Macrophages restrict the nephrogenic field and promote endothelial connections during kidney development
Figures
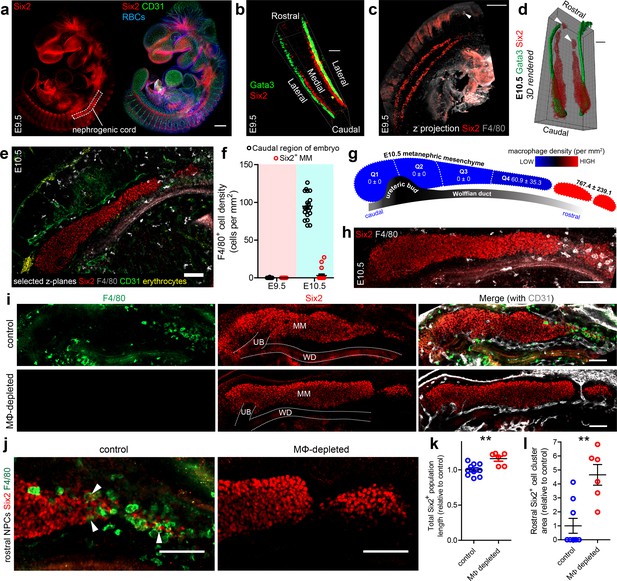
Macrophages and the initiation of kidney development.
(a) Nephrogenic cord location in the E9.5 embryo. (b) E9.5 nephrogenic cord and Wolffian duct arrangement. (c) Z-projection of E9.5 caudal part of mouse embryo and nephrogenic cords. White arrowhead shows an F4/80+ macrophage. (d) 3D rendering of E10.5 Wolffian ducts (Gata3) and nephrogenic progenitors (Six2). White arrowheads show isolated clusters of rostral nephrogenic cells. (e) F4/80+ macrophage localisation relative to metanephric mesenchyme at E10.5. (f) Macrophage density at E9.5 and E10.5 (n = 20 fields of view from two cleared E9.5 embryos and 20 fields from two cleared E10.5 embryos). MM, metanephric mesenchyme. (g–h) Macrophage density along the rostro-caudal axis of the E10.5 metanephric mesenchyme and within isolated groups of rostral nephrogenic cells (n = 4 kidneys from two embryos). Image in (h) shows a representative image of an E10.5 kidney. (i) Representative control and macrophage (MΦ)-depleted E11.5 metanephric mesenchyme populations (n for controls = 10 kidneys from six embryos; n for MΦ-depleted = 6 kidneys from three embryos). (j) Rostral nephrogenic progenitor cells (NPCs) in control and MΦ-depleted embryos. White arrowheads in left panel show Six2+ nuclei within cell bodies of F4/80+ macrophages. (k) Six2+ populations are extended in MΦ-depleted embryos compared to controls (two-tailed t-test; p=0.0031). (l) Rostral Six2+ clusters are larger in area in MΦ-depleted embryos compared to controls (two-tailed Mann-Whitney test; p=0.0042). Scale bars = 100 µm.
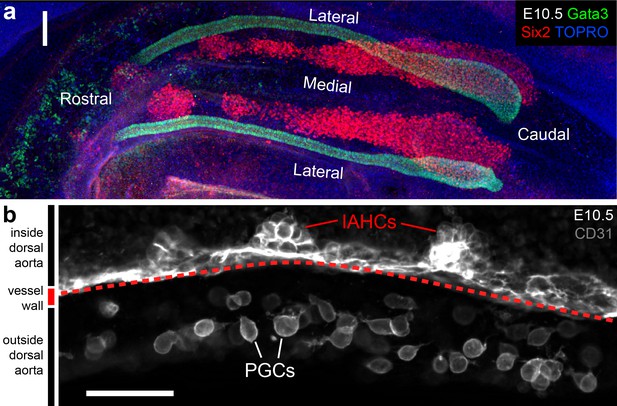
E10.5 immunostaining.
(a) E10.5 nephron progenitor cell arrangement. (b) CD31 marks intra-aortic hematopoietic cluster (IAHC) cells and primitive germ cells (PGCs) in the caudal part of the E10.5 mouse embryo. Scale bars: a = 100 µm; b = 50 µm.
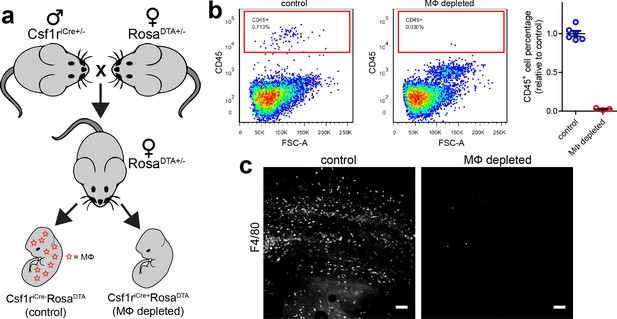
Macrophage depletion system and its consequences.
(a) Outline of breeding strategy used to deplete macrophages. (b) Flow cytometry data showing CD45+ cell numbers in control and macrophage depleted embryos. (c) Immunostaining data showing F4/80+ cells in caudal regions of control and macrophage depleted embryos (identical microscope settings were used). Scale bars = 50 µm.
