Mask family proteins ANKHD1 and ANKRD17 regulate YAP nuclear import and stability
Figures
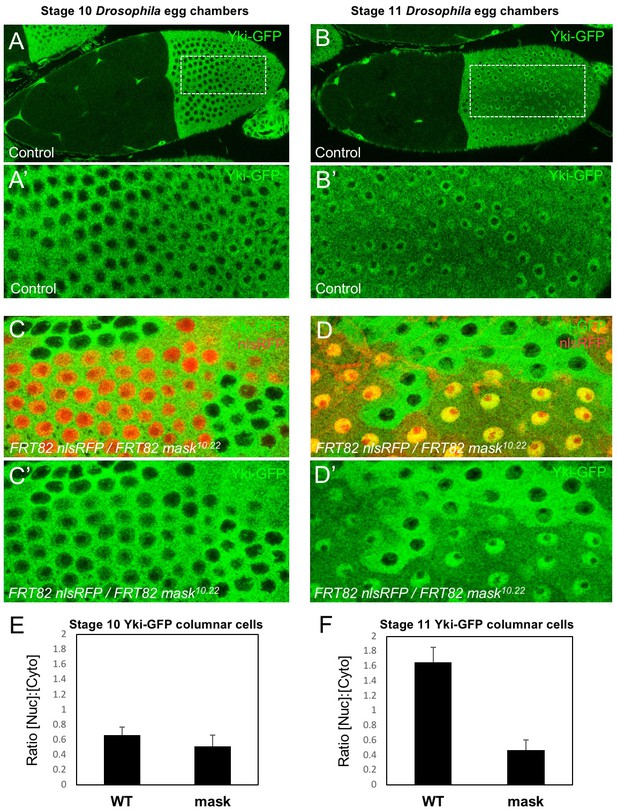
Mask is required to promote nuclear localisation of Yki in Drosophila follicle cells.
(A) Stage 10 Drosophila egg chamber with endogenously tagged Yki-GFP (green) localised to the nucleus of stretch cells (anterior) and cytoplasm of columnar cells (posterior). (A’) Magnification of columnar cells. (B) Stage 11 Drosophila egg chamber with endogenously tagged Yki-GFP (green) localised to the nucleus of stretch cells (anterior) and nucleus of flattening columnar cells caused by growth of the oocyte (posterior). (B’) Magnification of flattening columnar cells. (C) Stage 10 Drosophila egg chamber containing null mutant clones of mask (marked by absence of nuclear RFP, red) display cytoplasmic Yki-GFP. (C’) Yki-GFP single channel. (D) Stage 11 Drosophila egg chamber containing null mutant clones of mask (marked by absence of nuclear RFP, red) display cytoplasmic Yki-GFP. (D’) Yki-GFP single channel. (E) Quantification of nuclear:cytoplasmic ratio of Yki-GFP in (C) n = 7 clones. (F) Quantification of nuclear:cytoplasmic ratio of Yki-GFP in (D) n = 12 clones.
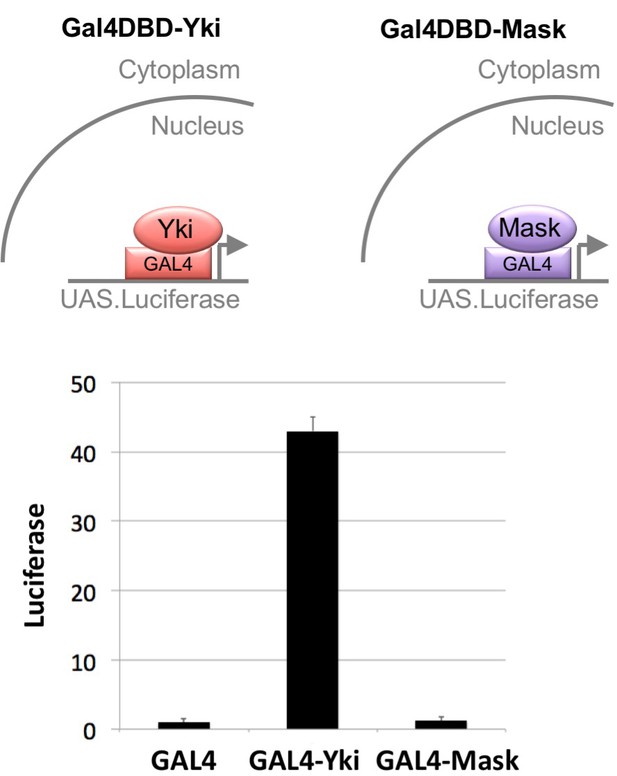
Mask has no intrinsic transcriptional co-activator activity in a GAL4-UAS reporter assay.
