ENaC-mediated sodium influx exacerbates NLRP3-dependent inflammation in cystic fibrosis
Figures
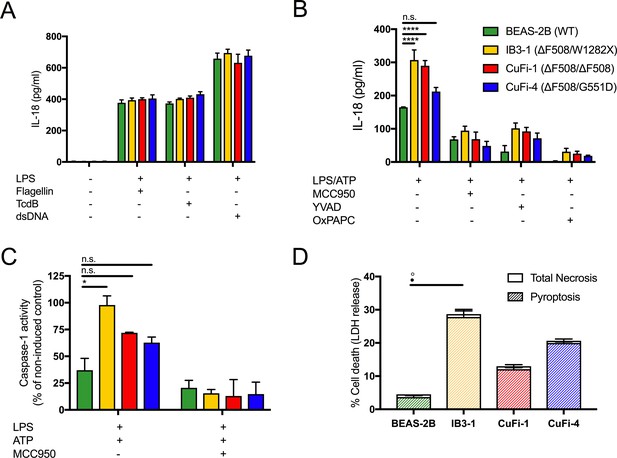
LPS-induced IL-18 secretion in human bronchial epithelial cells is higher in cells with CF-associated mutations and is NLRP3 inflammasome dependent.
Human bronchial epithelial cell (HBEC) lines (BEAS-2B (WT), IB3-1 (ΔF508/W1282X), CuFi1 (ΔF508/ ΔF508), CuFi4 (ΔF508/G551D) (n = 3 independent experiments) were unstimulated or stimulated with Lipopolysaccharide, from Escherichia coli K12 (LPS Ultrapure), which specifically targets TLR4 (10 ng/mL) for 4 hr before being stimulated for 4 hr with Flagellin (10 ng/mL with Lipofectamine 2000) for NLRC4 inflammasome, TcdB (10 ng/mL) for Pyrin inflammasome or poly(dA:dT) dsDNA (1 μg/mL with Lipofectamine 2000) for AIM2 inflammasome. ELISA assays were used to detect (A) IL-18. To monitor NLRP3 inflammasome activation, HBEC (n = 3 independent experiments) were pre-incubated with MCC950 (15 μM), OxPAPC (30 μg/mL) and YVAD (2 μg/mL) for 1 hr before a stimulation with LPS (10 ng/mL, 4 hr), and ATP (5 mM) for the final 30 min. ELISA assays were used to detect (B) IL-18 and (D) colourimetric assay used to detect caspase-1 activity in protein lysates for LPS/ATP and LPS/ATP/MCC950). (D) Necrosis and pyroptosis are represented as superimposed bar charts. Total necrosis was measured using LDH release assay. For pyroptotic cell death, each sample/condition was repeated in parallel with a caspase-1 inhibitor (YVAD (2 mg/mL, 1 hr)) pre-treatment. The total necrosis level was then taken away from the caspase-1 inhibited sample, or ‘caspase-1 independent’ necrosis, with the remaining LDH level termed ‘caspase-1 dependent necrosis’ or pyroptosis. Cells were then stimulated with LPS (10 ng/mL, 4 hr), and ATP (5 mM) for final 30 min. The assay was performed with HBEC lines (n = 3 independent experiments). (◦) Significance for Total Necrosis (●) Significance for Pyroptosis. A 2-way ANOVA with Tukey’s multiple comparison test was performed (p values * =< 0.05, ** =< 0.01, *** =< 0.001 and **** =< 0.0001).
