Induction of Sertoli-like cells from human fibroblasts by NR5A1 and GATA4
Figures
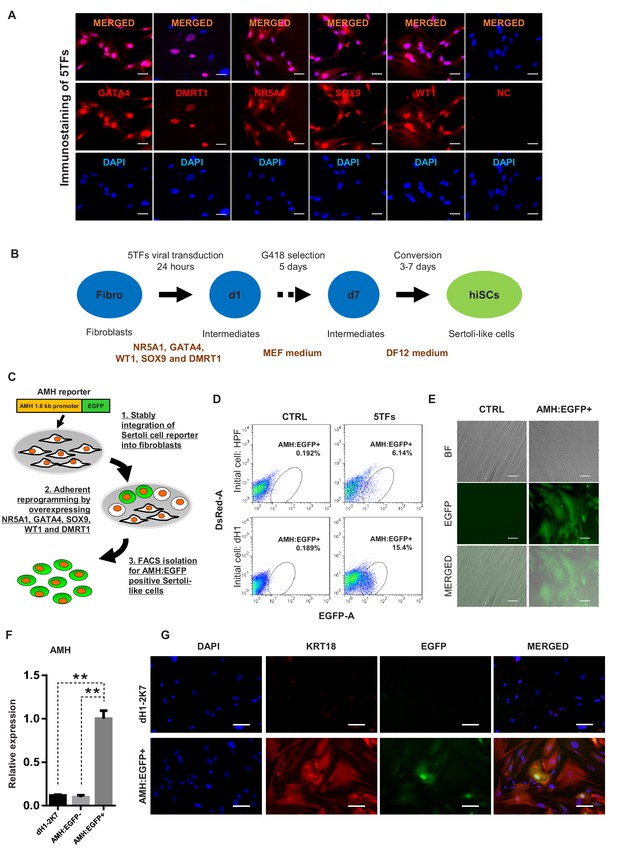
Induction of Sertoli-like cells (hiSCs) from human fibroblasts.
(A) Immunostaining of NR5A1, GATA4, WT1, SOX9 and DMRT1 (red signal) in fibroblasts (HPF) 3 days post infection. DAPI (blue signal) was used to indicate nucleus. Scale bar = 25 μm. NC represents HPF stained with only secondary antibody. (B) Experimental design for the reprogramming of human Sertoli-like cells (hiSCs). Human fibroblasts were infected by lentivirus carrying human transcription factors: NR5A1, GATA4, WT1, SOX9 and DMRT1 (5TFs). The cells were cultured in MEF medium for 1 day after infection and followed by G418 selection for 5 days, then changed to DMEM/F12 medium. hiSCs were characterized 10–14 days post viral transduction. (C) Schematic protocol for the enrichment of Sertoli-like cells (hiSCs). An AMH:EGFP reporter was integrated to the fibroblasts for hiSCs quantification and isolation. (D) FACS analysis of AMH:EGFP+ cell at day 10 after 5TFs infection. The percentage of EGFP+ cell was used to determine the reprogramming efficiency. Two types of human fibroblasts (HPF and dH1) were tested. Cells infected by p2k7 empty virus were used as negative control (CTRL). (D) Morphology of re-plated AMH:EGFP+ hiSCs after FACS. Scale bar = 20 μm. (F) The mRNA level of AMH was enriched in AMH:EGFP+ hiSCs, as compared to dH1 infected by p2k7 empty virus (dH1-2K7) and AMH:EGFP-. n = 3, technical replicates of ~5 × 104 cells were collected by FACS. All data are presented as means ± SD. *p<0.05, **p<0.01, Student’s t-test.>3 independent experiments were carried out. (G) Immunofluorescent staining of KRT18 in AMH:EGFP+ hiSCs and dH1 infected by p2k7 empty virus (dH1-2K7) after FACS. Scale bar = 100 μm.
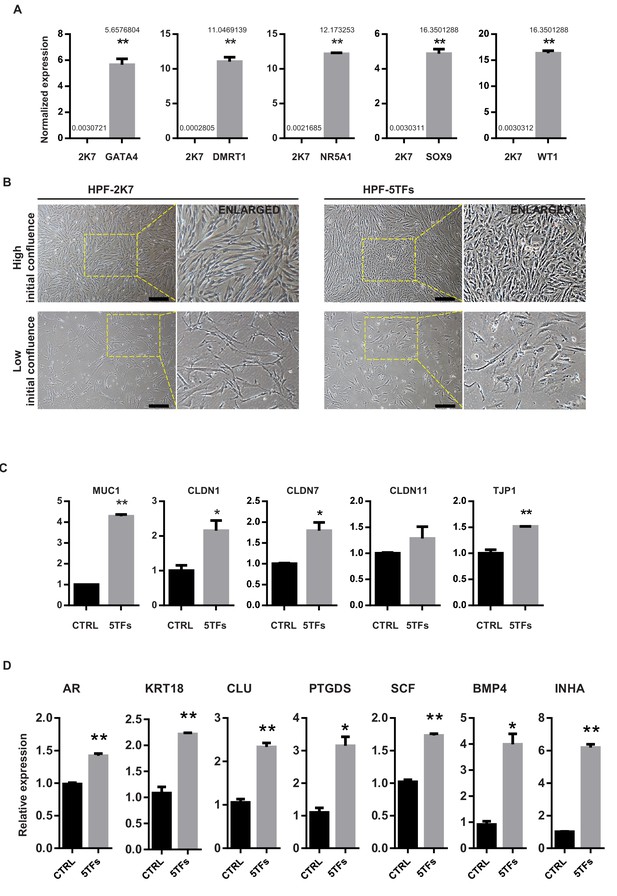
Reprogramming effect of 5F on human primary fibroblasts.
(A) Quantitative PCR result showing the mRNA level of GATA4, DMRT1, NR5A1, SOX9 and WT1 was overexpressed in 293FT cells after virus infection. ACTIN was used as the housekeeping gene for normalization. n = 2 (technical replicates of ~4 × 105 cells were collected). All data are presented as means ± SD, *p<0.05, **p<0.01, Student’s t-test.>2 independent experiments were carried out. (B) hiSCs showed epithelial morphology change after 5 TFs overexpression. Magnified images were shown on the right side. Two types of cell density were tested. Scale bars = 400 μm. (C) Quantitative PCR result showing that the mRNA level of epithelial markers increased after overexpression of NR5A1, GATA4, SOX9, WT1 and DMRT1 (5TFs). Fold expression was normalized to the group infected by empty virus p2k7 (CTRL). ACTIN was used as the housekeeping gene for normalization. All data are presented as means ± SD. *p<0.05, **p<0.01, Student’s t-test. n = 2 (technical replicates of ~4 × 105 cells were tested). (D) Quantitative PCR result showing the mRNA levels of Sertoli cell markers. Relative expression of 5TFs groups was normalized to the level of HPF cells infected by empty virus 2K7 (CTRL). All data are presented as means ± SD, asterisks indicate statistical significance, (*p<0.05, **p<0.01, Student’s t-test).
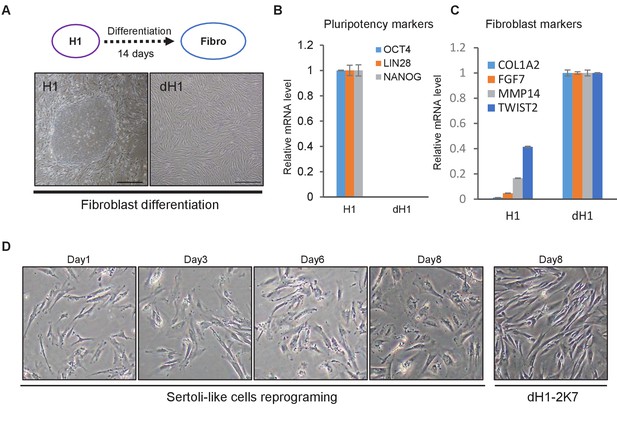
Reprogramming effect of 5F on differentiated hESC H1.