Drosophila S2 cells were transfected with a UAS.Luciferase reporter gene and various GAL4 linked constructs, including GAL4 DNA binding domain (DBD) alone, GAL4-DBD-Yki and GAL4-DBD-Mask. No effect of Mask on luciferase reporter activity was measured, while Yki had a very strong transcriptional activity on the UAS.Luciferase reporter gene.
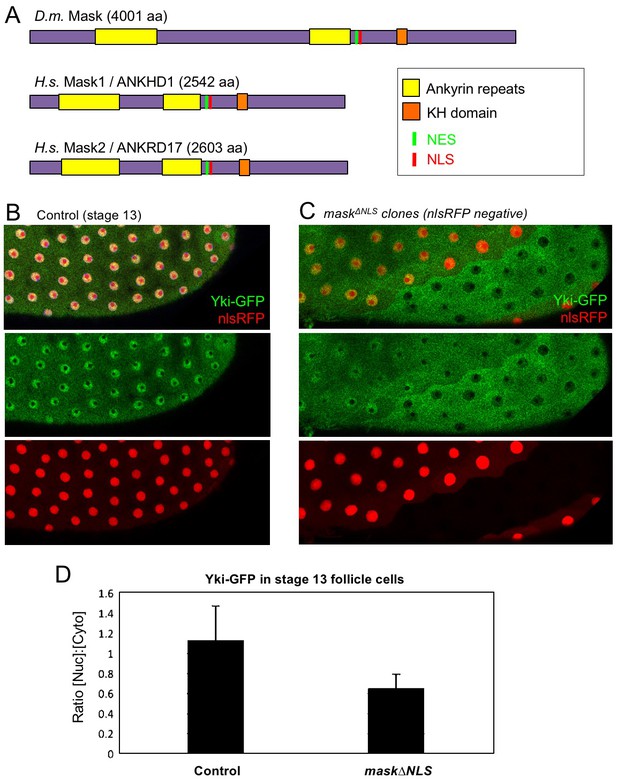
CRISPR-knockout of the Mask nuclear localisation signal leads to cytoplasmic accumulation of Yki.
(A) Schematic diagram of the Mask family proteins and their conserved domains and motifs. (B) Stage 13 Drosophila egg chamber with endogenously tagged Yki-GFP (green) localised to the nucleus of flattening columnar cells. Nuclear localised RFP is also shown (red). (C) Stage 13 Drosophila egg chamber with endogenously tagged Yki-GFP (green) containing homozygous mutant clones for a CRISPR-knockin mask∆NLS mutant allele, marked by the absence of nuclear RFP (red). Note that Yki-GFP is no longer localised to the nucleus of flattening columnar cells in the mask∆NLS mutant clone. (D) Quantification of nuclear:cytoplasmic ratio of Yki-GFP in multiple mask∆NLS mutant clones as shown in (C) n = 22 clones.
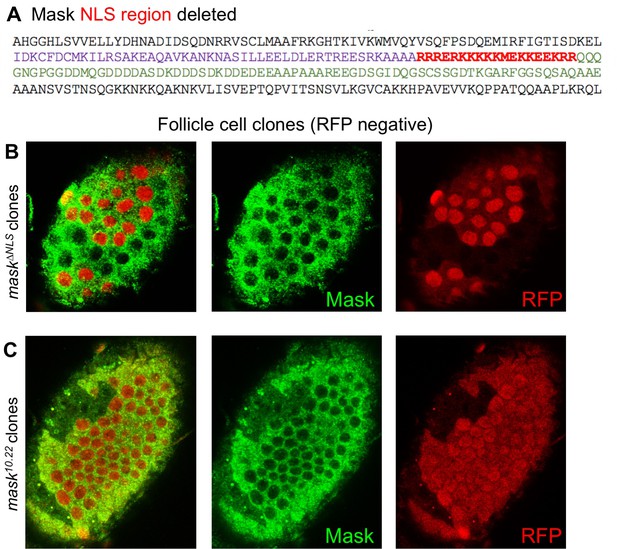
The Mask∆NLS protein is expressed at normal levels and is localised to the cytoplasm.
