Identification of an emphysema-associated genetic variant near TGFB2 with regulatory effects in lung fibroblasts
Figures
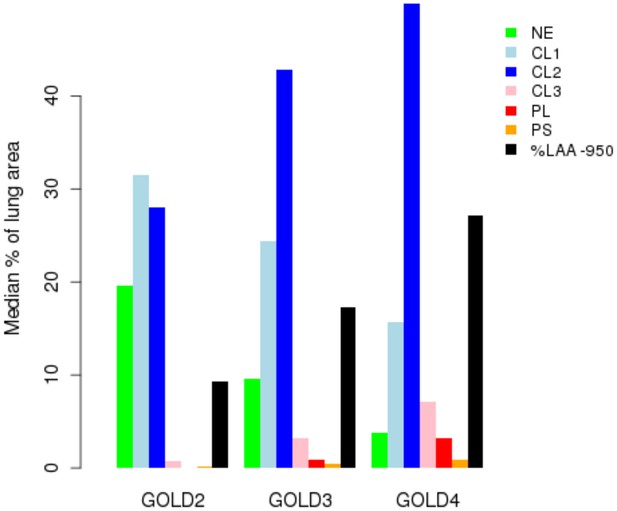
Percentage of each LHE-based emphysema pattern by Global Obstructive Lung Disease (GOLD) stage in ECLIPSE.
NE = Non-emphysematous lung. CL1 = Mild centrilobular. CL2 = Moderate centrilobular. CL3 = Severe centrilobular. PL = Panlobular. PS = Paraseptal. %LAA-950 = emphysema based on −950 Hounsfield unit threshold.
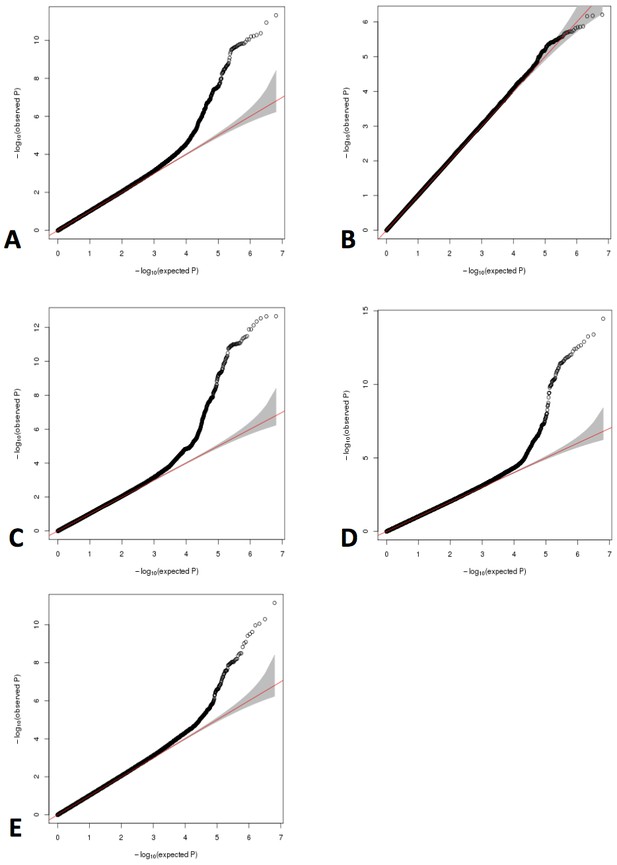
QQ plots for GWAS meta-analyses of non-transformed LHE phenotypes.
(A) Nonemphysematous lung. (B) Mild centrilobular pattern. (C) Moderate centrilobular. (D) Severe centrilobular. (E) Panlobular.
