The Caenorhabditis elegans Tubby homolog dynamically modulates olfactory cilia membrane morphogenesis and phospholipid composition
Figures
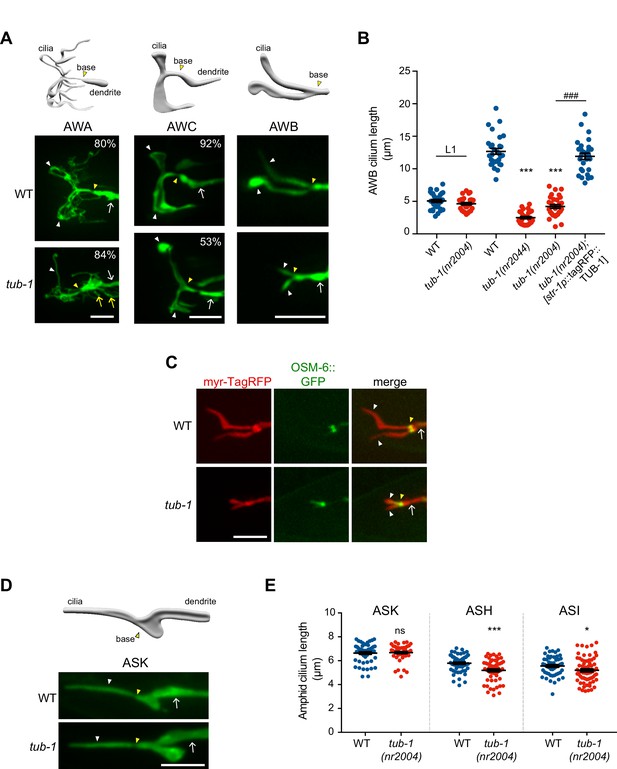
TUB-1 is necessary for membrane morphogenesis in wing cilia.
(A) Representative images of AWA, AWC and AWB cilia in wild-type and tub-1(nr2004) mutants. Cartoons of cilia morphologies are shown at top. Numbers at top right (in AWA and AWC image panels) indicate the percentage of neurons exhibiting the shown phenotype; n > 45 neurons each. (B) Quantification of total AWB cilia lengths in animals of the indicated genotypes. Animals were adult hermaphrodites unless indicated otherwise. The str-1 promoter drives expression primarily in AWB (Troemel et al., 1997). Each dot represents the combined AWB cilia lengths from a single neuron. *** and ### indicate different from wild-type or tub-1 mutant respectively, at the comparable developmental stage at p<0.001 (ANOVA with Tukey’s post-hoc test). (C) Representative images of wild-type and tub-1(nr2004) animals co-expressing the indicated fusion proteins. (D) (Left) Cartoon and representative images of ASK cilia in wild-type and tub-1(nr2004) mutants. (Right) Quantification of ASK, ASH, and ASI cilia lengths in adult hermaphrodites of the indicated genotypes. Each dot is cilia length from a single neuron. * and *** indicate different from wild-type at p<0.05 and 0.001, respectively (t-test); ns – not significant. In all images, yellow and white arrowheads indicate the cilia base and cilia, respectively; arrow indicates the dendrite. Scale bars: 5 μm. In all scatter plots, horizontal line is mean; error bars are SEM.
-
Figure 1—source data 1
Data for Figure 1B and E, and Figure 1—figure supplement 1.
- https://doi.org/10.7554/eLife.48789.004
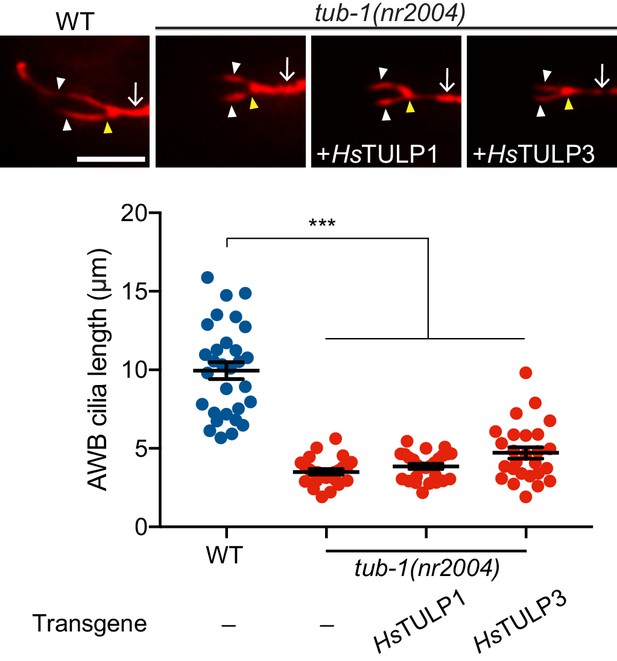
TUB-1 regulates AWB cilia morphology.
Representative images (top) and quantification (bottom) of AWB cilia lengths in animals of the indicated genotypes. Human (Hs - Homo sapiens) GFP::TULP1 and GFP::TULP3-encoding sequences were expressed in AWB under the str-1 promoter. Both proteins were expressed and localized to the AWB cilia base. The AWB neuron is marked via expression of str-1p::mCherry. Each dot represents measurements from a single AWB neuron. Horizontal line is mean; error bars are SEM. *** indicates different at p<0.001 between the indicated values (ANOVA and post-hoc Tukey’s test). Yellow and white arrowheads indicate the cilia base and cilia, respectively; arrow indicates the dendrite. Scale bars: 5 μm.
