Synergy and remarkable specificity of antimicrobial peptides in vivo using a systematic knockout approach
Figures
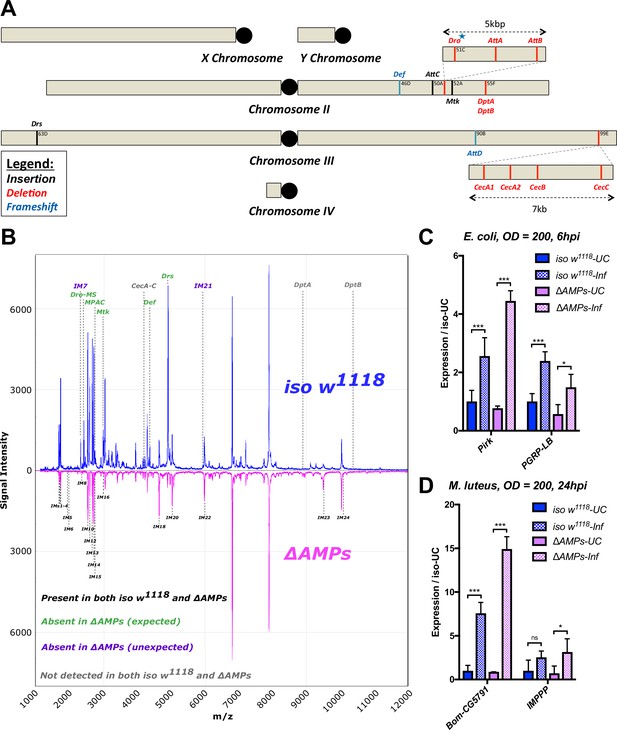
Description of AMP mutants.
(A) Chromosomal locations of AMP genes that were deleted. Each mutation is color-coded with the mutagenic agent: black, a Minos insertion or homologous recombination, red, CRISPR-CAS9-mediated deletion, and blue CRISPR CAS9 mediated indel causing a nonsense peptide. (B) A representative MALDI-TOF analysis of hemolymph samples from immune-challenged (1:1 E. coli and M. luteus at OD600 = 200) iso w1118 and ΔAMPs flies as described in Uttenweiler-Joseph et al. (1998). No AMP-derived products were detected in the hemolymph samples of ΔAMPs flies. No signals for IM7, nor IM21 were observed in the hemolymph samples of ΔAMPs mutants suggesting that these uncharacterized immune-induced molecules are the products of AMP genes. The Imd pathway (C) and Toll pathway (D) are functional and respond to immune challenge in ΔAMPs flies. We used alternate readouts to monitor the Toll and Imd pathways: pirk and PGRP-LB for Imd pathway and CG5791 (Bomanin) and IMPPP for Toll signaling (De Gregorio et al., 2002; Hanson et al., 2016). UC = unchallenged, Inf = infected. hpi = hours post-infection. Expression normalized with iso w1118-UC set to a value of 1.
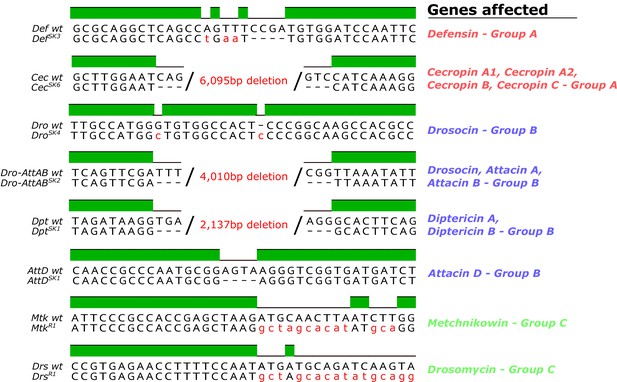
Genetic description of mutations generated in this study.
The MtkR1 and DrsR1 mutations entirely replaced the CDS with an insert from the piHR vector. Non-synonymous nucleotides in mutants are given in red. Mutations are listed according to groups in Figures 3–6 (discussed later).
