Schwann cells, but not Oligodendrocytes, Depend Strictly on Dynamin 2 Function
Figures
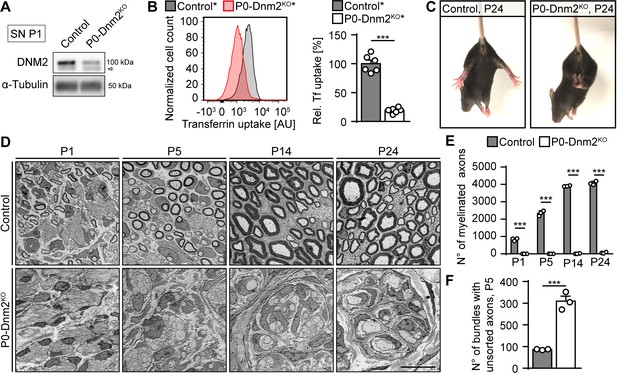
Lack of dynamin 2 in Schwann cells results in major radial sorting defects and failure of sciatic nerve myelination.
(A) Representative immunoblot of DNM2 in sciatic nerve (SN) extracts derived from Dnm2loxP/loxP (control) and MpzCre:Dnm2loxP/loxP (P0-Dnm2KO) mice at P1. N = 4 mice/genotype. Arrowhead: unspecific signal. Quantification in Figure 1—figure supplement 1B, full-length blots in Supplementary file 1A. (B) Representative experimental set (left) and quantified sets (right) of FACS analyses of transferrin (Tf) uptake of YFP+ primary mouse SCs derived from MpzCre:Rosa26-stoploxP/loxP-YFP (control*) and MpzCre:Dnm2loxP/loxP:Rosa26-stoploxP/loxP-YFP (P0-Dnm2KO*) P1 SNs. P0-Dnm2KO* SCs show reduced Tf uptake. Control average was set to 100. N = 6 independent SC preparations/genotype, two-tailed unpaired Student’s t-test. (C) Tail-suspension test showing hind paw clasping in P0-Dnm2KO, but not in control littermate at P24 (consistent phenotype observed in 30 mice/genotype). (D) Exemplary electron microscopy (EM) images showing ultrastructure of control and P0-Dnm2KO SNs at indicated time points (cross-sections). N = 3 mice/genotype and time point. Scale bar = 10 μm, referring to the entire panel. Quantifications referring to panel D: (E) Number of myelinated fibers/SN cross-section. N = 3 mice/genotype and time point. Two-Way ANOVA with Sidak’s multiple comparisons test. (F) Number of bundles with axons > 1 μm/SN cross-section at P5. N = 3 mice/genotype, two-tailed unpaired Student’s t-test. Results in graphs represent means ±s.e.m.; ***p<0.001.
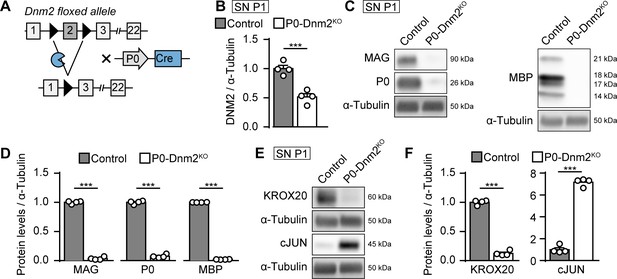
Dynamin 2 ablation in Schwann cells inhibits their differentiation and myelination.
(A) Schematic representation of Dnm2 ablation in SCs. Exon 2 of Dnm2 is flanked by LoxP sites and excised upon expression of Cre recombinase under the control of the SC-specific Mpz promoter. (B) Quantification referring to Figure 1A and Supplementary file 1A (full-length blots). Relative amounts of DNM2 protein in SN extracts derived from Dnm2loxP/loxP (control) and MpzCre:Dnm2loxP/loxP (P0-Dnm2KO) mice at P1. Control average is set to 1. N = 4 mice/genotype, two-tailed unpaired Student’s t-test. (C) Representative immunoblot of the myelin proteins MAG, P0, and MBP in SN lysates from P1 control and P0-Dnm2KO mice. N = 4 mice/genotype. Full-length blots in Supplementary file 1D. (D) Relative amounts of myelin proteins in P1 SNs of control and P0-Dnm2KO mice. Control average is set to 1. N = 4 mice/genotype, two-tailed unpaired Student’s t-test. (E) Representative immunoblot analysis of the differentiation markers KROX20 and cJUN in SN lysates from P1 control and P0-Dnm2KO mice. N = 4 mice/genotype, full-length blots in Supplementary file 1D. (F) Relative amounts of KROX20 and cJUN in P1 SNs of control and P0-Dnm2KO mice. Control average is set to 1. N = 4 mice/genotype, two-tailed unpaired Student’s t-test. Results in graphs represent means ±s.e.m.; ***p<0.001.
