B cell receptor and Toll-like receptor signaling coordinate to control distinct B-1 responses to both self and the microbiota
Figures
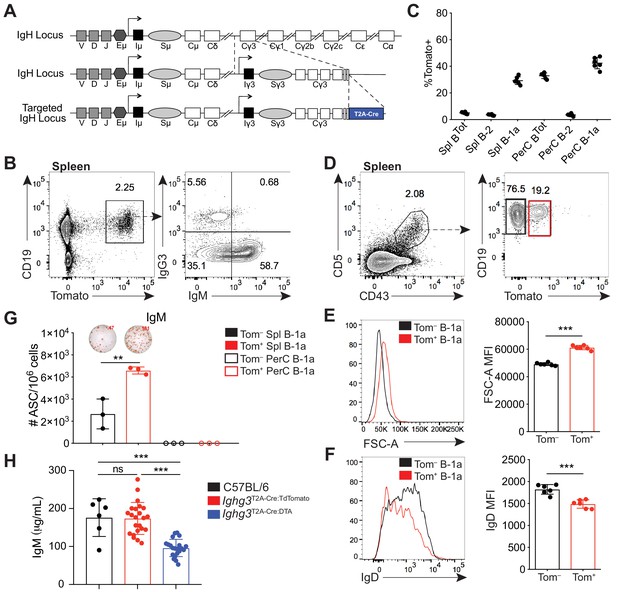
A reporter mouse marks activated B-1a cells.
(A) Schematic of targeted insertion of T2A-Cre into the Ighg3 (Iγ3) heavy chain locus to generate the Ighg3T2A-Cre reporter mouse. (B) Representative flow cytometry plot of IgG3 and IgM expression on pregated CD19+Tomato+ splenocytes from 6 wk old Ighg3T2A-Cre mice crossed to Rosa26STOP-flox-TdTomato mice (Ighg3T2A-Cre:TdTomato). (C) Percentage Tomato expression in B cell subsets from the peritoneal cavity (PerC) and spleen (Spl) of 6 wk old Ighg3T2A-Cre:TdTomato mice by flow cytometry. Total B cells defined as CD19+; B-2 cells defined as CD19+CD23+CD43–CD5–; B-1a cells defined as CD19+CD23–CD43+CD5+. (D) Representative flow cytometry gating of Tomato– and Tomato+ splenic B-1a cells from 3 wk old Ighg3T2A-Cre:TdTomato mice, pregated on Live/CD19+CD23– cells. (E) Representative histogram and quantification of FSC-A and (F) IgD expression on pre-gated Tomato– (black) and Tomato+ (red) splenic B-1a cells. (G) Number of IgM+ antibody secreting cells (ASCs)/106 present in purified Tomato– (black) or Tomato+ (red) splenic (closed circles) or peritoneal cavity (PerC) (open circles) B-1a cells from 6 wk old Ighg3T2A-Cre:TdTomato mice, as measured by ELISpot. (H) Serum IgM titers of 7 wk old C57BL/6 (black), Ighg3T2A-Cre:TdTomato (red) and Ighg3T2A-Cre mice crossed to Rosa26STOP-flox-DTA (Ighg3T2A-Cre:DTA) (blue) mice. Error bars indicate the mean (± SEM). *p<0.05, **p<0.01, and ***p<0.001 (unpaired two-tailed Student's t-test). Each data point represents an individual mouse (C, E-H). Data are representative of at least three independent experiments (B-G) or pooled from five independent experiments (H).
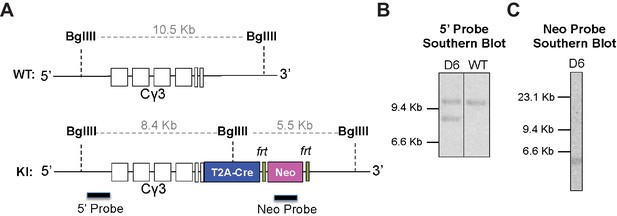
Validation of correct targeting of knock-in mouse.
(A) Southern blot strategy to identify targeted insertion of T2A-Cre after the last transmembrane exon of Ighg3 (Iγ3) using DNA probes 5’ of Ighg3 (5‘ probe) and to the neomycin-resistance gene (Neo probe). (B) Southern blot of BglII restriction-digested ES cell DNA from clone D6, which was used to generate the Ighg3T2A-Cre mouse using a 5’ probe to detect 10.5 Kb band corresponding to WT DNA and a 8.4 Kb band corresponding to targeted insertion of T2A-Cre. (C) Southern blot of BglII restriction-digested ES cell DNA from clone D6 using a Neo probe to detect a single 5.5 Kb band indicating a single copy, targeted insertion. Data are representative of at least two independent experiments.
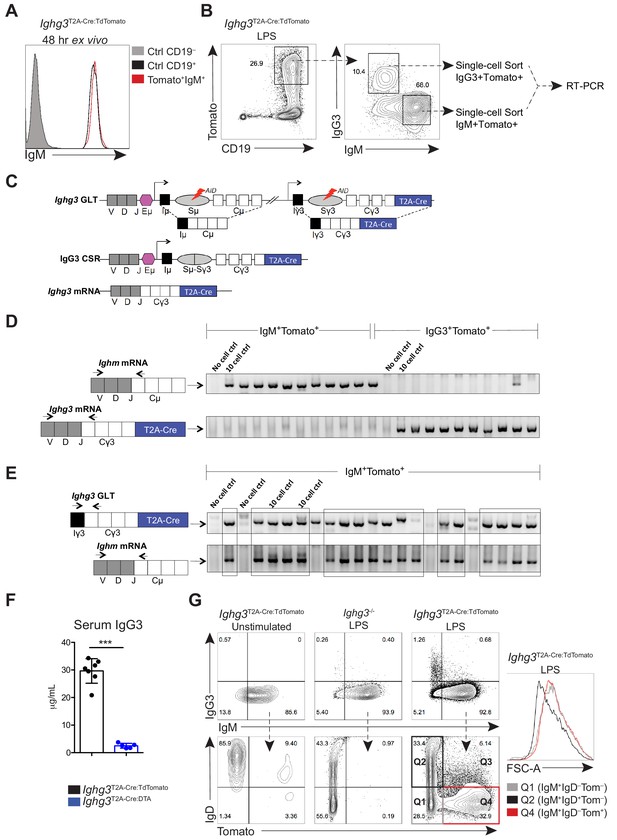
TdTomato expression in IgM+ cells correlates with Ighg3 germ-line transcription in Ighg3 reporter mouse.
