The Hox transcription factor Ubx stabilizes lineage commitment by suppressing cellular plasticity in Drosophila
Figures
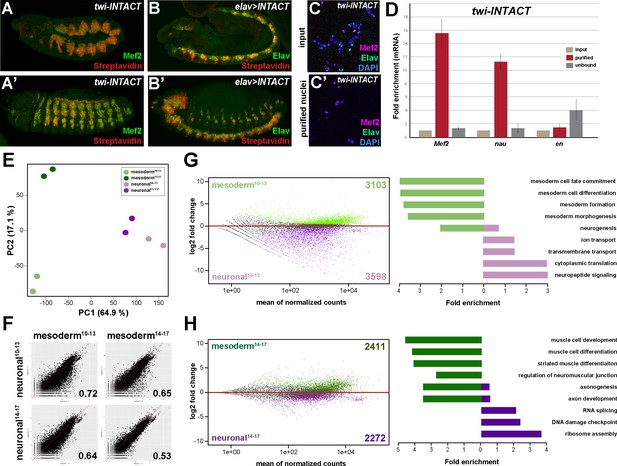
Comparative profiling of the mesodermal and neuronal transcriptomes in Drosophila embryos.
(A, A’) Lateral views of stage 11 (A) and stage 14 (A’) twi-INTACT Drosophila embryos stained for the muscle differentiation marker Mef2 (green) and Streptavidin (red). (B, B’) Lateral view of stage 11 (B) and stage 14 (B’) elav >INTACT Drosophila embryos stained for the pan-neuronal marker Elav (green) and Streptavidin (red). (C, C’) Nuclei obtained from twi-INTACT embryos stained for Mef2 (red), Elav (green) and DAPI (blue) before (C) and after (C’) INTACT purification. (D) Expression of mesodermal (Mef2, nau) and ectodermal (en) mRNA transcripts in nuclei purified from twi-INTACT embryos as measured by RT-PCR using RNA from total/input (10%), purified and unbound nuclei. (E) Principle Component Analysis (PCA) applied to all RNA-seq samples identifies three clusters: one corresponding to mesodermal specification events (mesoderm10-13), one to mesodermal differentiation events (mesoderm14-17) and the neuronal cluster sharing similar expression signatures. The two mesodermal cluster are distinct from the neuronal and result in the mesodermal group. (F) Pearson correlation analysis highlights the differences and similarities between the mesodermal and neuronal RNA-seq datasets. (G, H) Left: MA analysis identifies genes differentially expressed in the mesodermal and neuronal lineages at two different time windows, embryonic stage 10–13 (G) and embryonic stage 14–17 (H). Genes differentially expressed in the mesodermal lineage are indicated as green dots, those in the neuronal lineage as purple dots. Right: Bar diagram displaying fold enrichment of gene ontology terms of genes differentially expressed in the mesodermal or neuronal lineages at the two different time windows.
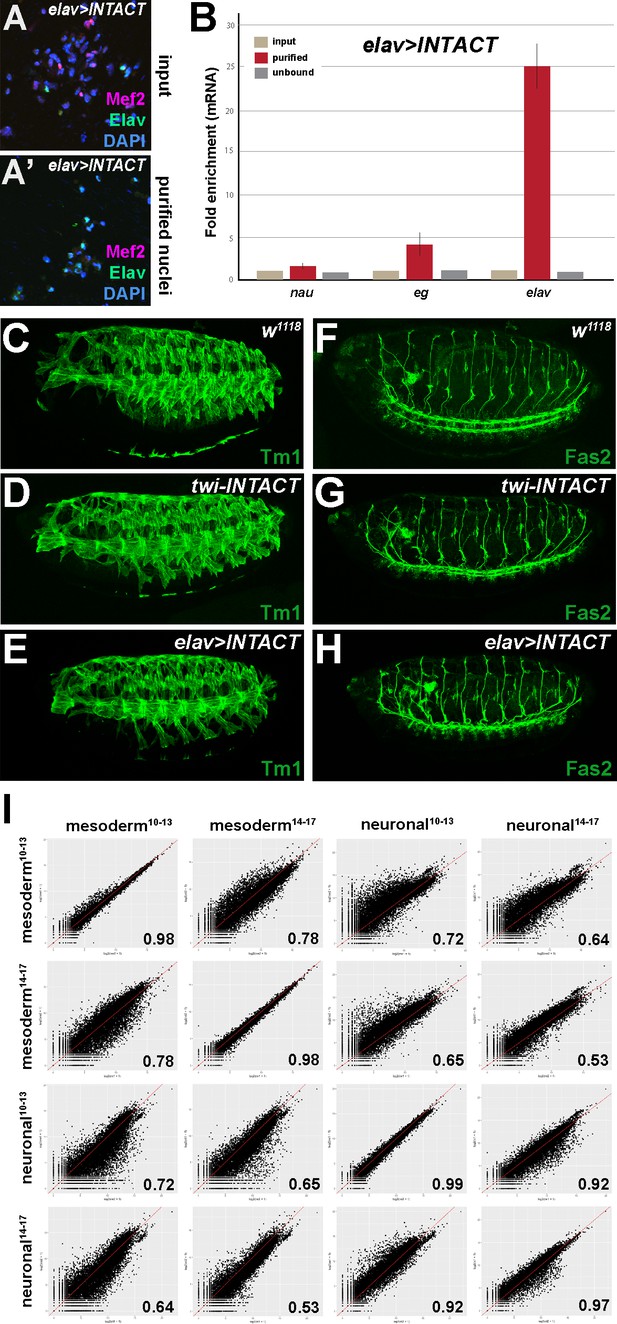
Comparative profiling of the mesodermal and neuronal transcriptomes in Drosophila embryos.
(A, A’) Nuclei obtained from elav-GAL4; UAS-INTACT (elav >INTACT) embryos stained for Mef2 (red), Elav (green) and DAPI (blue) before (A) and after (A’) INTACT purification. (B) Expression of mesodermal (nau) and neuronal (eg, elav) mRNA transcripts in nuclei purified from elav-INTACT embryos as measured by RT-qPCR using RNA from input (10%), purified and unbound nuclei. (C–H) Lateral view of stage 15 wild type control (w1118) (C, F), twi-INTACT (D, G) and elav >INTACT (E, H) Drosophila embryos stained for Tropomyosin 1 (Tm1) (C, D, E) labelling differentiated muscles and for Fasciclin 2 (Fas2) (F, G, H) labelling motoneurons. The INTACT flies display the same muscle and neuronal pattern as the wild type control (w1118) (C, F). (I) Pearson correlation analysis highlights the differences and similarities between the mesodermal and neuronal RNA-Seq datasets.