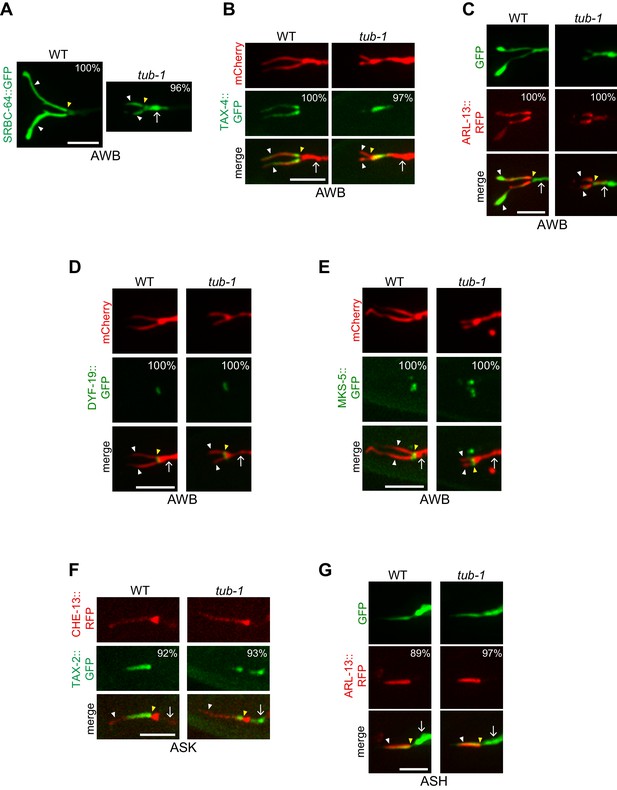
TUB-1 regulates ciliary targeting of transmembrane proteins in AWB.
(A–G) Representative images of localization patterns of the indicated fusion proteins in wild-type (WT) and tub-1(nr2004) mutants in AWB (A–E), ASK (F) and ASH (G). The AWB, ASK, and ASH neurons and/or cilia are marked via expression of str-1p::mCherry (B, D, E), str-1p::GFP (C), srbc-66p::CHE-13::RFP (F), or sra-6p::GFP (G). Numbers at top right indicate percentage of animals exhibiting the phenotype; n ≥ 20 neurons each. Localization patterns were analyzed only in animals expressing the fusion protein. Approximately similar percentages of wild-type and mutant animals expressed the fusion protein with the exception of animals expressing TAX-4::GFP (83% and 68% of wild-type and tub-1 mutants expressed TAX-4::GFP, respectively). Yellow and white arrowheads indicate the cilia base and cilia, respectively; arrow indicates the dendrite. Scale bars: 5 μm.
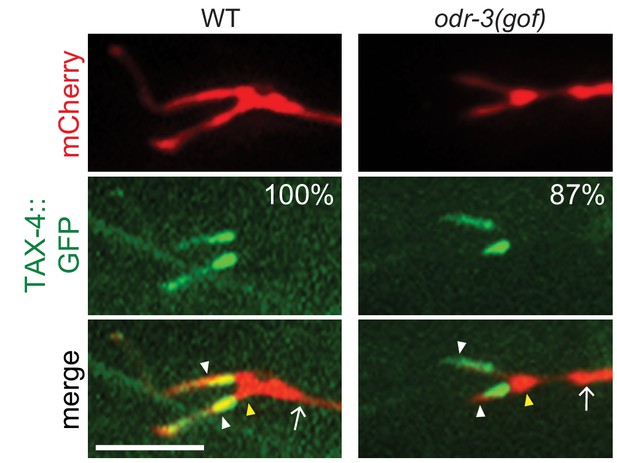
TAX-4 localization is unaltered in odr-3(gof) mutants.
Representative images of str-1p::mCherry and str-1p::TAX-4::GFP in AWB neurons of wild-type and odr-3(Q206L)XS (Roayaie et al., 1998) animals. The AWB neuron is marked via expression of str-1p::mCherry. Number at top right indicates the percentage of animals exhibiting the phenotype; n ≥ 29 neurons. Localization patterns were analyzed only in animals expressing the fusion protein. 83% and 30% of wild-type and odr-3(gof) animals expressed TAX-4::GFP, respectively. Yellow and white arrowheads indicate the cilia base and cilia, respectively; arrow indicates the dendrite. Scale bars: 5 μm.