(A) IgM expression on sorted IgM+Tomato+ splenocytes (red histogram) compared to Tomato–CD19+ splenocytes (black histogram) after 48hr ex vivo culture, as measured by flow cytometry. (B) Splenocytes from 3 wk old Ighg3T2A-Cre:TdTomato mice were stimulated with LPS for 72 hr, followed by single-cell sorting and subsequent RT-PCR. (C) Schematic of murine heavy chain locus depicting the generation of Ighg3 (Iγ3) germ-line transcript (GLT) prior to AID-mediated class switch recombination from IgM to IgG3. (D) RT-PCR of single-cell sorted IgG3–IgM+Tomato+ or IgG3+IgM–Tomato+ cells, as described in (B), for Ighm mRNA and Ighg3 mRNA, visualized by agarose gel electrophoresis. Arrows indicate primer binding sites. (E) Single-cell RT-PCR of Ighg3 germ-line transcript (GLT) and Ighm mRNA of IgG3–IgM+Tomato+ as described in (B), visualized by agarose gel electrophoresis. Arrows indicate primer binding sites. (F) Serum IgG3 titers of 7 wk old Ighg3T2A-Cre:TdTomato (black) and Ighg3T2A-Cre mice crossed to Rosa26STOP-flox-DTA (Ighg3T2A-Cre:DTA) (blue) mice.(G) Expression of IgM versus IgG3 protein on 3-day in vitro LPS-stimulated splenocytes from Ighg3T2A-Cre:TdTomato and Ighg3-/- mice (top panel), as measured by flow cytometry. IgD and Tomato expression on pregated IgM+ in vitro stimulated B cells (bottom panel). FSC-A of pregated IgM+IgD+Tomato– (gray histogram), IgM+IgD+Tomato– (black histogram), and IgM+IgD–Tomato+ (red histogram) LPS-stimulated Ighg3T2A-Cre:TdTomato splenocytes. Error bars indicate the mean (± SEM). *p<0.05, **p<0.01, and ***p<0.001 (unpaired two-tailed Student's t-test). Data are representative of at least two independent experiments.

Single-cell RT-PCR Primers.
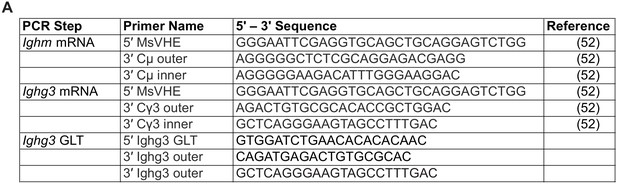
(A) Table summarizing the forward (5’) and reverse (3’) primers names and sequences for single-cell semi-nested RT-PCR of Ighm mRNA, Ighg3 mRNA, and Ighg3 germ-line transcript (GLT).
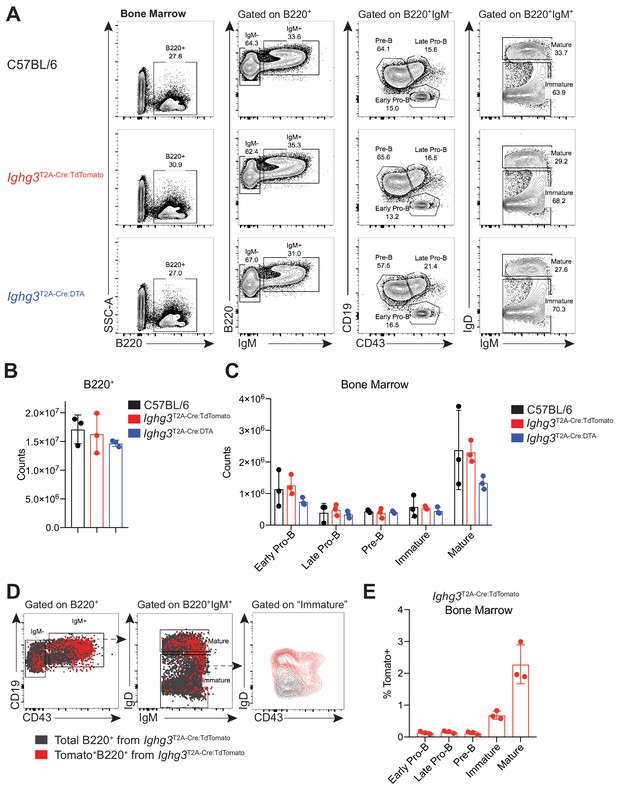
B cell development in bone marrow is unaltered in reporter mouse.