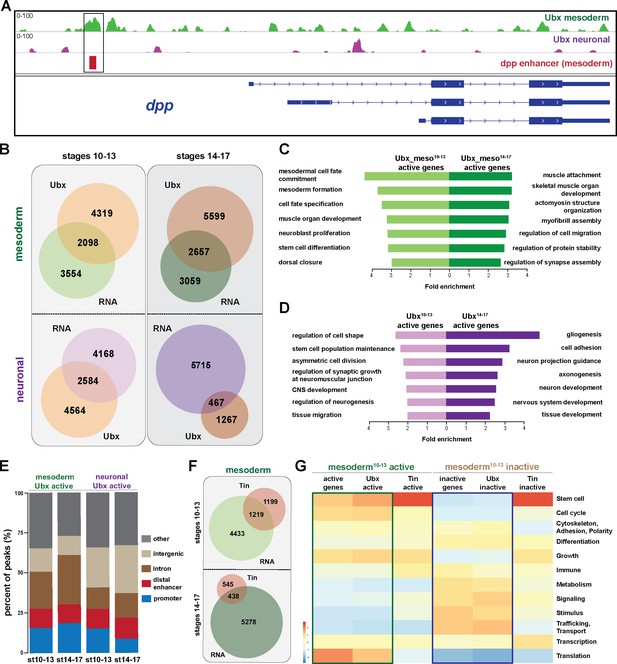
Ubx comprehensively controls tissue-specific transcriptional programs.
(A) ChIP-seq binding profiles of Ubx at the dpp genomic locus in mesodermal (green) and neuronal (purple) nuclei. The different isoforms of the dpp gene are shown in blue, the known dpp visceral enhancer in red. The box highlights Ubx binding to the dpp visceral enhancer in mesodermal but not in neuronal cells. (B) Venn diagrams representing the overlaps between the mesodermal (green) and neuronal (purple) transcriptomes and Ubx bound genes (orange) at two different stages. (C, D) Fold enrichment of gene ontology terms of genes expressed and targeted by Ubx in the mesoderm (C) or the neuronal (D) tissues, respectively. (E) Comparison of the localisation of peaks called within unique genomic regions of Ubx. Locations are classified as promoters (−1000- + 10 bp from TSS, 5’ UTR), distal enhancers (−2000 to −1000 from TSS, 3’ UTR, downstream), intron (intronic regions), intergenic (distal intergenic) and other regions (including exons). (F) Venn diagrams representing the overlaps between the mesodermal (green) transcriptomes and Tin bound genes (red) at two different stages. (G) Heat-map displaying presence of genes belonging to higher-order categories in the different gene classes. The colour range corresponds to the centred and scaled (per column) fraction of genes annotated to the category that also appear in the sample: red colour represents high values, blue colours low fractions of genes in the category, which are also present in the sample. Rows and columns are hierarchically clustered using Euclidean distance with complete linkage. The blue and green boxes highlight the distributions of functional terms among the expressed and Ubx targeted as well as the inactive and Ubx targeted genes.
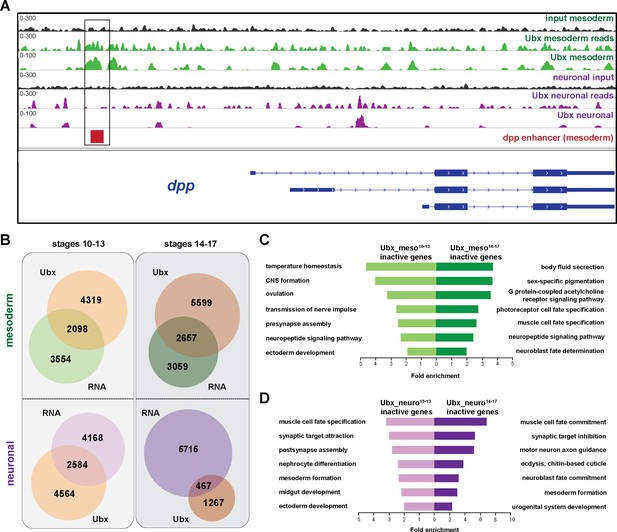
Ubx comprehensively controls tissue-specific transcriptional programs.
(A) Normalized ChIP-seq binding profiles of Ubx at the dpp genomic locus in mesodermal (green, mesodermal Ubx) and neuronal (purple, neuronal Ubx) nuclei. The Input reads are shown in black for the mesodermal and neuronal lineages, respectively, as well as the non-normalized ChIP-seq reads (mesodermal Ubx reads, green; neuronal Ubx reads, purple). The different isoforms of the dpp gene are shown in blue, the known dpp visceral enhancer in red. The box highlights Ubx binding to the dpp visceral enhancer in mesodermal but not in neuronal nuclei. (B) Venn diagrams representing the overlaps between the mesodermal (green) and neuronal (purple) transcriptomes and Ubx bound genes (orange) at two different stages. (C, D) Fold enrichment of gene ontology terms of inactive genes targeted by Ubx in the mesoderm (C) or the neuronal (D) tissues, respectively.
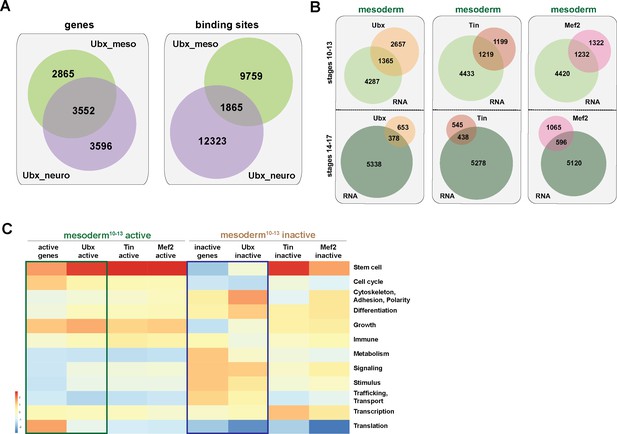
Ubx is a general regulator of mesoderm development.
(A) Left panel: Venn diagram representing the overlaps between Ubx bound genes in mesodermal (green) and neuronal (purple) nuclei in stage 10–13 embryos. Right panel: Venn diagram representing the overlaps between chromatin regions bound by Ubx in mesodermal (green) and neuronal (purple) nuclei. (B) Venn diagrams representing the overlaps between the mesodermal transcriptome (green) and Ubx (orange), Tin (light red) and Mef2 (pink) bound genes at two different stages. (C) Heat-map displaying presence of genes belonging to higher-order categories in the different gene classes. The colour range corresponds to the centered and scaled (per column) fraction of genes annotated to the category that also appear in the sample: red colour represents high values, blue colours low fractions of genes in the category, which are also present in the sample. Rows and columns are hierarchically clustered using Euclidean distance with complete linkage. The blue and green boxes highlight the distributions of functional terms among the expressed and Ubx targeted as well as the inactive and Ubx-targeted genes.