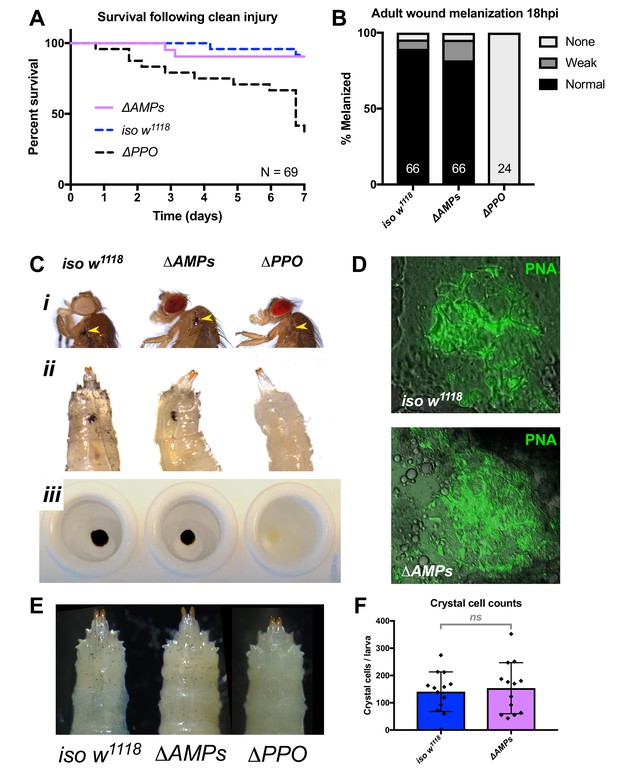
ΔAMPs flies have otherwise wild-type immune reactions.
(A) ΔAMPs flies survive clean injury like wild-type flies, while ΔPPO mutants deficient for melanization have reduced survival over time. (B) ΔAMPs flies melanize the cuticle similar to wild-type flies following pricking (χ2 = 2.14, p=0.34). Melanization categories (None, Weak, Normal) were as described in Dudzic et al (Dudzic et al., 2018). Sample sizes (n) are included in each bar. (C) Melanization in iso w1118, ΔAMPs, and ΔPPO flies of the cuticle in adults (i, yellow arrowheads), larvae (ii, melanized wounds), and larval hemolymph (iii). (D) To investigate clotting ability, we used the hanging drop assay (Scherfer et al., 2004) with ΔAMPs larval hemolymph and visualized clot fibers with PNA staining (green). Both iso w1118 and ΔAMPs hemolymph produced visible clot fibres measured after 20 min. Hemocyte populations are normal in ΔAMPs flies, including crystal cell distribution (E) and number (F).
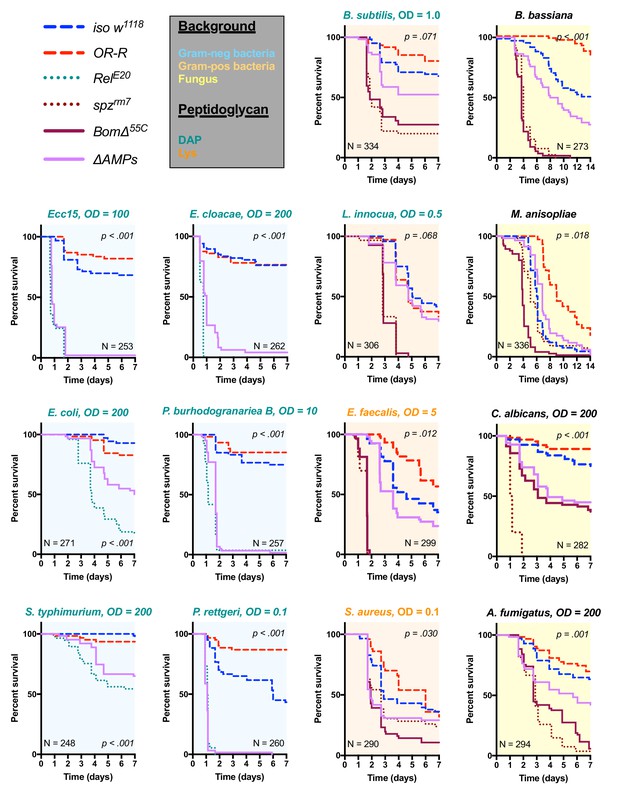
Survival of ΔAMPs flies to diverse microbial challenges.