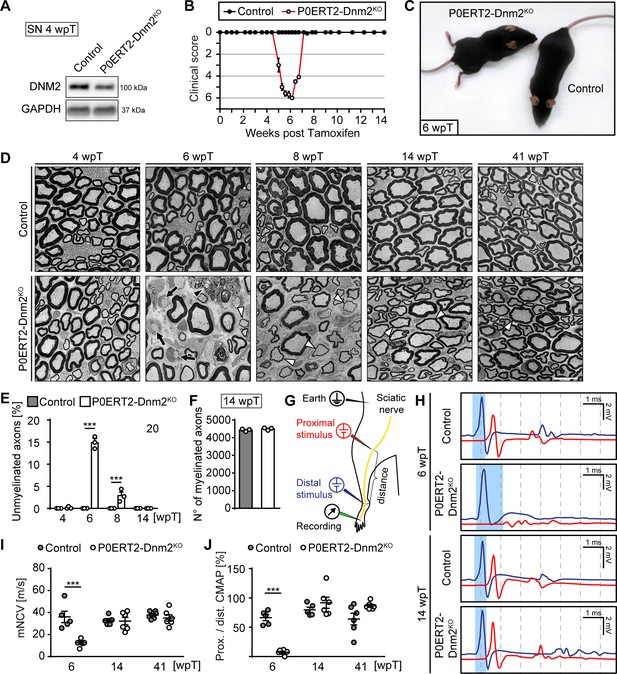
Schwann cell-specific ablation of dynamin 2 in adult mice causes a transient neuropathy.
(A) Immunoblot analysis of DNM2 in SN extracts of Dnm2loxP/loxP (control) and MpzCreERT2:Dnm2loxP/loxP (P0ERT-Dnm2KO) mice at 4 weeks post tamoxifen (wpT). N = 3 mice/genotype. Quantification in Figure 2—figure supplement 1B, full-length blots in Supplementary file 1B. (B) Time line of functional impairments and recovery of P0ERT-Dnm2KO mice after tamoxifen treatment based on clinical assessments over 14 weeks. Clinical score: 0 = normal, 1 = less lively, 2 = impaired righting reflex, 3 = absent righting reflex, 4 = ataxic gait, 5 = mild paraparesis, 6 = moderate paraparesis, 7 = severe paraparesis. N = 5 mice/genotype. (C) Exemplary picture showing moderate paraparesis of a P0ERT-Dnm2KO mouse, but not of a control littermate at 6 wpT. (D) Exemplary EM images showing ultrastructure of control and P0ERT2-Dnm2KO SN cross-sections at the indicated time points. N = 3 mice/time point and genotype. Black arrows: Unmyelinated axons; white arrowheads: Thinly melinated axons associated with remyelinating SCs; black arrowhead points to a macrophage. Scale bar = 10 μm, referring to entire panel. (E) Quantification of unmyelinated axons relative to total number of axons with calibers > 1 μm. At least 200 axons/sample were analyzed (random EM fields). N = 3 mice/time point and genotype, Two-Way ANOVA with Sidak’s multiple comparisons test. (F) Quantification of myelinated fibers/SN cross-section at 14 wpT. N = 3 mice/genotype, two-tailed unpaired Student’s t-test. (G) Schematic drawing of the setup for electrophysiological measurements. (H) Representative traces of electrophysiological findings in control and P0ERT2-Dnm2KO mice SNs at 6 wpT and 14 wpT. At 6 wpT, impulse conduction appears almost completely blocked upon proximal stimulation in P0ERT2-Dnm2KO mice and only a small and dispersed compound muscle action potential (CMAP) was recorded. These deficits were restored at 14 wpT. The blue-shaded columns illustrate the slowing in nerve conduction at 6 wpT in a P0ERT-Dnm2KO mouse and the recovery to normal values at 14 wpT. Blue profile: CMAP upon distal stimulation, red profile: CMAP upon proximal stimulation. (I) Motor nerve conduction velocity (mNCV) in control and P0ERT-Dnm2KO mice at the indicated time points. mNCV is reduced in P0ERT-Dnm2KO mice at 6 wpT, but is restored at 14 wpT. N = 5–6 mice/group, Two-Way ANOVA with Sidak´s multiple comparisons test. (J) Conduction block is indicated by severe reduction of the proximal to distal CMAP ratio in P0ERT-Dnm2KO mice at 6 wpT. This is resolved by 14 wpT. N = 5–6 mice/group, Two-Way ANOVA with Sidak´s multiple comparisons test. Results in graphs represent means ±s.e.m.; ***p<0.001.
-
Figure 2—source data 1
Phenotypical assessment of control and P0ERT-Dnm2KO mice over a period of 14 weeks after tamoxifen treatment.
Refers to panel B.