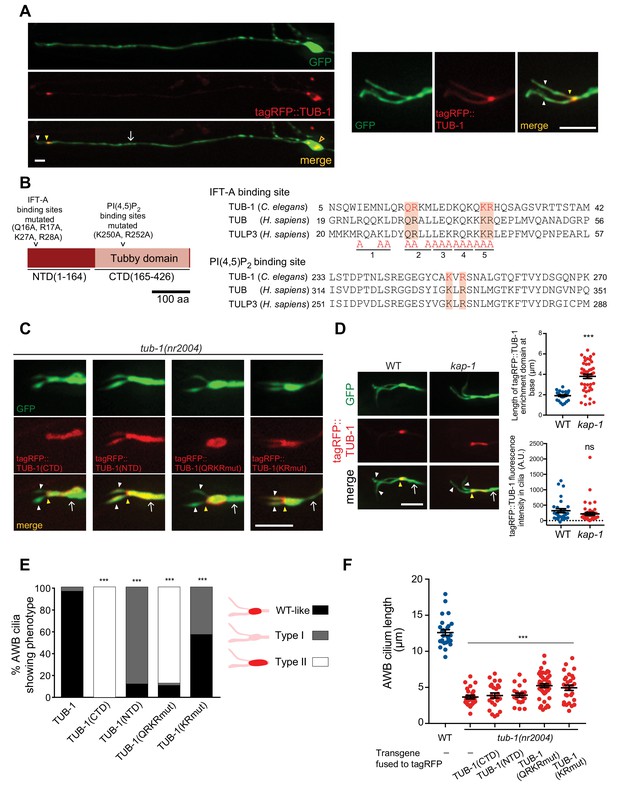
TUB-1 acts both at the AWB PCMC and within cilia to regulate AWB cilia morphology.
(A) Representative images of tagRFP::TUB-1 localization in AWB indicating cytoplasmic localization in the soma (open orange arrow) and enrichment at the dendritic ends (yellow arrowheads; left and right images). tagRFP::TUB-1 is also present at lower levels in AWB cilia (white arrowheads; left and right images). The AWB neurons were visualized via expression of str-1p::GFP. Scale bars: 5 μm. (B) (Left) Cartoon of TUB-1 domains. Residues mutated in the IFT-A binding site in the N-terminal domain (NTD) and PI(4,5)P2 binding site in the C-terminal Tubby domain (CTD) are indicated. (Right) Alignment of the predicted IFT-A binding domains in C. elegans TUB-1 and human TUB and TULP3 proteins. Residues in TULP3 which when replaced with Alanine abolishes IFT-A binding are shown; shown mutations in any of the five underlined regions abolish binding (Mukhopadhyay et al., 2010). Red shading indicates conserved residues mutated in this work. (C) Representative images of the indicated fusion proteins in tub-1(nr2004) mutants in AWB. AWB was visualized via expression of str-1p::GFP. (D) Representative images (left) and quantification of tagRFP::TUB-1 localization at the cilia base (top right) and cilia (bottom right) in wild-type and kap-1(ok676) mutants. The AWB neurons were visualized via expression of str-1p::GFP. *** indicates different from wild-type at p<0.001 (t-test); ns – not significant. (E) Percentage of AWB cilia exhibiting the indicated localization patterns of the shown fusion proteins in tub-1(nr2004) animals. Proteins were visualized via fusion with tagRFP. n ≥ 20 neurons each. *** indicates different from wild-type at p<0.001 (Fisher’s exact test). (F) Quantification of total AWB cilia lengths in wild-type or tub-1(nr2004) animals expressing the indicated fusion proteins. *** indicates different from wild-type at p<0.001 (ANOVA with Tukey’s post-hoc test). In all images, yellow and white arrowheads indicate the cilia base and cilia, respectively; arrow indicates the dendrite. Scale bars: 5 μm. In all scatter plots, each dot represents measurements from a single AWB neuron. Horizontal line is mean; error bars are SEM.
-
Figure 3—source data 1
Data for Figure 3D,E,F, and Figure 3—figure supplement 1A.
- https://doi.org/10.7554/eLife.48789.009
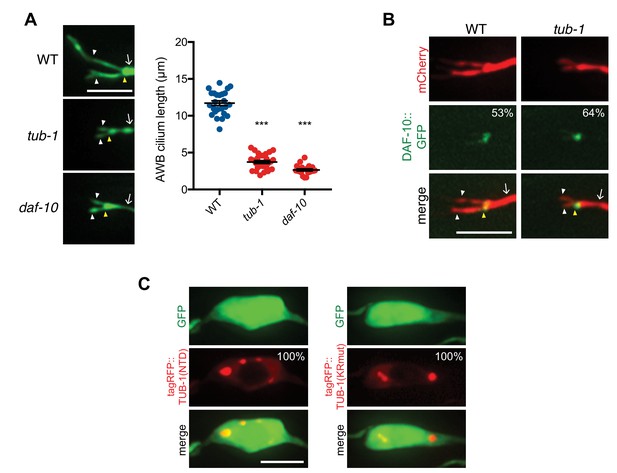
tub-1 shares a subset of phenotypes with daf-10.
(A) Representative images (left) and quantification of AWB cilia lengths (right) in animals of the indicated genotypes. Alleles used were tub-1(nr2004) and daf-10(e1387). *** indicates different from wild-type at p<0.001 (ANOVA with Tukey’s post-hoc test). Each dot in the scatter plot represents measurements from a single AWB neuron. Horizontal line is mean; error bars are SEM. (B) Representative images of DAF-10::GFP localization in AWB cilia in wild-type and tub-1(nr2004) mutants. Localization patterns were analyzed only in animals expressing the fusion protein. Approximately similar percentages of wild-type and tub-1 animals expressed DAF-10::GFP; n ≥ 39 neurons each. (C) Nuclear exclusion of the indicated fusion proteins in AWB soma. AWB neurons were visualized via expression of str-1p::GFP; n ≥ 36 neurons each. In all images, yellow and white arrowheads indicate the cilia base and cilia, respectively; arrow indicates the dendrite. Scale bars: 5 μm.