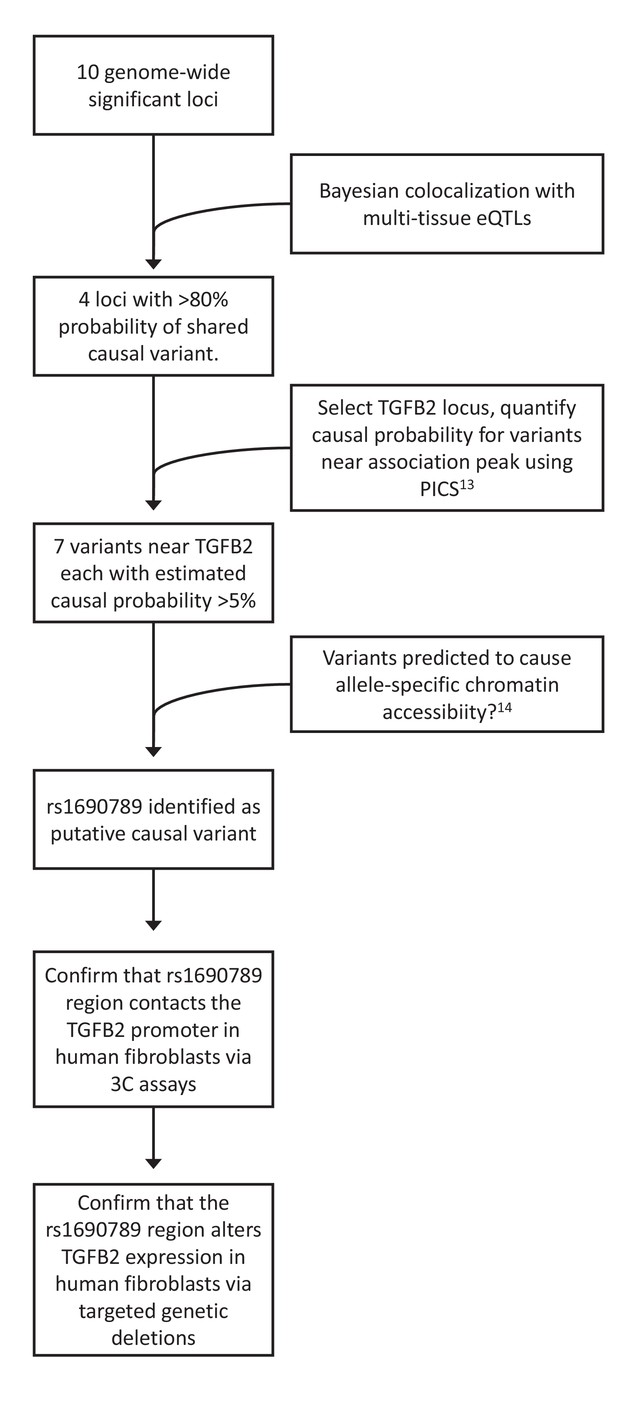
Overview of integrative analyses to prioritize genome-wide significant emphysema-associated loci for functional studies.
https://doi.org/10.7554/eLife.42720.006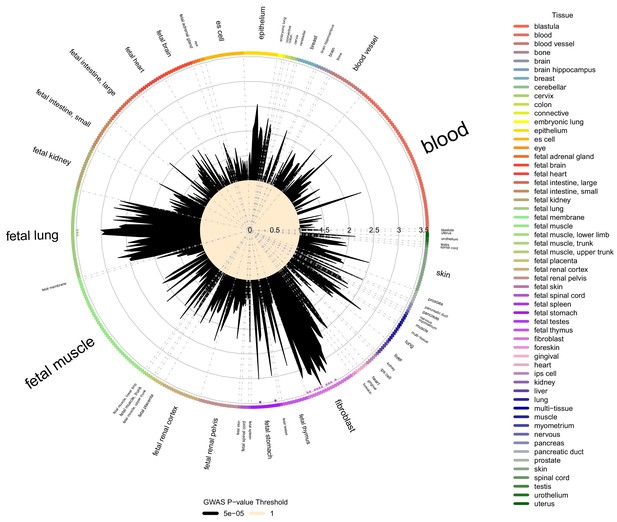
Cell type and tissue enrichment for moderate centribloular emphysema GWAS signals.