- https://doi.org/10.7554/eLife.42404.006
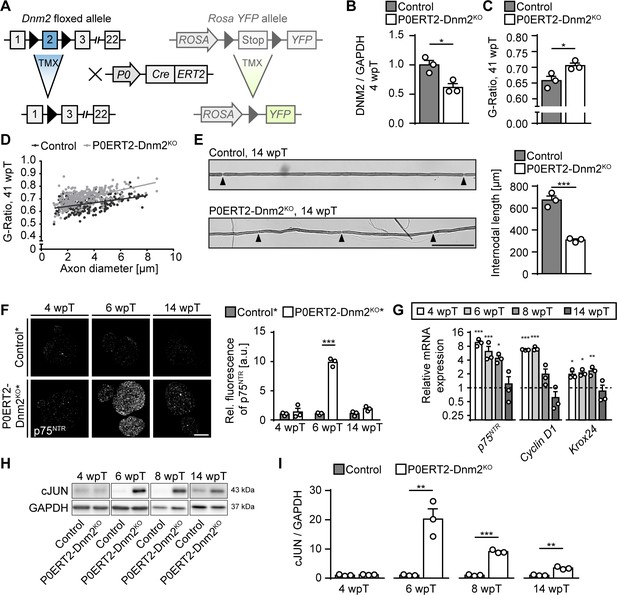
Dynamin 2 ablation in adult Schwann cells induces transient cellular dedifferentiation.
(A) Schematic representation of inducible Dnm2 ablation and YFP expression in adult SCs. The inducible Cre-ERT2 fusion recombinase is expressed under the SC-specific Myelin Protein Zero (Mpz) promoter. Upon tamoxifen injection, the recombinase shuttles into the nucleus leading to excision of floxed exon 2. The Cre-dependent RosaYFP-reporter allele was used in some experiments to track recombination in SCs. In this case, the transcription stop signal flanked by loxP sites is excised upon tamoxifen administration leading to YFP expression. (B) Quantification referring to Figure 2A, full-length blots in Supplementary file 1B. Relative amounts of DNM2 protein in sciatic nerve (SN) extracts derived from Dnm2loxP/loxP (control) and MpzCreERT2:Dnm2loxP/loxP (P0ERT2-Dnm2KO) mice at 4 wpT. Control average was set to 1. N = 3 mice/genotype, two-tailed unpaired Student’s t-test. (C) Average g-ratios of control and P0ERT2-Dnm2KO SNs indicate a slight hypomyelination of mutant mice at 41 wpT. At least 90 axons/sample were analyzed (random EM fields). N = 3 mice/genotype, two-tailed unpaired Student´s t-test. (D) G-ratios as function of axon diameters. (E) (left) Exemplary picture of internodes of teased osmicated fibers from SNs of control and P0ERT2-Dnm2KO mice at 14 wpT. (right) Quantification of internodal lengths. 100 internodes/animal were measured, n = 3 mice/genotype, two-tailed unpaired Student´s t-test. Black arrowheads: Nodes of Ranvier. Scale bar = 100 μm (for both panels). (F) (left) Immunostaining of MpzCreERT2:Rosa26-stoploxP/loxP-YFP (control*) and MpzCreERT2:Dnm2loxP/loxP:Rosa26-stoploxP/loxP-YFP (P0ERT2-Dnm2KO*) SN cross-sections for p75NTR at different time points. There was a pronounced, but transient increase of p75NTR staining at 6 wpT in P0ERT2-Dnm2KO* mice. (right) Quantification of relative signal intensity of p75NTR immunostaining/SN cross-section. N = 3 mice/genotype, Two-Way ANOVA with Sidak’s multiple comparisons test. Scale bar = 200 μm. (G) qRT-PCR of P75NTR, cyclinD1 and Krox24 from control and P0ERT2-Dnm2KO SNs at the indicated time points. Transcript levels were normalized to GAPDH. Control values were set to one for each individual transcript at each time point. N = 3 mice/genotype, Two-Way ANOVA with Sidak’s multiple comparisons test. (H) Exemplary picture of immunoblot analysis of cJUN in SN extracts from control and P0ERT-Dnm2KO mice at the indicated time points. N = 3 mice/genotype. Full-length blots in Supplementary file 1E. (I) Quantification referring to Figure 2H, Supplementary file 1E. Control average was set to one for each time point individually. N = 3 mice/genotype, two-tailed unpaired Student’s t-test. Results in graphs represent means ±s.e.m.; *p<0.05, **p<0.01, ***p<0.001.
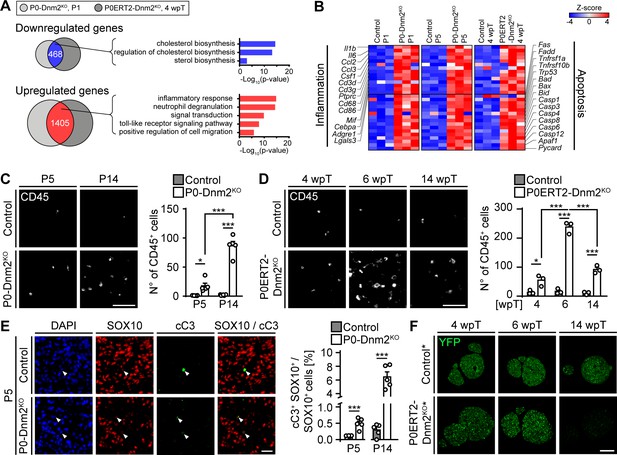
Mice lacking dynamin 2 in Schwann cells display characteristics of inflammation and Schwann cell death.