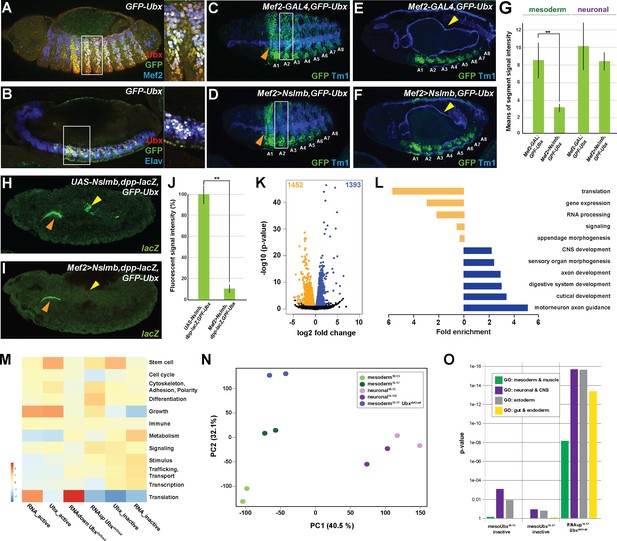
Ubx directly represses alternative fate genes in a lineage-specific manner.
(A, B) Lateral view of stage 14 GFP-Ubx Drosophila embryos stained for the muscle differentiation marker Mef2 (blue) (A), the pan-neuronal marker Elav (blue) (B), Ubx (red) and GFP (green). Boxes indicate the location of the close-ups on the right panel. (C–F) Lateral view of stage 15 Mef2-GAL4,GFP-Ubx and Mef2 >Nslmb,GFP-Ubx Drosophila embryos stained for Tm1 (blue) to indicate the differentiated muscles and GFP (green) to highlight GFP-Ubx expression. The boxes in (C, D) mark GFP expression in muscle cells of the 2nd abdominal segment (A2), which is lost when Ubx is degraded, while the ectodermal expression in the 1st abdominal segment (A1) is unaffected (marked by orange arrowheads). In (E, F), GFP expression in the CNS and the visceral mesoderm (marked by yellow arrowheads) is shown. GFP expression in the visceral mesoderm is lost in Mef2 >Nslmb,GFP-Ubx Drosophila embryos (F), leading to a loss of the second midgut constriction. (G) Quantification of GFP signal intensity in the mesoderm and the CNS of Mef2-GAL4,GFP-Ubx and Mef2 >Nslmb,GFP-Ubx Drosophila embryos, showing that GFP is strongly decreased in the mesoderm (**=p < 0.01). (H, I) lacZ mRNA expression in stage 14 UAS-Nslmb,dpp-lacZ,GFP-Ubx and Mef2 >Nslmb,dpp-lacZ,GFP-Ubx Drosophila embryos, highlighting that mesoderm-specific depletion of Ubx leads to a loss of dpp-lacZ enhancer activity (indicated by yellow arrowheads), a known and direct target of Ubx control (Capovilla and Botas, 1998; Manak et al., 1994). The orange arrowheads highlight unspecific enhancer activity in the salivary glands. (J) Quantification of lacZ signal intensity in UAS-Nslmb,dpp-lacZ,GFP-Ubx and Mef2 >Nslmb,dpp-lacZ,GFP-Ubx embryos, showing that lacZ expression is strongly decreased (**=p < 0.01). (K) Volcano plot displaying the differentially expressed genes between UAS-Nslmb,GFP-Ubx and Mef2 >Nslmb,GFP-Ubx INTACT-sorted mesodermal nuclei. The y-axis corresponds to the mean expression value of log10 (p-value), and the x-axis displays the log2 fold change value. The orange dots represent transcripts whose expression is down-regulated (padj-value <0.1), the blue dots represent the up-regulated expressed transcripts (padj-value <0.1) between UAS-Nslmb,GFP-Ubx and Mef2 >Nslmb,GFP-Ubx INTACT-sorted mesodermal nuclei. (L) Fold enrichment of gene ontology terms of down- (orange) and up-regulated (blue) genes in Mef2 >Nslmb,GFP-Ubx vs. UAS-Nslmb,GFP-Ubx mesodermal nuclei. (M) Heat-map displaying presence of genes belonging to higher-order categories in the different gene classes. The colour range corresponds to the centred and scaled (per column) fraction of genes annotated to the category that also appear in the sample: red colour represents high values, blue colours low fractions of genes in the category, which are also present in the sample. Rows and columns are hierarchically clustered using Euclidean distance with complete linkage. (N) PCA applied to all RNA-seq samples identifies the separation of the UbxdeGrad dataset from the two mesodermal as well as the neuronal datasets. (O) Multiple testing of higher-order GO-terms related to different lineages among different gene classes. The y-axis corresponds to the p-value, the x-axis displays the different categories of tested genes. Neuronal and ectodermal as well as gut/endoderm related GO-terms are significantly enriched in the tested samples.
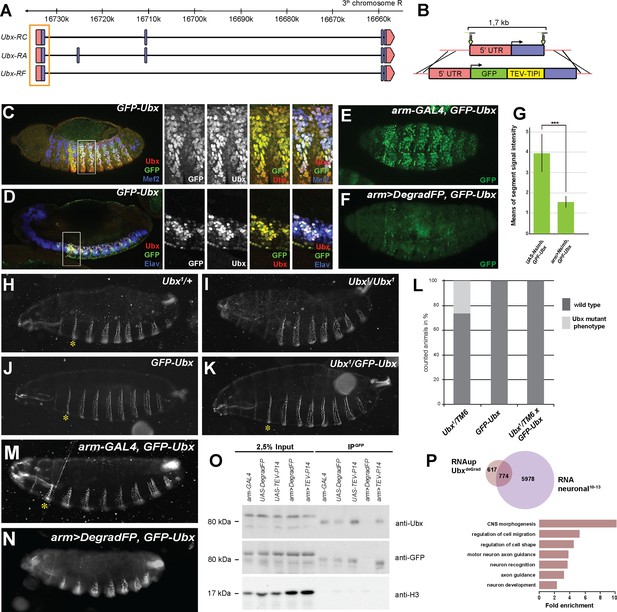
Generation and validation of CRIPR/Cas9 generated GFP-Ubx flies.