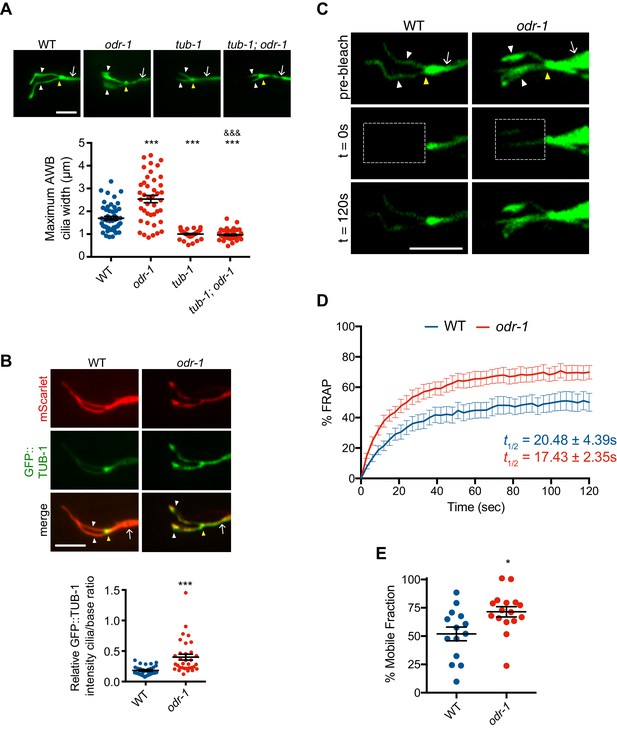
Reduced sensory signaling increases TUB-1 localization within AWB cilia.
(A) (Top) Representative images of AWB cilia morphologies in animals of the indicated genotypes. Alleles used were tub-1(nr2004) and odr-1(n1936). The AWB neurons were visualized via expression of str-1p::myr-GFP. (Bottom) Quantification of maximum AWB cilia widths. *** and &&& indicate different from wild-type and odr-1, respectively, at p<0.001 (ANOVA and post-hoc Tukey’s test). (B) (Top) Representative images of GFP::TUB-1 localization in wild-type or odr-1(n1936) mutants. The AWB neurons were visualized via expression of str-1p::GFP::TUB-1::SL2::mScarlet. (Bottom) Quantification of relative GFP::TUB-1 fluorescence intensity within cilia vs the cilia base. *** indicates different from wild-type at p<0.001 (t-test). (C) Representative images of AWB expressing GFP::TUB-1 pre-bleach, and at 0 s and 120 s post-bleach. The bleached area is indicated by a dotted box. (D) Mean fluorescence recovery following photobleaching normalized to pre-photobleached intensity at 0 s in wild-type and odr-1(n1936) animals. Errors are SEM. n ≥ 14 animals; three independent experiments. (E) Percent mobile fraction of GFP::TUB-1 in AWB cilia in wild-type and odr-1 animals calculated from data shown in D. * indicates different from wild-type at p<0.05 (t-test). In all images, yellow and white arrowheads indicate the cilia base and cilia, respectively; arrow indicates the dendrite. Scale bars: 5 μm. In all scatter plots, each dot represents measurements from a single AWB neuron. Horizontal line is mean; error bars are SEM.
-
Figure 4—source data 1
Data for Figure 4A,B,D,E, and Figure 4—figure supplement 1.
- https://doi.org/10.7554/eLife.48789.012
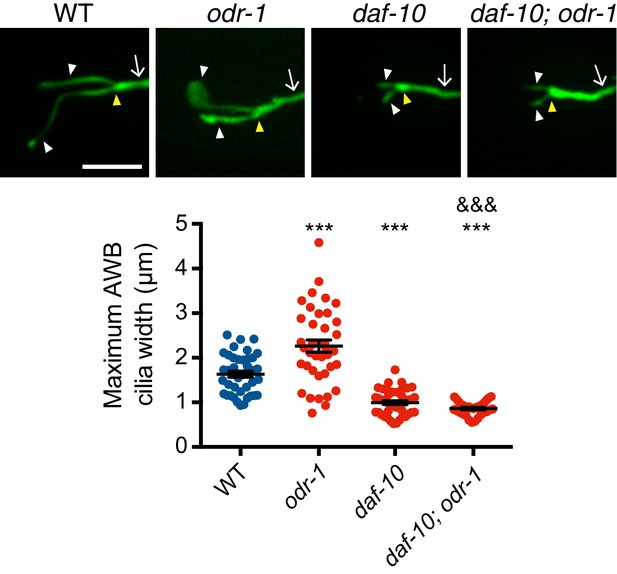
Loss-of-function mutations in daf-10 suppress the expanded ‘fan’ phenotype of odr-1 mutants.