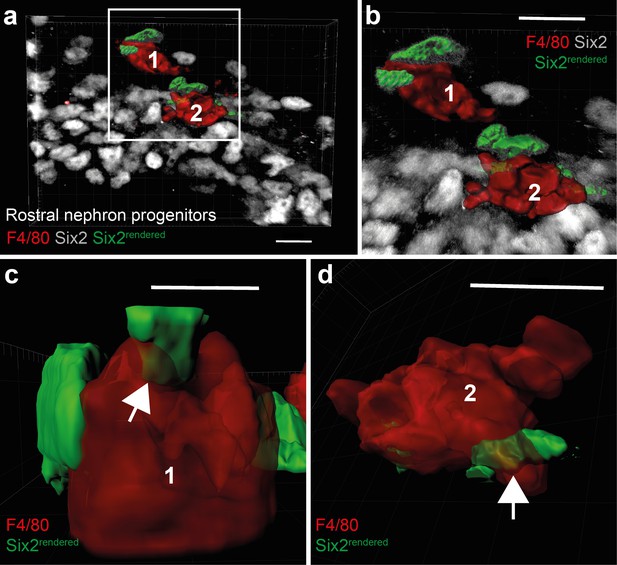
Rostral nephrogenic progenitor cell engulfment by macrophages.
(a) Overview of a region containing rostral Six2+ nephron progenitors and macrophages at E10.5. The Six2+ cells rendered in green were in contact with the two macrophages. (b) Shows a zoomed in image of the region covered by the white box in (a). (c–d) Shows Six2+ nuclei of rostral nephron progenitor cells partially surrounded by macrophage membranes (white arrowheads). Scale bars = 10 µm.
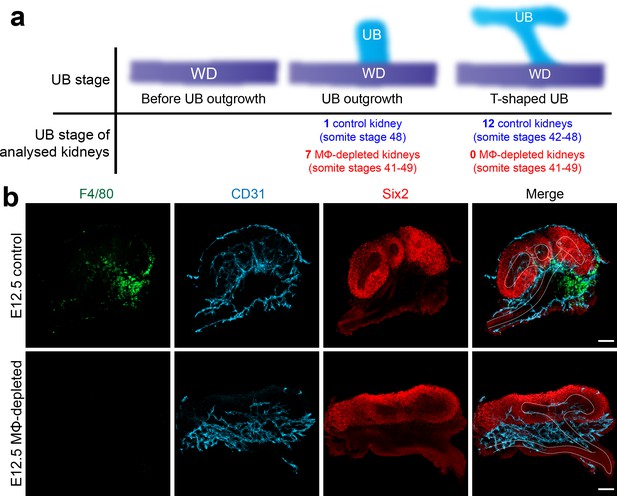
Ureteric bud development is delayed in E11.5-E12.5 macrophage-depleted embryos.
(a) Ureteric bud development in ~E11.5 macrophage-depleted kidneys relative to littermate control embryos (somite pairs of macrophage depleted embryos = 42–49 [n = 7 kidneys, four embryos]; somite pairs of control embryos = 41–48 [n = 13 kidneys, seven embryos]). UB, ureteric bud; WD, Wolffian Duct. (b) Ureteric bud development in an E12.5 macrophage-depleted embryo and littermate control embryo. White dashed lines in ‘Merge’ panels indicate position of the ureteric bud. Identical microscope settings were used for images in (b). Images in (b) are representative of n = 2 E12.5 macrophage-depleted embryos from two litters; n = 7 E12.5 control embryos from two litters. Scale bars = 100 µm.
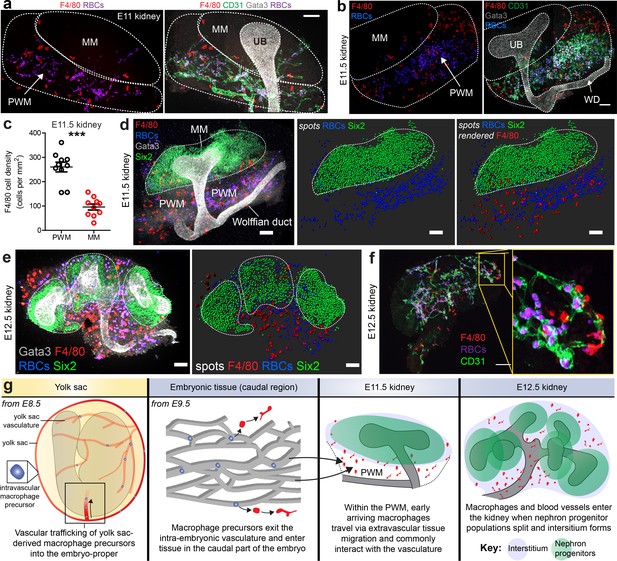
Arrival of macrophages in the early kidney.
