046 | 0.00147 | 0.01805 | 0.00003 | 0 |
| background dN/dS | 0.03172 | 1 | 0.03172 | 1 | |||
| foreground dN/dS | 0.03172 | 1 | 1 | 1 | |||
| H1 −39407 | proportion of sites | 0.98046 | 0.00147 | 0.01805 | 0.00003 | ||
| background dN/dS | 0.03172 | 1 | 0.03172 | 1 | |||
| foreground dN/dS | 0.03172 | 1 | 1 | 1 | |||
| amtB | H0 −68211 | proportion of sites | 0.99157 | 0.00673 | 0.00169 | 0.00001 | 0 |
| background dN/dS | 0.01992 | 1 | 0.01992 | 1 | |||
| foreground dN/dS | 0.01992 | 1 | 1 | 1 | |||
| H1 −68211 | proportion of sites | 0.9918 | 0.00677 | 0.00143 | 0.00001 | ||
| background dN/dS | 0.01992 | 1 | 0.01992 | 1 | |||
| foreground dN/dS | 0.01992 | 1 | 1 | 1 | |||
| glnA | H0 −65885 | proportion of sites | 0.99799 | 0.00141 | 0.0006 | 0 | 0 |
| background dN/dS | 0.01727 | 1 | 0.01727 | 1 | |||
| foreground dN/dS | 0.01727 | 1 | 1 | 1 | |||
| H1 −65885 | proportion of sites | 0.99799 | 0.00141 | 0.0006 | 0 | ||
| background dN/dS | 0.01727 | 1 | 0.01727 | 1 | |||
| foreground dN/dS | 0.01727 | 1 | 1 | 1 | |||
| glsF | H0 −249371 | proportion of sites | 0.98865 | 0.00686 | 0.00445 | 0.00003 | 0 |
| background dN/dS | 0.02306 | 1 | 0.02306 | 1 | |||
| foreground dN/dS | 0.02306 | 1 | 1 | 1 | |||
| H1 −249371 | proportion of sites | 0.98865 | 0.00686 | 0.00445 | 0.00003 | ||
| background dN/dS | 0.02306 | 1 | 0.02306 | 1 | |||
| foreground dN/dS | 0.02306 | 1 | 1 | 1 | |||
| napA | H0 −23014 | proportion of sites | 0.97128 | 0.01248 | 0.01604 | 0.00021 | 0 |
| background dN/dS | 0.01004 | 1 | 0.01004 | 1 | |||
| foreground dN/dS | 0.01004 | 1 | 1 | 1 | |||
| H1 −23014 | proportion of sites | 0.97122 | 0.01246 | 0.01611 | 0.00021 | ||
| background dN/dS | 0.01004 | 1 | 0.01004 | 1 | |||
| foreground dN/dS | 0.01004 | 1 | 1 | 1 | |||
| narB | H0 −38790 | proportion of sites | 0.97596 | 0.02404 | 0 | 0 | 40 |
| background dN/dS | 0.02939 | 1 | 0.02939 | 1 | |||
| foreground dN/dS | 0.02939 | 1 | 1 | 1 | |||
| H1 −38770 | proportion of sites | 0.9577 | 0.02346 | 0.01839 | 0.00045 | ||
| background dN/dS | 0.02762 | 1 | 0.02762 | 1 | |||
| foreground dN/dS | 0.02762 | 1 | 1 | 1 | |||
| moaA | H0 −22961 | proportion of sites | 0.90692 | 0.03565 | 0.05526 | 0.00217 | 0 |
| background dN/dS | 0.03704 | 1 | 0.03704 | 1 | |||
| foreground dN/dS | 0.03704 | 1 | 1 | 1 | |||
| H1 −22961 | proportion of sites | 0.90692 | 0.03565 | 0.05526 | 0.00217 | ||
| background dN/dS | 0.03704 | 1 | 0.03704 | 1 | |||
| foreground dN/dS | 0.03704 | 1 | 1 | 1 | |||
| focA | H0 −17695 | proportion of sites | 0.97605 | 0.01411 | 0.00969 | 0.00014 | 0 |
| background dN/dS | 0.01425 | 1 | 0.01425 | 1 | |||
| foreground dN/dS | 0.01425 | 1 | 1 | 1 | |||
| H1 −17695 | proportion of sites | 0.97605 | 0.01411 | 0.0097 | 0.00014 | ||
| background dN/dS | 0.01425 | 1 | 0.01425 | 1 | |||
| foreground dN/dS | 0.01425 | 1 | 1 | 1 | |||
| nirA type I | H0 −52349 | proportion of sites | 0.97292 | 0.01243 | 0.01446 | 0.00018 | 0 |
| background dN/dS | 0.04333 | 1 | 0.04333 | 1 | |||
| foreground dN/dS | 0.04333 | 1 | 1 | 1 | |||
| H1 −52349 | proportion of sites | 0.97292 | 0.01243 | 0.01446 | 0.00018 | ||
| background dN/dS | 0.04333 | 1 | 0.04333 | 1 | |||
| foreground dN/dS | 0.04333 | 1 | 1 | 1 | |||
| nirA type II | H0 −52358 | proportion of sites | 0.97139 | 0.01216 | 0.01625 | 0.0002 | 0 |
| background dN/dS | 0.0442 | 1 | 0.0442 | 1 | |||
| foreground dN/dS | 0.0442 | 1 | 1 | 1 | |||
| H1 −52358 | proportion of sites | 0.97139 | 0.01216 | 0.01625 | 0.0002 | ||
| background dN/dS | 0.0442 | 1 | 0.0442 | 1 | |||
| foreground dN/dS | 0.0442 | 1 | 1 | 1 |
-
Table 5—source data 1
Compressed tar archive (zip format) containing example codeml control files, codon alignments (phylip format), and tree files (newick format) used for branch-site model tests of adaptive evolution in the LLI clade.
- https://doi.org/10.7554/eLife.41043.027
Additional files
-
Supplementary file 1
Tab-delimited table containing the genomes and associated IMG accession numbers used in the final data set.
Clade indicates the respective Synechococcus subcluster or Prochlorococcus clade. Results from the narB PCR screening assay are presented as a binary (0 = negative; 1 = positive; n.d = not determined). Estimated genome recovery was determined using checkM (Parks et al., 2015) and the presence/absence of annotated reductase and transporter genes for nitrite (nirA and focA) and nitrate (narB and napA) in each genome assembly are as given as a binary (0 = absent; 1 = present).
- https://doi.org/10.7554/eLife.41043.031
-
Transparent reporting form
- https://doi.org/10.7554/eLife.41043.032