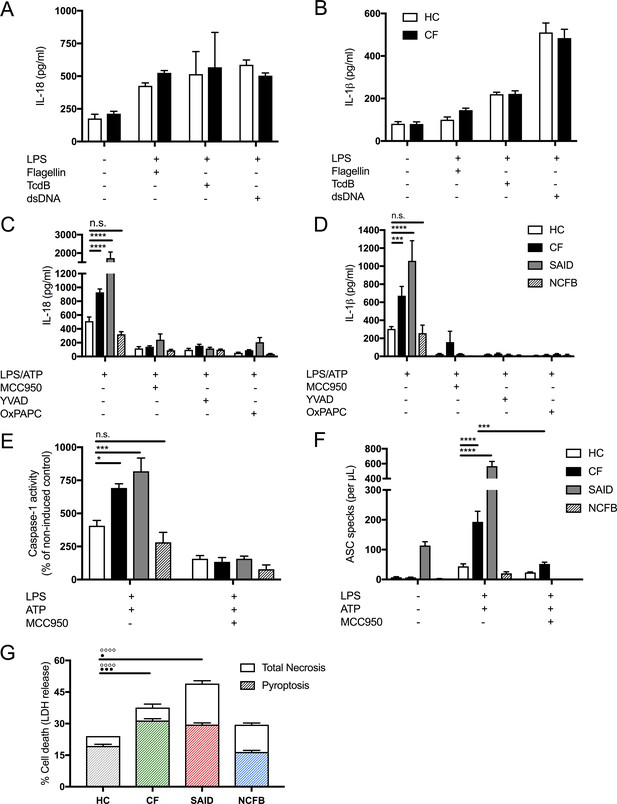
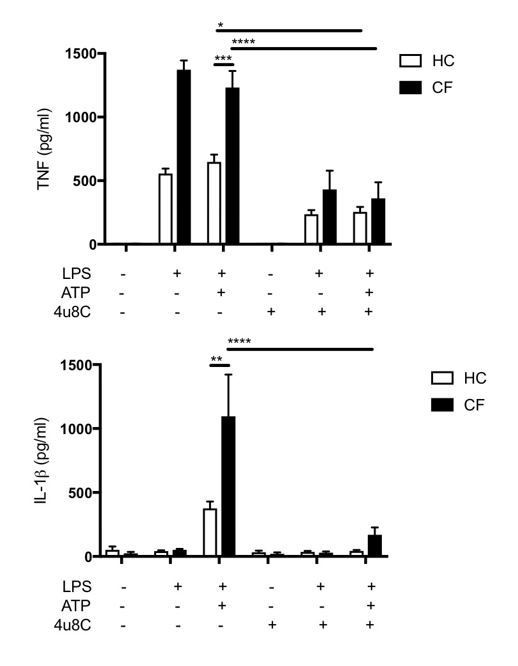
LPS-induced IL-1β/IL-18 secretion in human monocytes is higher in CF and is NLRP3 inflammasome dependent.
Primary monocytes from HC and CF (HC, n = 10; CF, n = 10) were unstimulated or stimulated with LPS which specifically targets TLR4 (10 ng/mL) for 4 hr before being stimulated for 4 hr with Flagellin (10 ng/mL with Lipofectamine 2000) for NLRC4 inflammasome, or TcdB (10 ng/mL) for Pyrin inflammasome or poly(dA:dT) dsDNA (1 μg/mL with Lipofectamine 2000) for AIM2 inflammasome. ELISA assays were used to detect (A) IL-18 and (B) IL-1β cytokine secretion in supernatants. To monitor NLRP3 inflammasome activation, primary monocytes from HC, CF, SAID and NCFB (HC, n = 10; CF, n = 10; SAID, n = 4; NCFB, n = 4) were pre-incubated with MCC950 (15 μM), OxPAPC (30 μg/mL) and YVAD (2 μg/mL) for 1 hr before a stimulation with LPS (10 ng/mL, 4 hr), and ATP (5 mM) for the final 30 min. ELISA assays were used to detect (C) IL-18 and (D) IL-1β cytokine secretion in supernatants and (E) a colourimetric assay was used to detect caspase-1 activity in protein lysates (HC, n = 10; CF, n = 10; SAID, n = 4; NCFB, n = 4). (F) Flow cytometry was used to detect ASC specks in supernatants of primary monocytes from HC, CF, SAID and NCFB (HC, n = 10; CF, n = 10; SAID, n = 6; NCFB, n = 4) for ±LPS/ATP and (HC, n = 5; CF, n = 5) for MCC950 with LPS/ATP. (G) Necrosis and pyroptosis are represented as superimposed bar charts. Total necrosis was measured using LDH release assay. For pyroptotic cell death, each sample/condition was repeated in parallel