(a–b) E11 and E11.5 kidneys: Macrophages localise in high numbers within the vascularised peri-Wolffian mesenchyme (PWM) but not within the metanephric mesenchyme (MM). Staining shows macrophages (F4/80), ureteric bud (UB; Gata3), erythrocytes (heme autofluorescence), and vasculature (CD31) (stitched confocal z-stacks; a = 2×1 tiles, b = 2×2 tiles). (c) Macrophage density is higher in the peri-Wolffian mesenchyme than the metanephric mesenchyme (macrophage density in peri-Wolffian mesenchyme (PWM) = 260.20 ± 20.08 per mm2 [mean ±SEM]; macrophage density in metanephric mesenchyme (MM) = 95.86 ± 12.29 per mm2 [mean ±SEM]; two-tailed paired t-test [testing whether densities were the same]; p<0.0001; n = 10 kidneys from five embryos). (d) At E11.5, nephron progenitors (Six2) gather as a single population, which is predominantly devoid of macrophages (F4/80) and red blood cells (RBCs; heme autofluorescence). (e) By E12.5, nephron progenitor cell populations have split, creating fissures in the mesenchyme that are occupied by macrophages and erythrocytes. The dashed white lines represent the boundaries of Six2+ nephron progenitor populations. Images are 3D z-projections; as a result, macrophages and RBCs appearing within Six2+ populations may in fact localise outside (in 3D). (f) At E12.5, many kidney macrophages are perivascular. (g) Working model for macrophage arrival in the kidney (based on our results and data from Stremmel et al., 2018). Scale bars = 100 µm (except e), which = 70 µm).
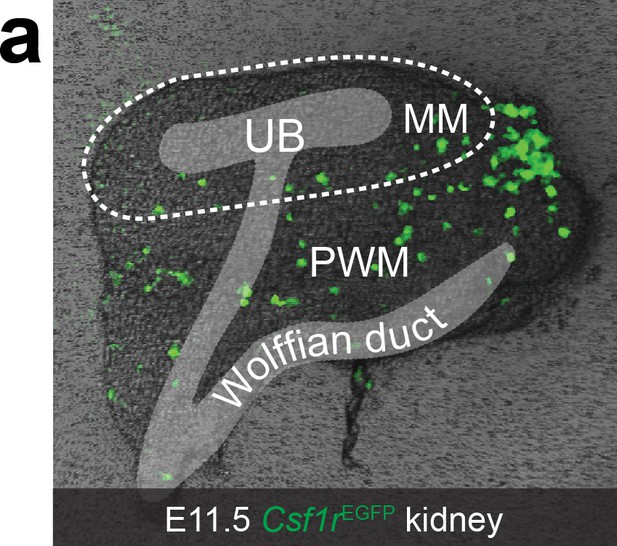
E11.5 Csf1rEGFP kidney.
(a) White dashed line indicates boundary of metanephric mesenchyme (MM). The location of the ureteric bud (UB) and Wolffian duct have been added in grey. PWM, peri-Wolffian mesenchyme.
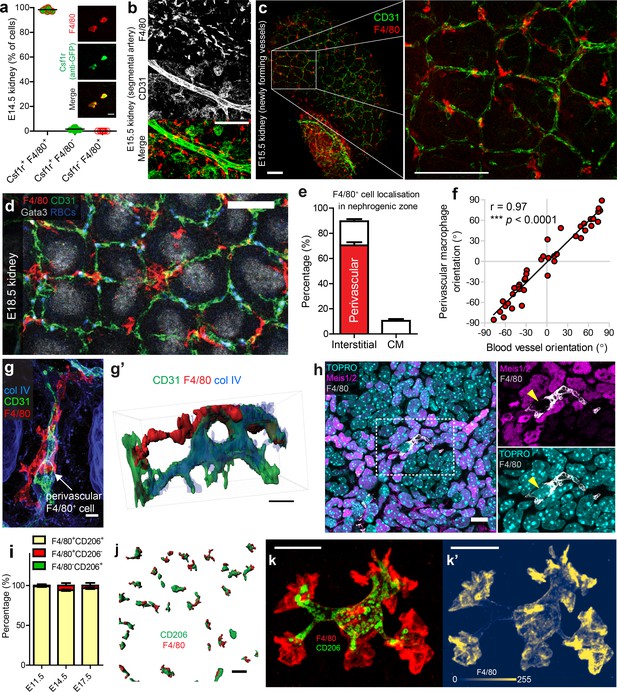
Most F4/80highCD206high kidney macrophages are interstitial perivascular cells.
(a) All F4/80+ cells co-express Csf1r (anti-GFP+) in the developing kidney (n = 730 cells from 8 z-slices of a cleared E14.5 ‘MacGreen’ kidney). Inset boxes show example Csf1r+F4/80+ cells. (b) Macrophages arrange parallel to major vessels in the E15.5 kidney, but rarely contact their endothelia. (c–d) Macrophages interact with newly forming vessels in the cortical nephrogenic zone (c = E15.5 and d = E18.5). (e) Most kidney macrophages in the E17.5 nephrogenic zone are interstitial perivascular cells (n = 10 z-stacks from four kidneys). The red bar shows the percentage of interstitial macrophages that are perivascular. CM, cap mesenchyme. (f) Macrophages align with their underlying blood vessels (p for correlation <0.0001; n = 40 macrophage/blood vessel pairs from n = 8 kidneys at E17.5). (g–g’) Example of a perivascular macrophage in the nephrogenic zone, with intra-, peri-, and extra-vascular regions (g’ is a 3D rendered version of g). (h) Macrophages do not express the renal interstitial cell markers, Meis1/2. (i) Percentage of cells singly or doubly positive for CD206 and F4/80 in the E11.5 kidney and the cortical nephrogenic zone of E14.5 and E17.5 kidneys. Discernible co-staining at E11.5, 98.8 ± 0.71 [mean ±SEM], n = 5 kidneys; E14.5, 94.1 ± 1.0% of cells, n = 10; at E17.5, 96.5 ± 1.0% of cells, n = 10. (j) Representative image of CD206+F4/80+ cells (3D rendered) in nephrogenic zone. (k–k’) High-resolution 3D image of an F4/80+CD206+ macrophage. (k’) shows cellular intensity of F4/80 using colourmap developed by Nuñez et al. (2018). Scale bars: b-c = 100 µm; (d) j = 50 µm; g = 5 µm; g’) =, h, k-k’=10 µm.