(A) Morphology of human embryonic stem cells (H1) in feeder cell system (left) and the fibroblast-like cells (dH1, right) derived from H1 by spontaneous differentiation in KSR + K-DMEM medium for 14 days. Scale bar = 400 μm. (B) Expression of hESC pluripotency markers in H1 and dH1 at day 14 after spontaneous differentiation, measured by qPCR. Expression level in dH1 was normalized to the level of undifferentiated H1. All data are presented as means ± SD, asterisks indicate statistical significance, *p<0.05; **p<0.01, Student’s t-test, versus control. (C) Expression of fibroblast markers in H1 and dH1 at day 14 after spontaneous differentiation, measured by qPCR. Expression level in dH1 was normalized to the level of undifferentiated H1.All data are presented as means ± SD, asterisks indicate statistical significance, *p<0.05; **p<0.01, Student’s t-test, versus control. (D) Cell morphology of reprogrammed dH1 at day1 to day8 after induction. dH1 transduced with p2k7 lenivirus was used as a control.
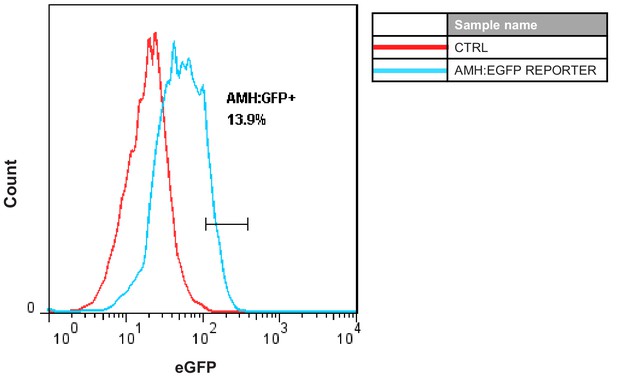
FACS analysis of AMH:EGFP+ population in adult Sertoli cells 10 days after AMH:EGFP reporter virus infection.
Cells infected by p2k7 empty virus were used as a negative control (CTRL).
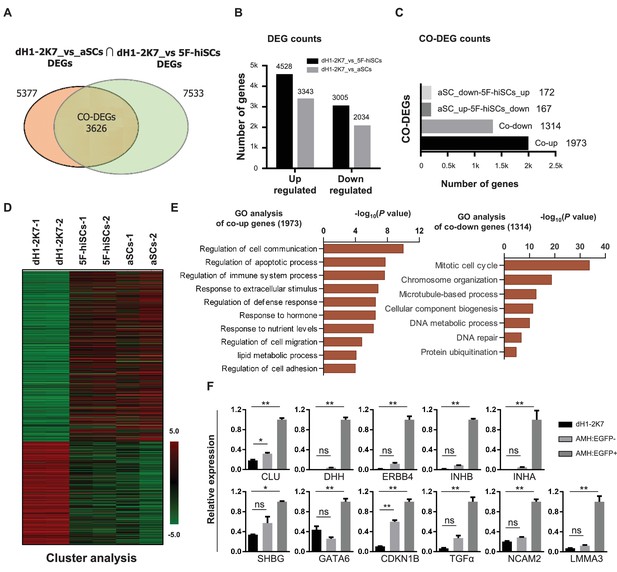
Whole genome transcriptional profiling of AMH:EGFP+ hiSCs.
(A) Venn diagram to display differentially expressed genes (DEGs, FPKM value, fold change >2) between dH1-2K7_vs_aSCs (n = 2) and dH1-2K7_vs_5F-hiSCs (n = 2). Intersection part represents the number of co-differentially expressed genes (CO-DEGs). (B) Summary of DEGs in (A). X axis represents upregulated DEGs and downregulated DEGs. Y axis represents DEGs numbers. Comparisons of different sample sets were represented by different colors. Black bar represents dH1-2K7_vs_5F-hiSCs. Gray bar represents dH1-2K7_vs_aSCs. (C) Summary of CO-DEGs in (A). X axis represents CO-DEGs number. Black bar represents co-upregulated DEGs. Gray bar represents co-downregulated DEGs. Dark gray bar and light gray bar represent CO-DEGs in opposite trend, respectively. (D) Heat map of gene expression of dH1-2K7, 5F-hiSCs and aSCs (n = 2 in each group). Red indicates upregulated expression, green indicates downregulated expression. (E) Functional enrichment analysis, biological processes of 1973 and 1314 differentially expressed genes were showed. (F) The mRNA level of Sertoli cell markers in dH1-2K7, AMH:EGFP- cells and AMH:EGFP+ cells, as measured by qPCR. Relative expression was normalized to dH1-2K7. GAPDH was used as the housekeeping gene. All data are presented as means ± SD, n = 2, *p<0.05, **p<0.01, Dunnett’s test. three independent experiments were carried out.
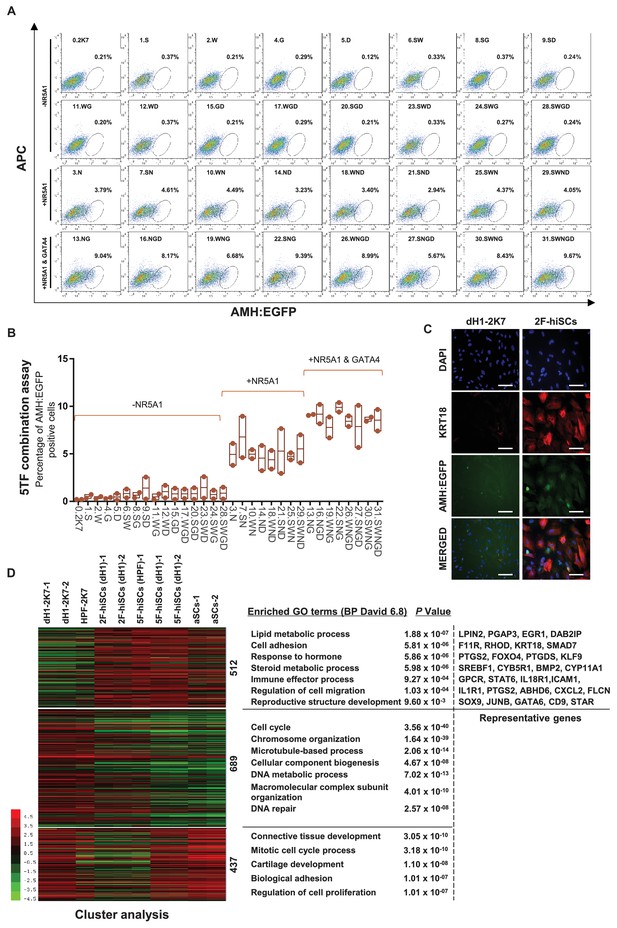
NR5A1 and GATA4 are sufficient to derive hiSCs.
(A) Representative FACS results of different combinations of NR5A1, GATA4, WT1, SOX9 and DMRT1 for the induction of AMH:EGFP+ cells. dH1 fibroblasts were transduced with the indicated factors and reprogrammed for 10 days. The combinations were divided into three groups: -NR5A1, combinations without NR5A1; +NR5A1, combinations with NR5A1 but without GATA4; +NR5A1 and GATA4, combinations with both NR5A1 and GATA4. (B) Quantitative data of EGFP+ cells in (A). n = 2, biological replicates, error bar indicates SD, three independent combination experiments were conducted. ~104 cells were analyzed in each experiment. (C) Immunofluorescent staining of KRT18 in 2F-hiSCs and dH1-2K7 after FACS. Scale bar = 100 μm. (D) Heat map of RNA-seq data illustrating differentially co-expressed genes. Red indicates upregulated expression, whereas green indicates downregulated expression. The genes were divided into three groups: Genes upregulated in 2F-hiSCs (dH1), 5F-hiSCs (dH1) and aSCs; genes upregulated in 5F-hiSCs (dH1) and aSCs but downregulated in 2F-hiSCs (dH1); genes downregulated in 2F-hiSCs (dH1), 5F-hiSCs (dH1) and aSCs; all compared to dH1-2K7. The gene number of each group was listed next to the map. Functional enrichment terms of each group and the representative genes were shown on the right side.