with a caspase-1 inhibitor (YVAD (2 mg/mL, 1 hr)) pre-treatment. The total necrosis level was taken away from the caspase-1 inhibited sample, or ‘caspase-1 independent’ necrosis, with the remaining LDH level termed ‘caspase-1 dependent necrosis’ or pyroptosis. Cells were then stimulated with LPS (10 ng/mL, 4 hr), and ATP (5 mM) for final 30 min. The assay was performed with primary monocytes from HC, CF, SAID and NCFB (HC, n = 10; CF, n = 10; SAID, n = 4; NCFB, n = 4). (◦) Significance for Total Necrosis (●) Significance for pyroptosis. A 2-way ANOVA statistical test was performed, with Tukey post-hoc correction (p values * =< 0.05, ** =< 0.01, *** =< 0.001 and **** =< 0.0001; error bars ± SEM). Inhibitor treatments in panels a-c were found to significantly reduce cytokine secretion and caspase-1 activity to **p =< 0.01 or less, for CF and SAID groups respectively. Significance values not displayed on the graph.
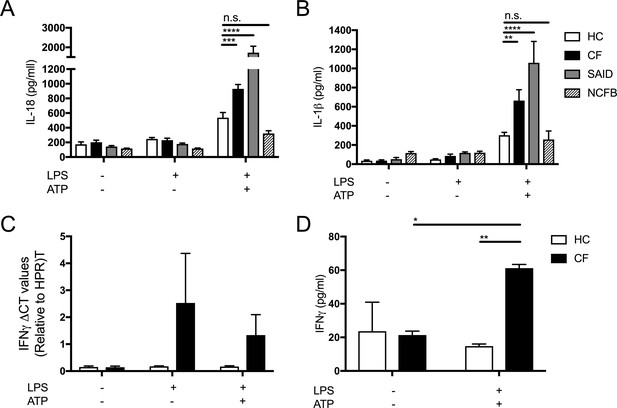
Primary monocytes from HC, CF, SAID and NCFB (HC n = 9, CF n = 9, SAID n = 4, NCFB n = 4) were unstimulated or stimulated with LPS (10 ng/ml, 4 hr) or LPS (10 ng/ml, 4 hr) and ATP (5 mM) for the final 30 min.
ELISAs were used to detect (A) IL-18, and (B) IL-1β cytokine secretion in supernatants. (C) PBMCs were unstimulated or stimulated with LPS (10 ng/ml, 4 hr) or LPS (10 ng/ml, 4 hr) and ATP (5 mM) for the final 30 min. Taqman RT-qPCR was used to measure IFNγ gene expression and (D) Luminex was used to measure IFNγ secretion from peripheral blood mononuclear cells (PBMC) populations from HCs and patients with CF-associated mutations (n = 10).