Control lines for survival experiments included two wild-types (w;Drosdel (iso w1118) and Oregon R (OR-R) as an alternate wild-type), mutants for the Imd response (RelE20), mutants for Toll signaling (spzrm7), and mutants for Bomanins (BomΔ55C). ΔAMPs flies are extremely susceptible to infection with Gram-negative bacteria (blue backgrounds). Unexpectedly, ΔAMPs flies were not markedly susceptible to infection with Gram-positive bacteria (orange backgrounds), while BomΔ55C flies were extremely susceptible, often mirroring spzrm7 mutants. This pattern of BomΔ55C susceptibility held true for fungal infections (yellow backgrounds). ΔAMPs flies are somewhat susceptible to fungal infections, but the severity shifts with different fungi. Pellet densities are reported for all systemic infections in OD at 600 nm. p-Values are given for ΔAMPs flies compared to iso w1118 using a Cox-proportional hazards model. N = total number of flies in experiments. A full description of p-values relative to iso w1118 can be found in Figure 2—source data 1.
-
Figure 2—source data 1
p-Values from Figure 2A relative to iso w1118.
- https://doi.org/10.7554/eLife.44341.008
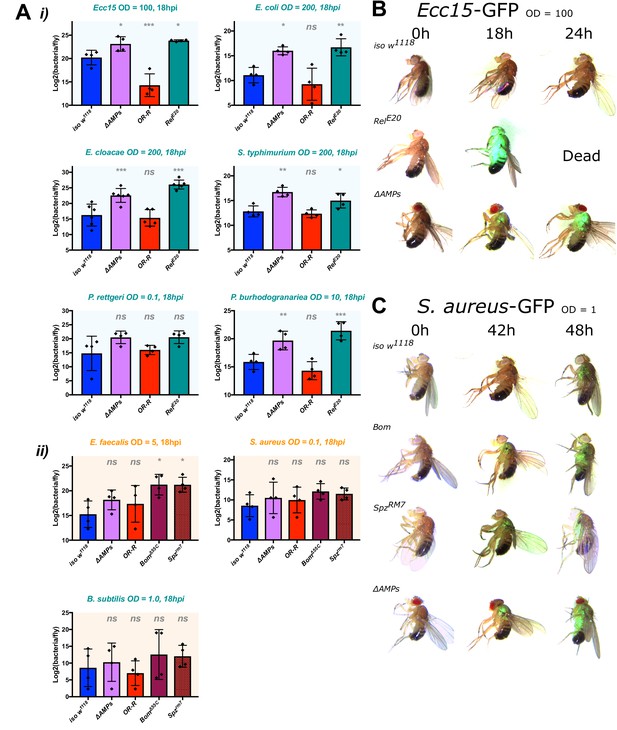
ΔAMPs flies fail to suppress Gram-negative bacterial growth.
Colony counts were performed on pooled samples (five flies) for bacteria amenable to LB agar, a medium that avoids overnight growth of the host microbiota. (A) For Gram-negative bacterial infections, ΔAMPs flies have significantly higher bacterial loads compared to iso w1118 at 18 hr post-infection (hpi) (i). This is not true for any of the Gram-positive bacteria tested (ii), while spzrm7 mutants carried higher bacterial loads, significantly so in E. faecalis infections. Gram-negative (B) and Gram-positive (C) infections with GFP-labelled bacteria spread from the wound site systemically in all genotypes tested. Thus, ΔAMPs fly mortality is likely not due to tissue-specific colonization by invading bacteria, but rather a failure to suppress bacterial growth first locally, and then systemically. One-way ANOVA: not significant = ns, p<0.05 = *, p<0.01 = **, and p<0.001 = *** relative to iso w1118.
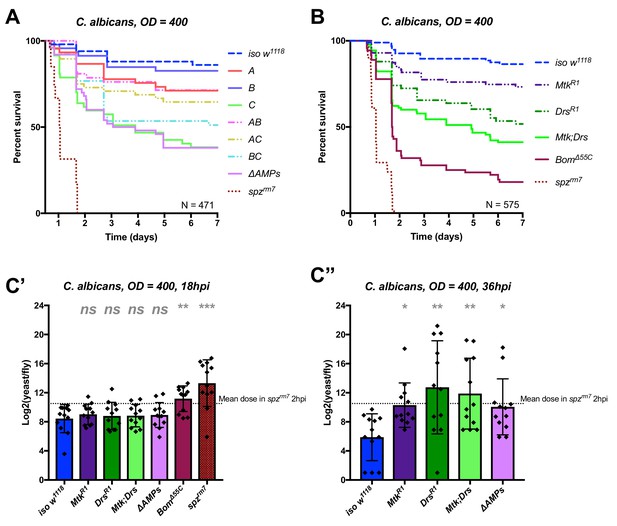
Identification of AMPs involved in the susceptibility of ΔAMPs flies to C. albicans.