Representative images (top) and quantification of maximum AWB cilia widths in animals of the indicated genotypes. *** and &&& indicate different from wild-type and odr-1, respectively, at p<0.001 (ANOVA with Tukey’s post-hoc test). Yellow and white arrowheads indicate the cilia base and cilia, respectively; arrow indicates the dendrite. Scale bars: 5 μm. Each dot in the scatter plot represents measurements from a single AWB neuron. Horizontal line is mean; error bars are SEM.
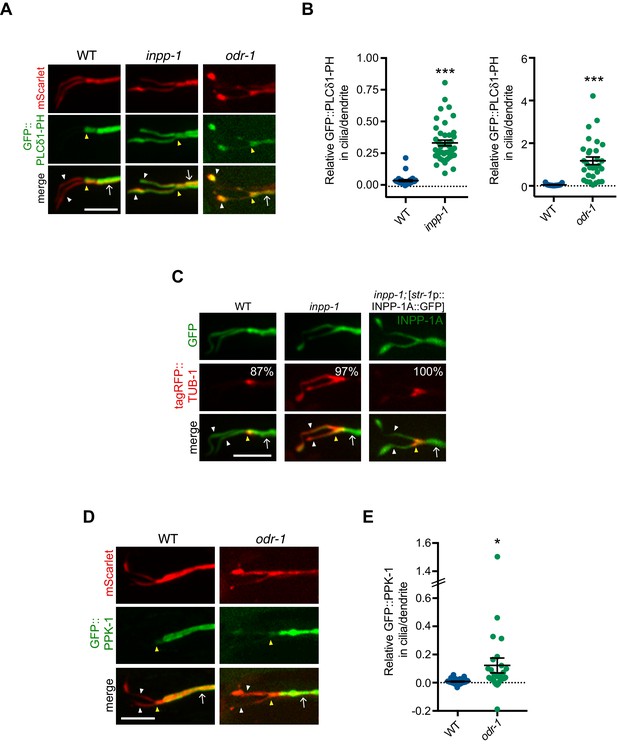
Levels of PI(4,5)P2 and PPK-1 are increased within AWB cilia in odr-1 mutants.
(A, C, D) Representative images of GFP::PLCδ1-PH (A), tagRFP::TUB-1 (C) and GFP::PPK-1 (D) localization in AWB cilia of animals of the indicated genotypes. Alleles used were odr-1(n1936) and inpp-1(gk3262). AWB neurons were visualized via expression of str-1p::GFP::PLCδ1-PH::SL2::mScarlet (A), str-1p::GFP (C), str-1p::INPP-1A::GFP (C, right column) or GFP::PPK-1::SL2::mScarlet (D). Numbers at top right in C indicate percentage of animals exhibiting the phenotype; n ≥ 23 neurons. Yellow and white arrowheads indicate the cilia base and cilia, respectively; arrow indicates the dendrite. Scale bars: 5 μm. (B, E) Quantification of relative GFP::PLCδ1-PH (B) and GFP::PPK-1 (E) fluorescence intensities in AWB cilia vs dendrites in animals of the indicated genotypes. * and *** indicate different from wild-type at p<0.05 and p<0.001, respectively (t-test). Each dot represents measurements from a single AWB neuron. Horizontal line is mean; error bars are SEM.
-
Figure 5—source data 1
Data for Figures 5B and E, and Figure 5—figure supplement 1.
- https://doi.org/10.7554/eLife.48789.015
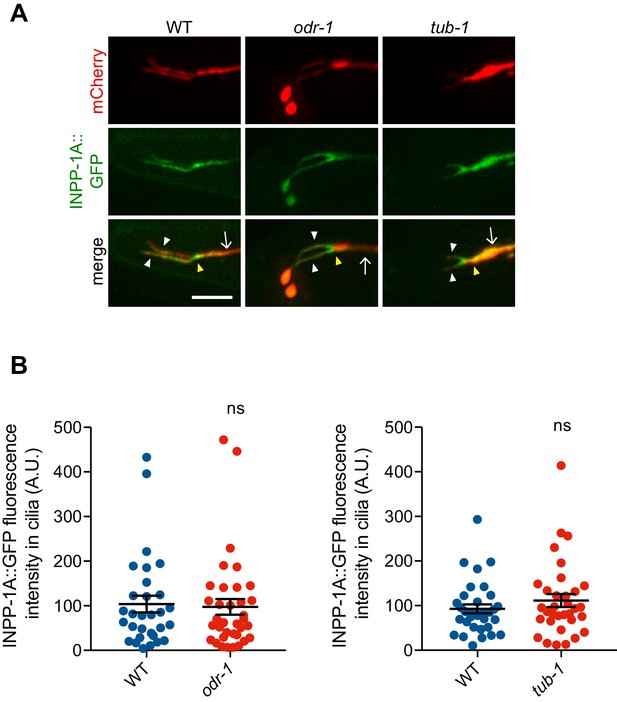
INPP-1 localization is unaltered in tub-1 and odr-1 mutants.
(A) Representative images of INPP-1A::GFP localization in AWB cilia of animals of the indicated genotypes. Yellow and white arrowheads indicate the cilia base and cilia, respectively; arrow indicates the dendrite. Scale bar: 5 μm. Alleles used were odr-1(n1936) and tub-1(nr2004). (B) Quantification of INPP-1A::GFP intensity in AWB cilia in animals of the indicated genotypes. Each dot represents measurements from a single AWB neuron. Horizontal line is mean; error bars are SEM.