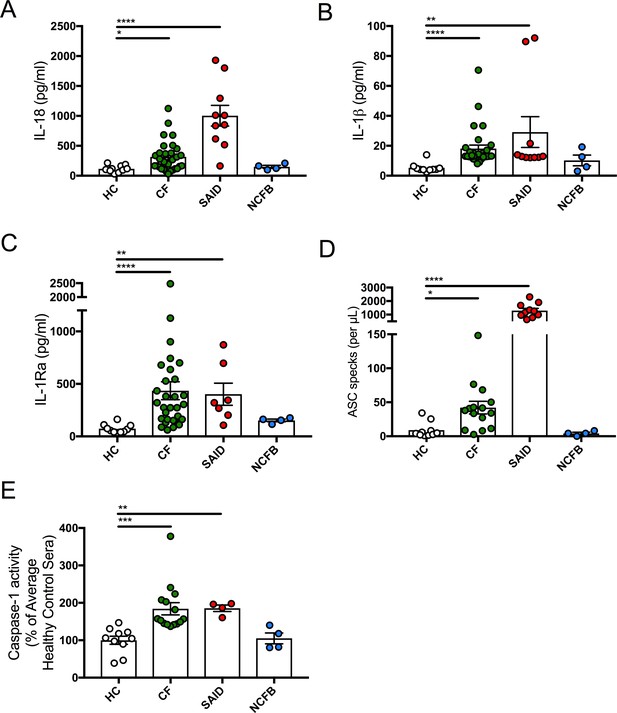
Inflammatory serum cytokine signature in CF.
(A) ELISA assays were used to detect IL-18 (HC, n = 10, CF, n = 30, SAID, n = 10, NCFB, n = 4), (B) IL-1β (HC, n = 10, CF, n = 30, SAID, n = 10, NCFB, n = 4), (C) IL-1Ra (HC, n = 10, CF, n = 30, SAID, n = 7, NCFB, n = 4) in patient sera. Outliers in SAID group for IL-1β and IL-1Ra correspond to HIDS one and A20 deficiency (D) Flow cytometry was used to detect ASC specks (HC, n = 10, CF, n = 15, SAID, n = 10, NCFB, n = 4) in patient sera. (E) A colorimetric assay to detect caspase-1 activity in sera of patients with CF, SAID and NCFB as a percentage of HC (HC, n = 10, CF, n = 15, SAID, n = 4, NCFB, n = 4). Of note, an undetermined amount of detected IL-1Ra is attributed to circulating Anakinra (recombinant IL-1Ra) specifically in the SAID cohort. The Kruskal-Wallis non-parametric test, with Dunn’s multiple comparison test, was performed (p values * =< 0.05, ** =< 0.01, *** =< 0.001 and **** =< 0.0001; error bars ± S.E.M).
-
Figure 3—source data 1
Serum cytokine levels for all patient groups.
- https://doi.org/10.7554/eLife.49248.008
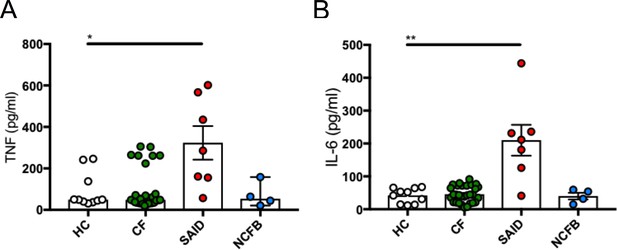
Inflammatory serum cytokine signature in CF.
ELISA assays were used to detect (A) TNF and (B) IL-6 (HC n = 10, CF n = 30, SAID n = 7, NCFB = 4) in patient serum. The Kruskal-Wallis non-parametric test, with Dunn’s multiple comparison test, was performed (p values * = 0.05, ** = 0.01, *** = 0.001 and **** = 0.0001; error bars ± S.E.M).
-
Figure 3—figure supplement 1—source data 1
Serum cytokine levels for all patient groups.
