Philosophy of Biology: The meanings of 'function' in biology and the problematic case of de novo gene emergence
Figures
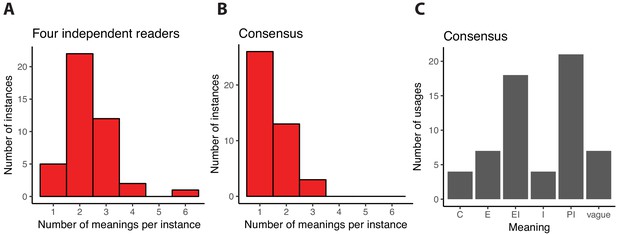
Interpreting the word function in scientific abstracts related to de novo gene birth.
We analyzed a sample of 20 abstracts containing 42 instances where the word function or one of its derivatives was used to describe DNA, RNA or protein objects. First, each of us read the abstracts independently and assigned one or several of the meanings of function as defined in the Pittsburgh model to each of these instances. The distribution of the number of distinct meanings that we assigned to the 42 instances is shown in panel (A). For only 5 instances did all of us independently assign the same unique meaning, suggesting that function is most often interpreted in multiple ways by independent readers. Next, we discussed each instance to see if we could reach consensus assignments based on the textual evidence. Consensus was built through conversations and agreement between the readers, rather than majority opinion. The distribution of the number of unique meanings assigned after consensus agreement to each of the 42 instances is shown in panel (B). Most (26/42) instances are now assigned to a single meaning. When more than one meaning remains, the readers agreed that the textual evidence supported multiple meanings except for one instance where consensus could not be reached and three meanings were assigned to reflect all the differing interpretations of our team members. In panel C, we show the number of times each of the five meanings of function defined in the Pittsburgh model is assigned to an instance of function.
-
Figure 1—source data 1
Independent and consensus assignments.
This table lists the results of our textual analyses for each instance of function.
- https://doi.org/10.7554/eLife.47014.004
Tables
The Pittsburgh model of function.
The hierarchical order of the meanings did not directly derive from our textual analysis, but was inspired from a reductionist interpretation of the flow of genetic information over time and space. It also reflects a possible ordering of the series of properties that must be acquired by a locus to undergo de novo gene birth.
| Meanings | Definitions |
|---|---|
| Evolutionary Implications | The object's influence on population dynamics over successive generations, as enabled by its physiological implications and their interplay with environmental pressures |
| Physiological Implications | The object's involvement in biological processes as enabled by a set of its capacities, interactions and expression patterns, independent of cross-generational considerations |
| Interactions | Physical contacts, direct or indirect, between the object under investigation and the other components of a system, including contacts that mediate chemical transformations |
| Capacities | Intrinsic physical properties of the object under investigation; the necessity of the object's behavior given an environment (eg., structural constraints) |
| Expression | The presence or amount of the object under investigation (RNA or protein object), or the presence or amount of its transcription or translation products (DNA object) |
| Vague | Sufficient evidence was not found to infer one or more meanings of function within this model, nor to derive a new meaning |
References for 20 abstracts analyzed in our study.
Countries (based on affiliations of all authors) and model organisms are included to display the diversity of the abstracts.
| Papers | Countries | Model Organisms |
|---|---|---|
| Keese, P. K., and Gibbs, A. (1992). Origins of genes: ‘big bang’ or continuous creation? PNAS 89:9489–9493. | Australia | Cellular life, Viruses |
| Kastenmayer, J. P., Ni, L., Chu, A., Kitchen, L. E., Au, W. C., Yang, H.,. .. and Basrai, M. A. (2006). Functional genomics of genes with small open reading frames (sORFs) in S. cerevisiae. Genome Research 16:365–373. | USA | S. cerevisiae |
| Levine, M. T., Jones, C. D., Kern, A. D., Lindfors, H. A., and Begun, D. J. (2006). Novel genes derived from noncoding DNA in Drosophila melanogaster are frequently X-linked and exhibit testis-biased expression. PNAS 103:9935–9939. | USA | D. melanogaster |
| Stepanov, V. G., and Fox, G. E. (2007). Stress-driven in vivo selection of a functional mini-gene from a randomized DNA library expressing combinatorial peptides in Escherichia coli. Molecular Biology and Evolution 24:1480–1491. | USA | E. coli |