(A) Survival of mutants for groups of AMPs reveals that loss of only Toll-responsive Group C peptides (Metchnikowin and Drosomycin) is required to recapitulate the susceptibility of ΔAMPs flies. Co-occurring loss of groups A and C has a net protective effect (A*C: HR = −1.71, p=0.002). (B) Further dissection of Group C mutations reveals that both Metchnikowin and Drosomycin contribute to resist C. albicans survival (p=0.008 and p<0.001, respectively). The interaction of Metchnikowin and Drosomycin was not different from the sum of their individual effects (Mtk*Drs: HR = −0.80, p=0.116). (C) Fungal loads of individual flies at 18 hpi. At this time point, BomΔ55C mutants and spzrm7 flies have already failed to constrain C. albicans growth (C’). Fungal titres at 36hpi (C’’), a time point closer to mortality for many AMP mutants, show that some AMP mutants fail to control fungal load, while wild-type flies consistently controlled fungal titre. One-way ANOVA: not significant = ns, p<0.05 = *, p<0.01 = **, and p<0.001 = *** relative to iso w1118.
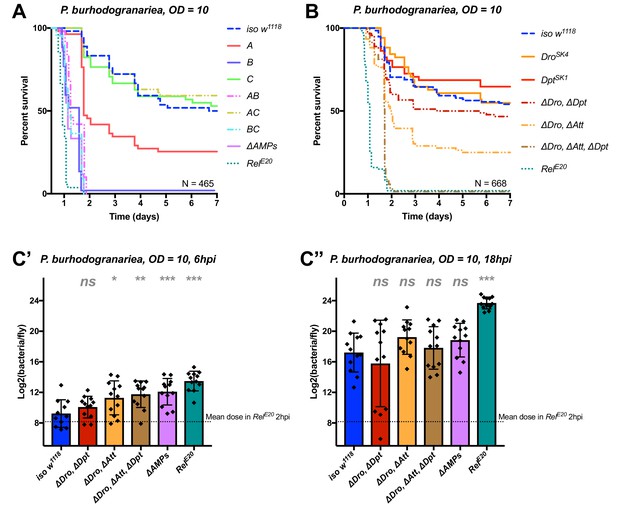
Identification of AMPs involved in the susceptibility of ΔAMPs flies to P. burhodogranariea.
(A) Survival of mutants for groups of AMPs reveals that loss of Imd-responsive Group B peptides (Drosocin, Attacins, and Diptericins) recapitulates the susceptibility of ΔAMPs flies. Loss of Group A peptides also resulted in strong susceptibility (p<0.001) due to additive effects of Defensin and Cecropins (Figure 4—figure supplement 4). (B) Further dissection of AMPs deleted in Group B reveals that only the loss of all Drosocin, Attacin, and Diptericin gene families leads to susceptibility similar to ΔAMPs flies. Simultaneous loss of Attacins and Diptericins results in a synergistic loss of resistance (ΔAtt*ΔDpt: HR =+1.45, p<0.001). (C) Bacterial loads of individual flies at 6 hpi (C’). At this time point, most AMP mutants had significantly higher bacterial loads compared to wild-type flies. At 18 hpi (C’’), differences in bacterial load are reduced, likely owing to the high chronic load P. burhodogranariea establishes even in surviving flies (Duneau et al., 2017). Meanwhile RelE20 flies succumb ~18 hr earlier than ΔAMPs flies in survival experiments, and already have significantly higher loads. One-way ANOVA: not significant = ns, p<0.05 = *, p<0.01 = **, and p<0.001 = *** relative to iso w1118.
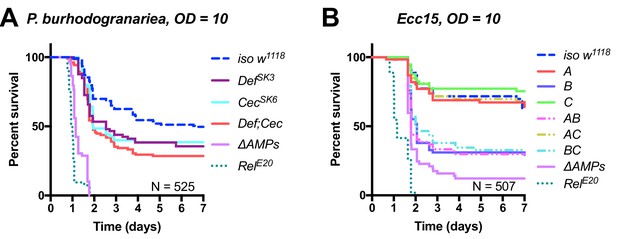
Further dissecting effects of AMP groups.
(A) Dissection of the susceptibility of Group A flies lacking Defensin and Cecropins reveals that combined mutants have an additive loss of resistance (Def*Cec, HR =+0.36, p=0.342). (B) Upon infection with the Gram-negative Ecc15, Group B peptides (Drosocin, Attacins and Diptericins) explain the bulk of mortality, but additional loss of other peptides in ΔAMPs flies leads to increased mortality (Log-Rank p=0.013).