(A) The Nuclear Localisation Signal (NLS) region of the wild-type Mask protein is indicated in red. This region was deleted in the mask∆NLS allele. (B) Clones homozygous for the FRT mask∆NLS allele were induced by flp-FRT recombination over an FRT nlsRFP marker chromosome. Loss of nlsRFP indicates the homozygous cells. Note normal expression and cytoplasic localization of the Mask∆NLS protein. (C) Clones homozygous for the null mutant allele of FRT82 mask10.22 show a loss of Mask protein.
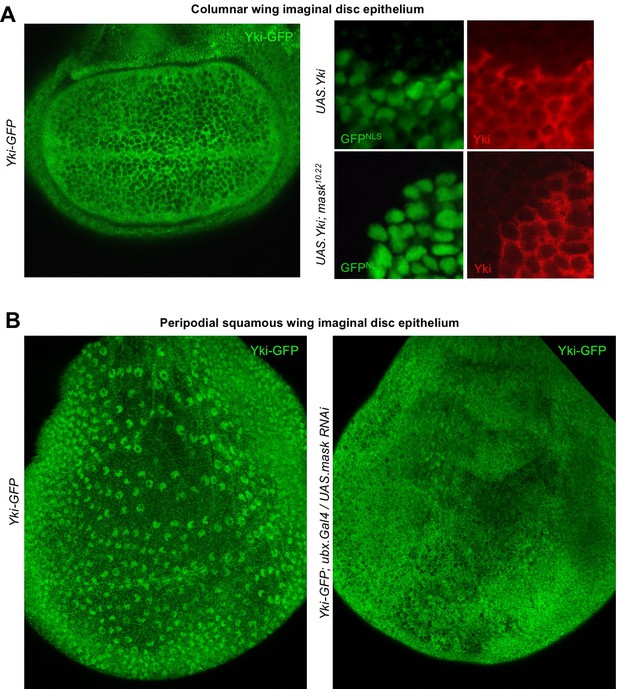
Mask is required to promote nuclear localisation of Yki in developing Drosophila wings.
(A) Columnar epithelium of the wing imaginal disc from third instar larva showing mostly cytoplasmic localisation of endogenously tagged Yki-GFP (left). MARCM clones expressing Yki with the GAL4/UAS system show both nuclear and cytoplasmic Yki localisation, as detected by Yki antibody staining (nlsGFP+). MARCM clones for a null mask mutant allele show reduced nuclear Yki (nlsGFP+). n = 6 clones. (B) Squamous peripodial epithelium from the wing imaginal disc from third instar larva showing mostly nuclear localisation of endogenously tagged Yki-GFP (left). Silencing of Mask expression by ubx.Gal4 driven UAS.mask-IR (inverted repeat hairpin RNAi) in the peripodial epithelium is sufficient to reduce nuclear Yki-GFP localisation.
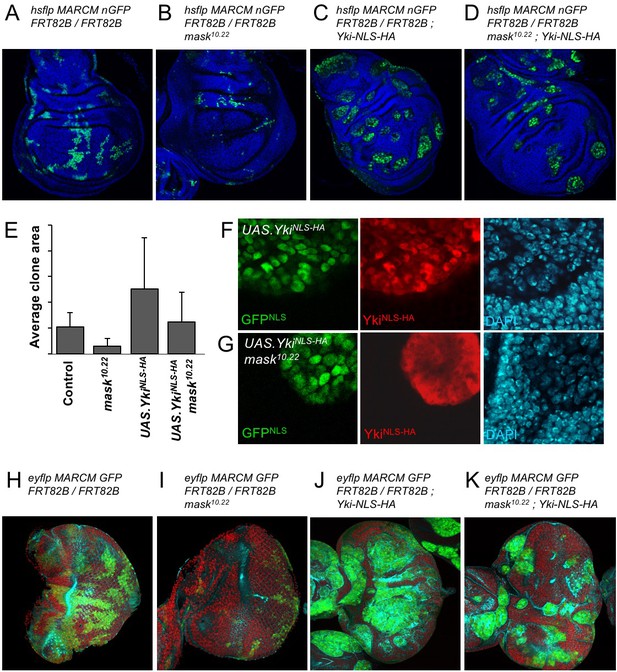
Addition of an NLS to Yki enables it to enter the nucleus and drive clonal growth in mask mutant cells.