(A) Representative flow cyometry gating of B cell subsets in the bone marrow of 7 wk old C57BL/6 (black), Ighg3T2A-Cre:TdTomato (red), and Ighg3T2A-Cre:DTA (blue) mice. Pre-B, Late Pro-B and, Early Pro-B cells were pregated as B220+IgM– and identified according to their expression of CD19 and CD43. Mature and Immature B cells were pregated as B220+IgM+ and identified according to their expression of IgM and IgD. (B) Absolute counts of B220+ cells in the bone marrow from one femur in 7 wk old C57BL/6 (black), Ighg3T2A-Cre:TdTomato (red), and Ighg3T2A-Cre:DTA (blue) mice. (C) Absolute counts of Early Pro-B, Late Pro-B, Pre-B, Immature, and Mature B cells in the bone marrow from one femur in 7 wk old C57BL/6 (black), Ighg3T2A-Cre:TdTomato (red), and Ighg3T2A-Cre:DTA (blue) mice. (D) Representative flow cytometry dot plot showing the expression of CD19 and CD43 on pregated B220+ bone marrow cells (gray) overlayed with pregated Tomato+B220+ bone marrow cells (red) from Ighg3T2A-Cre:TdTomato mice (left); representative flow cytometry dot plot showing the expression of IgD and IgM on pregated B220+IgM+ bone marrow cells (gray) overlayed with pregated Tomato+B220+IgM+ bone marrow cells (red) from Ighg3T2A-Cre:TdTomato mice (middle); representative flow cytometry dot plot showing the expression of IgD and CD43 on pregated ‘immature’ B220+IgM+IgD– bone marrow cells (gray) overlayed with pregated ‘immature’ Tomato+B220+IgM+IgD– bone marrow cells (red) from Ighg3T2A-Cre:TdTomato mice (right). (E) Percentage Tomato expression in various bone marrow B cell subsets in 7 wk old Ighg3T2A-Cre:TdTomato mice. Each data point represents an individual mouse. Data are representative of two independent experiments.
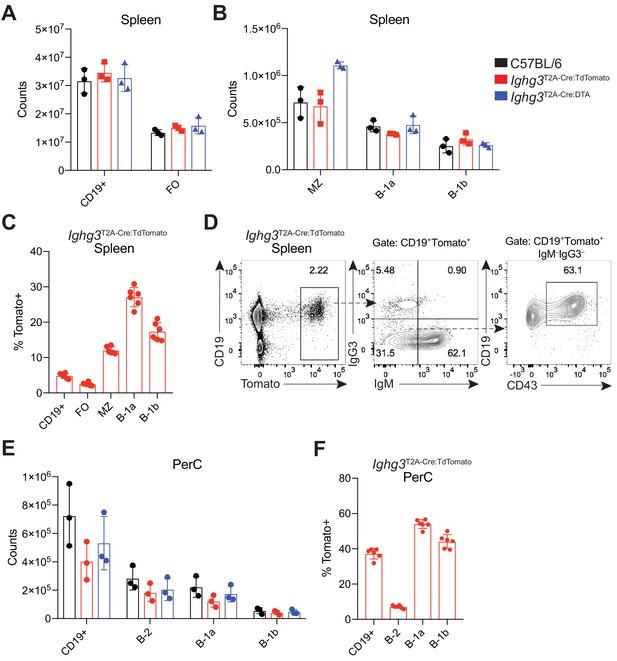
Characterization of B cell subsets in the spleen and peritoneal cavity of reporter mice.
(A) Absolute counts of CD19+ and CD19+CD23+ Follicular (FO) B cells from the spleens of 7 wk old C57BL/6 (black), Ighg3T2A-Cre:TdTomato (red), and Ighg3T2A-Cre:DTA (blue) mice. (B) Absolute counts of marginal zone B (MZ) (CD19+CD23–CD21+), B-1a (CD19+CD23–CD23–CD43+CD5+), and B-1b (CD19+CD23–CD23–CD43+CD5–) cells in the spleens of 7 wk old C57BL/6 (black), Ighg3T2A-Cre:TdTomato (red), or Ighg3T2A-Cre:DTA (blue) mice. (C) Percentage Tomato expression in various B cell subsets, as described in (B), in the spleens of 7 wk old Ighg3T2A-Cre:TdTomato mice. (D) Representative flow cytometry plot characterizing expression of CD43 and CD19 on pregated Tomato+CD19+IgM–IgG3– splenocytes from a 7 wk old Ighg3T2A-Cre:TdTomato mice. (E) Absolute counts of CD19+, B-1a (CD19+CD23–CD23–CD43+CD5+), and B-1b (CD19+CD23–CD23–CD43+CD5–) cells in the peritoneal cavities (PerC) of 7 wk old C57BL/6 (black), Ighg3T2A-Cre:TdTomato (red), or Ighg3T2A-Cre:DTA (blue) mice. (F) Percentage Tomato expression in various B cell subsetsm as described in (E), in the peritoneal cavity (PerC) of 7 wk old Ighg3T2A-Cre:TdTomato mice. Each data point represents an individual mouse. Data are representative of two (A, B, E) or at 5 (C, D, F) independent experiments.
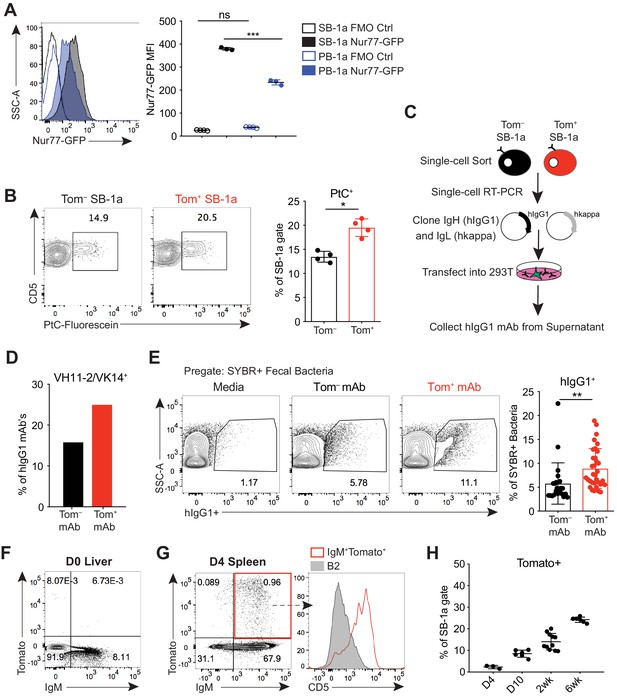
B-1a cells have a history of B cell receptor-mediated activation.