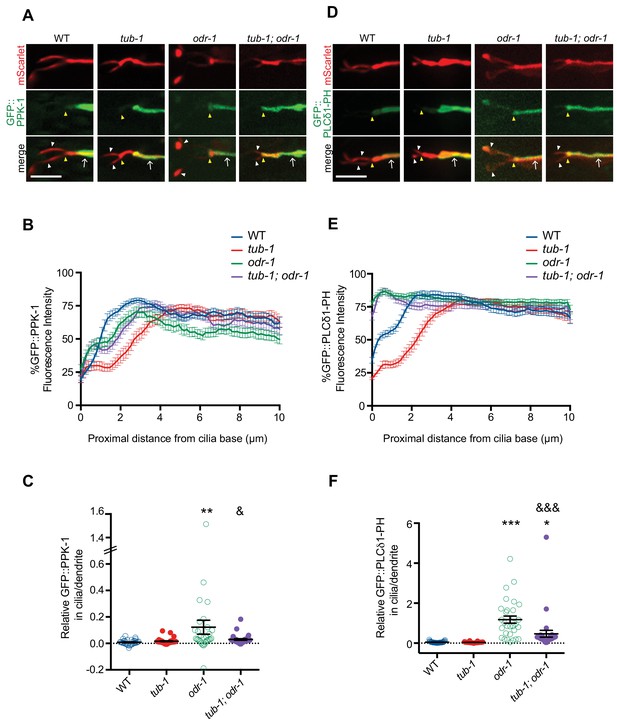
TUB-1 regulates PPK-1 localization and PI(4,5)P2 distribution in AWB dendrites and cilia.
(A, D) Representative images of GFP::PPK-1 (A) and GFP::PLCδ1-PH (D) localization in animals of the indicated genotypes. Alleles used were tub-1(nr2004) and odr-1(n1936). The AWB neuron is marked via expression of str-1p::GFP::PPK-1::SL2::mScarlet or GFP::PLCδ1-PH::SL2::mScarlet (A, D, respectively). Yellow and white arrowheads indicate the cilia base and cilia, respectively; arrow indicates the dendrite. Scale bars: 5 μm. (B, E) Line scans of GFP::PPK-1 (B) and GFP::PLCδ1-PH (E) intensities in AWB dendrites of animals of the indicated genotypes. Zero indicates the cilia base/dendritic tip. Fluorescence intensities were normalized to the maximum intensity for each individual animal across the measured region, and the percent of this maximum intensity was calculated at each location to create individual line scans. Error bars are the SEM; n ≥ 30 neurons each. (C, F) Relative fluorescence intensities of GFP::PPK-1 (C) and GFP::PLCδ1-PH (F) in AWB cilia vs dendrites in animals of the indicated genotypes. Open circles indicate data repeated from Figure 5. *, ** and *** indicate different from wild-type at p<0.05, p<0.01, and p<0.001; & and &&& indicate different from odr-1 at p<0.05 and p<0.001, (ANOVA and post-hoc Tukey’s test). Each dot represents measurements from a single AWB neuron. Horizontal line is mean; error bars are SEM.
-
Figure 6—source data 1
Data for Figure 6B,C,E,F, Figure 6—figure supplement 1, and Figure 6—figure supplement 2B and C.
- https://doi.org/10.7554/eLife.48789.019
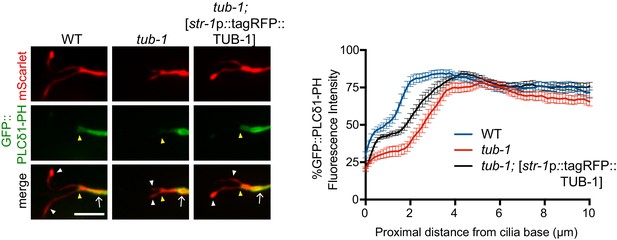
TUB-1 may act cell-autonomously to regulate GFP::PLCδ1-PH distribution in AWB.
Representative images (left) and line scans (right) of GFP::PLCδ1-PH distribution in AWB cilia and dendrites in animals of the indicated genotypes. Yellow and white arrowheads indicate the cilia base and cilia, respectively; arrow indicates the dendrite. Scale bar: 5 μm. The str-1 promoter drives expression primarily in AWB. Line scans of GFP::PLCδ1-PH intensity in AWB dendrites of animals of the indicated genotypes. Zero indicates the cilia base. Error bars are the SEM; n ≥ 27 neurons each.