(A) Gene ontology analysis of jointly downregulated or upregulated genes in P1 P0-Dnm2KO and 4 wpT P0ERT2-Dnm2KO SNs compared to controls. (B) Heat map showing representative differentially expressed genes and their GO categories in controls versus Dnm2KO SNs or P0CreERT2-Dnm2KO SNs at the indicated time points. N = 3 mice/genotype. (C) (Left) Immunostainings of controls and P0-Dnm2KO SN cross-sections for the inflammatory cell marker CD45 at P5 and P14. (Right) Quantification of CD45+ cells/SN cross-sections. N = 5 mice/time point and genotype. Two-Way ANOVA with Holm-Sidak’s multiple comparisons test. Scale bar = 50 μm for entire panel. (D) (Left) Immunostainings of controls and P0ERT2-Dnm2KO SN cross-sections for the inflammatory cells marker CD45 at 4 wpT, 6 wpT, and 14 wpT. (Right) Quantification of CD45+ cells/SN cross-sections. N = 3 mice/time point and genotype. Two-Way ANOVA with Holm-Sidak’s multiple comparisons test. Scale bar = 50 μm for entire panel. (E) (Left) Immunostainings of controls and P0-Dnm2KO SN cross-sections for cleaved-caspase 3 (cC3) in combination with the SC marker SOX10 at P5. (Right) Percentage of cC3+ SOX10+ SCs at P5 and P14. N = 5 mice/time point and genotype, two-tailed unpaired Student’s t-test. Scale bar = 25 μm for entire panel. (F) Immunostainings for YFP (recombined cells) on SN cross-sections derived from MpzCreERT2:Rosa26-stoploxP/loxP-YFP (control*) and MpzCreERT2:Dnm2loxP/loxP:Rosa26-stoploxP/loxP-YFP (P0ERT2-Dnm2KO*) mice at 4 wpT, 6 wpT, and 14 wpT. YPF+ cells were no longer detectable at 14 wpT in P0ERT2-Dnm2KO mice. N = 3 mice/time point and genotype. Scale bar = 200 μm for the entire panel. Results in graphs represent means ±s.e.m. *p<0.05, **p<0.01, ***p<0.001.
-
Figure 3—source data 1
Transcriptomes of sciatic nerves from P0-Dnm2KO and control mice at P1 and P5, and from P0ERT2-Dnm2KO and control mice at 4 wpT.
Refers to panels A and B.
- https://doi.org/10.7554/eLife.42404.013
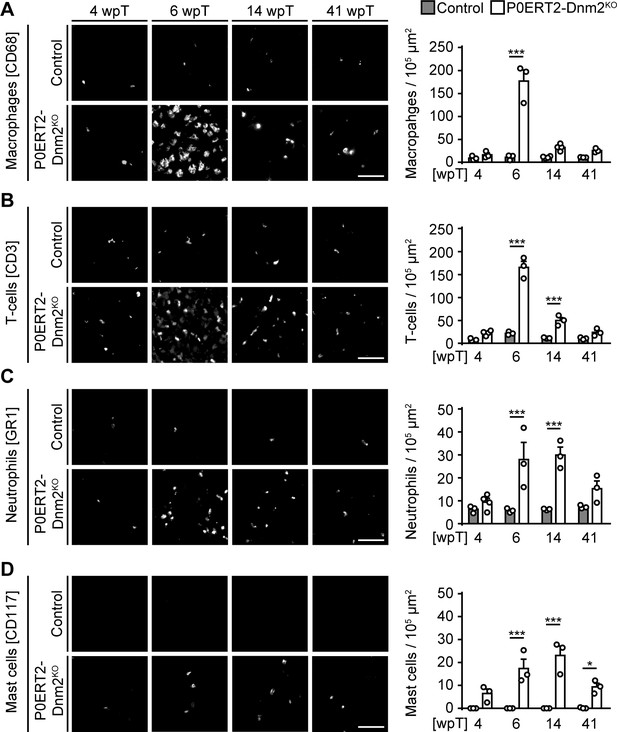
Various immune cells invade sciatic nerves after dynamin 2 depletion in adult Schwann cells.