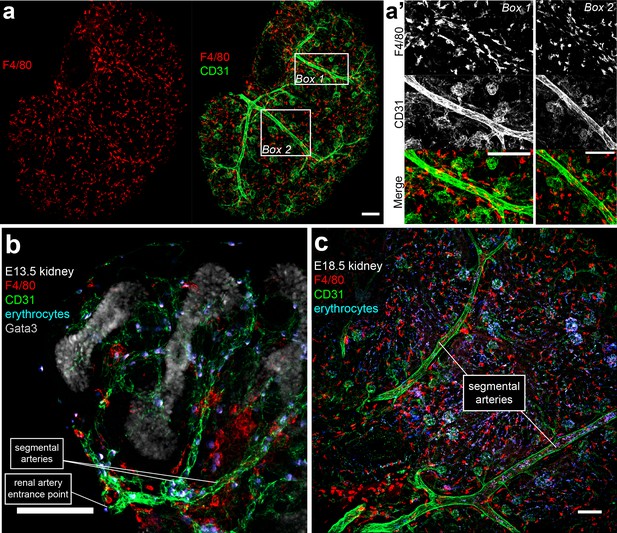
Macrophage arrangement relative to major renal blood vessels.
(a–c) Macrophages arrange parallel to segmental arteries in (a–a’) E15.5, (b) E13.5, and (c) E18.5 kidneys. Scale bars = 100 µm.
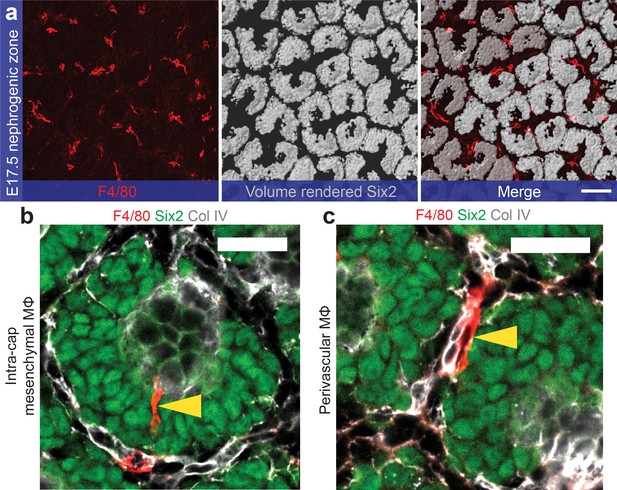
F4/80+macrophages predominantly avoid nephron progenitor populations in the nephrogenic zone.
(a) Macrophage localisation around nephron progenitor populations in the E17.5 nephrogenic zone (Six2+ populations are computer rendered). (b) A small percentage of macrophages localise inside the nephron progenitor population (intra-cap mesenchymal MΦs). (c) Most interstitial macrophages in the nephrogenic zone are perivascular. Scale bars: a = 100 µm; b), c = 20 µm.
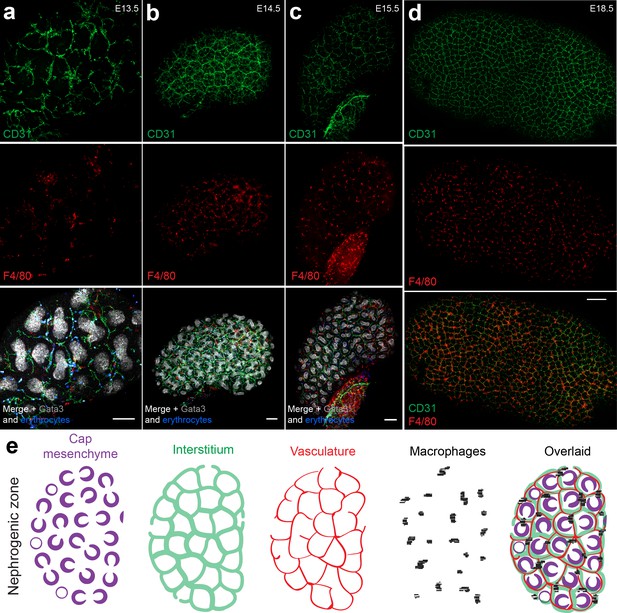
Most F4/80+macrophages localise around the interstitial vasculature in the nephrogenic zone.