- https://doi.org/10.7554/eLife.49248.007
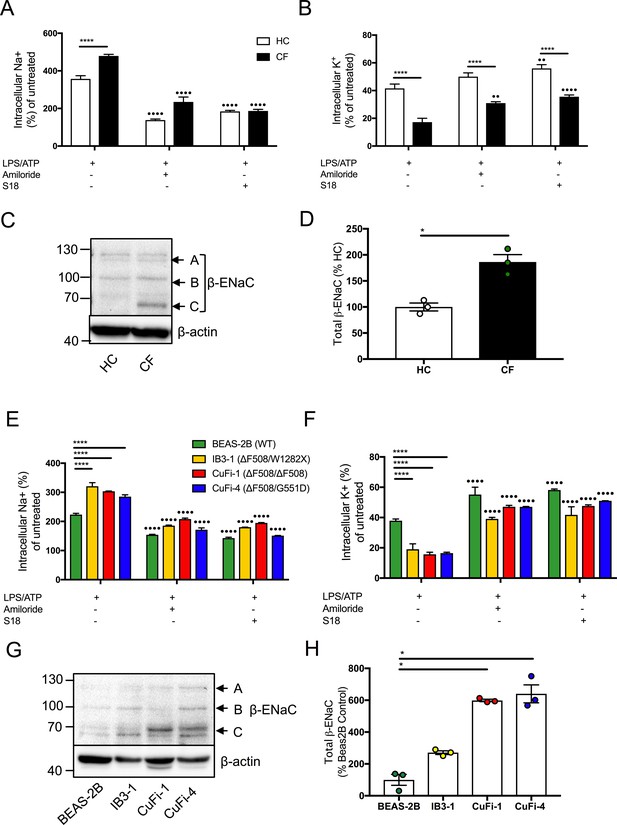
Dysregulated Na+ and K+ in cells with CF-associated mutations can be modulated with ENaC inhibitors.
Intracellular Na+ was detected using an AM ester of sodium indictor SBFI (S-1263) and (B, D) intracellular K+ was detected using an AM ester of potassium indictor PBFI (P-1266); changes in fluorescence were measured by fluorimeter post-stimulation with 5 mM ATP in (A, B) monocytes (HC = 7, CF = 7) (E, F) HBECs (n = 3 independent experiments). Cells were pre-treated with the following: amiloride (100 μM), S18 derived peptide (25 μM, 4 hr) with LPS (10 ng/mL, 4 hr) and ATP (5 mM) for the final 30 min. A 2-way ANOVA with Tukey’s multiple comparison test was performed (p values * =< 0.05, ** =< 0.01, *** =< 0.001 and **** =< 0.0001) (*) indicate significance when comparing HC with CF-associated mutants. (•) indicate significance between treatments within the same cell line. (C) Endogenous β-ENaC protein expression was detected using western blot in BEAS-2B HBEC, HC and CF monocytes (C) and densitometry analysis of total β-ENaC (bands A, B, C indicated on blot) was quantified in (D) for CF relative to HC (n = 3 independent experiments). (G) BEAS-2B, IB3-1, CuFi-1 and CuFi-4 HBEC lines and densitometry analysis of total β−ENaC (bands A, B, C indicated on blot) was quantified in (H) (n = 3 independent experiments). Band A represents complex N-Glycosylation, 110 kDa β-ENaC (found when associated as ENaC complex); Band B represents Endo-H sensitive N-Glycosylation, 96 kDa β-ENaC; Band C represents immature non-glycosylated, 66 kDa β−ENaC. The Mann-Whitney non-parametric test was performed (p values * =≤ 0.05).
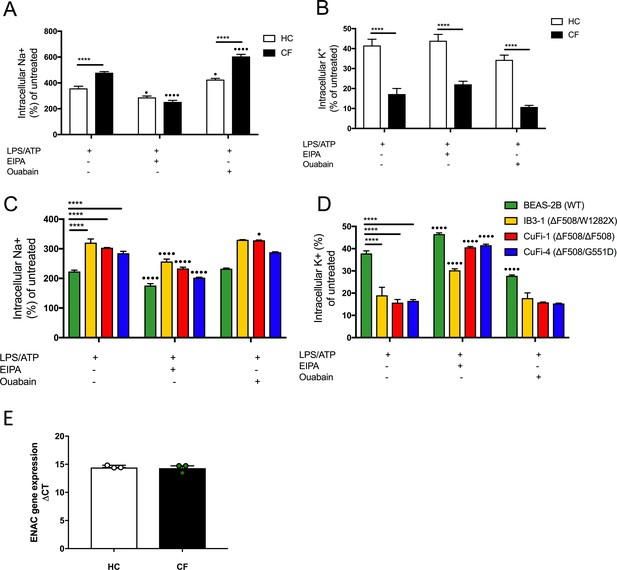
Increased sodium influx in cells with CF-associated mutations.