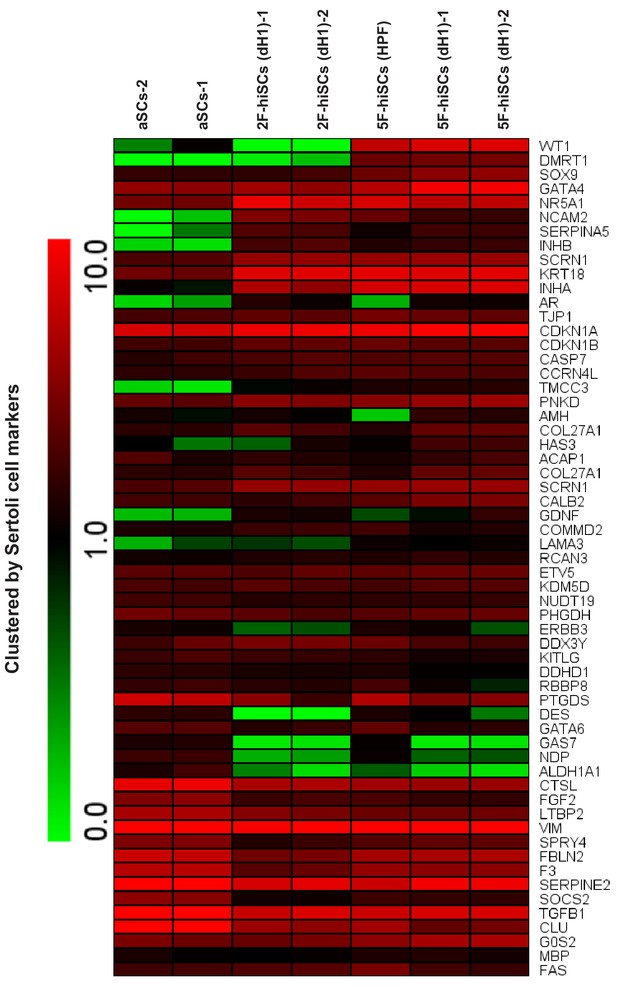
Heat map clustered by the expression of Sertoli cell markers in 2F-hiSCs, 5F-hiSCs and aSCs cells.
Red indicates higher expression and green indicates lower expression.
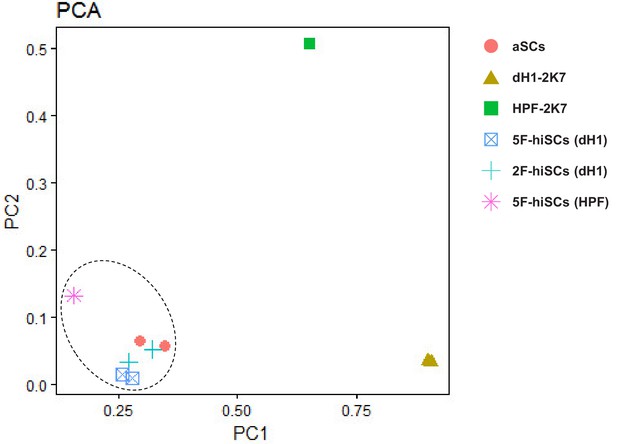
PCA plot of differentially co-expressed genes in 2F-hiSCs (dH1), 5F-hiSCs (dH1) , 5F-hiSCs (HPF) and aSCs, compared to dH1-2K7.
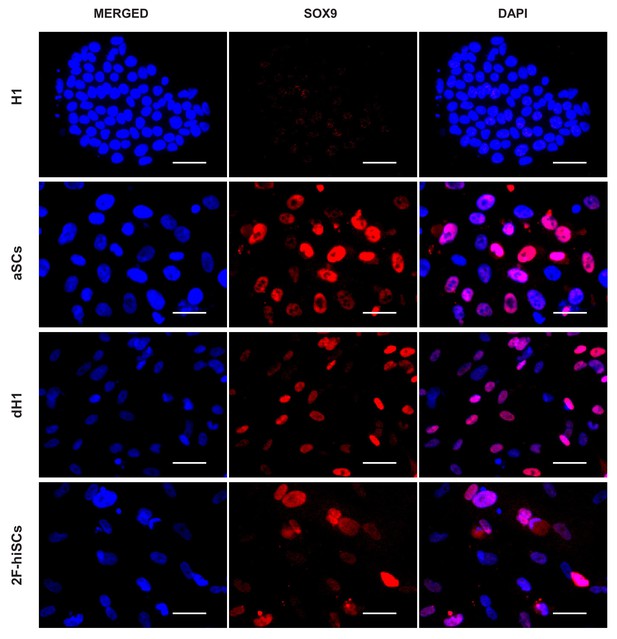
Immunofluorescent staining of SOX9 (red) in dH1, 2F-hiSCs, hESCs (H1) and adult Sertoli cells (aSCs).
DAPI (blue) was used to indicate the nucleus. Scale bar = 50 μm.
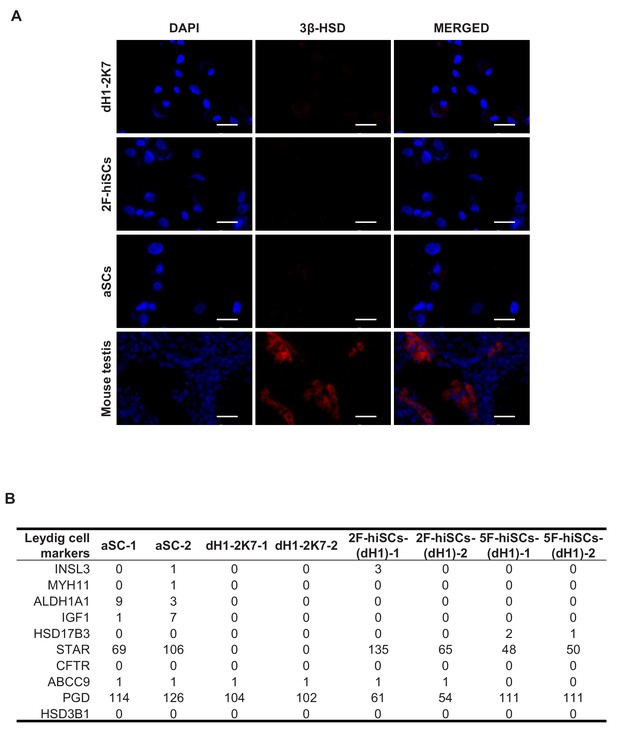
hiSCs and aSCs do not express Leydig cell specific markers.
(A) Cytospin immunostaining of Leydig cell specific marker 3β-HSD in fibroblasts (dH1-2K7), 2F-hiSCs and adult Sertoli cells (aSCs). 18 dpp mouse testis freezing section was used as positive control cells. Scale bar = 50 μm. (B) FPKM values of some reported Leydig cell markers in dH1-2K7, 2F-hiSCs (dH1), 5F-hiSCs (dH1) and aSCs, according to RNA-seq data presented in Figure 2.