(a–d) Macrophages interact with vascular plexuses in the nephrogenic zone of (a) E13.5, (b) E14.5, (c) E15.5, and (d) E18.5 kidneys. (e) Schematic depicting macrophage localisation within the nephrogenic zone (ureteric tips are not shown for illustrative purposes). Scale bars = 100 µm (except d), where the scale bar represents 200 µm).
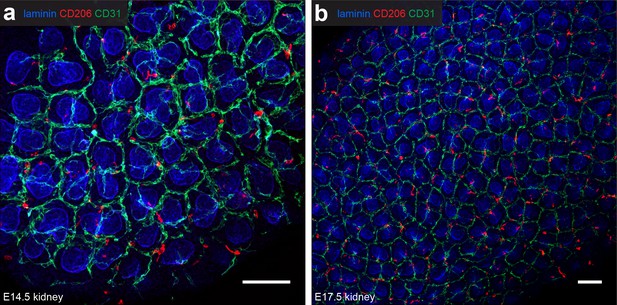
CD206+macrophages in the nephrogenic zone.
(a–b) Most CD206+ macrophages are perivascular in the nephrogenic zone of (a) E14.5 and (b) E17.5 kidneys. Scale bars = 100 µm.
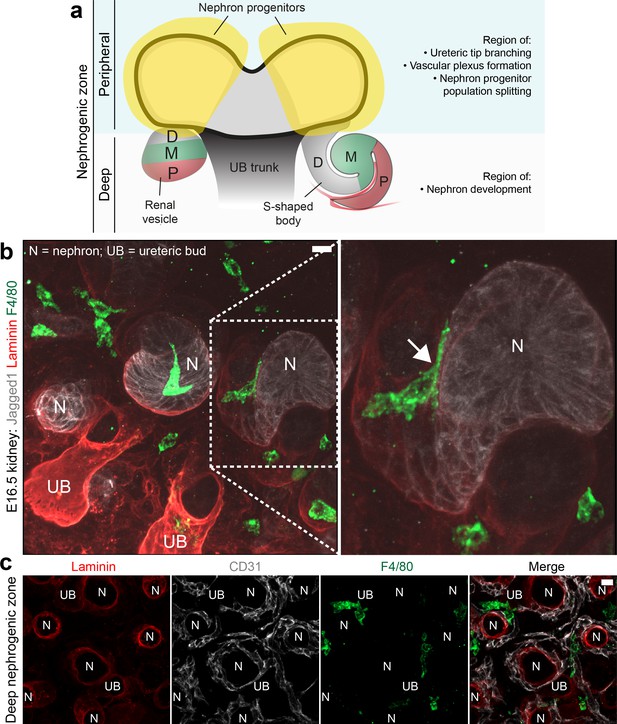
Macrophages interact with nephrons and their vasculature in the deep nephrogenic zone.
(a) Cartoon representing the different regions of the nephrogenic zone and the different processes that occur in the deep and peripheral regions. D, distal; M, medial; P, proximal. (b) Macrophages (F4/80) interact with the epithelial tubules of developing nephrons (Jagged1). (c) Macrophages interact with the blood vessels (CD31) adjacent to developing nephrons (laminin, showing basement membrane). N, nephron; UB, ureteric bud. Scale bars = 10 µm.
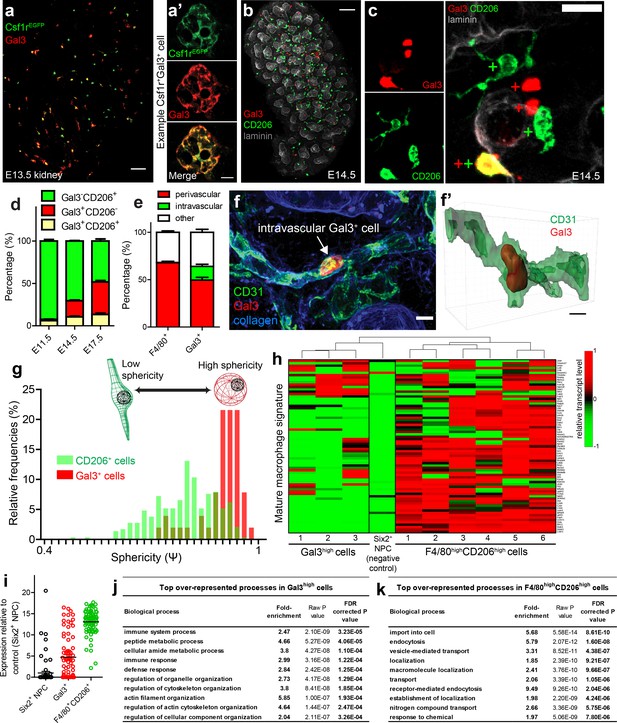
Characterisation of Gal3+cells in kidney development.