(A) Representative flow cytometry histogram (left) and quantification of mean fluorescence intensity (right) of GFP expression on splenic (SB-1a) (black) and peritoneal cavity (PB-1a) (blue) B-1a cells (CD19+CD23–CD43+CD5+) in 3 wk old Nur77-GFP reporter mice (filled histogram, closed circles) or reporter-negative mice as controls (FMO Ctrl) (unfilled histogram, open circles). (B) Representative flow cytometry plots (left) and quantification (right) of the percentage of Tomato– and Tomato+ splenic B-1a cells from 6 wk old Ighg3T2A-Cre:TdTomato mice stained with fluorescein-labeled phosphatidylcholine liposomes. (C) Schematic showing the generation of hIgG1 recombinant monoclonal antibodies (mAb) from single-cell sorted Tomato– or Tomato+ splenic B-1a cells (SB-1a) from 6wk old Ighg3T2A-Cre:TdTomato mice. There are two source files associated with this figure with a comprehensive description of all of the mAbs generated. (D) The percentage VH11-2/VK14 gene usage in monoclonal antibodies (mAbs) generated from Tomato– (black) and Tomato+ (red) splenic B-1a (SB-1a) cells from 6 wk old Ighg3T2A-Cre:TdTomato mice, as described in (C); Tom+ mAb n= 48; Tom– mAb n= 37. (E) Representative flow cyometry plots (left) and quantification (right) of pregated SYBR+ fecal bacteria bound by recombinant hIgG1 monoclonal antibodies generated as described in (C); PtC-reactive VH11-2/VK14 expressing mAb’s were excluded from this analysis; Tom+ mAb n= 32; Tom– mAb n= 23. (F) Representative flow cytometry plot of IgM versus Tomato expression in pre-gated CD19+ D0 liver cells or (G) CD19+ D4 spleen cells from Ighg3T2A-Cre:TdTomato mice; representative flow cytometry histogram of CD5 expression on pre-gated CD19+IgM+Tomato+ D4 spleen cells (red, unfilled) compared to CD19+IgM+CD23+CD43–CD5– B2 cells (gray, filled) (G, right). (H) Quantification of percentage Tomato expression in B-1a cells from the spleens (SB-1a) of D4, D10, 2 wk, and 6 wk old Ighg3T2A-Cre:TdTomato mice by flow cytometry. Error bars indicate the mean (± SEM). *p<0.05, **p<0.01, and ***p<0.001 (unpaired two-tailed Student's t-test (B, E) or one-way ANOVA (A)). Each data point represents an individual mouse (A,B,H) or monoclonal antibody (E). Data are representative of at least three independent experiments (A-B, E-H).
-
Figure 2—source data 1
Summary of monoclonal antibodies generated from Tomato– spleen B-1a cells.
The variable heavy chain gene (VH gene), joining heavy chain gene (JH), heavy chain CDR3 peptide sequence, variable kappa chain gene (VH gene), joining kappa chain gene (JH gene), and kappa chain CDR3 peptide sequence for monoclonal antibodies generated from single-cell sorted Tomato– splenic B-1a cells from 6 wk old Ighg3T2A-Cre:TdTomato mice (n = 38).
- https://doi.org/10.7554/eLife.47015.009
-
Figure 2—source data 2
Summary of monoclonal antibodies generated from Tomato+ spleen B-1a cells.
The variable heavy chain gene (VH gene), joining heavy chain gene (JH), heavy chain CDR3 peptide sequence, variable kappa chain gene (VH gene), joining kappa chain gene (JH gene), and kappa chain CDR3 peptide sequence for monoclonal antibodies generated from single-cell sorted Tomato+ splenic B-1a cells from 6 wk old Ighg3T2A-Cre:TdTomato mice (n = 48).
- https://doi.org/10.7554/eLife.47015.010
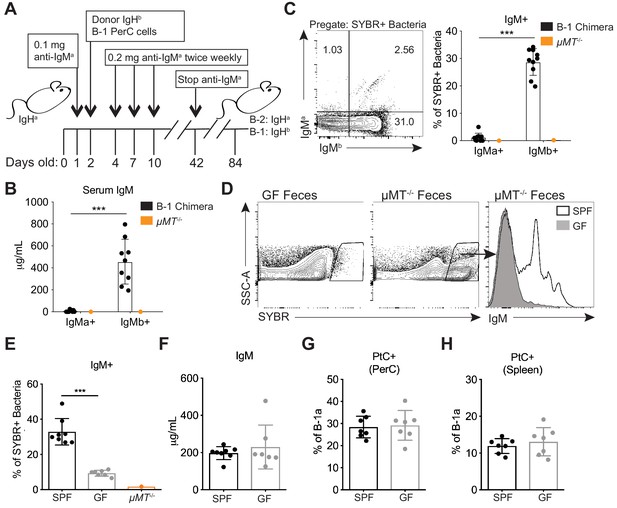
Colonization of a microbiota is required for B-1 derived microbiota-reactive IgM.
(A) Schematic showing the generation of B-1 disparate allotype chimera mice. (B) Serum titers of B-2-derived IgMa and B-1-derived IgMb in 14 wk old B-1 chimeras, as described in (A). (C) Representative flow cytometry plot (left) and quantification (right) of percentage of SYBR+ fecal bacteria bound by B-1-derived serum IgMb and B-2-derived serum IgMa from 14 wk old B-1 chimeras, as described in (A); μMT-/- mouse serum included as a negative control (orange). (D) Representative SYBR staining of germ-free (GF) (left) or SPF μMT-/- (middle) feces, as measured by flow cytometry. Representative flow cytometry histogram plot showing IgM staining of SYBR+μMT-/- fecal bacteria with serum from 7 wk old SPF (black line, no fill) or germ free (GF) mice (gray line, filled) (right). (E) Quantification of percentage of SYBR+μMT-/- fecal bacteria bound by serum IgM from seven wk old SPF (black), GF (gray), or μMT-/- control (orange) mice (right). (F) Serum titers of IgM in 7 wk old SPF (black) or GF (gray) mice, as measured by ELISA. (G-H) Percentage fluorescein-labeled PtC-liposome positive (PtC+) B-1a cells (CD19+CD23–CD43+CD5+) in the (G) peritoneal cavity (PerC) and (H) spleen of 7 wk old SPF (black) or GF (gray) mice, as measured by flow cytometry. Each data point represents an individual mouse (B-C, E-H). Error bars indicate the mean (± SEM). *p<0.05, **p<0.01, and ***p<0.001 (unpaired two-tailed Student's t-test). Data are representative of at least three independent experiments (B-H).