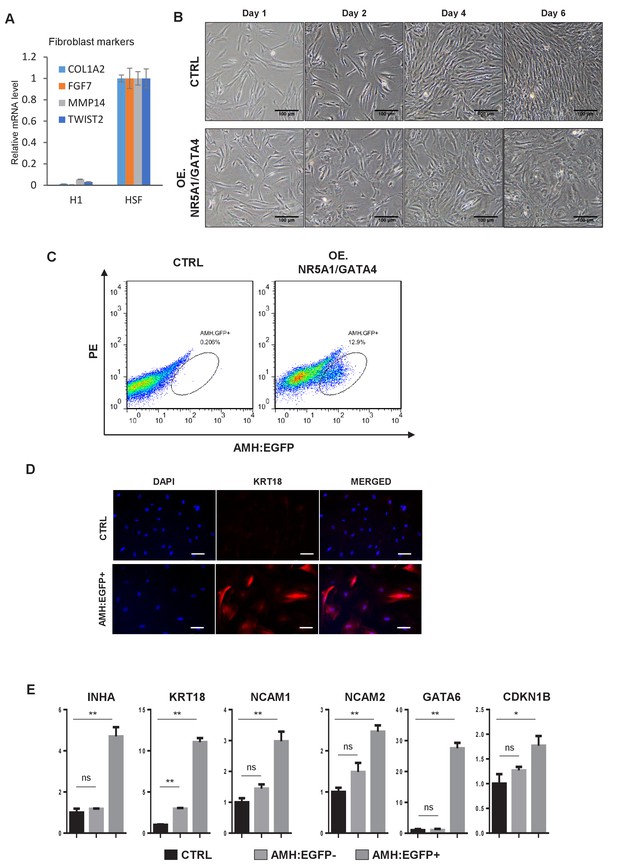
Induction of hiSCs from primary human skin fibroblasts.
(A) Expression of fibroblast markers in primary human skin fibroblasts (HSF) as measured by qPCR. Expression level in dH1 was normalized to the level of undifferentiated H1. All data are presented as means ± SD. (B) Cell morphology of reprogrammed HSF at day1, day2, day4 and day6 after infection by NR5A1 and GATA4. HSF transduced with p2k7 lenivirus was used as a control. Scale bar = 200 μm. (C) FACS analysis of AMH:EGFP+ cells at day 10 after NR5A1 and GATA4 virus infection, the percentage of EGFP+ cell was used to determine the induction efficiency. HSF transduced with p2k7 lentivirus was used as a control. (D) Immunofluorescent staining of KRT18 in AMH:EGFP+ hiSCs and HSF infected by p2k7 empty virus (CTRL) after FACS. Scale bar = 100 μm. (E) The mRNA level of Sertoli cell markers in AMH:EGFP- cells and AMH:EGFP+ cells as measured by qPCR. Fold expression was normalized to the levels of HSF infected by empty virus p2k7 (CTRL). ACTIN was used as the housekeeping gene for normalization. All data are presented as means ± SD, n = 2, *p<0.05, **p<0.01, dunnett-t test, three independent experiments were carried out.
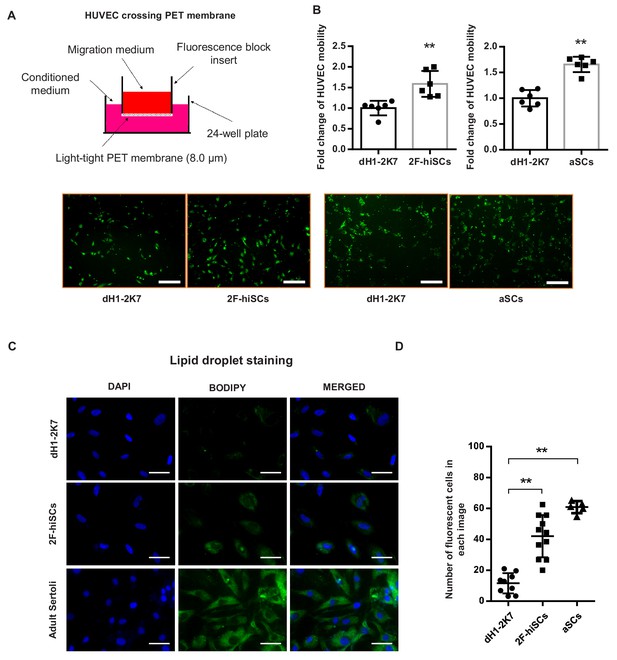
2F-hiSCs attract endothelial cells and accumulate lipid droplets.
(A) Mobility comparison of HUVECs incubated in conditioned medium collected from dH1-2K7, 2F-hiSCs and aSCs. Top, experimental diagram using the fluoroblok 24-multiwell insert plates with 8.0 μm pores. For observation, HUVEC were treated with 2.5 μM Calcein-AM fluorescent dye 1 hr prior to seeding. Down, fluorescent images showing cells passed through the pores. Conditioned medium collected from dH1-2K7 was used as control. Scar bar = 300 μm. (B) Summary of HUVEC cells crossing through the trans-well membrane. Each data point on the graph represents the number of cells in one separate image observed under 10 × fold microscope, area = 1690 × 1690 (μm). Error bars represent standard deviation of all the counted data points, n = 6, technical replicates, *p<0.05, **p<0.01, Student’s t-test. two independent experiments were carried out. (C) BODIPY staining of lipid droplets in dH1-2K7, 2F-hiSCs and adult Sertoli cells (aSCs). All cells were fixed with 4% PFA and then stained with BODIPY for lipid droplets and DAPI for nucleus. Scar bar = 50 μm. (D) Each data point on the graph represents number of BODIPY+ cells in one separate image observed under 20 × fold microscope, area = 275 × 275 (μm). Error bars represent standard deviations of all the counted data points, *p<0.05, **p<0.01, Student’s t-test. two independent experiments were carried out.
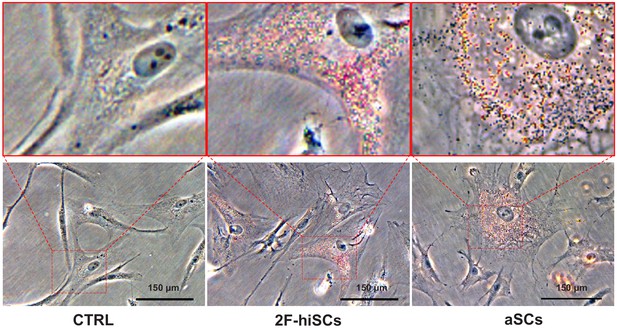
Oil-red O staining showing lipid droplets (red) in dH1-2K7 (CTRL), aSCs and 2F-hiSCs.
All cells were fixed with 4% PFA and then stained with Oil-red O for lipid droplets. Bright field picture. Scar bar = 150 μm.
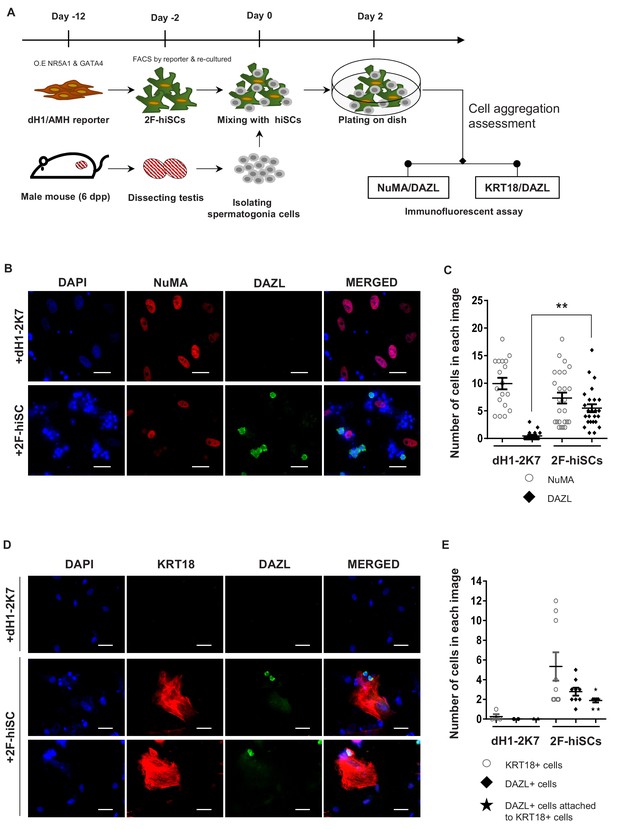
2F-hiSCs sustain the viability of mouse spermatogonia cells.