(a–a’) Gal3 is expressed by a subset of Csf1rEGFP+ cells. (a’) shows an example Gal3+Csf1r+ cell. (b) Gal3+ and CD206+ cells in the E14.5 kidney. (c) Examples of Gal3+CD206-, Gal3-CD206+, and Gal3+CD206+ cells in the E14.5 kidney. (d) The proportion of Gal3+ myeloid cells increases over developmental time, while the relative proportion of CD206+ cells decreases. (e) Quantification of Gal3+ and F4/80+ macrophage localisation in E17.5 nephrogenic zone (n = 14 fields of view for F4/80 localisation and n = 15 fields of view for Gal3 localisation). The only intravascular cells were Gal3+. (f–f’) Example of an intravascular Gal3+ cell at E17.5. 3D rendering of the intravascular Gal3+ cell from f). (g) Gal3+ cells are more spherical than CD206+ cells (p<0.0001; two-tailed t-test; n = 290 CD206+ cells; n = 51 Gal3+ cells from E14.5 nephrogenic zone). (h) Expression of mature macrophage signature genes by individual F4/80highCD206high cells, Gal3high cells, and a nephron progenitor cell (negative control). (i) Average mRNA transcript levels for genes associated with mature macrophage status (n = 6 F4/80highCD206high cells; n = 3 Gal3high cells; n = 1 Six2high nephron progenitor cell [NPC]). Each data point represents the average expression of a single relevant gene. (j) Top 10 over-represented biological processes in Gal3high cells. (k) Top 10 over-represented biological processes in F4/80highCD206high cells. Scale bars: a = 10 µm; a’ and f = 5 µm; b = 100 µm; c = 20 µm.
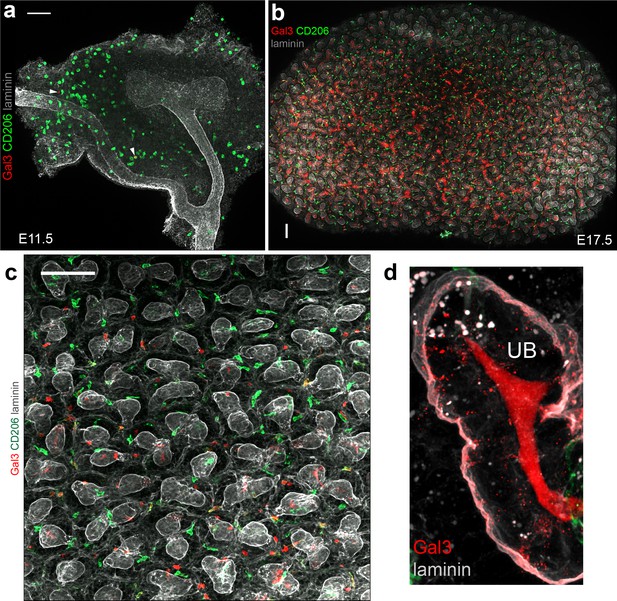
Gal3 expression in the developing kidney.
(a–b) Representative overview images of Gal3+ and CD206+ cells in the (a) E11.5 and (b) E17.5 kidney. White arrowheads in a) show Gal3+CD206+ cells. (c) Gal3+ and CD206+ cells in a region of the nephrogenic zone of the E17.5 kidney. (d) Gal3+ is also expressed in tubular lumens in E17.5 kidneys. Scale bars = 100 µm.
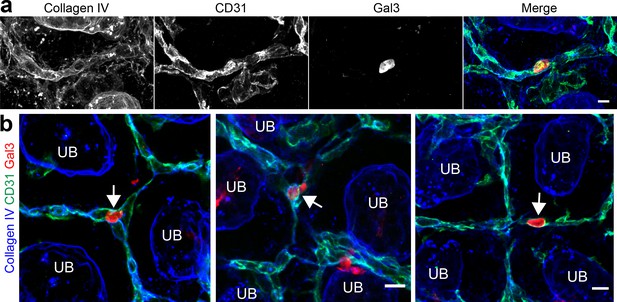
Examples of Gal3+cells being carried within blood vessels of the developing kidney.
(a) Individual channels for the image shown in Figure 4f–f’. (b) Other examples of Gal3 +cells being carried within the renal vasculature in the nephrogenic zone. Scale bars = 10 µm.
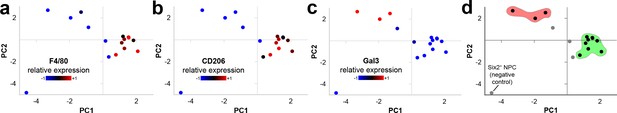
Macrophage single-cell RNA sequencing gene expression signatures.
(a–d) F4/80+CD206+ cells separate distinctly from Gal3+ cells. PC, principle component. Each dot represents a single-cell. The green group in d) shows F4/80+CD206+ cells and the pink group shows Gal3+ cells.
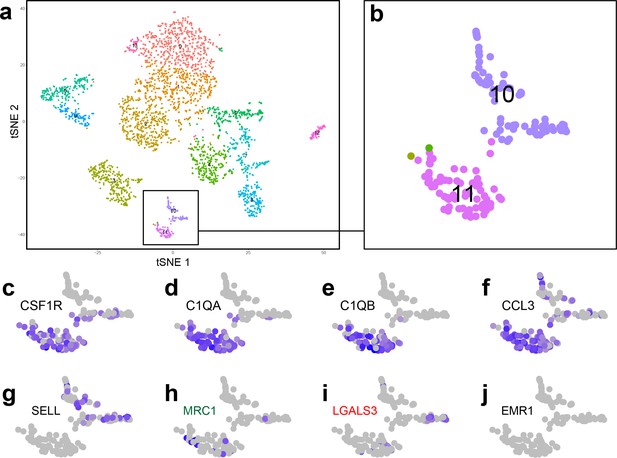
Analysis of single-cell RNA sequencing data from human foetal kidney cells.