(A) Genomic region of Ubx gene, exons are illustrated in boxes (red: non-coding regions, blue: coding regions) and introns in lines, orange box highlights the 1st exon. (B) close up of the 1st exon targeted by guide RNAs (green arrows) for CRISPR/Cas9, below: construct used for homologous recombination, recombination event illustrated by the crossed lines. Red: non-coding region, blue: coding region, green: GFP coding region, yellow: spacer containing a TEV cleavage site with TIPI-degron (TEV protease- induced protein inactivation) (C, D) Lateral view of stage 14 GFP-Ubx Drosophila embryos stained for the muscle differentiation marker Mef2 (blue) (C), the pan-neuronal marker Elav (blue) (D), Ubx (red) and GFP green). Boxes indicate the location of the close-ups on the right panel. (E, F) Lateral view of stage 15 arm-GAL4,GFP-Ubx and arm >Nslmb,GFP-Ubx Drosophila embryos stained for GFP (green) to highlight Ubx expression. The GFP staining is mostly lost in the ectoderm, mesoderm and neuronal system when Ubx is degraded (F) as compared to the control (E). (G) Quantification of GFP signal intensity in the whole arm-GAL4,GFP-Ubx and arm >Nslmb,GFP-Ubx Drosophila embryos, showing that GFP is strongly decreased (***=p < 0.001). (H–K) Lateral view of first instar larva cuticles, heterozygous (H) and homozygous Ubx mutants (I), homozygous GFP-Ubx (J), heterozygous Ubx mutant over GFP-Ubx (K), asterisk indicates the 1st abdominal segment (A1), homozygous Ubx mutants show a loss of the denticle belts in A1 compared to the heterozygous control and the GFP-Ubx line, the Ubx mutant phenotype can be rescued with the GFP-Ubx. (L) Quantification of the rescue experiment, showing that no Ubx phenotype is detachable in the Ubx1/TM6 with GFP-Ubx cross, indicating that GFP-Ubx is fully functional during embryonic and larval development. (M, N) Lateral view of first instar larva cuticles, arm-GAL4,GFP-Ubx (M) and arm >Nslmb,GFP-Ubx (N), asterisk indicates A1, arm >Nslmb,GFP-Ubx larvae (N) shows a loss of the A1 denticle similar to the homozygous Ubx mutant (I) when compared to arm-GAL4,GFP-Ubx (M). (O) GFP protein immunoprecipitation (IP) experiments in Drosophila embryos, comparison of GFP-nanobody (deGrad) and TEV-TIPI protein degradation efficiency, control lines: arm-GAL4,GFP-Ubx (arm-GAL4), UAS-Nslmb,GFP-Ubx (UAS-Nslmb), UAS-TEV-P14;GFP-Ubx (UAS-TEV-P14), crosses: arm-GAL4,GFP-Ubx x UAS-Nslmb,GFP-Ubx (arm >Nslmb), arm-GAL4,GFP-Ubx x UAS-TEV-P14;GFP-Ubx (arm >TEV P14), expected size of GFP-Ubx 80 kDa, IPs are probed with antibodies indicated on the right, lane 1–5 are 2.5% input of each sample, lane 6–10 GFP-trap bead IP-experiments for each sample, control samples indicate bands for GFP-Ubx using GFP or Ubx antibodies, arm >Nslmb shows no band for GFP-Ubx, whereas arm >TEV P14 displays the same band as the control samples, indicating that the degradation of the GFP-Ubx with GFP-nanobody technology is more efficient than with the TEV-TIPI system. (P) Upper panel: Venn diagram representing the overlap between the neuronal stage 10–13 transcriptome (purple) and the genes up-regulated in mesodermal nuclei after Ubx depletion (RNAup UbxDeGrad) (pink). Lower panel: Fold enrichment of gene ontology terms of the 578 genes expressed in neuronal cells and up-regulated in mesodermal cells after Ubx depletion.
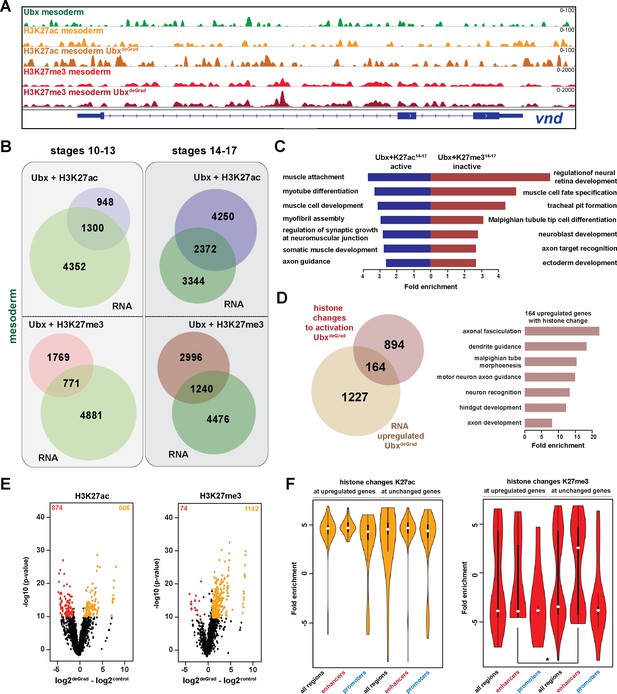
Ubx represses alternative fate genes by organizing the epigenetic landscape.