(A) Timeline and steps of co-culturing hiSCs with mouse germ cells. (B) Immunofluorescent staining of germ cell marker, DAZL (green), and human specific marker, NuMA (red), in co-cultured mouse spermatogonia cells with 2F-hiSCs or dH1-2K7 48 hr after plating. DAPI (blue) was used to indicate the nucleus. Scale bar = 50 μm. (C) Summary of cell numbers on the co-cultured plate in (B). Each data point indicates the number of cells counted in one separate image observed under 20 × fold microscope, area = 330 × 330 (μm). Error bars are the standard deviations of all the counted data points. dH1-2K7 (n = 18, technical replicates from two separated experiments), 2F-hiSCs (n = 25, technical replicates from two separated experiments). *p<0.05, **p<0.01, Student’s t-test. (D) Immunofluorescent staining of DAZL (green) and KRT18 (red) in co-cultured mouse spermatogonia cells and 2F-hiSCs 48 hr after plating. DAPI (blue) was used to indicate the nucleus. Scale bar = 50 μm. (E) Summary of cell numbers on the co-cultured plate in (D). Each data point indicates the number of cell counted in one separate image observed under 20 × fold microscope, area = 282 × 330 (μm). Error bars are the standard deviations of all the counted data points. dH1-2K7 (n = 4, technical replicates from two separated experiments), 2F-hiSCs (n = 9, technical replicates from two separated experiments). *p<0.05, **p<0.01, Student’s t-test.
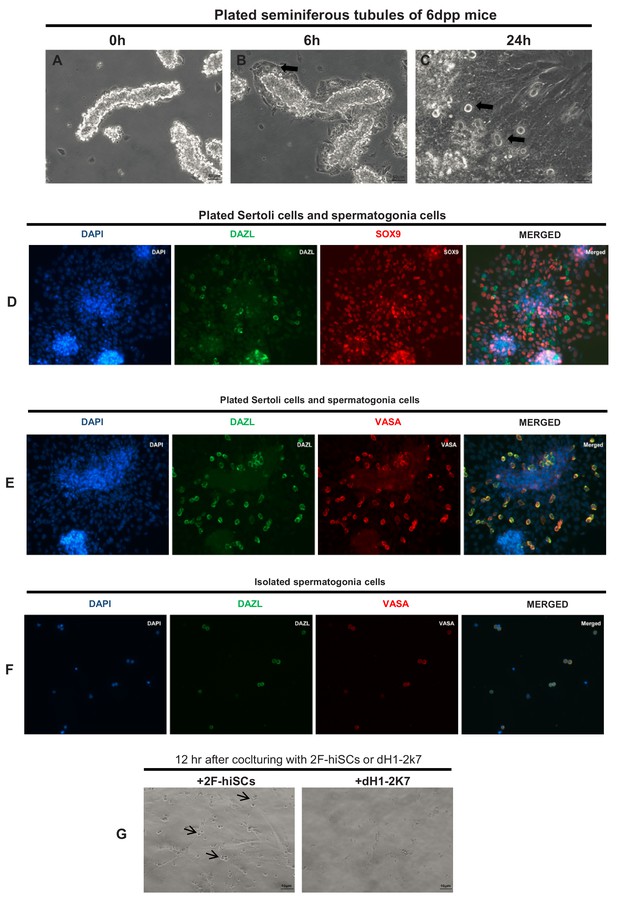
Isolation of mouse spermatogonia cells and co-culturing with hiSCs.
(A–C) Bright field images of mouse seminiferous tubules after plating on culture plate. Arrow indicates location of spermatogonia cells attached to Sertoli cells. (D) Immunostainning of germ cell marker, DAZL, and Sertoli cell marker, SOX9, in cultured mouse seminiferous tubules. (E) Immunostainning of germ cell marker, DAZL and VASA, in cultured mouse seminiferous tubules. (F) Isolated spermatogonia cells immunostained with DAZL and VASA. (G) Isolated mouse spermatogonia cells plated on 2F-hiSCs or dH1-2K7. Bright field image was taken after 12 hr culturing. More round and healthy cells (arrow heads) attached to hiSCs than dH1-2K7 cells. Scale bar = 10 μm.
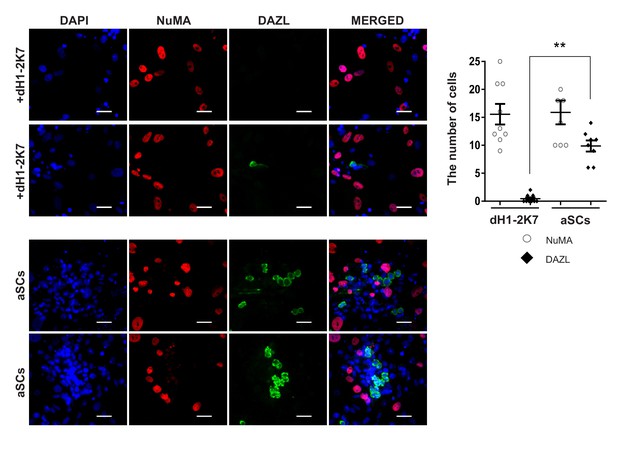
Immunofluorescent staining of germ cell marker, DAZL (green), and human specific marker, NuMA (red), in co-cultured mouse spermatogonia cells with aSCs or dH1-2K7 48 hr after plating.
DAPI (blue) was used to indicate the nucleus. Scale bar = 50 μm. Summary of cell numbers on the co-cultured plates was shown on the left. Each data point indicates the number of cell counted in one separate image observed under 20 × fold microscope, area = 282 × 330 (μm). Error bars are the standard deviations of all the counted data points.
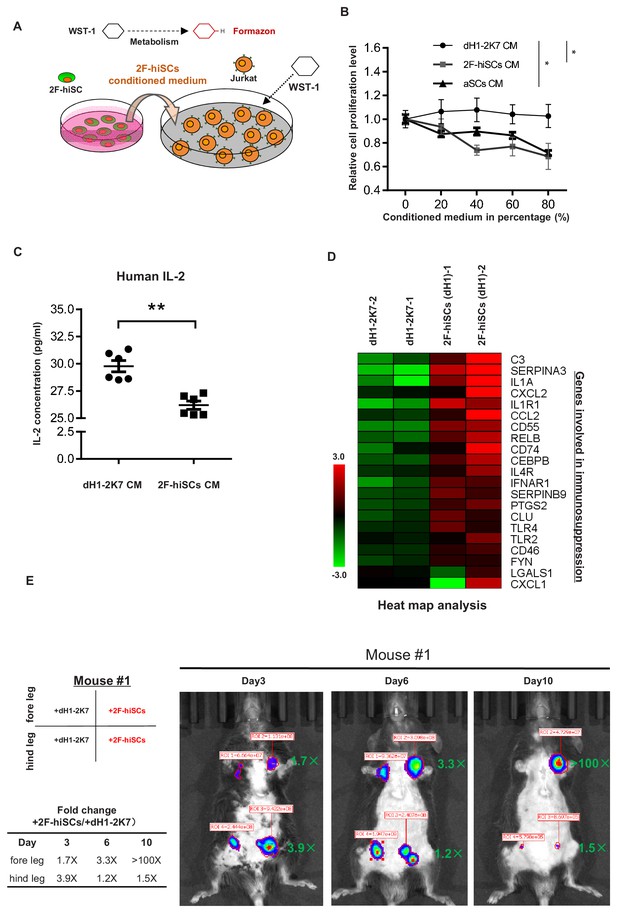
2F-hiSCs exhibit immunosuppressive function and protect human cells in xenotransplantations.