(A) Wing imaginal disc containing control MARCM clones (nlsGFP+, green). (B) Wing imaginal disc containing null mutant mask MARCM clones (nlsGFP+, green) are almost eliminated. (C) Wing imaginal disc containing control MARCM clones also expressing nlsYkiHA (nlsGFP+, green) proliferate more strongly than controls. (D) Wing imaginal disc containing null mutant mask MARCM clones also expressing nlsYkiHA (nlsGFP+, green) are rescued for their growth and survival to a level similar to wild-type clones. (E) Quantification of average clone sizes in A-D. (F) Close-up view of control MARCM clones expressing nlsYkiHA in red (marked by nlsGFP+, green). DAPI in blue. (G) Close-up view of null mutant mask MARCM clones expressing nlsYkiHA in red (marked by nlsGFP+, green). DAPI in blue. (H) Eye imaginal disc containing control MARCM clones (nlsGFP+, green). (I) Eye imaginal disc containing null mutant mask MARCM clones (nlsGFP+, green) are almost eliminated. (J) Eye imaginal disc containing control MARCM clones also expressing nlsYkiHA (nlsGFP+, green) proliferate more strongly than controls. (K) Eye imaginal disc containing null mutant mask MARCM clones also expressing nlsYkiHA (nlsGFP+, green) are rescued for their growth and survival to a level similar to wild-type clones.
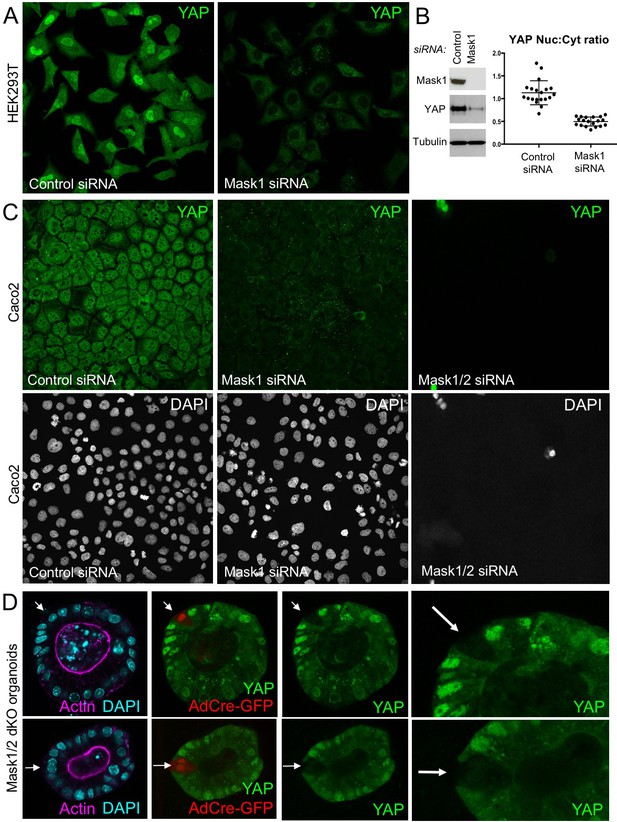
Mammalian Mask1/2 are required for nuclear localisation and stability of YAP.
(A) HEK293T cells display nuclear YAP localisation after transfection with control scrambled siRNA, and low levels of cytoplasmic YAP localisation after transfection with Mask1 siRNA oligonucleotides. (B) Western analysis of Mask1 and YAP after transfection of HEK293T cells with control scrambled or Mask1 siRNA oligonucleotides (left). Quantification of the nuclear:cytoplasmic ratio of YAP subcellular localisation after transfection of HEK293T cells with control scrambled or Mask1 siRNA oligonucleotides as shown in A (right). (C) Caco2 intestinal epithelial cells transfected with control or Mask1 or Mask1/2 siRNA oligonucleotides and immunostained for YAP or stained with DAPI to mark nuclei. (D) Intestinal organoids freshly cultured from Mask1/2 double conditional knockout mice, in which Ad-Cre was used to induce clonal knockout of both Mask1 and Mask2 in single cells (marked by Ad-Cre-GFP in red). Note the dramatic reduction in YAP protein levels (n = 12 organoids).