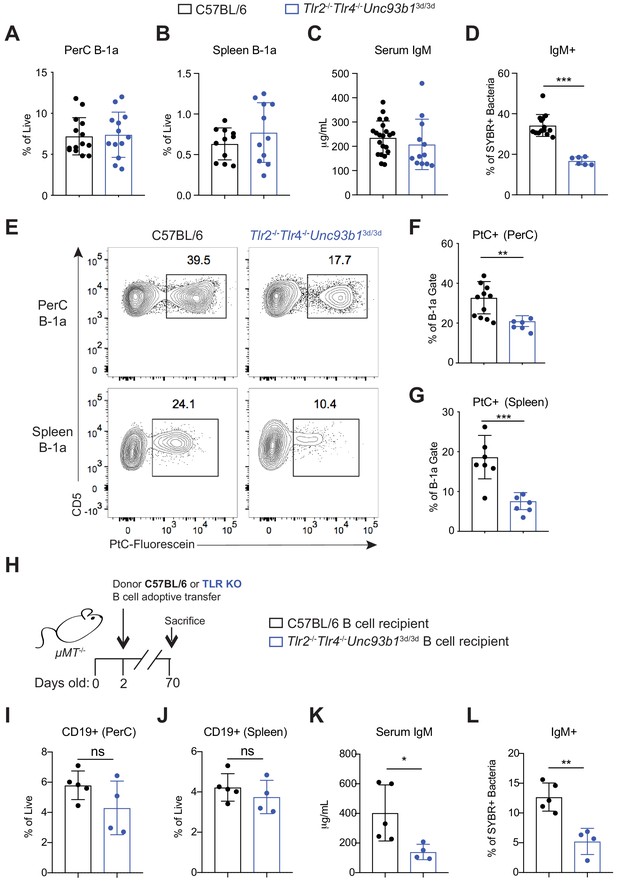
Loss of Toll-like receptor signaling results in reduced B-1a responses to both phosphatidylcholine and the microbiota.
(A) Frequency of live B-1a cells in 7 wk old WT (black) or Tlr2-/-Tlr4-/-Unc93b13d/3d (blue) mice in the peritoneal cavity (PerC) and (B) spleen, as measured by flow cytometry. B-1a cells defined as CD19+CD23–CD43+CD5+. (C) Serum IgM titers in 7 wk old WT (black) or Tlr2-/-Tlr4-/-Unc93b13d/3d (blue) mice, as measured by ELISA. (D) Percentage of SYBR+μMT-/- fecal bacteria bound by serum IgM from WT (black) or Tlr2-/-Tlr4-/-Unc93b13d/3d (blue) mice, as measured by flow cytometry. (E) Representative flow cytometry plot and quantification (F-G) of percentage of B-1a cells in the (F) peritoneal cavity (PerC) and (G) spleen bound by fluorescein-labeled phosphatidylcholine liposomes in 7 wk old WT (black) or Tlr2-/-Tlr4-/-Unc93b13d/3d (blue) mice. (H) Schematic showing adoptive transfer of CD19+ pooled bone marrow and spleen cells (B cells) from either C57BL/6 (WT) (black) or Tlr2-/-Tlr4-/-Unc93b13d/3d (blue) into B cell deficient μMT-/- recipeint mice at 2 days of age and sacrificed at 10 weeks of age. (I-J) Frequency of live B-1a cells in 7 wk old WT (black) or Tlr2-/-Tlr4-/-Unc93b13d/3d (blue) B cell recipient mice in the (J) peritoneal cavity (PerC) and (J) spleen, as measured by flow cytometry. (K) Serum IgM titers in 7 wk old WT (black) or Tlr2-/-Tlr4-/-Unc93b13d/3d (blue) B cell recipient mice, as measured by ELISA. (L) Percentage of SYBR+ fecal bacteria bound by serum IgM from WT (black) or Tlr2-/-Tlr4-/-Unc93b13d/3d (blue) B cell recipient mice, as measured by flow cytometry. Error bars indicate the mean (± SEM). *p<0.05, **p<0.01, and ***p<0.001 (unpaired two-tailed Student's t-test). Each data point represents an individual mouse. Data are pooled from at least three independent experiments (A-G; I-L).
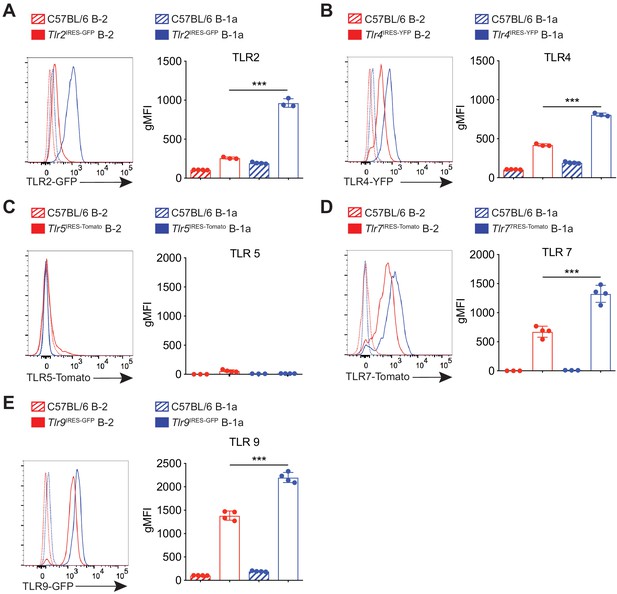
TLR expression on B cell subsets using TLR reporter mice.