(A) Schematic illustration of the WST-1 assay to measure the inhibiting effect of 2F-hiSCs on Jurkat-E6 cell proliferation. Cleavage of the tetrazolium salt WST-1 by metabolically active cells results in the formation of formazon. The level of formazon is directly proportional to the proliferation level of the Jurkat cells. (B) Measurement of proliferation level in Jurkat cells treated with dH1-2K7, 2F-hiSCs or adult Sertoli cell (aSCs) conditioned medium. X axis represents the indicated proportion of conditioned medium tested in experiment. Conditioned medium from dH1-2K7 was used as a control. Values are presented as means ± SD and are from two separate experiments with each concentration tested in triplicate; Dunnett’s test was carried out among samples at 80% CM, *p<0.05, **p<0.01. (C) Histogram showing the production of human IL-2 in lymphocytes treated by dH1-2K7 or 2F-hiSCs conditioned medium, as measured by ELISA. Error bars represent standard deviation of three technical duplicates from two separate experiments, *p<0.05, **p<0.01, Student’s t-test. (D) A cluster of known immune-modulating genes for Sertoli cell immunoprotection. Relative gene expression level was indicated as red (upregulated) or green (downregulated). (E) Live imaging of luciferase-tracking assay 3 days to 10 days after transplantation. ~1.3 × 106 293 FT cells were subcutaneously injected into the indicated sites of the mouse #1 with 2 × 105 dH1-2K7 or 2F-hiSCs cells. Cell type and location of transplantation are indicated at left. Numbers in red represent primary readings of luciferase activity. The normalized fold change of luciferase cell activity between hiSCs and dH1-2K7 in each time point was summarized in the table.
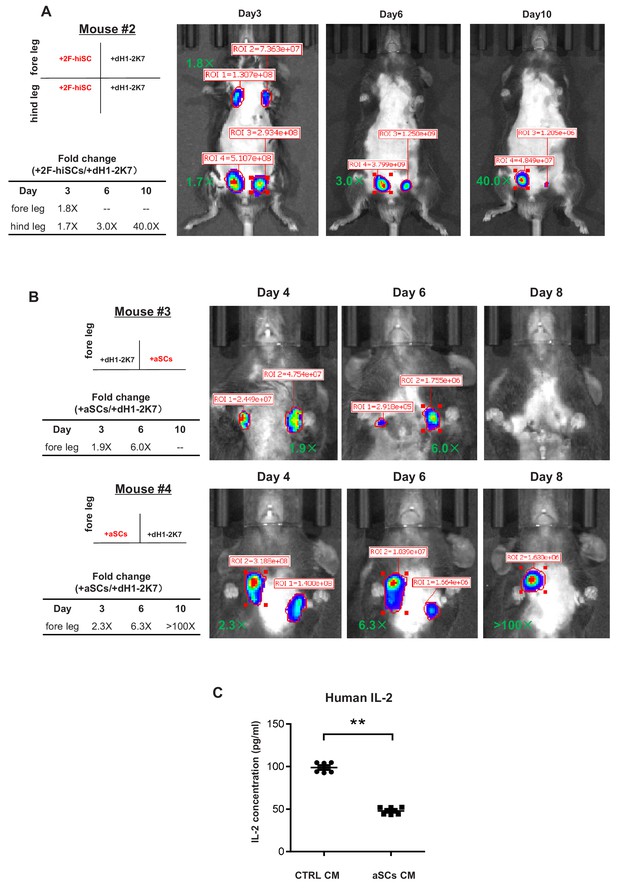
hiSCs protect the survival of human cells 293FT during xenotransplantation.
(A) Live imaging of luciferase tracking assay at day3, day6 and day10 after transplantations in mouse #2. 1.3 × 106 293 FT cells were subcutaneously injected with 2 × 105 dH1-2K7 or 2F-hiSCs into the indicated sites of the mouse. Cell type and location of transplantation is indicated on the left. Numbers in red indicate primary readings of luciferase activity. The normalized fold change of luciferase cell activity between hiSCs and dH1-2K7 in each time point was summarized in the table. (B) Live imaging of luciferase-tracking assay at day4, day6 and day8 after transplantations in mouse #3 (male) and mouse #4 (female).~1.0 × 106 293 FT cells were subcutaneously injected with 2 × 105 dH1-2K7 or adult Sertoli cells (aSCs) into the indicated sites of the mouse. Cell type and location of transplantation is indicated on the left. Numbers in red indicate primary readings of luciferase activity. The normalized fold change of luciferase cell activity between aSCs and dH1-2K7 in each time point was summarized in the table. (C) Graph showing the production of human IL-2 in lymphocytes treated by CTRL(dH1-2K7) or aSCs conditioned medium, as measured by ELISA. Error bars represent standard deviation of three technical duplicates from two separate experiments, *p<0.05, **p<0.01, Student’s t-test.
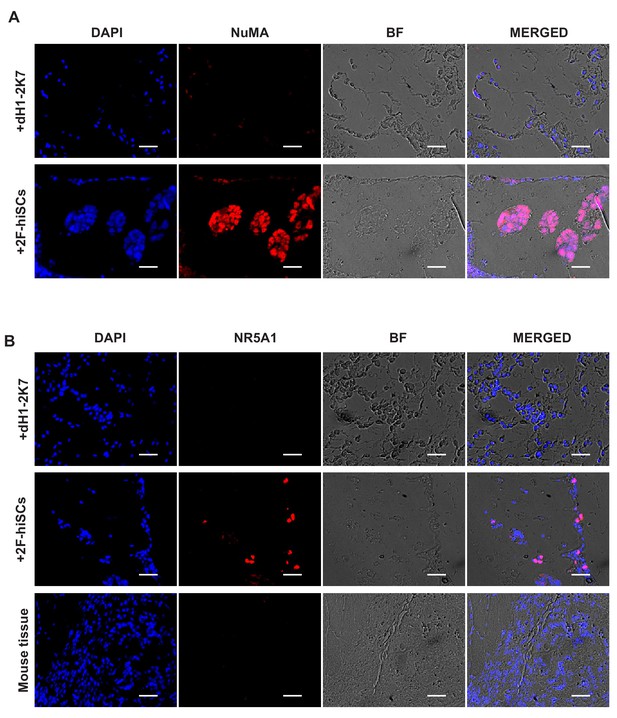
Survival of human 293FT cells with hiSC in xenograft site.
(A–B) Immunofluostaining of human NuMA and NR5A1, respectively in xenograft site of 293FT+dH1-2K7 or 293FT+2F-hiSCs collected at day10. Scale bar = 50 μm.
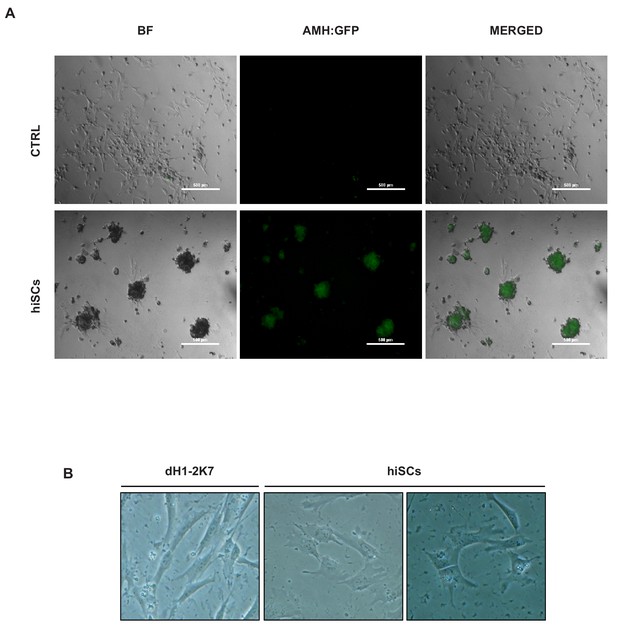
hiSCs form spherical cellular aggregates in culture medium.