(A) (Left) Immunostainings for the macrophage marker CD68 on cross-sections of control and P0ERT2-Dnm2KO SNs at 4 wpT, 6 wpT, 14 wpT, and 41 wpT. (Right) Quantification of CD68+ cells (random fields). N = 3 mice/genotype for 4, 6, 41 wpT; N = 4 mice/genotype for 14 wpT. (B) (Left) Immunostainings for the T-cell marker CD3 on cross-sections of control and P0ERT2-Dnm2KO SNs at 4 wpT, 6 wpT, 14 wpT, and 41 wpT. (Right) Quantification of CD3+ cells (random fields). N = 3 mice/genotype for 6, 14, and 41 wpT. At 4 wpT, N = 3 controls and N = 4 P0ERT2-Dnm2KO mice. (C) (Left) Immunostainings for the neutrophil marker GR1 on cross-sections of control and P0ERT2-Dnm2KO SNs at 4 wpT, 6 wpT, 14 wpT, and 41 wpT. (Right) Quantification of GR1+ cells (random fields). N = 3 mice/genotype for 6, 14, and 41 wpT. At 4 wpT, N = 3 controls and N = 4 P0ERT2-Dnm2KO mice. (D) (Left) Immunostainings for the mast cell marker CD117 on cross-sections of control and P0ERT2-Dnm2KO SNs at 4 wpT, 6 wpT, 14 wpT, and 41 wpT. (Right) Quantification of CD117+ cells (random fields). N = 3 mice/time point and genotype. Two-Way ANOVA with Sidak´s multiple comparisons test for all statistical analyses. Scale bars = 50 μm for all panels. Results in graphs represent means ±s.e.m.; *p<0.05, ***p<0.001.
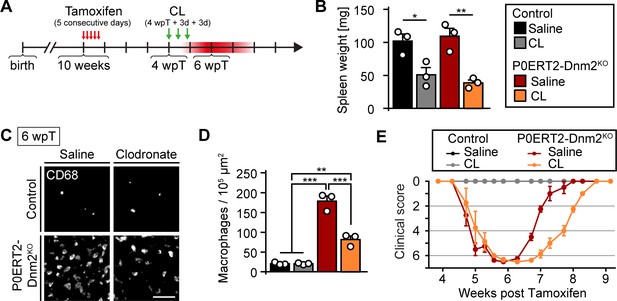
Macrophages are beneficial for recovery after demyelination due to loss of dynamin 2 in adult Schwann cells.
(A) Schematic drawing of the experimental timeline. 10 weeks-old control (Dnm2loxP/loxP) and P0ERT2-Dnm2KO (MpzCreERT2:Dnm2loxP/loxP) mice were injected intraperitoneally with tamoxifen on five consecutive days. At 4 wpT, the mice were injected intravenously with clodronate liposomes (CL) or saline (Sal) three times (4 wpT, 4 wpT +3 d, 4 wpT +6 d). (B) Spleen weights in controls and P0ERT2-Dnm2KO at 6 wpT are consistent with effective macrophage deprivation after CL treatment in both genotypes. N = 3 mice/genotype and treatment, One-way ANOVA with Tukey´s multiple comparisons test. (C) Immunostainings for the macrophage marker CD68 on SN cross-sections derived from control and P0ERT2-Dnm2KO mice, treated with CL or Sal, at 6 wpT. N = 3 mice/genotype and treatment. Scale bar = 50 μm for entire panel. (D) Quantification of (C) (random fields). N = 3 mice/condition (color code as in B), One-Way ANOVA with Tukey´s multiple comparisons test. (E) CL- and Sal-treated control and P0ERT2-Dnm2KO mice were phenotypically assessed over a period of 5 weeks. All P0ERT2-Dnm2KO developed a transient moderate paraparesis around 6 wpT. CL-treated P0ERT2-Dnm2KO recovered more slowly compared to Saline-treated P0ERT2-Dnm2KO mice. N = 3 controls and N = 4 P0ERT2-Dnm2KO mice. Clinical score: 0 = normal, 1 = less lively, 2 = impaired righting, 3 = absent righting, 4 = ataxic gait, 5 = mild paraparesis, 6 = moderate paraparesis, 7 = severe paraparesis. Results in graphs represent means ±s.e.m.; *p<0.05, **p<0.01, ***p<0.001.
-
Figure 3—Figure supplement 2—Source data 1
Phenotypical assessment of control and P0ERT-Dnm2KO mice treated with Clodronate or Saline from day 27 to day 64 post tamoxifen.
Refers to panel E.
- https://doi.org/10.7554/eLife.42404.010
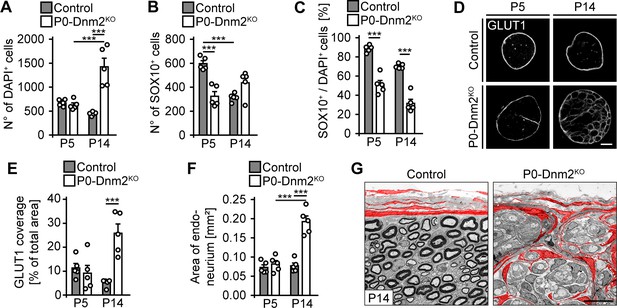
Lack of dynamin 2 in Schwann cells of developing nerves causes Schwann cell loss and invasion of perineurial cells.