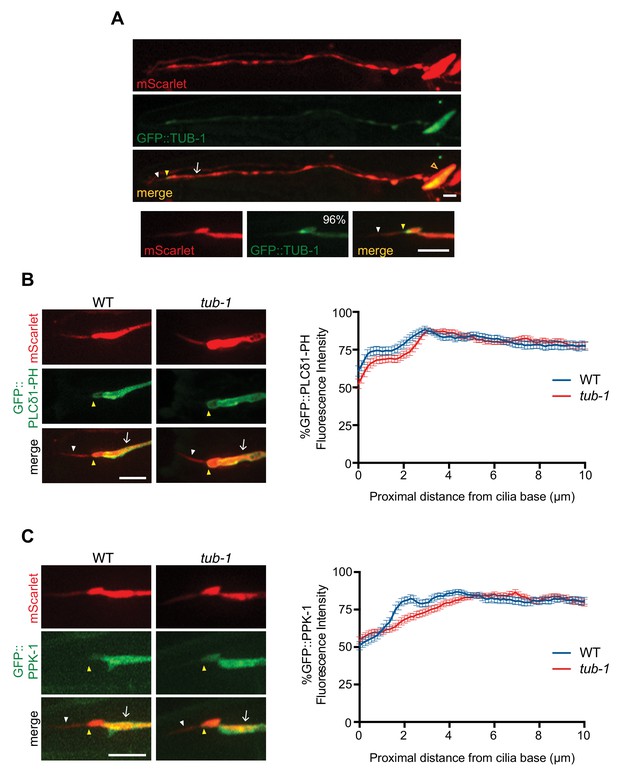
TUB-1 does not regulate PPK-1 and PI(4,5)P2 distribution in ASK dendrites and cilia.
(A) Representative images of GFP::TUB-1 localization in the ASK neurons and enrichment at the dendritic ends (yellow arrowheads). ASK soma is indicated by an open orange arrowhead. GFP::TUB-1 is also present at lower levels in ASK cilia (white arrowheads). Number at top right indicates the percentage of animals exhibiting the phenotype; n = 52 neurons. Scale bar: 5 μm. (B, C) (Left) Representative images of GFP::PLCδ1-PH and GFP::PPK-1 localization in ASK neurons of wild-type and tub-1(nr2004) mutants. Yellow and white arrowheads indicate the cilia base and cilia, respectively; arrow indicates the dendrite. Scale bar: 5 μm. (Right) Line scans of GFP::PLCδ1-PH (B) GFP::PPK-1 (C) intensities in ASK dendrites of animals of the indicated genotypes. Zero indicates the cilia base. Error bars are the SEM; n ≥ 30 neurons each.
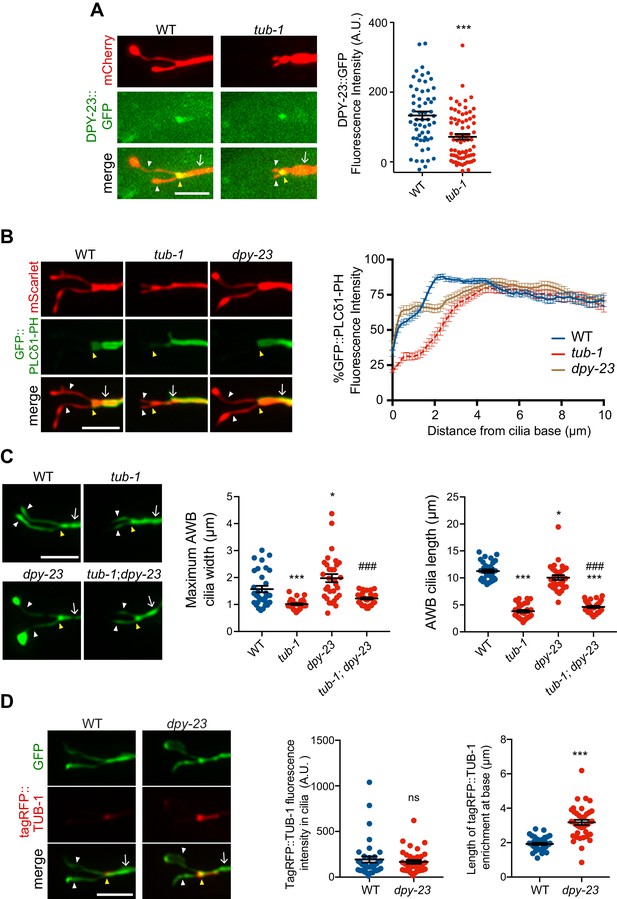
TUB-1 acts in part via the DPY-23 AP-2 μ subunit to localize PPK-1 at the PCMC.
(A, B) (Left) Representative images of DPY-23::GFP (A) and GFP::PLCδ1-PH (B) localization in AWB cilia of animals of the indicated genotypes. AWB cilia were visualized via expression of str-1p::mCherry (A) or of str-1p::GFP::PLCδ1-PH::SL2::mScarlet (B). (A, Right) Quantification of DPY-23::GFP intensity in a 1 μm2 area at the cilia base in A. *** indicates different from wild-type at p<0.001 (t-test). (B, right) Line scans of GFP::PLCδ1-PH intensity in AWB dendrites of animals of the indicated genotypes. Zero indicates the cilia base/dendritic tip. Dashed lines indicate data repeated from Figure 6. Error bars are the SEM; n ≥ 29 neurons each. Alleles used were tub-1(nr2004) and dpy-23(e840). (C) (Left) Representative images of AWB cilia morphology in animals of the indicated genotypes. AWB cilia were visualized via expression of str-1p::gfp. (Middle) Quantification of total AWB cilia lengths and (right) maximum cilia widths of individual animals of the indicated genotypes. *** and ### indicate different from wild-type and dpy-23(e840) mutants, respectively, at p<0.001 (ANOVA with Tukey’s post-hoc test). (D) (Left) Representative images of tagRFP::TUB-1 localization in AWB cilia of wild-type and dpy-23(e840) animals. AWB cilia were visualized via expression of str-1p::gfp. (Middle) Quantification of tagRFP::TUB-1 intensity within cilia, and (right) localization at the AWB cilia base in wild-type and dpy-23(e840) animals. *** indicates different from wild-type at p<0.001, ns – not significant (t- test). In all images, yellow and white arrowheads indicate the cilia base and cilia, respectively; arrow indicates the dendrite. Scale bars: 5 μm. In all scatter plots, each dot represents measurements from a single AWB neuron. Horizontal line is mean; error bars are SEM.