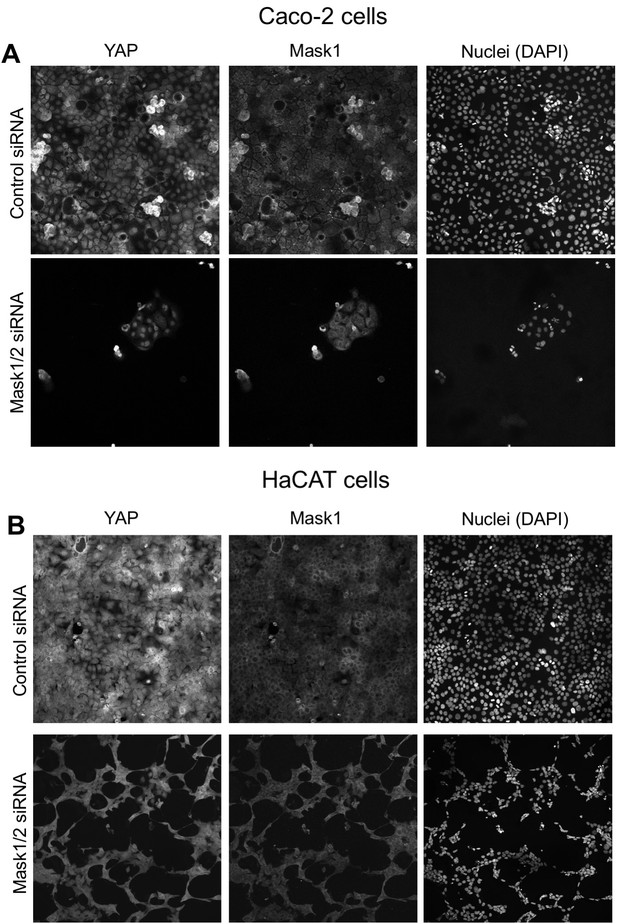
Mask1 and two double siRNA transfected cells undergo cell death.
(A) Caco2 intestinal epithelial cells are eliminated following transfection of both Mask1/2 siRNAs. No YAP-negative or Mask1-negative cells remain. (B) HaCat skin epithelial cells are eliminated following transfection of both Mask1/2 siRNAs. No YAP-negative or Mask 1-negative cells remain.
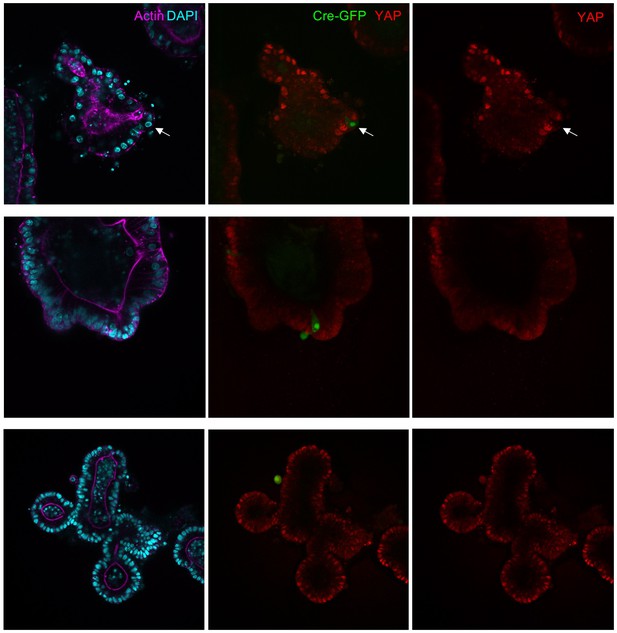
Mask1 and 2 dKO cells induced by Adenoviral-Cre infection tend to be extruded from intestinal organoids.
Mouse intestinal organoids derived from Mask1/2 double floxed conditional knockout mice were cultured in Matrigel and infected with Ad-Cre-GFP (green) before immunostaining for YAP (red) and phalloidin (purple)/DAPI (blue). Note that Ad-Cre-GFP clones do not grow larger than single cells and tend to be extruded from the epithelium. Representative examples of n = 25 organoids are shown.