(a) tSNE plot showing individual cells from the cortex of the developing human kidney. (b) Two clusters (10 and 11) were identified as immune cells. (c–f) Cluster 11 cells are macrophages, which express macrophage markers such as (c) CSF1R, Colony stimulating factor one receptor; (d) C1QA, Complement C1q subcomponent subunit A; (e) C1QB, Complement C1q subcomponent subunit B; (f) CCL3, macrophage inflammatory protein-1α. (g) Cluster 10 cells are a distinct leukocyte cell type, which specifically express L-selectin (SELL). (h–i) MRC1+ cells are mostly found within cluster 11 (macrophage group), whereas LGALS3+ cells are distributed within clusters 10 and 11. (j) EMR1, the human homolog of F4/80, is not expressed by human kidney macrophages.
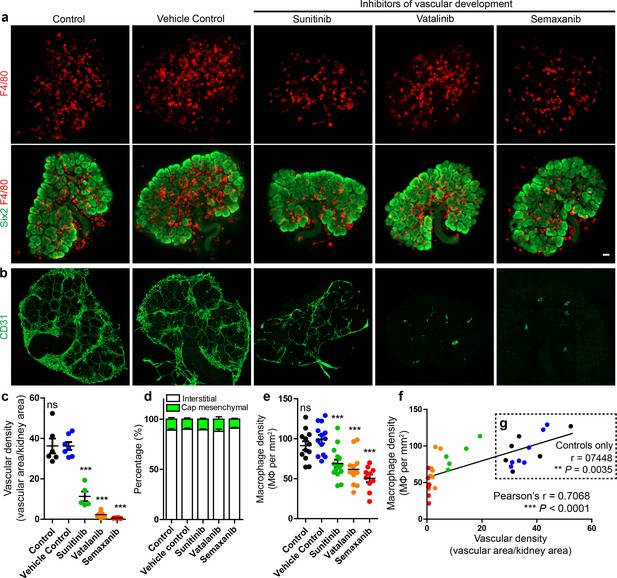
Macrophage localisation in the interstitium is not dependent on renal endothelial cells in culture.
(a) Representative images of macrophages (F4/80) and nephron progenitor populations (Six2) in control and treated kidneys. (b–c) Endothelial cells are robustly depleted in sunitinib, vatalinib, and semaxanib treated kidneys (representative images; stitched confocal z-stacks, 3 × 3 tiles; p for overall ANOVA < 0.0001; n = 5–8 per group). (d) Macrophage localisation is unaltered between groups (p for overall ANOVA = 0.71; n = 4–9 kidneys per group). (e) Macrophage density is decreased in vascular-depleted kidneys (p for overall ANOVA < 0.0001; n = 10–15 kidneys per group). (f–g) Macrophage density positively correlated with vascular density in all groups (f): r = 0.71, 0.48–0.85 95% CI; p<0.0001; n = 32 kidneys) and in control groups only (g): r = 0.74, 0.33–0.92 95% CI; p=0.0035; n = 13 kidneys). In f–g), the data point colour represents its group (e.g. green dots represent sunitinib-treated kidneys). Data are from two identical experiments (different antibodies were used for post-fixation sample staining). Macrophage localisation data in d) were determined from the first experimental run. Vascular density data used in c), (f), and g) were obtained from the second experimental run. Macrophage density data in e) includes data from both experimental runs. Scale bar = 100 µm.
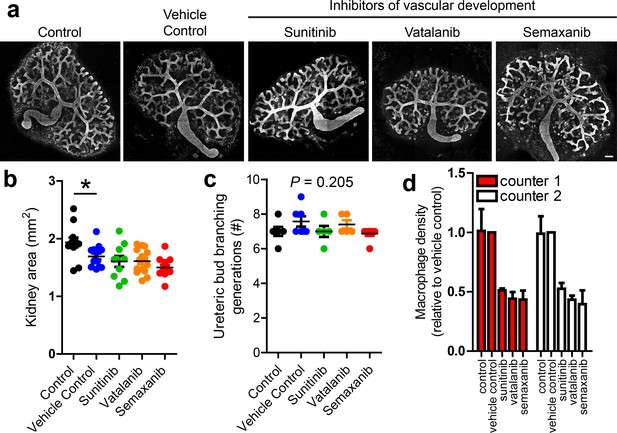
Influence of vascular inhibition on other aspects of kidney development.
(a) Representative images of collecting duct trees in different groups. (b) Kidney area in control and treatment groups (n = 10–15 per group; one-way ANOVA; p=0.0003). (c) Ureteric bud branching generations did not differ between groups (n = 5–8 per group; one-way ANOVA; p=0.205). (d) Accuracy of macrophage density counts was confirmed by a second blinded counter (n = 3 per group; two-way ANOVA; p for counter differences = 0.842). Scale bar = 100 µm.