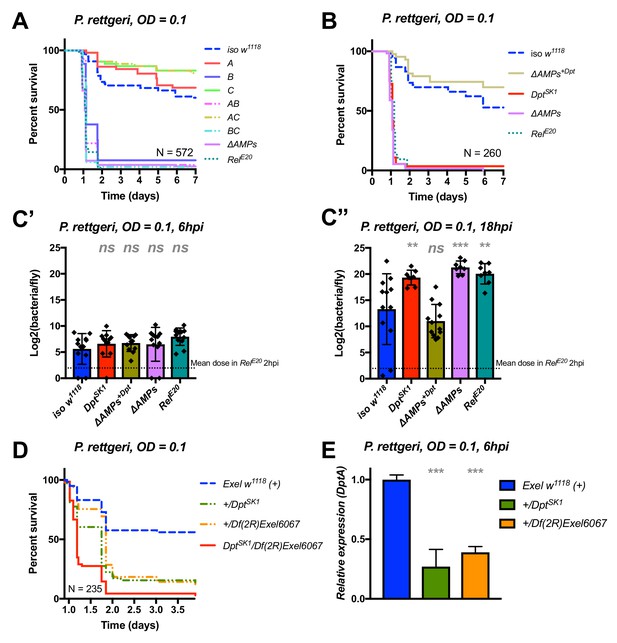
Identification of AMPs involved in the susceptibility of ΔAMPs flies to P. rettgeri.
(A) Survival of mutants for groups of AMPs reveals that only loss of Imd-responsive Group B peptides (Drosocin, Attacins, and Diptericins) recapitulates the susceptibility of ΔAMPs flies. (B) Further dissection of the mutations affected in Group B reveals that only the loss of Diptericins (DptSK1) leads to susceptibility similar to ΔAMPs flies. Remarkably, flies lacking all other AMPs (ΔAMPs+Dpt) resist as wild-type. (C) Bacterial loads of individual flies are similar at 6hpi (C’), but by 18hpi (C’’), Dpt mutants and RelE20 flies have all failed to control P. rettgeri growth. (D) Heterozygote flies for DptSK1 and a deficiency including the Diptericins and flanking genes (Df(2R)Exel6067) recapitulates the susceptibility of Diptericin mutants. Intriguingly, heterozygotes with one functional copy of the Diptericins (+/DptSK1 or +/Df(2R)Exel6067) are nonetheless highly susceptible to infection. (E) Diptericin A transcriptional output is strongly reduced in heterozygotes 6 hpi compared to wild-type flies. One-way ANOVA: not significant = ns, p<0.05 = *, p<0.01 = **, and p<0.001 = *** relative to iso w1118.
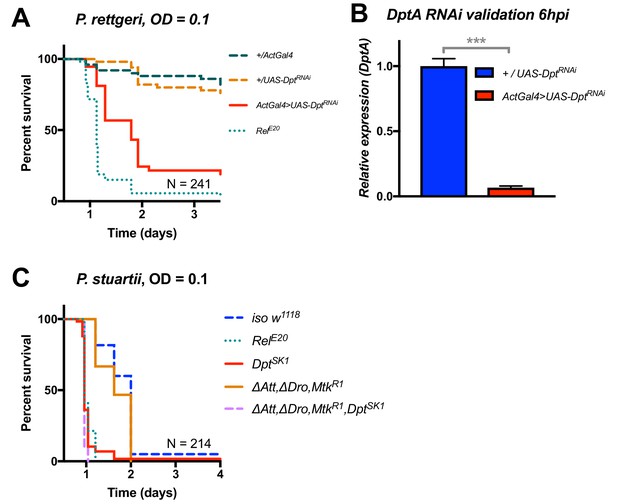
Additional validation of the role of Diptericin in resistance to Providencia.
(A) Silencing of Diptericin by RNAi leads to higher susceptibility to P. rettgeri infection (p<0.001). (B) Validation of the Diptericin RNAi construct 6 hpi. C) Mutants lacking multiple peptides (Attacins, Drosocin, and Metchnikowin) succumb to P. stuartii infection as wild-type (‘ΔAtt, ΔDro, MtkR1’), while Diptericin mutation alone (DptSK1) or combined (‘ΔAtt, ΔDro, MtkR1, DptSK1’) leads to a susceptibility similar to RelE20 mutants. This pattern of survival was similar to the pattern observed with P. rettgeri. One-way ANOVA: p<0.001 = ***.