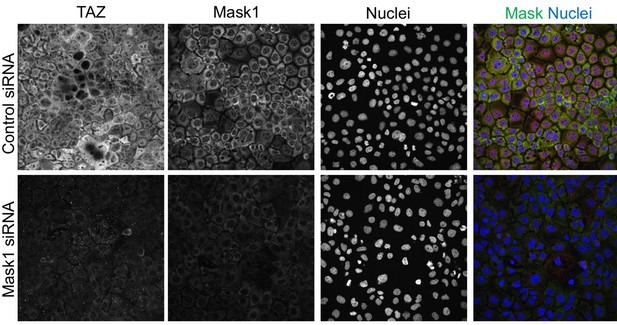
Mask1 regulates TAZ similarly to YAP in Caco2 cells.
Human Caco2 epithelial cells were transfected with scrambled siRNA (control) or specific siRNAs targeting Mask1 and then immunostained for TAZ, Mask1 and Nuclei (with DAPI). Silencing of Mask1 reduces the expression level of both Mask1 and TAZ proteins, with TAZ also showing a cytoplasmic localization in the absence of Mask1.
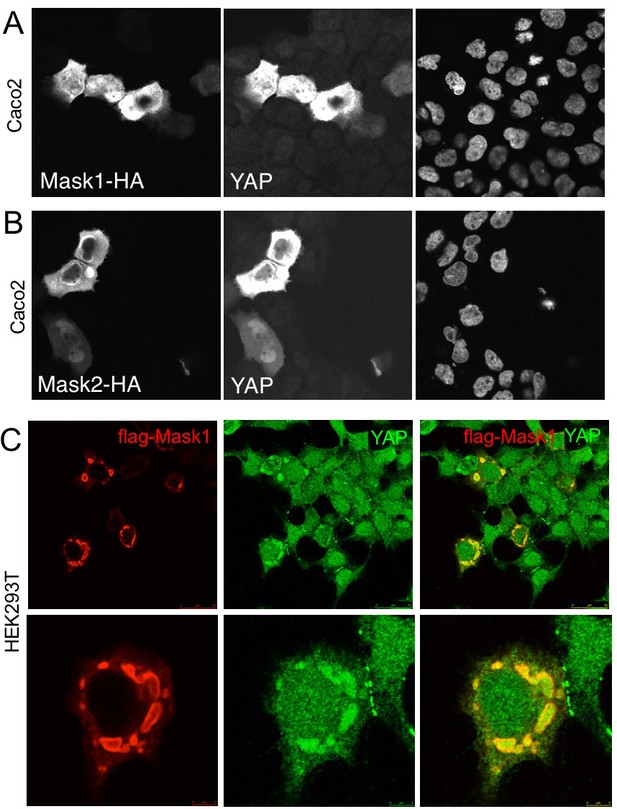
Overexpression of Mask1/2 stabilizes YAP and can induce phase separation of YAP.
(A) Caco2 intestinal epithelial cells overexpressing HÁ-tagged Mask1, which induces stabilisation of endogenous YAP. DAPI marks nuclei in the right-hand panel. (B) Caco2 intestinal epithelial cells overexpressing HÁ-tagged Mask2, which also induces stabilisation of endogenous YAP. DAPI marks nuclei in the right-hand panel. (C) HEK293T cells overexpressing HÁ-tagged Mask1, which induces stabilisation of endogenous YAP and formation of large liquid-like droplets with smooth boundaries containing both proteins – suggestive of colloidal phase separation.
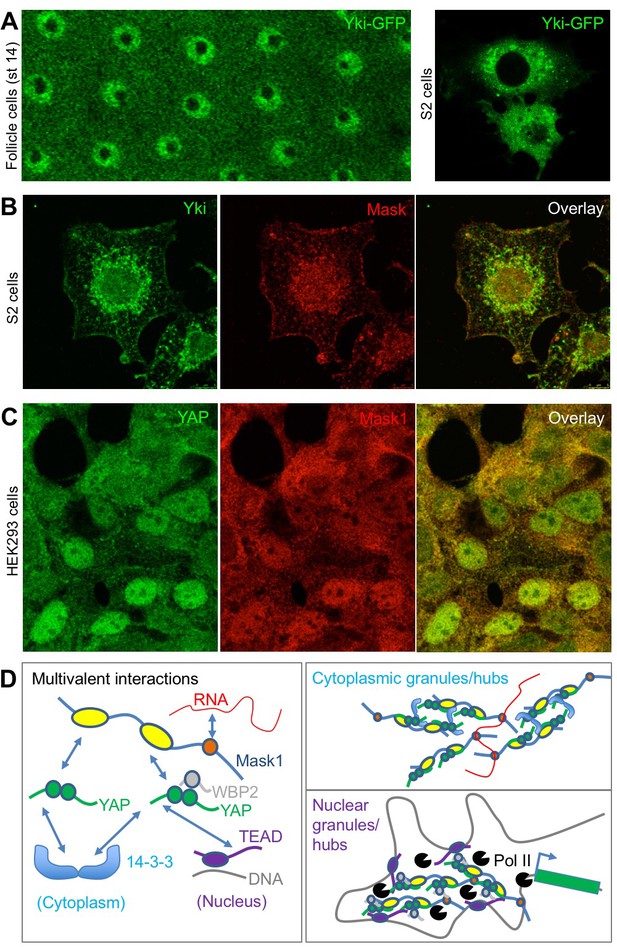
Endogenous Yki/YAP and Mask co-localise in intracellular granules in both nucleus and cytoplasm.