| Cai, J., Zhao, R., Jiang, H., and Wang, W. (2008). De novo origination of a new protein-coding gene in Saccharomyces cerevisiae. Genetics 179:487–496. | China | S. cerevisiae |
| Zhou, Q., Zhang, G., Zhang, Y., Xu, S., Zhao, R., Zhan, Z.,. .. and Wang, W. (2008). On the origin of new genes in Drosophila. Genome Research 18:1446–1455. | China | Drosophila |
| Xiao, W., Liu, H., Li, Y., Li, X., Xu, C., Long, M., and Wang, S. (2009). A rice gene of de novo origin negatively regulates pathogen-induced defense response. PLoS One 4:e4603. | China, USA | rice |
| Carvunis, A. R., Rolland, T., Wapinski, I., Calderwood, M. A., Yildirim, M. A., Simonis, N.,. ..and Vidal M. (2012). Proto-genes and de novo gene birth. Nature 487:370–374. | Belgium, France, USA | S. cerevisiae |
| Ding, Y., Zhou, Q., and Wang, W. (2012). Origins of new genes and evolution of their novel functions. Annual Review of Ecology, Evolution, and Systematics 43:345–363. | China, USA | |
| Tautz, D., Neme, R., and Domazet-Lošo, T. (2013). Evolutionary Origin of Orphan Genes. In: Encyclopedia of Life Sciences. John Wiley & Sons. DOI: https://doi.org/10.1002/9780470015902.a0024601 | Croatia, Germany | |
| Reinhardt, J. A., Wanjiru, B. M., Brant, A. T., Saelao, P., Begun, D. J., and Jones, C. D. (2013). De novo ORFs in Drosophila are important to organismal fitness and evolved rapidly from previously non-coding sequences. PLoS Genetics 9:e1003860. | USA | D. melanogaster |
| Wissler, L., Gadau, J., Simola, D. F., Helmkampf, M., and Bornberg-Bauer, E. (2013). Mechanisms and dynamics of orphan gene emergence in insect genomes. Genome Biology and Evolution 5:439–455. | Germany, USA | Insects |
| Brylinski, M. (2013). Exploring the ‘dark matter’ of a mammalian proteome by protein structure and function modeling. Proteome Science 11:47. | USA | M. musculus |
| Li, D., Yan, Z., Lu, L., Jiang, H., and Wang, W. (2014). Pleiotropy of the de novo-originated gene MDF1. Scientific Reports 4:7280. | China | S. cerevisiae |
| Wirthlin, M., Lovell, P. V., Jarvis, E. D., and Mello, C. V. (2014). Comparative genomics reveals molecular features unique to the songbird lineage. BMC Genomics 15:1082. | USA | Songbirds |
| Suenaga, Y., Islam, S. R., Alagu, J., Kaneko, Y., Kato, M., Tanaka, Y.,. .. and Nakagawara, A.(2014). NCYM, a Cis-antisense gene of MYCN, encodes a de novo evolved protein that inhibits GSK3β resulting in the stabilization of MYCN in human neuroblastomas. PLoS Genetics 10:e1003996. | Japan | Human |
| Arendsee, Z. W., Li, L., and Wurtele, E. S. (2014). Coming of age: Orphan genes in plants. Trends in Plant Science 19:698–708. | USA | A. thaliana |
| Ruiz-Orera, J., Hernandez-Rodriguez, J., Chiva, C., Sabidó, E., Kondova, I., Bontrop, R.,. .. and Albà, M. M. (2015). Origins of de novo genes in human and chimpanzee. PLoS Genetics 11:e1005721. | Spain, The Netherlands | Human, Chimpanzee |
| Couso, J. P., and Patraquim, P. (2017). Classification and function of small open reading frames.Nature Reviews Molecular Cell Biology 18:575–589. | Spain, UK | D. melanogaster |
| Luis Villanueva-Cañas, J., Ruiz-Orera, J., Agea, M. I., Gallo, M., Andreu, D., and Albà, M. M. (2017). New genes and functional innovation in mammals. Genome Biology and Evolution 9:1886–1900. | Spain | Mammals |
Examples of each meaning of function as assigned to instances of usage.
Underlined portions of sentences serve as the contextual evidence used to assign the ‘code’, or meaning, to the bolded instances analyzed.
| Reference | Instance of function usage | Consensus meanings |
|---|---|---|
| Wirthin et al., 2014 | ‘Here we performed a comparative analysis of 48 avian genomes to identify genomic features that are unique to songbirds, as well as an initial assessment of function by investigating their tissue distribution and predicted protein domain structure.’ | Expression, Capacities |
| Brylinski, 2013 | A subsequent structure-based function annotation of small protein models exposes 178,745 putative protein-protein interactions with the remaining gene products in the mouse proteome, 1,100 potential binding sites for small organic molecules and 987 metal-binding signatures. | Interaction |
| Li et al., 2014 | ‘Therefore, MDF1 functions in two important molecular pathways, mating and fermentation, and mediates the crosstalk between reproduction and vegetative growth.’ | Physiological Implications |
| Ruiz-Orera et al., 2015 | ‘In general, these transcripts show little evidence of purifying selection, suggesting that many of them are not functional’ | Evolutionary Implications |
Additional files
-
Transparent reporting form
- https://doi.org/10.7554/eLife.47014.007