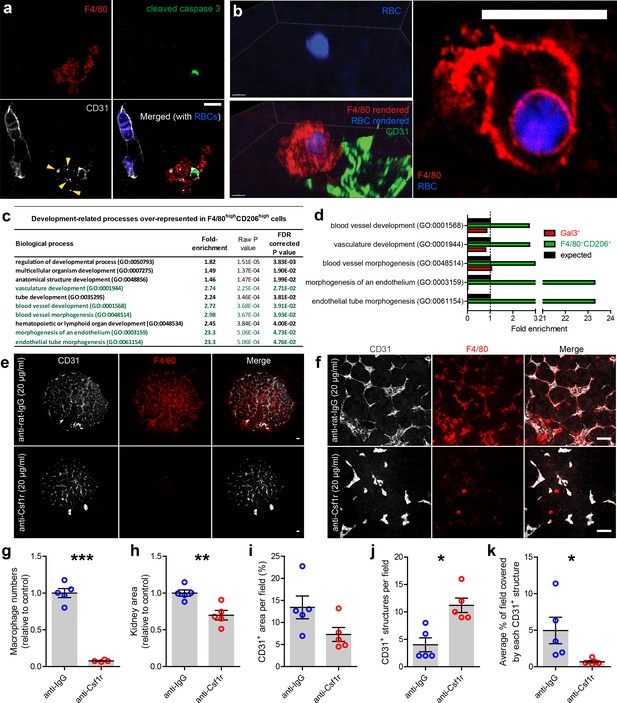
Kidney macrophages facilitate vascular development.
(a) Apoptotic endothelial cell (cleaved caspase 3+CD31+) in the cortical nephrogenic zone being phagocytosed by an F4/80+ kidney macrophage. Yellow arrowheads show CD31+ cell debris within the macrophage cell body. (b) F4/80+ kidney macrophage phagocytosing an erythrocyte (RBC; heme autofluorescence) in the E13.5 kidney. (c–d) Genes associated with vascular development are significantly over-represented in the top 1% of expressed genes in F4/80highCD206high macrophages, but not in the top 1% of the Gal3high myeloid cells. Biological processes highlighted in green in a) are all related to vascular development. (e–f) Representative (e) overview images and (f) fields of view of anti-rat IgG (control) and anti-Csf1r (blocking antibody) treated kidney explants. (g–k) Anti-Csf1r treated kidneys had (g) fewer macrophages (p<0.0001), (h) reduced kidney area (p=0.0046), (j) more isolated endothelial structures (p=0.011), and (k) smaller endothelial structures (p=0.046) relative to control treated kidneys (n = 5 biological replicates; 1 field of view per kidney). (i) Vascular density (CD31+ area per field) did not significantly differ between groups (p=0.077). Data that were normally distributed (Kolmogorov-Smirnov normality testing) were compared using two-tailed t-tests. Only data from j) were not normally distributed and were compared using a non-parametric Mann-Whitney test. Scale bars: a) and b = 10 µm; e) and f = 100 µm.
Videos
E9.5 3D caudal-part mouse embryo (Six2, F4/80, and CD31).
Few F4/80+ macrophages are present in the embryo-proper at this stage.
E9.5 and E10.5 caudal-part mouse embryos z-plane scroll (Six2, F4/80, CD31, and erythrocytes [heme autofluorescence]).
https://doi.org/10.7554/eLife.43271.009Time-lapse showing macrophages and isolectin-B4-labelled blood vessels within cultured E11.5 Csf1rEGFP kidneys.
Isolectin-B4 also labels macrophages. White arrows present in the video between 18–40 s show examples of perivascular macrophages.
E13.5 and E14.5 MacGreen kidneys (F4/80, GFP, and Gata3).
https://doi.org/10.7554/eLife.43271.019E15.5 and E18.5 kidney macrophages (F4/80, Gata3, erythrocytes, and CD31).
https://doi.org/10.7554/eLife.43271.020Macrophage contact with epithelial tubules of developing nephrons (F4/80, Jagged 1, Laminin).
https://doi.org/10.7554/eLife.43271.021F4/80+ macrophages engulfing an erythrocyte (heme autofluorescence) and an apoptotic endothelial cell in kidney development (scroll through individual confocal z-planes).
https://doi.org/10.7554/eLife.43271.030Additional files
-
Supplementary file 1
Genes used for single-cell RNA sequencing PCA graphs.
- https://doi.org/10.7554/eLife.43271.031
-
Supplementary file 2
Biological processes enriched in the top 1% of genes expressed by Gal3high cells.
- https://doi.org/10.7554/eLife.43271.032
-
Supplementary file 3
Biological processes enriched in the top 1% of genes expressed by F4/80high CD206high cells.
- https://doi.org/10.7554/eLife.43271.033
-
Supplementary file 4
Cellular components enriched in the top 1% of genes expressed by Gal3high cells.
- https://doi.org/10.7554/eLife.43271.034
-
Supplementary file 5
Cellular components enriched in the top 1% of genes expressed by F4/80high CD206high cells.
- https://doi.org/10.7554/eLife.43271.035
-
Supplementary file 6
Antibodies used in this study.
- https://doi.org/10.7554/eLife.43271.036
-
Transparent reporting form
- https://doi.org/10.7554/eLife.43271.037