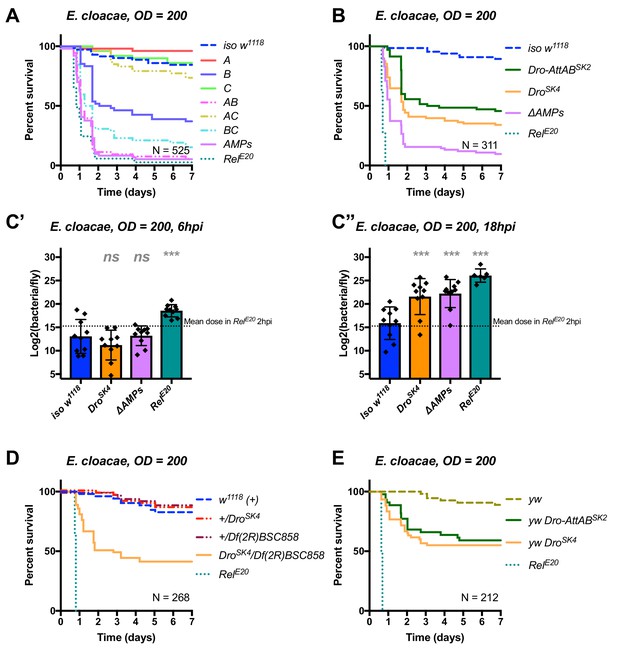
Identification of AMPs involved in the susceptibility of ΔAMPs flies to E.cloacae.
(A) Survival of mutants for groups of AMPs reveals that loss of Imd-responsive Group B peptides (Drosocin, Attacins, and Diptericins) results in a strong susceptibility to infection (p<0.001), while loss of Group A or C peptides alone resists as wild-type (p>0.1 each). Group AB flies were as susceptible as ΔAMPs flies, and we observed a synergistic interaction between Group A and B mutations (A*B: HR =+2.55, p=0.003). (B) Further dissection of the mutations in Group B revealed that loss of Drosocin alone (DroSK4), or a deficiency lacking both Drosocin and Attacins AttA and AttB (Dro-AttABSK2) recapitulates the susceptibility of Group B flies. (C) By 18hpi, bacterial loads in individual Drosocin mutants or RelE20 flies are significantly higher than wild-type. (D) Heterozygote flies for DroSK4 and Df(2R)BSC858 (a deficiency removing Drosocin, Attacins AttA and AttB, and other genes) are strongly susceptible to E. cloacae infection. (E) Drosocin mutants in an alternate genetic background (yw) are susceptible to E. cloacae. One-way ANOVA: not significant = ns, and p<0.001 = *** relative to iso w1118.
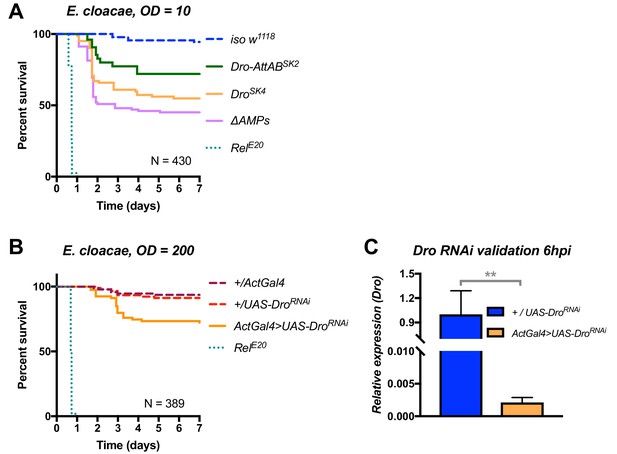
Additional validation of the role of Drosocin in defence against E. cloacae.
(A) Drosocin mutant susceptibility remains even at a lower dose (OD = 10,~7000 bacteria/fly), while RelE20 flies succumb rapidly regardless of initial dose. (B) Silencing of Drosocin by RNAi leads to significant mortality from E. cloacae infection (p<0.001). (C) Validation of the Drosocin RNAi construct 6hpi.
Additional files
-
Supplementary file 1
Primers used in this study.
- https://doi.org/10.7554/eLife.44341.016
-
Transparent reporting form
- https://doi.org/10.7554/eLife.44341.017