(A) Quantification of total nuclei on full cross-sections of control and P0-Dnm2KO SNs at P5 and P14 (P5-representative picture in Figure 3E). N = 5 mice/time point and genotype, Two-Way ANOVA with Sidak’s multiple comparisons test. (B) Quantification of total SOX10+ SCs on cross-sections of control and P0ERT2-Dnm2KO SNs at P5 and P14 (P5-representative picture in Figure 3E). N = 5 mice/time point and genotype, Two-Way ANOVA with Sidak’s multiple comparisons test. (C) Percentage of SOX10+ SCs among total DAPI+ cells in controls and P0ERT2-Dnm2KO SNs at P5 and P14. N = 5 mice/time point and genotype, Two-Way ANOVA with Sidak’s multiple comparisons test. (D) Immunostainings of GLUT1 on cross-sections of control and P0ERT2-Dnm2KO SNs at P5 and P14. Scale bar = 100 μm for entire panel. (E) Quantification of (D). Relative GLUT1-immunopositive area/total area of SN cross-sections of control and P0ERT2-Dnm2KO mice. N = 5 mice/time point and genotype, Two-Way ANOVA with Sidak’s multiple comparisons test. (F) Quantification of (D). SN endoneurial area (mm2). N = 5 mice/time point and genotype, Two-Way ANOVA with Sidak’s multiple comparisons test. (G) Exemplary EM images showing ultrastructural organization of control and P0-Dnm2KO SNs at P14. Perineurial cells are false colored in red. N = 3 mice/genotype. Scale bar = 10 μm for entire panel. Results in graphs represent means ±s.e.m.; ***p<0.001.
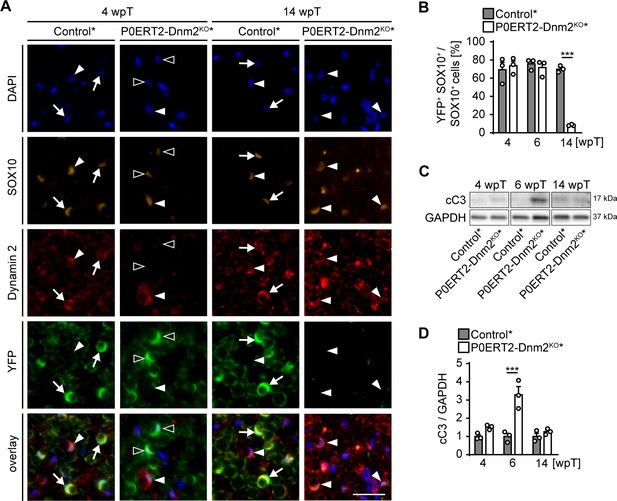
Adult dynamin 2-depleted Schwann cells are replaced by dynamin 2-positive Schwann cells in remyelinated nerves.
(A) Immunostainings for DNM2, SOX10 (SCs) and YFP (reporter-recombined SCs) on SN cross-sections of control* (MpzCreERT2:Rosa26-stoploxP/loxP-YFP) and P0ERT2-Dnm2KO* (MpzCreERT2:Dnm2loxP/loxP: Rosa26-stoploxP/loxP-YFP) mice at 4 wpT and 14 wpT. White arrows: YFP-recombined SCs that express DNM2 (in controls at 4 wpt and 14 wpt); black arrowheads: YFP-recombined SCs lacking DNM2 (in mutants at 4 wpt); white arrowheads: Non-recombined SCs expressing DNM2 (in controls and mutants at 4 wpt and 14 wpt). Scale bar = 25 μm for entire panel. (B) Quantification related to (A). Percentage of YFP+ cells among SOX10+ SCs/SN cross-sections at 4 wpT, 6 wpT, and 14 wpT. N = 3 mice/genotype, Two-Way ANOVA with Sidak’s multiple comparisons test. (C) Western blot analysis of cleaved caspase 3 (cC3) in SN lysates of control* and P0ERT2-Dnm2KO* mice at 4 wpT, 6 wpT and 14 wpT. Full-length blots in Supplementary file 1F. (D) Quantification referring to (C). Control average was set to 1. N = 3 mice/time point and genotype, Two-Way ANOVA with Sidak’s multiple comparisons test. Results in graphs represent means ±s.e.m.; ***p<0.001.
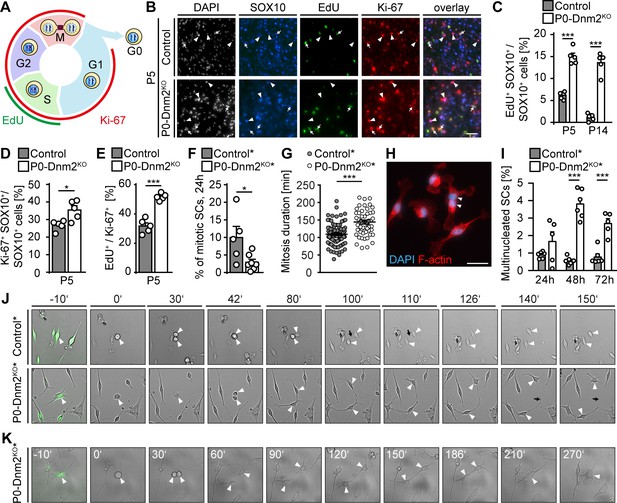
Schwann cells lacking dynamin 2 show impaired cell cycle progression, reduced mitosis rate, and cytokinesis defects.