(A) Comparison between fibroblasts and 2F-hiSCs in their capability to form spherical cellular aggregates when cultured with 10% FBS on 50% matrigel coated plate for 72 hr. Green channel represents the signal from AMH:EGFP reporter. Scale bar = 500 μm. (B) Comparison between dH1-2K7 fibroblast and 2F-hiSCs in their capability to form ring-like structures when cultured with 2% FBS for 72 hr on 10% Matrigel coated plate.
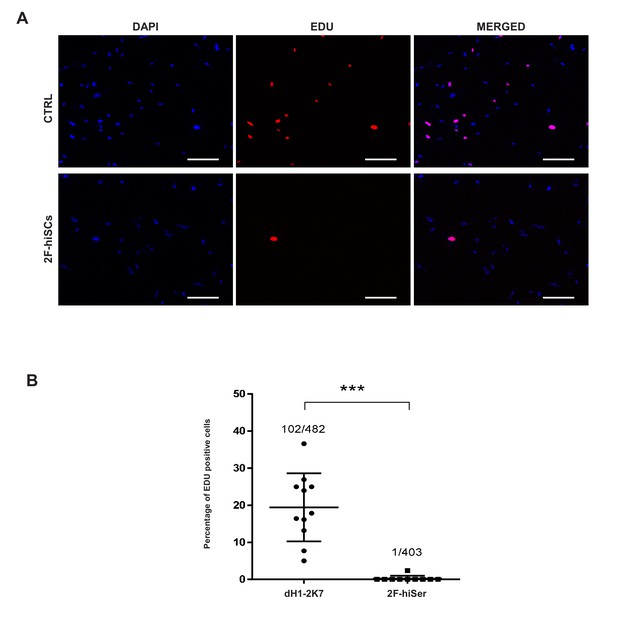
Proliferation assays showing hiSCs are not proliferative.
(A) EDU staining in dH1-2K7 (CTRL) and 2F-hiSCs. All cells were fixed with 4% PFA and then stained with EDU for lipid droplets and DAPI for nucleus. Scale bar = 200 μm. (B) Quantifications of EDU positive cells in each image. Image area = 1250×1250 (μm).
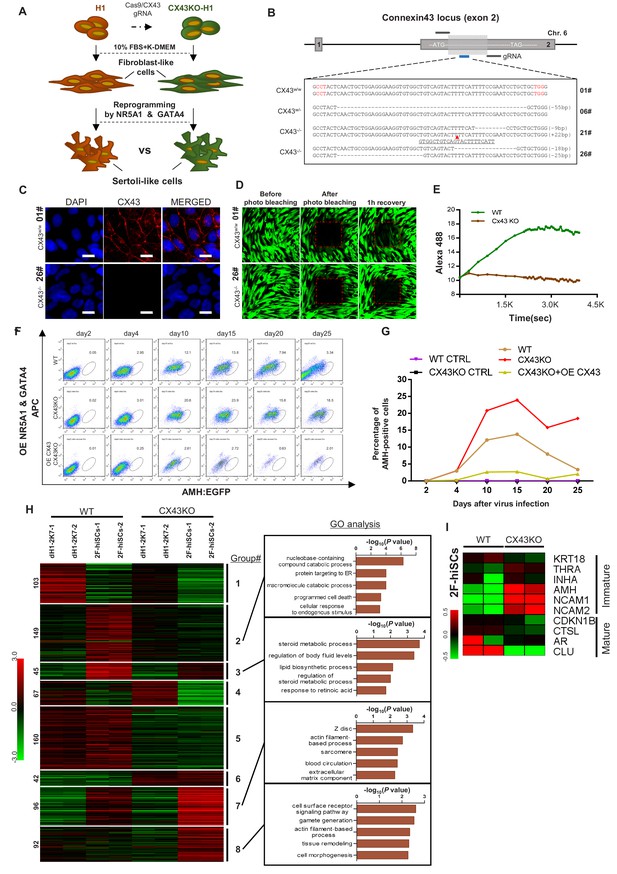
CX43 regulates maturation of Sertoli cells.
(A) Schematic diagram showing the comparison of 2F-hiSCs cells reprogrammed from CX43 knock out fibroblasts and wild type fibroblasts. (B) CX43 knockout design and the targeted sequences of indicated hESC lines. The knockout region was located in the second exon of the CX43 gene. Three knockout cell lines were correctly targeted and mutated: one was heterozygous (06#) and the other two were homozygous (21# and 26#). Dotted lines indicate deletion mutations, red triangles indicate insertion mutations. (C) Immunofluorescence analysis of CX43 (red) in wild type ES cell line (01#) and knock out cell line (26#). DAPI (blue) was used to indicate the nucleus. Scale bar = 20 μm. (D) Photo bleaching assay to test the Calcien AM transport ability of fibroblasts differentiated from wild type ES cell line (01#) and knock out cell lines (26#). (E) Measurement of Calcien AM signal over time after fluorescence bleaching. (F) Time course experiment during 2F-hiSCs reprogramming. The AMH:EGFP+ percentage resulted from three initial fibroblasts were tested: wild type dH1 (WT), CX43 knock out dH1 (CX43KO) and CX43 knock out plus CX43 overexpressed dH1 (OE CX43-CX43KO). (G) Summary of 2F-hiSCs reprogramming efficiency in (F). WT fibroblasts (WT) and CX43KO fibroblasts infected by p2k7 empty virus were also included. (H) Hierarchical clustering analysis using DEGs (FPKM value, fold change >2) in Figure 7—figure supplement 2B. Genes were classified into eight parts according to their expression pattern in CX43 knock out dH1, CX43KO 2F-hiSCs, wild type dH1 and wild type 2F-hiSCs. Gene Ontology analysis of genes in part 2, 3, 7, eight was shown on the right respectively. Other groups were shown in Figure 7—figure supplement 2C. (I) Heat map indicating expression level of immature and mature Sertoli cell markers in WT and CX43KO 2F-hiSCs. Relative gene expression level was indicated as red (upregulated) or green (downregulated).
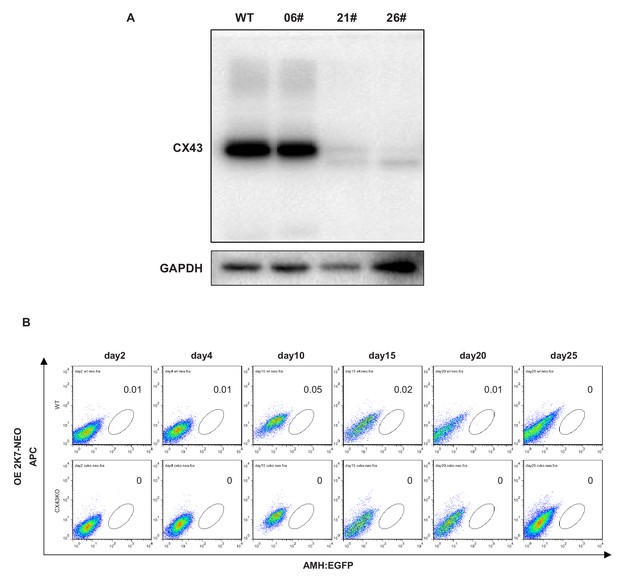
Effect of CX43 knock out on protein level and percentage of AMH-GFP.
(A) Western blot analysis of CX43 expression in wild type ES cell line (01#) and knock out cell lines 06#, 21# and 26#. The loading control is GAPDH. (B) FACS analysis of AMH:EGFP+ cells in control groups without overexpression factors (WT versus CX43KO cell lines), from day2 to day25. No GFP+ cell or very few GFP+ was detected using the same setting as in Figure 7f.