(A) ChIP-seq binding profiles of Ubx (green), H3K27ac (light +dark orange) and H3K27me3 (light +dark red) in control (UAS-Nslmb,GFP-Ubx) (light orange +red) and UbxDeGrad (Mef2 >Nslmb,GFP-Ubx) (dark orange +red) mesodermal nuclei at the vnd genomic locus (blue), a gene up-regulated in the absence of Ubx. (B) Venn diagrams representing the overlaps between genes expressed in the mesoderm (green), which are targeted by Ubx and simultaneously associated with H3K27ac (Ubx +H3K27ac, purple) or H3K27me3 (Ubx +H3K27me3, red) marks, respectively. (C) Fold enrichment of gene ontology terms among the genes expressed, targeted by Ubx and associated with H3K27ac marks (Ubx +K27ac14-17 active, blue) and the genes not expressed in the mesoderm, targeted by Ubx and associated with H3K27me3 marks (Ubx +K27me314-17 inactive, red). (D) Left panel: Venn diagram representing the overlap between genes up-regulated in Ubx depleted mesodermal nuclei (beige) and genes associated with histone mark changes indicative for gene activation (gain/increase in H3K27 acetylation, loss/reduction in H3K27 tri-methylation) (brown). Right panel: Fold enrichment of gene ontology terms of the 164 genes, which have histone changes to activation and are up-regulated in Ubx depleted mesodermal nuclei. (E) Volcano plot displaying the changes of H3K27ac and H3K27me3 marks between UAS-Nslmb,GFP-Ubx (control) and Mef2 >Nslmb,GFP-Ubx (degrade) INTACT-sorted mesodermal nuclei using differential binding analysis of ChIP-Seq peaks (DiffBind). The y-axis corresponds to the mean expression value of log10 (p-value), and the x-axis displays the difference of the log2 fold change value in UbxDegrad compared to the log2 of the control. The orange dots represent genomic regions containing an increase in H3K27ac or H3K27me3 (p-value<0.05), the red dots represent genomic regions containing a decrease in H3K27ac or H3K27me3 (p-value<0.05) between UAS-Nslmb,GFP-Ubx and Mef2 >Nslmb,GFP-Ubx INTACT-sorted mesodermal nuclei. (F) Comparison of H3K27ac or H3K27me3 fold enrichments at Ubx targeted genomic regions that experienced histone changes towards activation represented as Violin plots. Quantification of the histone changes in Ubx bound upregulated (164) and unchanged genes (894) by using the results of the differential binding analysis of ChIP-Seq peaks. The y-axis displays the fold change calculated by DiffBind. The x-axis shows all selected regions (upregulated and unchanged) and a sub-division of the regions in promoter (−1000 to +10 from TSS, 5’ UTR) and enhancer (distal enhancers (−2000 to −1000 from TSS), 3’ UTR, downstream, intronic regions, distal intergenic). Each sample contains a significant increase in H3K27ac levels (yellow), H3K27me3 levels (red) are decreased, but enhancer regions close to genes not changed in their expression have significantly higher H3K27me3 levels than enhancers close to genes up-regulated after Ubx depletion (*=p < 0.05).
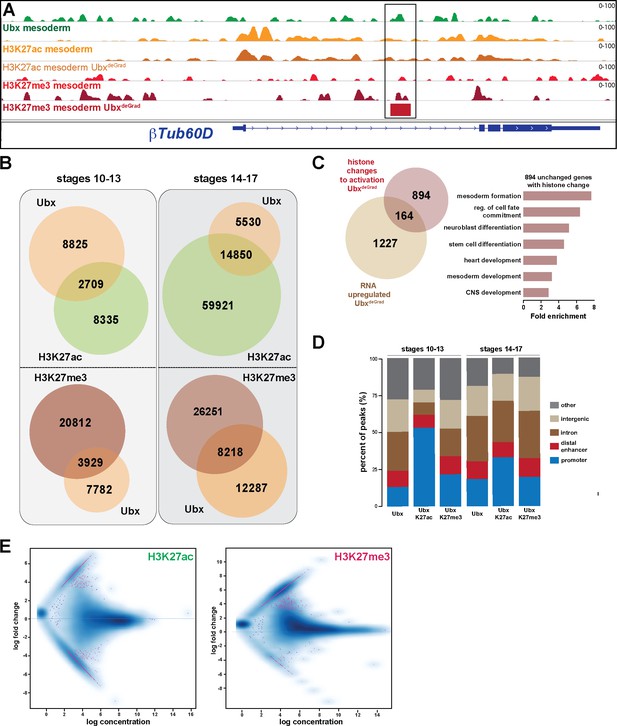
Ubx represses alternative fate genes by organizing the epigenetic landscape.
(A) ChIP-seq binding profiles of Ubx (green), H3K27ac (light +dark orange) and H3K27me3 (light +dark red) in control (Mef2-GAL4,GFP-Ubx) (light orange +red) and UbxDeGrad (Mef2 >Nslmb,GFP-Ubx) (dark orange +red) mesodermal nuclei at the βTub60D genomic locus (blue), a gene down-regulated in the absence of Ubx. (B) Venn diagrams representing the overlaps between genomic regions identified by Ubx (orange), H3K27ac (green) and H3K27me3 (red) ChIP-Seq experiments in the mesoderm at different developmental stages. (C) Left panel: Venn diagram representing the overlap between genes up-regulated in Ubx depleted mesodermal nuclei (beige) and genes associated with histone mark changes indicative for gene activation (gain/increase in H3K27 acetylation, loss/reduction in H3K27 tri-methylation) (brown). Right panel: Fold enrichment of gene ontology terms of the 894 genes, which have histone changes to activation and are unchanged in terms of expression in Ubx depleted mesodermal nuclei. (D) Location of peaks called within unique genomic regions of Ubx, Ubx-overlap with H3K27ac (Ubx K27ac), Ubx-overlap with H3K27me3 (Ubx K27me3) at different stages. Locations are classified as promoters (−1000- to +10 bp from TSS, 5’ UTR), distal enhancers (−2000 to −1000 from TSS, 3’ UTR, downstream), intron (intronic regions), intergenic (distal intergenic) and other regions (including exons). (E) MA plot displaying the changes of H3K27ac and H3K27me3 marks between UAS-Nslmb,GFP-Ubx (control) and Mef2 >Nslmb;GFP-Ubx (deGrade) INTACT-sorted mesodermal nuclei using differential binding analysis of ChIP-Seq peaks (DiffBind). Left panel: MA plot displaying the changes of H3K27ac histone mark between UAS-Nslmb,GFP-Ubx (control) and Mef2 >Nslmb,GFP-Ubx (deGrade) INTACT-sorted mesodermal nuclei (p-value<0.05).
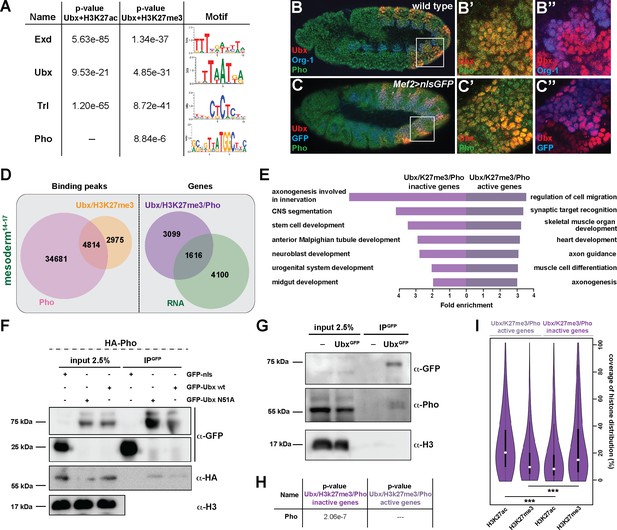
Ubx interacts with Pho at H3K27me3 marked inactive genes encoding alternative fate genes.