(A) Schematic representation of the cell cycle phases marked by Ki-67 and EdU. (B) EdU-labeling, combined with immunostainings for Ki-67 and SOX10 on control and P0-Dnm2KO SN cross-sections at P5. Arrows: EdU+ Ki-67+ SOX10+ SCs, arrowheads: Ki-67+ SOX10+ SCs. Scale bar = 25 μm for entire panel. (C) Quantification of (B). Percentage of EdU+ SCs/SN cross-sections. N = 5 mice/time point and genotype, Two-Way ANOVA with Sidak’s multiple comparisons test. (D) Quantification of (B). Percentage of Ki-67+ SCs/SN cross-sections at P5. N = 5 mice/genotype, two-tailed unpaired Student´s t-test. (E) Quantification of (B). Percentage of EdU+ among Ki-67+ SCs/SN cross-section at P5. N = 5 mice/genotype, two-tailed unpaired Student´s t-test. (F) Quantification of mitotic events in cultured mouse SCs isolated from MpzCre:Rosa26-stoploxP/loxP-YFP (control*) and MpzCre:Dnm2loxP/loxP:Rosa26-stoploxP/loxP-YFP (P0-Dnm2KO*) SNs at P1, monitored by time-lapse microscopy for 24 hr. Each data point represents one individual animal (at least 42 cells/animal analyzed). N = 5 controls and seven mutant mice, two-tailed unpaired Student´s t-test. (G) Quantification of mitosis duration (minutes) of SCs of control* and P0-Dnm2KO* mice, monitored by time-lapse microscopy for 24 hr. Each data point represents one cell derived from a total of 5 control and seven mutant mice. Cells derived from each animal were isolated and analyzed separately, but pooled in one graph; two-tailed unpaired Student´s t-test. (H) Exemplary picture of a multinucleated Dnm2KO SCs after 48 hr in culture. Scale bar = 25 μm. (I) Quantification of multinucleated cells in control and Dnm2KO SCs after 24 hr, 48 hr and 76 hr in culture. N = 6 mice/genotype for 48 hr; N = 6 control* and n = 5 P0-Dnm2KO* mice for 24 hr and 72 hr, two-Way ANOVA with Sidak’s multiple comparisons test. (J) Exemplary time-lapse images of the mitosis of control and Dnm2KO SCs (YFP+ cells). Arrowheads: cell body of dividing SCs, black arrows: cytokinesis site. (K) Representative time-lapse images of Dnm2KO SC (YFP+) failing cytokinesis. The SC is undergoing mitosis, but fails to divide (arrowheads point to the cell body). Results in graphs represent means ±s.e.m.; *p<0.05, ***p<0.001.
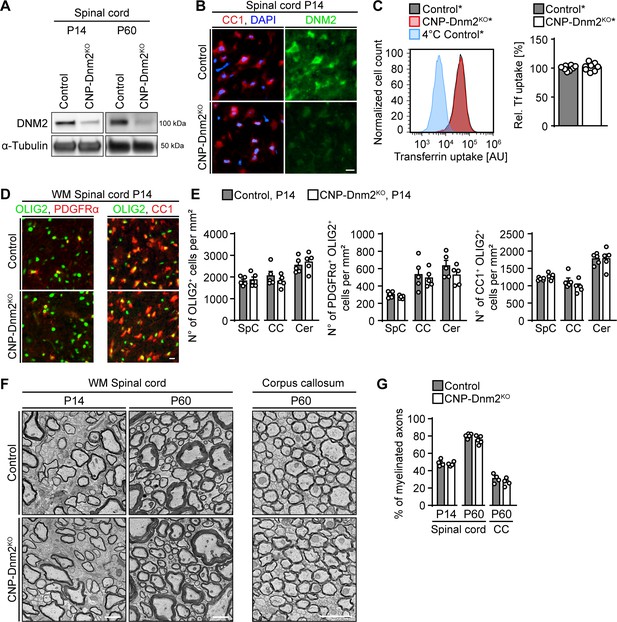
Oligodendrocytes can differentiate and myelinate without dynamin 2.