(A) Yki-GFP exhibits a granular appearance in both nucleus and cytoplasm of flattened Drosophila follicle cells. Yki-GFP also exhibits a granular appearance in the cytoplasm of S2 cells. (B) Co-localisation of endogenous Yki and Mask in nuclear and cytoplasmic granules of S2 cells. (C) Co-localisation of endogenous YAP and Mask1 in nuclear and cytoplasmic granules of HEK293 cells. (D) Schematic diagram of multivalent interactions between YAP binding partners and the possible formation of clusters (hubs/granules) in either the cytoplasm or nucleus.
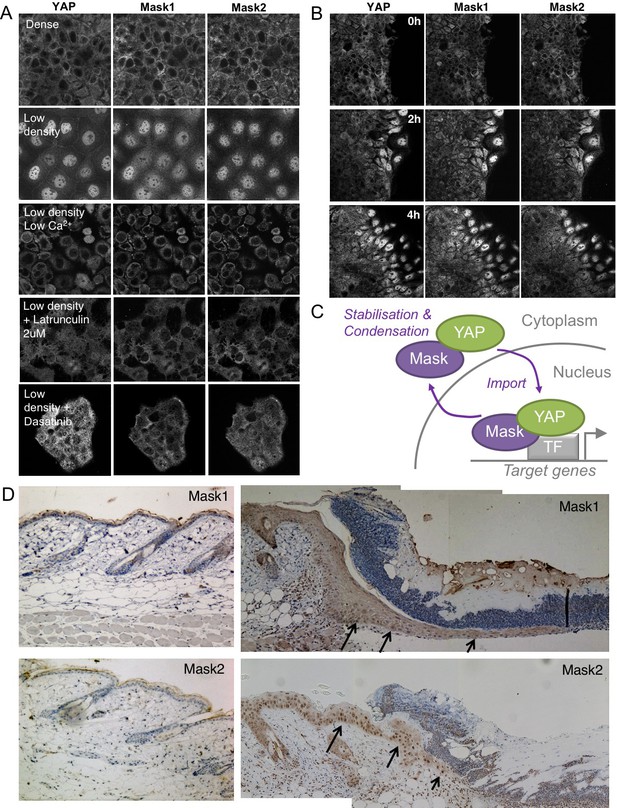
Co-regulation of YAP and Mask1/2 both in vitro and in vivo.
(A) Caco2 intestinal epithelial cells immunostained for YAP, Mask1 and Mask2 at different densities and following treatments to interrupt Integrin-Src mediated mechanotransduction (low Calcium, Latrunculin, or Dasatinib). (B) Caco2 intestinal epithelial cells immunostained for YAP, Mask1 and Mask2 at various times after scratch wounding of the epithelium. (C) Schematic diagram of Mask-YAP interactions in the nucleus and cytoplasm and their co-regulation during nucleo-cytoplasmic shuttling. (D) Mouse skin epithelium before (left) and after (right) wounding, showing upregulation and nuclear localisation of Mask1/2 in vivo.
Additional files
-
Source data 1
Primary data file.
This .xlsx file contains the primary data used to generate the graphs shown in the main figures.
- https://cdn.elifesciences.org/articles/48601/elife-48601-data1-v3.xlsx
-
Transparent reporting form
- https://cdn.elifesciences.org/articles/48601/elife-48601-transrepform-v3.pdf