(A) The top four hits from MEME-ChIP analysis using all Ubx targeted, H3K27ac marked genomic regions in the vicinity of expressed genes (Ubx +H3K27ac) or Ubx targeted, H3K27me3 marked genomic regions in the vicinity of inactive genes (Ubx +H3K27me3) as input. While the Ubx, Exd and Trl motifs were found over-represented in both datasets, the Pho motif was found over-represented only in the Ubx +H3K27me3 dataset. (B–B’’) Lateral view of stage 11 wild-type Drosophila embryos stained for the mesodermal marker Org-1 (blue), Ubx (red) and Pho (green). (B’) and (B’’) are dual colour images of (B), highlighting that Ubx and Pho are co-expressed in Org-1 labelled mesodermal cells. (C–C’’) Lateral view of stage 11 Mef2 >nlsGFP Drosophila embryos stained for GFP (blue), Ubx (red) and Pho (green). (C’) and (C’’) are dual colour images of (C). (D) Left: Venn diagram representing the overlap between Ubx targeted and H3K27me3 marked chromatin regions (orange) and Pho binding peaks (pink) in stage 14–17 mesodermal nuclei. Right: Venn diagram representing the overlap between genes targeted by Ubx +Pho and marked by H3K27me3 (purple) and the genes expressed in the mesoderm (green). (E) Fold enrichment of gene ontology terms of the genes targeted by Ubx and Pho, which are additionally marked by H3K27me3 and either not expressed (3578) or expressed (1137) in mesodermal cells. (F) Co-immunoprecipitation of exogenous GFP fusion proteins using GFP-trap beads after transfection of S2R + Drosophila cells with HA-Pho coupled with GFP-nls (negative control), GFP-Ubx WT or N51A (mutant of the homeodomain, Asparagine 51 replaced by Alanine residue). Western blots were probed with the indicated antibodies. Pho is detected in the immunoprecipitated fraction of GFP-Ubx WT and N51A, while it is absent in the GFP negative control. (G) Co-immunoprecipitation with the GFP-trap system of endogenous proteins performed on nuclear extract from embryos expressing (or not) the endogenous GFP-Ubx fusion protein. Pho co-immunoprecipitates with GFP-Ubx, while it is absent from w1118 fly line. (H) MEME-ChIP analysis using Ubx and Pho targeted chromatin regions marked by H3K27me3 located in the vicinity of active or inactive genes identifies the classical Pho motif over-represented only in the vicinity of inactive genes. (I) Quantification of H3K27ac and H3K27me3 distributions at Ubx and Pho targeted chromatin regions marked by H3K27me3 located either in the vicinity of active or inactive genes are represented as Violin plot. The y-axis displays the coverage of the histone mark distribution in percent, the x-axis indicates the histone mark that was analysed. Active genes display a higher coverage of H3K27ac, while inactive genes have a higher coverage of H3K27me3 (***=p < 0.001).
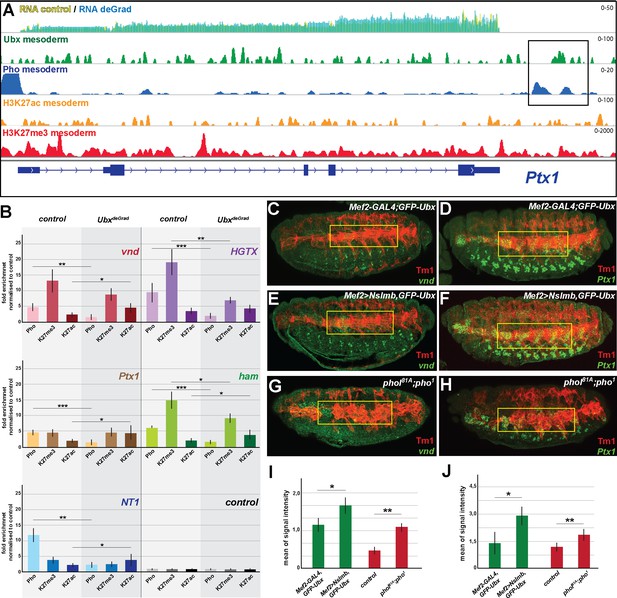
Ubx is required for stabilizing Pho binding to H3K27me3 chromatin regions.
(A) ChIP-seq binding profiles of Ubx (green), Pho (Pho), H3K27ac (orange) and H3K27me3 (red) in mesodermal nuclei as well as RNA-seq profiles in Mef2-GAL4,GFP-Ubx control (yellow) versus and Mef2 >Nslmb,GFP-Ubx (blue) mesodermal nuclei at the Ptx1 genomic locus. The Ptx1 gene are shown in blue. The box highlights the overlapping binding peaks of Ubx and Pho in the vicinity of the Ptx1 coding region. (B) ChIP-qPCR experiments for Pho, H3K27ac and H3K27me3 using chromatin regions close to the vnd, HGTX, Ptx1, ham and NT1 genes bound by Pho and Ubx using chromatin isolated from UAS-Nslmb,GFP-Ubx (light green) and Mef2 >Nslmb,GFP-Ubx (dark green) mesodermal nuclei. As control locus sites, a exon region in the scramb genes was used, which is not bound by Pho nor by Ubx. All loci show a significant reduction of Pho binding, H3K27me3 levels were reduced at the HGTX associated Ubx/Pho chromatin regions. In the vnd, Ptx1, ham and NT1 associated Ubx/Pho loci H3K27ac levels are significantly enriched (*=p < 0.05, **=p < 0.01, ***=p < 0.001). (C–H) Lateral view of stage 16 Mef2-GAL4,GFP-Ubx (C, D), Mef2 >Nslmb,GFP-Ubx (E, F) and phol81A;pho1 mutant (G, H) embryos stained for the muscle marker Tm1 (C–H) and for vnd (C, E, G) and Ptx1 (D, F, H) transcripts. The yellow boxes highlight the lateral muscles. (I, J) Quantification of vnd (I) and Ptx1 (J) signal intensities in the lateral muscles (as indicated by the yellow boxes) in Mef2-GAL4,GFP-Ubx, Mef2 >Nslmb,GFP-Ubx and phol81A;pho1 mutant embryos, showing that the expression of vnd and Ptx1 is significantly increased in comparison to control embryos (*=p < 0.05).