-
Figure 7—source data 1
Data for Figure 7A–D and Figure 7—figure supplement 1A–C.
- https://doi.org/10.7554/eLife.48789.022
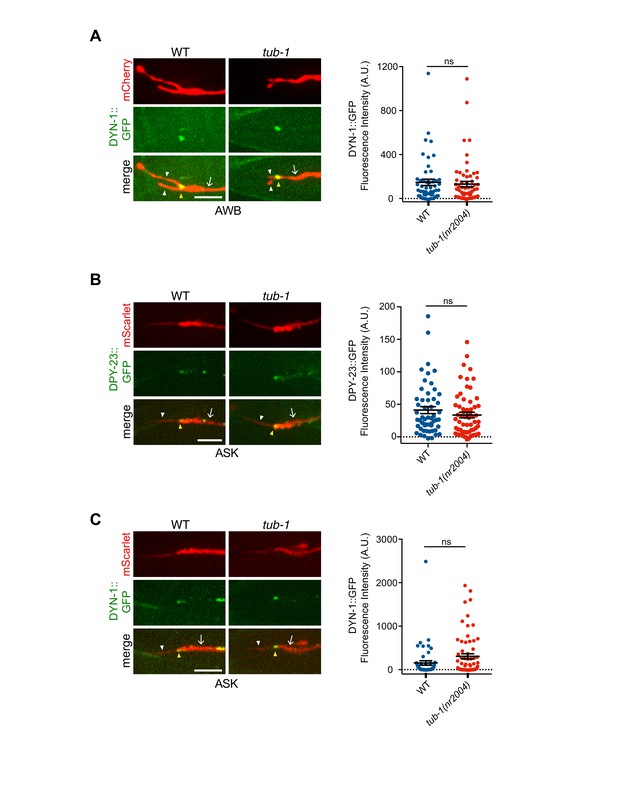
Localization patterns of endocytic proteins in AWB and ASK in tub-1 mutants.
(A–C) Representative images (left) and quantification of fluorescence intensities (right) of the indicated proteins at the PCMCs of AWB (A) and ASK (B and C). (Left images) Neurons are marked via expression of str-1p::mCherry (A) or sra-9p::mScarlet (B, C). In ASK, DPY-23::GFP and DYN-1::GFP puncta are also observed in the dendrites in addition to the PCMC. In all images, yellow and white arrowheads indicate the cilia base and cilia, respectively; arrow indicates the dendrite. Scale bars: 5 μm. In all scatter plots, each dot represents measurements from a single neuron. Horizontal line is mean; error bars are SEM.
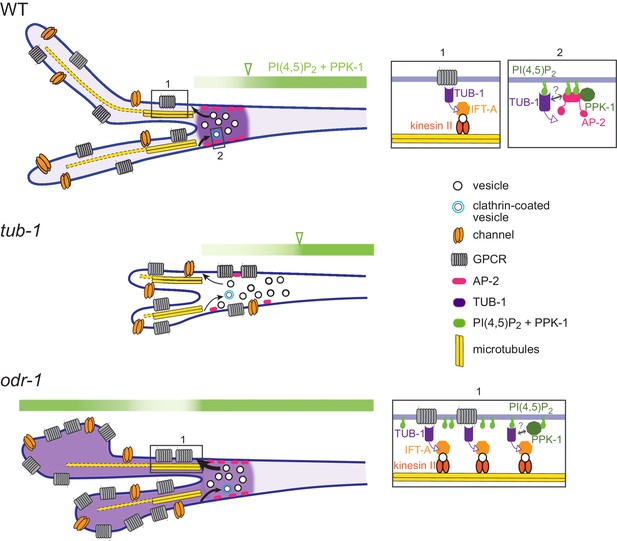
Model for TUB-1 function in regulating AWB olfactory cilia properties.
Cartoons of proposed TUB-1 functions at the PCMC and within AWB cilia. Dotted yellow bars indicate singlet microtubules present in the distal segments of AWB cilia; the origins of these singlet microtubules are unclear (Doroquez et al., 2014). Numbered boxes in cartoons at left correspond to expanded diagrams at right. Open green arrowheads indicate the distal boundary of enrichment of PI(4,5)P2 and PPK-1 in AWB dendrites. See text for additional details.
Additional files
-
Supplementary file 1
List of strains used in this work.
- https://doi.org/10.7554/eLife.48789.024
-
Supplementary file 2
List of plasmids used in this work.
- https://doi.org/10.7554/eLife.48789.025
-
Transparent reporting form
- https://doi.org/10.7554/eLife.48789.026