(A–E) Expression of TLR2, TLR4, TLR5, TLR7, and TLR9 on peritoneal cavity B-2 cells (CD19+CD23+) and B-1a cells (CD19+CD23–CD43+CD5+) using Tlr2IRES-GFP, Tlr4IRES-YFP, Tlr5IRES-TdTomato, Tlr7IRES-TdTomato, Tlr9IRES-GFP reporter mice, respectively, or C57BL/6 mice as controls. Each data point represents an individual mouse. Error bars indicate the mean (± SEM). *p<0.05, **p<0.01, and ***p<0.001 (unpaired two-tailed Student's t-test). Data are representative of at least two experiments.
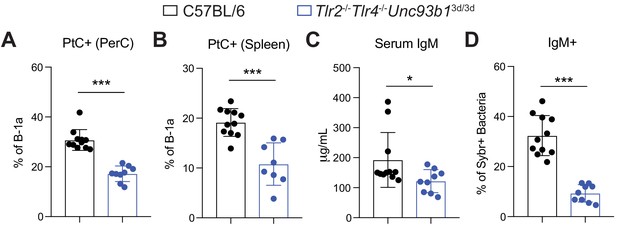
Loss of Toll-like receptor signaling results in reduced B-1a responses to both phosphatidylcholine and the microbiota in co-housed mice.
WT (black) and Tlr2-/-Tlr4-/-Unc93b13d/3d (blue) mice were cohoused for 4 weeks post weaning to normalize microbiota between mice. (A-B) Percentage of B-1a cells bound by fluorescein-labeled phosphatidylcholine liposomes (PtC+) in the (A) peritoneal cavity (PerC) and (B) spleen in 7 wk old WT (black) or Tlr2-/-Tlr4-/-Unc93b13d/3d (blue) mice, as measured by flow cytometry. B-1a cells defined as CD19+CD23–CD43+CD5+. (C) Serum IgM titers in 7 wk old WT (black) or Tlr2-/-Tlr4-/-Unc93b13d/3d (blue) mice, as measured by ELISA. (D) Percentage of SYBR+ fecal bacteria bound by serum IgM from WT (black) or Tlr2-/-Tlr4-/-Unc93b13d/3d (blue) mice, as measured by flow cytometry. Error bars indicate the mean (± SEM). *p<0.05, **p<0.01, and ***p<0.001 (unpaired two-tailed Student's t-test). Each data point represents an individual mouse. Data are pooled from four independent experiments.
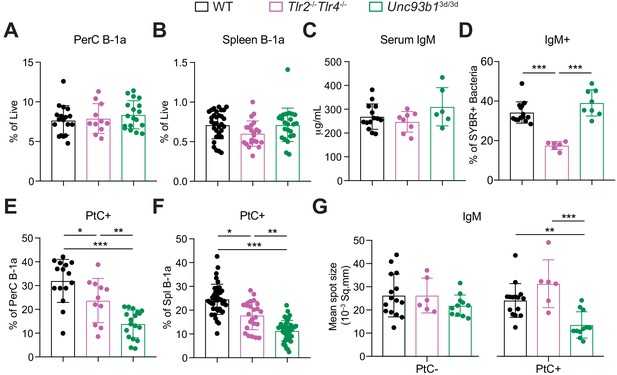
TLR2 and TLR4 regulate B-1a responses to the microbiota, whereas Unc93B1-dependent TLRs regulate phosphatidylcholine-reactive B-1a responses.
(A–B) Frequency of live B-1a cells in 7 wk old WT (black), Tlr2-/-Tlr4-/- (pink), or Unc93b13d/3d (green) mice in the (A) peritoneal cavity (PerC) and (B) spleen, as measured by flow cytometry. (C) Serum IgM titers in 7 wk old WT (black), Tlr2-/-Tlr4-/- (pink), or Unc93b13d/3d (green) mice, as measured by ELISA. (D) Percentage of SYBR+μMT-/- fecal bacteria bound by serum IgM from 7 wk old WT (black), Tlr2-/-Tlr4-/- (pink), or Unc93b13d/3d (green) mice, as measured by flow cytometry. (E-F) Percentage fluorescein-labeled phosphatidylcholine-liposome positive (PtC+) (E) peritoneal cavity (PerC) B-1a and (F) spleen (Spl) B-1a cells in 7 wk old WT (black), Tlr2-/-Tlr4-/- (pink), or Unc93b13d/3d (green) mice, as measured by flow cytometry. (G) Quantification of the mean spot size of IgM secreted by sorted PtC– (left) or PtC+ (right) splenic B-1a cells from WT (black), Tlr2-/-Tlr4-/- (pink), or Unc93b13d/3d (green) mice after 20 hr in culture, as measured by ELISpot. Error bars indicate the mean (± SEM). *p<0.05, **p<0.01, and ***p<0.001 (One-way ANOVA). Each data point represents an individual mouse. Data are pooled from at least three independent experiments (A-G).
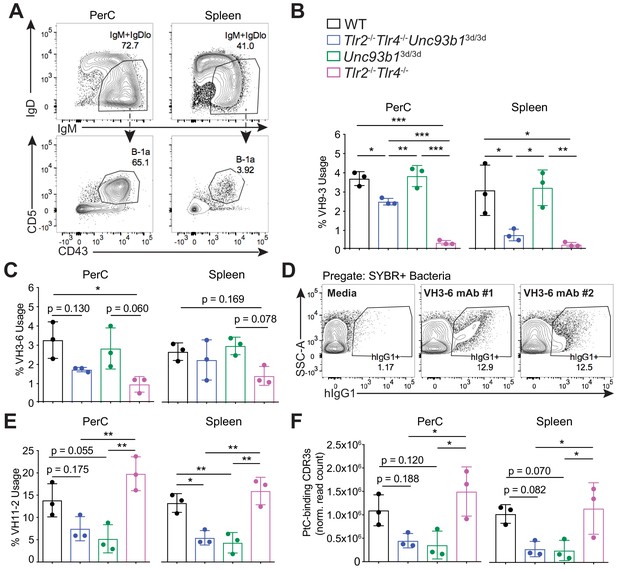
B-1a immunoglobulin repertoire analysis reveals unique regulation of a subset of heavy chain genes by distinct subsets of TLRs.