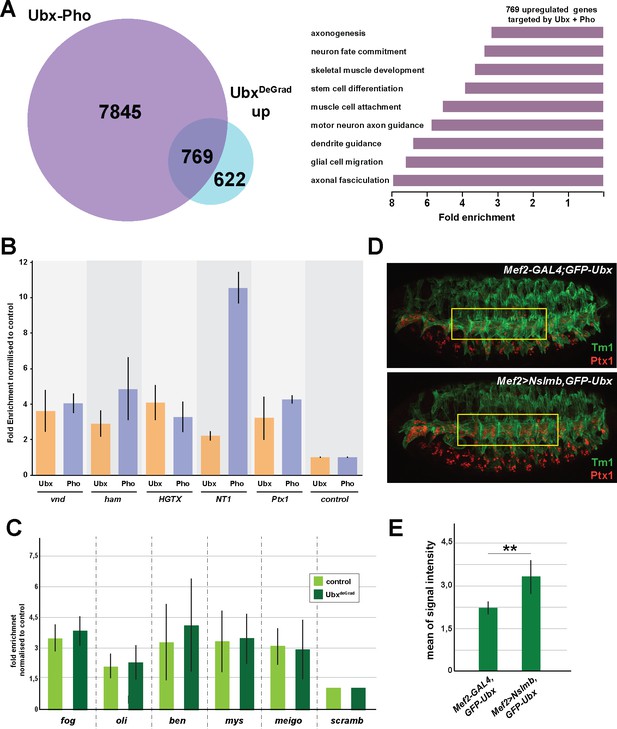
Ubx interacts with Pho at inactive genes encoding alternative fate genes.
(A) Left Panel: Venn diagram representing the overlap between genes up-regulated in Ubx depleted mesodermal nuclei (light blue) and genes associated with overlapping Ubx and Pho peaks (Ubx-Pho) (purple). Right panel: Fold enrichment of gene ontology terms of the 768 genes, which are bound by Ubx and Pho and are up-regulated in Ubx depleted mesodermal nuclei. (B) ChIP-qPCR experiments of Ubx and Pho using chromatin from stage 14–17 w1118 and twi-INTACT sorted nuclei, respectively. Genomic regions related to the genes vnd, HGTX, Ptx1, ham and NT1 are analyzed using the same primer pairs in close proximity (200 bp), the scramb1 locus is used as control. (C) ChIP-qPCR experiments of Pho detecting chromatin regions close to the fog, oli, ben, mys and meigo genes bound by Pho but not Ubx and chromatin isolated from UAS-Nslmb,GFP-Ubx (light green) and Mef2 >Nslmb,GFP-Ubx (dark green) mesodermal nuclei. As control locus, an exon region in the scramb genes was used, which is not bound by Pho or Ubx. (D) Lateral view of stage 16 Mef2-GAL4,GFP-Ubx and Mef2 >Nslmb,GFP-Ubx embryos stained for the muscle marker Tm1 and Ptx1 protein. The yellow boxes highlight the lateral muscles. (E) Quantification of Ptx1 signal intensities in the lateral muscles (as indicated by the yellow boxes) in Mef2-GAL4,GFP-Ubx and Mef2 >Nslmb,GFP-Ubx embryos, showing that the expression of Ptx1 is significantly increased in comparison to control embryos (**=p < 0.01).
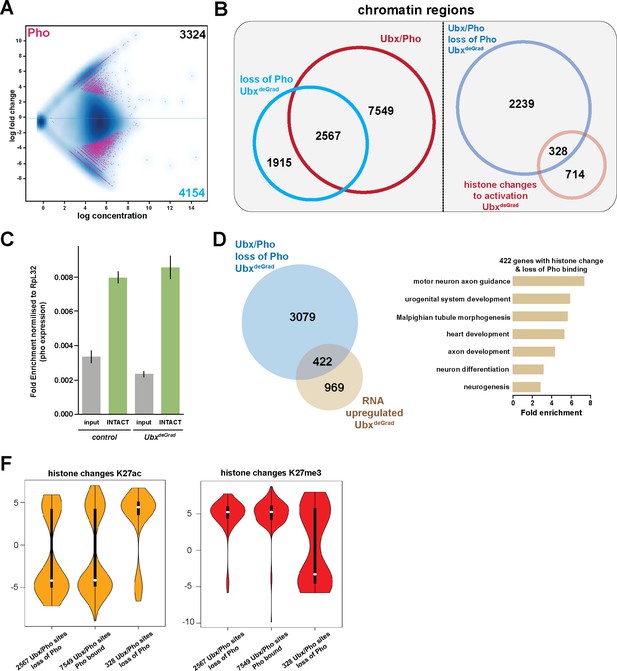
General relevance of Pho dependency on Ubx.
(A) MA plot displaying the changes of Pho binding between UAS-Nslmb,GFP-Ubx (control) and Mef2 >Nslmb;GFP-Ubx (UbxdeGrad) INTACT-sorted mesodermal nuclei using differential binding analysis of ChIP-Seq peaks. The y-axis corresponds to the log fold change, and the x-axis displays the log concentration. (B) Left panel: Venn diagram representing an overlap between genomic regions co-bound by Ubx and Pho (Ubx/Pho, red) and regions that lose Pho binding in the UbxdeGrad background. Right panel: Venn diagram illustrating the overlap between genomic regions co-bound by Ubx and Pho that experience a loss of Pho binding in the UbxdeGrad (light blue) and regions containing a histone mark change towards activation in the Ubx depleted mesodermal nuclei (light red). (C) Expression of pho in in UAS-Nslmb;GFP-Ubx (control) and Mef2 >Nslmb;GFP-Ubx (UbxdeGrad) mesodermal nuclei, fold change is normalized to Ribosomal protein L32 (RpL32). The expression of pho is not significantly changed in the mesodermal, INTACT- sorted nuclei (INTACT) upon Ubx depletion (UbxdeGrad) in comparison to the control sample. (D) Left panel: Venn diagram representing the overlap between genes up-regulated in Ubx depleted mesodermal nuclei (yellow) and genes associated with Ubx and Pho that experience a loss of Pho binding in the UbxdeGrad (blue). Right panel: Fold enrichment of gene ontology terms of the 422 genes, which contain Ubx and Pho binding, lose Pho binding and are up-regulated in Ubx depleted mesodermal nuclei. (F) Comparison of H3K27ac or H3K27me3 fold enrichments at Ubx and Pho targeted genomic regions that experienced a loss of Pho binding, represented as Violin plots. Quantification of the different regions: 1) Ubx/Pho sites that lose Pho binding (2597), 2) Ubx/Pho sites that maintain Pho binding (7549), 3) Ubx/Pho sites that lose Pho binding and experiences a histone mark change (328). The y-axis displays the fold change calculated by DiffBind. The x-axis shows all selected regions as disrobed 1), 2) and 3). Regions that lose Pho binding and experiences a histone mark change (328) experience a significant increase in H3K27ac levels (yellow) and a decrease in H3K27me3 levels (red).