(A) Immunoblot analysis of DNM2 in spinal cord extracts from Dnm2loxP/loxP (control) and CnpCre: Dnm2loxP/loxP (CNP-Dnm2KO) mice (P14, P60). N = 4 mice/genotype. Quantification in Figure 5—figure supplement 1B, full-length blot in Supplementary file 1C. (B) Exemplary transverse sections of ventral horn region of lumbar spinal cord from control and CNP-Dnm2KO mice at P14, double-stained for CC1 (red; mature oligodendrocyte (OL) marker) and DNM2 (green), showing DNM2 ablation in mutant OLs. N = 3 mice/genotype. Scale bar = 10 μm. (C) Representative experimental set (left) and quantified sets (right) of FACS analyses of transferrin (Tf) uptake on primary mouse OLs derived from CnpCre:Rosa26-stoploxP/loxP-YFP (control*) and CnpCre:Dnm2loxP/loxP:Rosa26-stoploxP/loxP-YFP (CNP-Dnm2KO*) P7 brain (14 days in culture). CNP-Dnm2KO OLs show no detectable difference in Tf uptake compared to controls (note the virtually complete overlap of the corresponding peaks). Control average is set to 100. N = 8 mice/genotype, two-tailed unpaired Student’s t-test. (D) Representative immunostainings of control and CNP-Dnm2KO in white matter (WM) ventral spinal cord cross-sections for OLIG2 and PDGFRα (left panel) or OLIG2 and CC1 (right panel) at P14. N = 5 mice/genotype. Scale bar = 10 μm. (E) Quantification of (D) (Spinal cord (SpC) ventrolateral WM hemisection, corpus callosum (CC) midbody, and cerebellum (Cer) lobes IV,V). Total OL lineage cells (OLIG2+), OPC (PDGFRα+ OLIG2+) and mature OL (CC1+ OLIG2+) in SpC ventrolateral WM, corpus callosum (CC), and cerebellum (Cer) of control and CNP-Dnm2KO mice. N = 5 mice/genotype, two-tailed unpaired Student’s t-test. (F) Exemplary EM images showing ultrastructure of control and CNP-Dnm2KO ventral SpC WM (P14, P60) and CC (P60). Scale bars = 2 μm. (G) Quantification of (F) (random fields). Percentages of myelinated fibers in SpC ventral WM (P14, P60) and corpus callosum (P60). At least 752 axons/animal were analyzed. N = 4–5 control and mutant mice at the indicated time points, two-tailed unpaired Student’s t-test. Results in graphs represent means ±s.e.m.
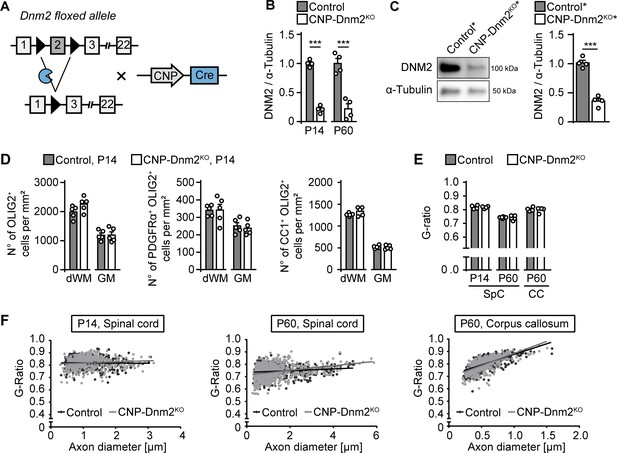
Dynamin 2-depleted oligodendrocytes do not show detectable defects in differentiation and myelination.
(A) Schematic representation of Dnm2 ablation in the OL lineage. Exon 2 of Dnm2 is flanked by LoxP sites and excised upon expression of Cre recombinase under the control of CNPase gene regulatory elements. (B) Quantification referring to Figure 5A, full-length blot Supplementary file 1C. Relative amount of DNM2 in spinal cord extracts from Dnm2loxP/loxP (control) and CnpCre:Dnm2loxP/loxP (CNP-Dnm2KO) mice (P14, P60). Control average was set to 1. N = 4 mice/genotype, two-tailed unpaired Student’s t-test. (C) (left panel) Exemplary immunoblot of DNM2 in extracts of primary mouse OLs derived from CnpCre:Rosa26-stoploxP/loxP-YFP (control*) and CnpCre:Dnm2loxP/loxP:Rosa26-stoploxP/loxP-YFP (CNP-Dnm2KO*) from P7 brain (after 14 day in culture). (right panel) Relative amounts of DNM2, full-length blot in Supplementary file 1G. N = 4 mice/genotype, two-tailed unpaired Student’s t-test. (D) Detailed quantification of Figure 5D (spinal cord hemisections). Total OL lineage cells (OLIG2+), OPC (PDGFRα+OLIG2+) and mature OL (CC1+ OLIG2+), in spinal cord dorsal white matter (dWM) and grey matter (GM) of control and CNP-Dnm2KO mice. N = 5 mice/genotype, two-tailed unpaired Student’s t-test. (E) Average g-ratios of control and CNP-Dnm2KO myelinated axons in spinal cord ventral WM (SpC; P14, P60) and CC (P60). At least 108 axons/sample were analyzed (random EM fields). N = 4 or 5 mice/time point and genotype, two-tailed unpaired Student´s t-test. (F) G-ratios as function of axon diameter. Results in graphs represent means ±s.e.m.; ***p<0.001.
Additional files
-
Supplementary file 1
Full-length western blot images.
- https://doi.org/10.7554/eLife.42404.017
-
Supplementary file 2
Statistic summary.
- https://doi.org/10.7554/eLife.42404.018
-
Transparent reporting form
- https://doi.org/10.7554/eLife.42404.019