(A) Representative flow cytometry gating strategy for sorting IgM+IgDloCD43+CD5+B-1a cells (pregated as CD19+/DAPI–) in the peritoneal cavity (PerC) (left) and spleen (right). Prior to sorting, splenocytes were depleted of CD3+CD4+CD8+F4/80+NK1.1+GR-1+ cells using biotinylated antibodies and streptavidin magnetic beads. (B-C) The percentage of heavy chain CDR3 nucleotide sequencing reads expressing the germline (B) VH9-3 allele or the (C) VH3-6 allele in peritoneal cavity (PerC) (left) and spleen (right) B-1a samples (% usage). (D) Percentage of pre-gated SYBR+ fecal bacteria bound by two individual recombinant hIgG1 monoclonal antibodies (#1 and #2) expressing the germline VH3-6 allele generated from splenic B-1a cells from 6 wk old mice, described in Figure 2C, as measured by flow cytometry. (E) The percentage of heavy chain CDR3 nucleotide sequencing reads expressing the germline VH11-2 allele in peritoneal cavity (PerC) (left) and spleen (right) B-1a samples (% usage). (F) The combined normalized read counts of PtC-binding CDR3 peptide sequences (MRYSNYWYFDV, MRYGSSYWYFDV, and MRYGNYWYFDV) in peritoneal cavity (PerC) (left) and spleen (right) B-1a samples. Black circles represent WT mice, blue circles represent Tlr2-/-Tlr4-/-Unc93b13d/3d mice, green circles represent Unc93b13d/3d mice, and pink circles represent Tlr2-/-Tlr4-/- mice (B, C, E, F). CDR3 frequencies were artificially scaled to 10 million reads to account for differences in read depth among samples (F). Error bars indicate the mean (± SEM). *p<0.05, **p<0.01, and ***p<0.001 (One-way ANOVA). Each data point represents an individual mouse (B, C, E, F). Data representative of 3 independent experiments (D). There are three source files associated with this figure.
-
Figure 6—source data 1
Top 10 highly recurring CDR3 sequences (peptide and V(D)J recombination) detected in peritoneal cavity B-1a samples from age-matched mice lacking different subsets of TLRs.
The CDR3 peptide sequence, variable heavy chain gene, joining heavy chain gene, total read counts (copy), normalized read counts (norm.copy), and the sum of normalized counts of PtC-binding CDR3 peptide sequences in peritoneal cavity (PerC) B-1a samples from 7 wk old WT (black), Tlr2-/-Tlr4-/-Unc93b13d/3d (blue), Tlr2-/-Tlr4-/- (pink), and Unc93b13d/3d (green) mice. PtC-binding CDR3 peptide sequences include MRYSNYWYFDV (highlighted in blue), MRYGSSYWYFDV (highlighted in orange), and MRYGNYWYFDV (highlighted in purple). CDR3 sequences in italic font denotes that they did not appear in the top 10 CDR3 sequences for that sample, but were included to be able to determine the sum of PtC-binding CDR3 sequencing reads. Sequencing reads were normalized by artificially scaling to 10 million reads to account for differences in read depth among samples. There are three biological replicates for each genotype in one experiment.
- https://doi.org/10.7554/eLife.47015.019
-
Figure 6—source data 2
Top 10 highly recurring CDR3 sequences (peptide and V(D)J recombination) detected in spleen B-1a samples from age-matched mice lacking different subsets of TLRs.
The CDR3 peptide sequence, variable heavy chain gene, joining heavy chain gene, total read counts (copy), normalized read counts (norm.copy), and the sum of normalized counts of PtC-binding CDR3 peptide sequences in spleen (Spl) B-1a samples from 7 wk old WT (black), Tlr2-/-Tlr4-/-Unc93b13d/3d (blue), Tlr2-/-Tlr4-/- (pink), and Unc93b13d/3d (green) mice. PtC-binding CDR3 peptide sequences include MRYSNYWYFDV (highlighted in blue), MRYGSSYWYFDV (highlighted in orange), and MRYGNYWYFDV (highlighted in purple). CDR3 sequences in italic font denotes that they did not appear in the top 10 CDR3 sequences for that sample, but were included to be able to determine the sum of PtC-binding CDR3 sequencing reads. Sequencing reads were normalized by artificially scaling to 10 million reads to account for differences in read depth among samples. There are three biological replicates for each genotype in one experiment.
- https://doi.org/10.7554/eLife.47015.020
-
Figure 6—source data 3
Variable heavy chain gene usage summary in spleen and peritoneal cavity B-1a cells in mice lacking different subsets of TLRs.
Table summarizing the variable heavy chain gene usage frequency in peritoneal cavity and spleen B-1a samples from 7 wk old WT (black), Tlr2-/-Tlr4-/-Unc93b13d/3d (blue), Tlr2-/-Tlr4-/- (pink), and Unc93b13d/3d (green) mice. There are three biological replicates for each genotype in one experiment.