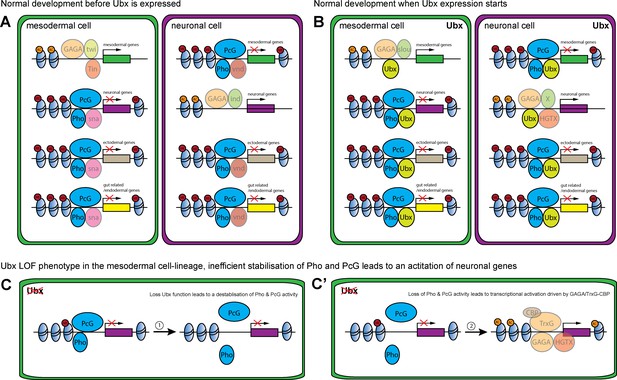
Model of Ubx and Pho combinatorial action in mesoderm development.
(A, B) Chromatin regions close to genes encoding different lineage functions are shown in mesodermal (green) and neuronal (purple) wild-type cells before (A) and after (B) Ubx is expressed. (A) Mesodermal determination and specification factors, like Twist (Twi) and Tinman (Tin), activate mesodermal genes, possibly in combination with Trl (GAGA) and/or other factors. In the same cell, alternative fate genes (grey: ectodermal genes, yellow: gut related genes; purple: neuronal) are repressed, possibly by the mesoderm-specific TF Snail (sna) with Pho and PcG (dark blue). In neuronal determination and specification, TFs like Intermediate neuroblasts defective (Ind) activate neuronal genes, possibly in combination with with Trl (GAGA), while alternative fate genes (grey: ectodermal genes, yellow: gut related genes; green: mesodermal) are repressed, for example via Ventral nervous system defective (Vnd) and Pho and PcG (dark blue). (B) When Ubx expression starts, determination and specification factors are turned off or their expression gets restricted. Now, Ubx ensures the continued activation of the proper set of lineage-specific genes, while at the same time maintaining the repression of alternative fate genes. (C, C’) Loss of Ubx in the mesodermal lineage destabilises Pho binding to regulatory regions of alternative fate genes (example for neuronal (purple)). (C) This results in a destabilisation of the PcG complex at these regions, leading to a loss/reduction of H3K23me3 marks. (C’) In the absence of Ubx and PcG, the sites can be bound by the TrxG complex through Trl (GAGA) in combination with CBP. Consequently, Lys 27 of histone three at these regions will get acetylated, resulting in the activation of gene expression.
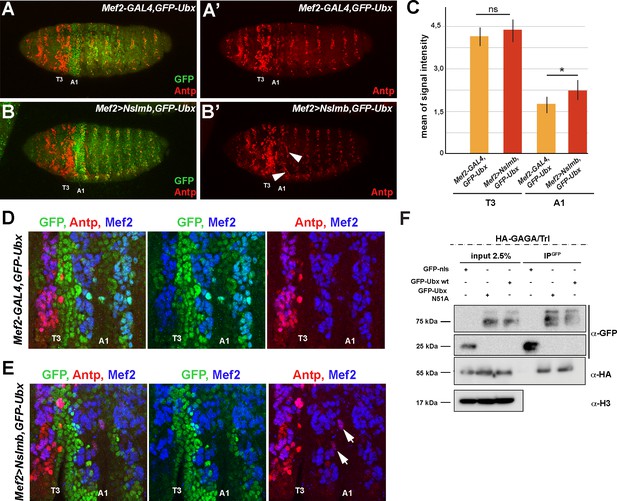
Antp is ectopically expressed in the mesoderm in the absence of Ubx.
(A, A’) Lateral view of stage 15 Mef2-GAL4,GFP-Ubx Drosophila embryos stained for GFP (green) and Antennapedia (Antp, red), (A’) single channel image of Antp (red). Third thoracic (T3) and first abdominal (A1) segments are indicated. (B, B’) Lateral view of stage 15 Mef2 >Nslmb,GFP-Ubx Drosophila embryos stained for GFP (green) and Antp (red), (B’) single channel for Antp (red). Arrowheads highlight Antp positive muscle cells in the first abdominal segment. (C) Quantification of Antp signal intensity in the third thoracic (T3) and first abdominal (A1) segment of Mef2-GAL4,GFP-Ubx and Mef2 >Nslmb,GFP-Ubx Drosophila embryos, showing an increase of Antp signal in A1 (*=p < 0.05). (D) Higher magnifications of lateral view of late stage 13 Mef2-GAL4,GFP-Ubx Drosophila embryos stained for GFP (green), Antp (red) and the and the muscle differentiation marker Mef2 (blue). Focus on third thoracic (T3) and first abdominal (A1) segments. (E) Higher magnifications of a lateral view of late stage 13 Mef2 >Nslmb,GFP-Ubx Drosophila embryos stained for GFP (green), Antp (red) and the and the muscle differentiation marker Mef2 (blue). Focus on third thoracic (T3) and first abdominal (A1) segments. Arrows mark Antp positive nuclei in the first abdominal segment. (F) Co-immunoprecipitation of exogenous GFP fusion proteins using GFP-trap beads after transfection of S2R + Drosophila cells with HA-Trl coupled with GFP-nls (negative control), GFP-Ubx WT or N51A (mutant of the homeodomain, Asparagine 51 replaced by Alanine residue). Western blots were probed with the indicated antibodies. Trl is detected in the immunoprecipitated fraction of GFP-Ubx WT and N51A, while it is absent in the GFP negative control.
Additional files
-
Supplementary file 1
Table of known genes expressed in the mesodermal or neuronal lineages.
- https://doi.org/10.7554/eLife.42675.017
-
Supplementary file 2
Primers used for qPCR experiments to test total RNA.
- https://doi.org/10.7554/eLife.42675.018
-
Supplementary file 3
Primers used for qPCR to test genomic loci identified by ChIP experiments.
- https://doi.org/10.7554/eLife.42675.019
-
Supplementary file 4
Collection of GO terms grouped into different categories for multiple GO term testing.
- https://doi.org/10.7554/eLife.42675.020
-
Transparent reporting form
- https://doi.org/10.7554/eLife.42675.021