(A, C) Intracellular Na+ was detected using an AM ester of sodium indictor SBFI (S-1263) and (B, D) intracellular K+ was detected using an AM ester of potassium indictor PBFI (P-1266); changes in fluorescence were measured by fluorimeter post-stimulation in (A, B) monocytes (HC = 7, CF = 7) (C, D) HBECs (n = 3 independent experiments). Cells were pre-treated with the following: EIPA (10 mM, 1 hr) and ouabain (100 nM, 24 hr) before a stimulation with LPS (10 ng/mL, 4 hr) and ATP (5 mM) for the final 30 min. A 2-way ANOVA with Tukey’s multiple comparison test was performed (p values * = 0.05, ** = 0.01, *** = 0.001 and **** = 0.0001) (*) indicate significance when comparing HC with CF-associated mutatns. (•) indicate significance between treatments within the same cell line. (E) Gene expression of b-ENaC in HC vs CF (n = 1), represented as DCT. The Mann-Whitney non-parametric test was performed (p values * = 0.05).
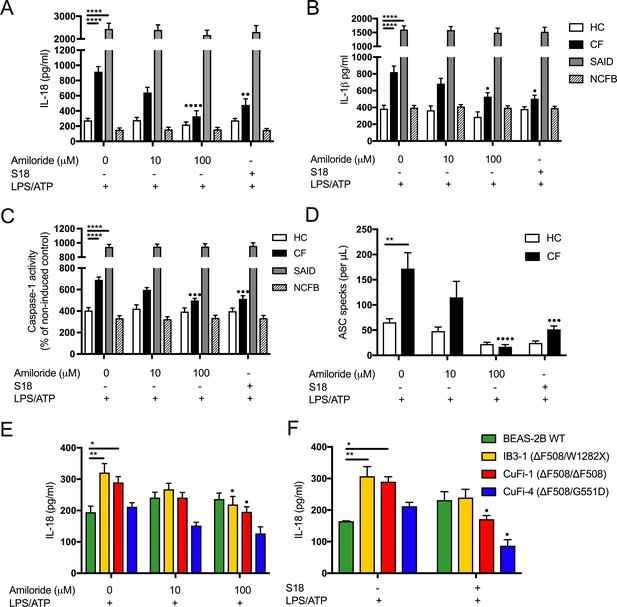
Inhibition of amiloride-sensitive sodium channels modulates inflammation in cells with CF-associated mutations.
ELISA assays were used to detect IL-18 (A) and (B) IL-1β in monocytes from HC (n = 9 amiloride, n = 10 S18), patients with CF (n = 10), SAID (n = 4) and NCFB (n = 4) and IL-18 (E) HBEC (n = 3, amiloride independent experiments) (F) HBEC (n = 3, S18 independent experiments). (C) Colourimetric assay was used to detect caspase-1 activity in protein lysates (HC n = 11, CF n = 11) and (D) flow cytometry was used to detect ASC specks in supernatant of primary monocytes (HC n = 5, CF n = 5). Cell stimulation was as follows: Amiloride (100 μM or 10 μM, 1 hr) or S18 derived peptide (25 μM, 4 hr) were used as a pre-treatment before a stimulation with LPS (10 ng/mL, 4 hr) and ATP (5 mM) for the final 30 min. (E, F) SCNN1B over-expression in BEAS-2B cells increases pro-inflammatory cytokine secretion. (E) BEAS-2B cells were transiently transfected with 10µg SCNN1B cDNA (+) or a pcDNA3.1 vector only control (-) for 48 hr then stimulated with LPS (10 ng/mL, 4 hr) and ATP (5 mM) for the final 30 min (n = 3 independent experiments). Cells were lysed and immunoblotted for β-ENaC and β-actin. (F) ELISA assays were used to detect IL-18 in the supernatant fraction. A 2-way ANOVA with Tukey’s multiple comparison test was performed (p values * =≤ 0.05, ** =≤ 0.01, *** =≤ 0.001 and **** =≤ 0.0001) (*) indicate significance, when comparing HC with CF. (•) indicate significance between treatments within the same cell line.