- https://doi.org/10.7554/eLife.47015.021
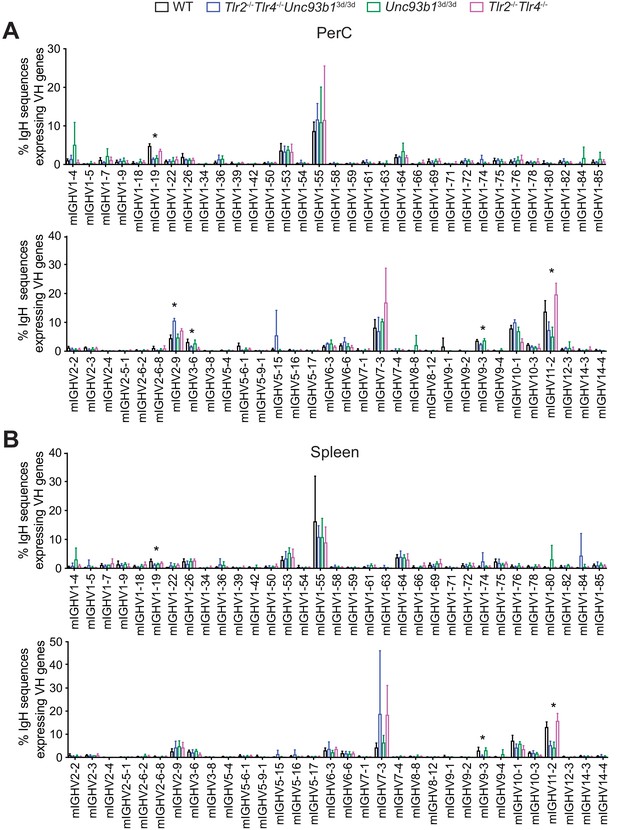
VH gene-usage distribution in splenic and peritoneal cavity B-1a samples.
(A–B) The percentage of heavy chain CDR3 nucleotide sequencing reads expressing different germline V-alleles in (A) peritoneal cavity (PerC) and (B) spleen B-1a samples from 7 wk old WT (black), Tlr2-/-Tlr4-/-Unc93b13d/3d (blue), Tlr2-/-Tlr4-/- (pink), or Unc93b13d/3d (green) mice. Each data point represents an individual mouse. Error bars indicate the mean (± SEM). *p<0.05, **p<0.01, and ***p<0.001 (One-way ANOVA). Asterisks denote a statistical difference in a unique Ig VH gene between at least two different genotypes.
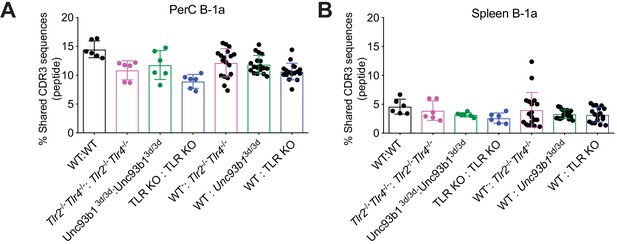
There is considerable variation in B-1a IgH CDR3 repertoire between different mice, independent of TLR signaling.
(A–B) CDR3 peptide pair-wise sharing analysis of IgH repertoire similarity between (A) peritoneal cavity (PerC) and (B) spleen B-1a samples from 7 wk old WT (black), Tlr2-/-Tlr4-/- (pink), Unc93b13d/3d (green), or TLR KO (blue) mice. CDR3 peptide pair-wise analysis was conducted between WT mice (WT/WT), Tlr2-/-Tlr4-/- mice (Tlr2-/-Tlr4-/-/Tlr2-/-Tlr4-/-), Unc93b13d/3d mice (Unc93b13d/3d/Unc93b13d/3d), TLR KO mice (TLR KO/TLR KO) and between WT vs Tlr2-/-Tlr4-/- mice (WT/Tlr2-/-Tlr4-/-), WT vs. Unc93b13d/3d mice (WT/Unc93b13d/3d), and WT vs TLR KO mice (WT/TLR KO). TLR KO mice refers to Tlr2-/-Tlr4-/-Unc93b13d/3d mice. Each dot represents the percentage of shared CDR3 peptide sequences between two mice. A total of 3 mice per genotype were included in the analysis, yielding six pair-wise comparisons within the same genotype ((% shared CDR3 in WT1 and WT2)/(Total CDR3 sequences in WT 1), (% shared CDR3 in WT1 and WT2)/(Total CDR3 sequences in WT 2), etc.) and 18 pair-wise comparisons when comparing two different genotypes. Data are presented as percentage of total CDR3 nucleotide sequencing reads. CDR3 frequencies were artificially scaled to 10 million reads to account for differences in read depth among samples, and a CDR3 frequency of 1000 was set as the minimum cutoff for pairwise-analysis. There are three source files associated with this figure.
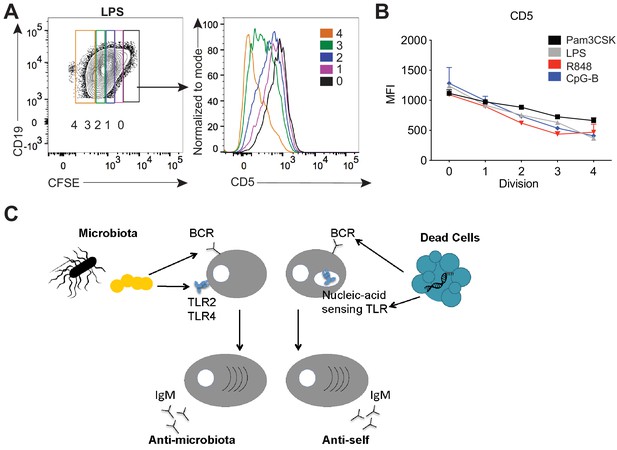
TLR stimulation results in CD5 downregulation on B-1a cells.
(A) Representative flow cytometry histogram plot and quantification of mean fluorescence intensity of CD5 expression (B) of in vitro Pam3CSK (black), LPS- (gray), R848- (red) and CpG-B- (blue) stimulated peritoneal cavity B-1a cells at 0, 1, 2, 3, and 4 cycles of division as determined by their CFSE dilution (n = 3, technical replicates). (C) Model depicting how TLR and BCR signaling integrate to control distinct B-1a responses. Data are representative of at least three independent experiments (A-B).
Additional files
-
Transparent reporting form
- https://doi.org/10.7554/eLife.47015.023