Using Garfield (Iotchkova et al., 2019) for enrichment analysis, we tested the enrichment of moderate centrilobular GWAS loci (harboring associations a p<5×10−5) in DNaseI peaks from 424 cell lines and cell types in ENCODE and Roadmap. Significant enrichments were observed in fetal lung, fetal stomach, and multiple fibroblast cell types. These significant enrichments are denoted by colored dots located just inside the boundary of the circle of cell types.
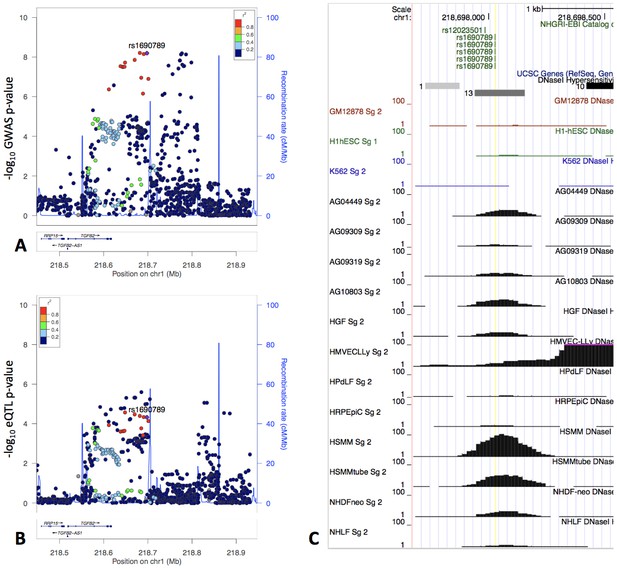
The locus zoom plot of GWAS p-values suggests two independent associations (Panel A), and the GWAS signal colocalizes with an eQTL signal in fibroblasts from GTEx (Panel B).
rs1690789 is located at one of these GWAS association peaks and lies within a context-specific DNaseI peak (Panel C). GM12878, H1hESC, and K562 cell lines are shown for reference, and the remaining cell types are those with DNaseI peaks that overlap rs1690789. Raw DNaseI data from only one experimental replicate are shown. GM12878 = lymphblastoid cell line. H1hESC = human embryonic stem cell. K562 = leukemia cell line. AG0449-AG10803 refer to fibroblasts from different subjects and sampling sites. HGF = gingival fibroblasts. HMVECLLy = lung derived microvascular endothelial cells. HPdLF - Periodontal ligament fibroblasts. HRPEpiC – retinal pigment epithelial cells. HSMM – skeletal muscle myoblasts. NHDF – dermal fibroblasts. NHLF – lung fibroblasts.
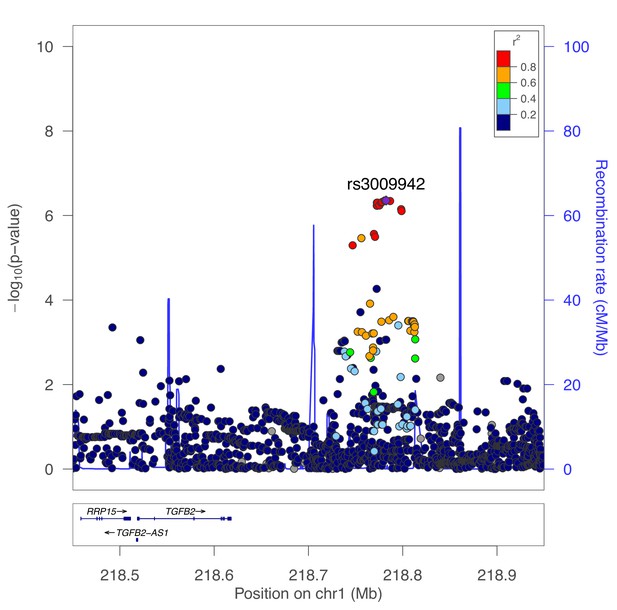
Secondary association for moderate centrilobular emphysema at the TGFB2 locus.