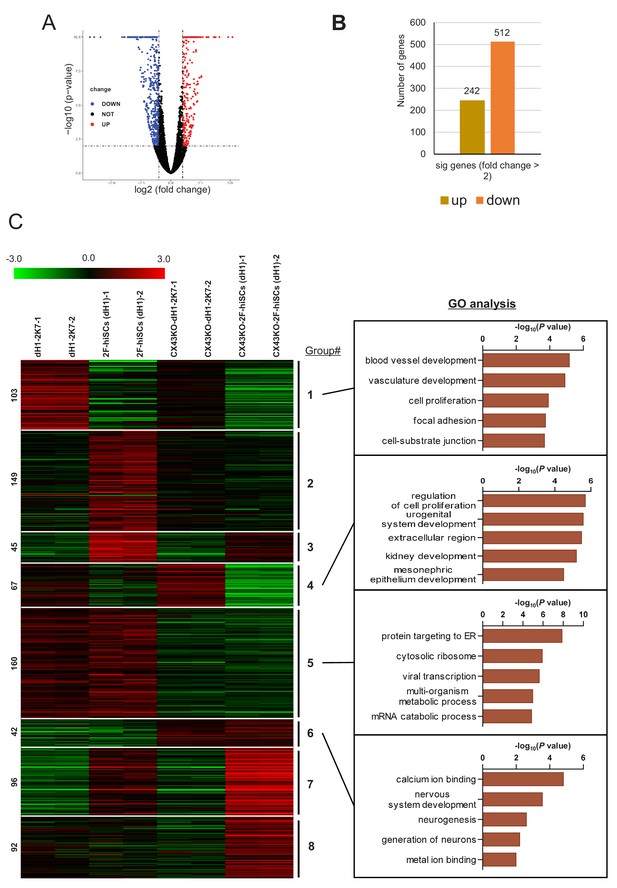
The effect of CX43 knock out on transcriptome of AMH-GFP cells.
(A) Volcano plot of DEGs between CX43 knock out 2F-hiSCs versus wild type 2F-hiSCs. Y axis represents -log10 of p-value, X axis represents log2 transformed fold change (FPKM, CXKO-2F-hiSCs/2F-hiSCs). Points in red represent significantly upregulated DEGs (fold change >2). Points in blue represent significantly downregulated DEGs (fold change <= 2). Black points represent non-DEGs. Cutoff for log2 (fold change) is 1. Cutoff for p-value is 0.01. (B) The number of significance DEGs (>2 folds) between CX43KO 2F-hiSCs and WT 2F-hiSCs. Brown bar indicates upregulated genes (242) and orange bar indicates downregulated genes (512). (C) Hierarchical clustering analysis using significance DEGs from (B) (>2 folds). Genes were classified into eight parts according to the expression pattern in CX43 knock out dH1, CX43 knock out 2F-hiSCs, wild type dH1 and wild type 2F-hiSCs. Gene Ontology analysis of genes in part 1, 4, 5, six was shown on the right, respectively.
Tables
| Reagent type (species) or resource | Designation | Source or reference | Identifiers | Additional information |
|---|---|---|---|---|
| Mouse | C57BL/6 | Vital River Laboratory Animal Technology | ||
| Cell line (Homo-sapiens) | 293FT cells | Thermo Fisher Scientific | Cat# R70007 | |
| Cell line (Homo-sapiens) | H1 ES cells | WiCell Research Institute | Cat# WA01 | |
| Cell line (Homo-sapiens) | Human Pulmonary Fibroblasts | National Infrastructure of Cell Line Resource | Cat# CCC-HPF-1 (PUMC, Beijing) | |
| Transfected construct (Homo-sapiens) | p2k7-EF1alpha-NR5A1 | This paper | Lentiviral construct to express target gene | |
| Transfected construct (Homo-sapiens) | p2k7-EF1alpha-GATA4 | This paper | Lentiviral construct to express target gene | |
| Transfected construct (Homo-sapiens) | p2k7-EF1alpha-SOX9 | This paper | Lentiviral construct to express target gene | |
| Transfected construct (Homo-sapiens) | p2k7-EF1alpha-WT1 | This paper | Lentiviral construct to express target gene | |
| Transfected construct (Homo-sapiens) | p2k7-EF1alpha-DMRT1 | This paper | Lentiviral construct to express target gene | |
| Transfected construct | p2k7-EF1alpha-luciferin | This paper | Lentiviral construct to express target gene | |
| Transfected construct (Homo-sapiens) | p2k7-EF1alpha-CX43 | This paper | Lentiviral construct to express target gene | |
| Recombinant DNA reagent | p2k7-AMH-GFP | This paper | Lentiviral construct for AMH reporter | |
| Recombinant DNA reagent | pX335-U6-Chimeric_BB-CBh-hSpCas9n(D10A) | Addgene #42335 | Expressing Cas9 and gRNA (Cong et al., 2013) | |
| Biological sample (Homo-sapiens) | Primary adult human Sertoli cells | Shanghai Jiao Tong University | Freshly isolated | |
| Antibody | anti-DAZL(Mouse polyclonal) | AbD Serotec | Cat#: MCA2336; RRID: AB_2292585 | IF(1:50) |
| Antibody | anti-NuMA (Rabbit polyclonal) | Abcam | Cat#: Ab97585; RRID: AB_GR27454-16 | IF(1:100) |
| Antibody | anti-human KRT18 (Rabbit polyclonal) | Proteintech | 10830–1-AP | IF(1:100) |
| Antibody | anti-SOX9 (Rabbit monoclonal) | Abcam | Cat#: Ab170660; RRID: AB_GR155689-1 | IF(1:100) |
| Antibody | anti-human CX43 (Rabbit polyclonal) | Cell Sigaling | Cat#: 3512S | IF(1:100) |
| Antibody | anti-human DMRT1 (Rabbit polyclonal) | Abcam | Cat#: Ab1786 | IF(1:100) |
| Antibody | anti-human WT1 (Rabbit polyclonal) | Proteintech | Cat#: 12609–1-AP | IF(1:100) |
| Antibody | anti-human GATA4 (Rabbit polyclonal) | Proteintech | Cat#: 19530–1-AP | IF(1:100) |
| Antibody | anti-human NR5A1 (Rabbit polyclonal) | Proteintech | Cat#: 18658–1-AP | IF(1:100) |
| Antibody | donkey anti-Mouse IgG (H+L) Highly Cross-Adsorbed Secondary Antibody, Alexa Fluor 488 | Invitrogen | Cat#: A-21202; RRID: AB_141607 | IF(1:1000) |
| Antibody | donkey anti-Mouse IgG (H+L) Highly Cross-Adsorbed Secondary Antibody, Alexa Fluor 555 | Invitrogen | Cat#: A-31572; RRID: AB_1567203 | IF(1:1000) |
| Strain, strain background (Escherichia coli) | TransStbl3 Chemically Competent Cell | TransGen Biotech | Cat#: CD521 | |
| Peptide, recombinant protein | bFGF, Recombinant Human FGF basic Protein | R and D Systems | Cat#: 233-FB-CF | |
| Chemical compound, drug | G418, Geneticin | Thermo Fisher Scientific | Cat#: 10131035 | |
| Chemical compound, drug | Blasticidin | Thermo Fisher Scientific | Cat#: R21002 | |
| Software, algorithm, website | ImageJ | NIH | https://imagej.nih.gov/ij/ | |
| Software, algorithm, website | FlowJo (v10.3) | BD | https://www.flowjo.com | |
| Software, algorithm, website | Prism7 (v7.0 a) | Graphpad Software | https://www.graphpad.com/scientific-software/prism/ | |
| Software, algorithm | DAVID Bioinformatics Resources (v6.8; GO) | https://david.ncifcrf.gov | ||
| Software, algorithm, website | Tophat2/cufflinks | (Kim et al., 2013) | http://ccb.jhu.edu/software/tophat | |
| Software, algorithm, website | R (v3.4.1; PCA, cluster and DEG) | https://www.R-project.org | ||
| Other | DAPI stain | Invitrogen | D1306 | (1 µg/mL) |