Results from genetic association meta-analysis conditioned on the lead SNP (rs796395) at this region.
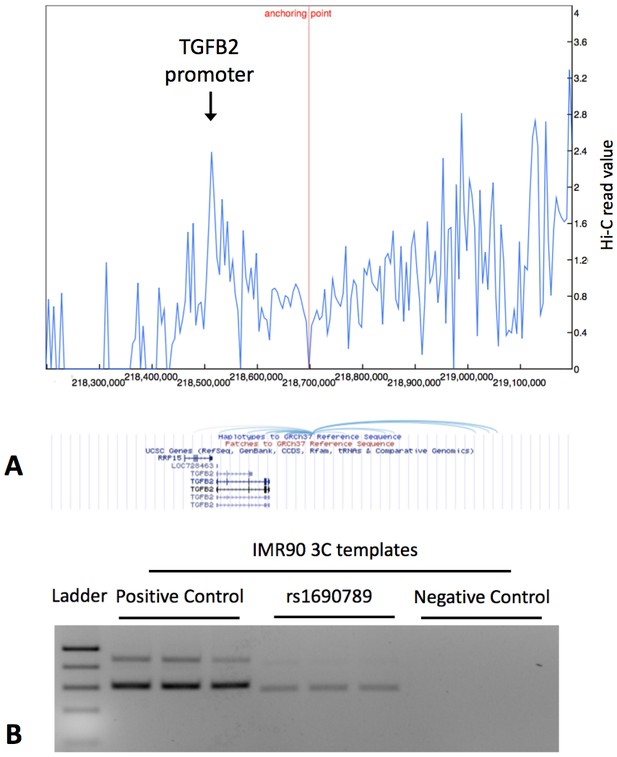
Publicly available chromatin conformation capture (4C) results in IMR90 cells show multiple peaks of interaction for the 10 kb region containing the context-specific regulatory element around rs1690789 (Panel A – blue spikes in top figure indicate regions of high interaction frequency and light blue curved lines in lower figure indicate chromosomal interactions with the 10 kb region containing rs1690789).
Newly generated 3C assays in IMR90 fibroblasts verify the interaction between the region containing rs1690789 and TGFB2 promoter (Panel B). Primer sequences are listed in Supplementary file 1 Table 10.
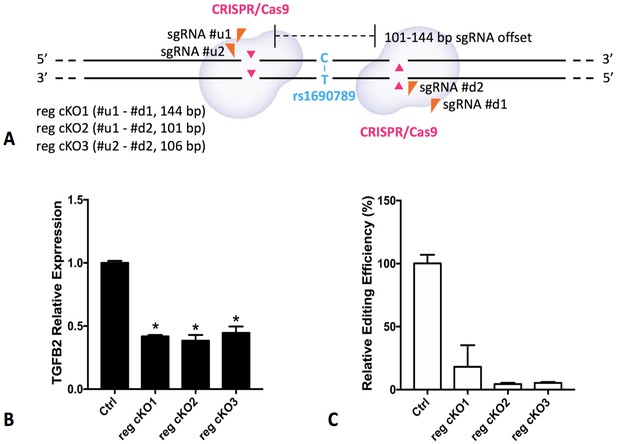
Regional knockout of rs1690789 in primary human lung fibroblasts using CRISPR/Cas9 editing.
Three pairs of sgRNAs were applied to delete a DNA region of ~100 bp spanning rs1690789 (A). The expression of TGFB2 is downregulated in rs1690789 knockout lung fibroblast cells with qPCR quantification, n = 4 (B). The editing efficiency is examined to confirm the effect of CRISPR/Cas9 regional knockout, n = 4 (C). *p<0.05 compared to control by unpaired t test.
Tables
Genome-wide significant meta-analysis results for local histogram emphysema phenotypes.
https://doi.org/10.7554/eLife.42720.005| LHE pattern | Lead SNP | Chromosome | Position (hg19) | EAF | Effect allele | P value meta | Effect meta | SE effect meta |
|---|---|---|---|---|---|---|---|---|
| Moderate Centrilobular | rs56077333 | 15 | 78899003 | 0.67 | a | 2.2E-13 | 0.016 | 2.2E-03 |
| rs17368582 | 11 | 102738075 | 0.14 | t | 8.1E-12 | 0.024 | 3.5E-03 | |
| rs56113850 | 19 | 41353107 | 0.59 | t | 1.5E-09 | −0.016 | 2.6E-03 | |
| rs796395 | 1 | 218681971 | 0.52 | a | 6.1E-09 | 0.013 | 2.2E-03 | |
| rs138641402 | 4 | 145445779 | 0.38 | a | 3.3E-08 | 0.014 | 2.5E-03 | |
| Nonemphysematous (Normal Lung) | rs138641402 | 4 | 145445779 | 0.38 | a | 4.2E-08 | −0.021 | 3.9E-03 |
| rs7170068 | 15 | 78912943 | 0.78 | a | 4.8E-12 | 0.028 | 4.0E-03 | |
| rs17368659 | 11 | 102742761 | 0.86 | t | 6.3E-11 | 0.036 | 5.4E-03 | |
| rs28929474* | 14 | 94844947 | 0.98 | t | 6.2E-09 | −0.071 | 1.2E-02 | |
| Panlobular | rs11852372 | 15 | 78801394 | 0.35 | a | 7.1E-12 | −0.003 | 5.0E-04 |
| rs76756075* | 11 | 112349844 | 0.02 | t | 2.0E-08 | −0.007 | 1.2E-03 | |
| rs78070126* | 11 | 6574608 | 0.97 | t | 2.8E-08 | 0.006 | 1.1E-03 | |
| rs145770770 | 2 | 152487808 | 0.99 | a | 3.8E-08 | −0.015 | 2.8E-03 | |
| Severe Centrilobular | rs9788721 | 15 | 78802869 | 0.38 | t | 3.4E-15 | −0.005 | 6.0E-04 |
| rs379123 | 17 | 30891814 | 0.40 | t | 3.7E-08 | −0.004 | 7.0E-04 |
-
LHE - local histogram emphysema.
EAF - effect allele frequency in 1000 Genomes CEU population.
-
*indicates novel association not previously associated in GWAS of COPD or emphysema (rs28929474 was associated to FEV1/FVC in smokers in Li et al., 2018, during preparation of this manuscript).
Genomewide significant LHE loci that colocalize with eQTL.
https://doi.org/10.7554/eLife.42720.007| GWAS | SNP | CHR | POS | Tissue | GENE |
|---|---|---|---|---|---|
| Moderate centrilobular | rs796395 | 1 | 218681971 | Fibroblasts | TGFB2 |
| rs56077333 | 15 | 78899003 | Fibroblasts, Testis | PSMA4, CHRNA5 | |
| Moderate centrilobular, normal | rs56113850 | 19 | 41353107 | Lung, Testis | CYP2A6, AKT2 |
| Panlobular | rs11852372 | 15 | 78801394 | Testis | CHRNA5 |
| All loci have colocalization probability > 80%, reflecting the estimated probability that the GWAS and eQTL association signals arise from a shared causal variant. LHE pattern – local histogram emphysema phenotype used for GWAS. SNP - lead GWAS variant in locus. Tissue - tissue of origin for gene expression data in eQTL analysis. Gene – eQTL targeted gene in specified GTEx tissue. | |||||
Additional files
-
Supplementary file 1
This file contains Tables 1-11.
- https://doi.org/10.7554/eLife.42720.013
-
Transparent reporting form
- https://doi.org/10.7554/eLife.42720.014