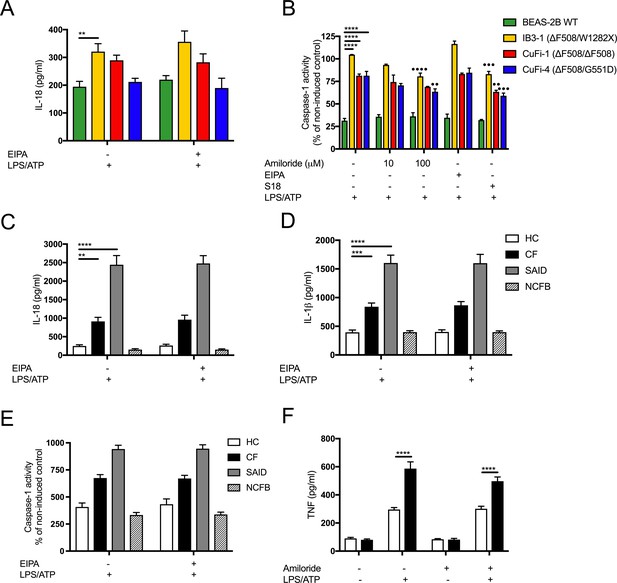
Inhibition of amiloride-sensitive sodium channels modulates inflammation in cells with CF-associated mutations.
(A) ELISA assays were used to detect IL-18 in HBECs (n = 3 independent experiments) and (C) IL-18 and (D) IL-1β and (F) TNF in monocytes from HC (n = 10), patients with CF (n = 10), SAID (n = 4) and NCFB (n = 4). Cell stimulation was as follows: (F) Amiloride (100 mM or 10 mM, 1 hr) (A, C, D, E), EIPA (10 mM, 1 hr), were used as a pre-treatment before a stimulation with LPS (10 ng/mL, 4 hr) and ATP (5 mM) for the final 30 min. (H) All the above stimulations were used to detect caspase-1 activity using a colorimetric assay in HBECs and for EIPA in monocytes (G). A 2-way ANOVA with Tukey’s multiple comparison test was performed (p values * = 0.05, ** = 0.01, *** = 0.001 and **** = 0.0001) (*) indicate significance when comparing HC with CF-associated mutants. (•) indicate significance between treatments within the same cell line.
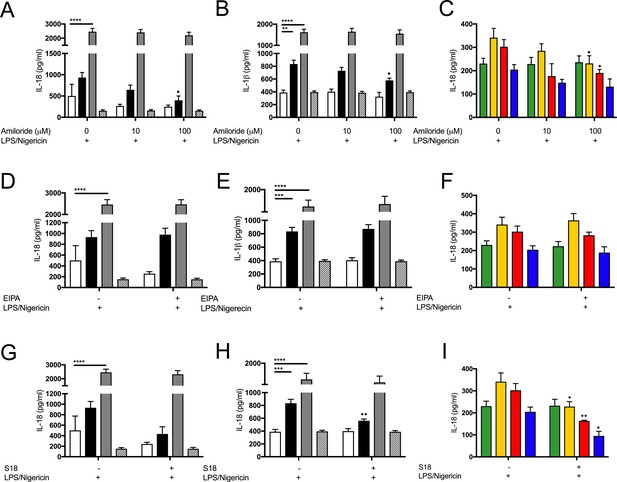
Inhibition of amiloride-sensitive sodium channels modulates inflammation in cells with CF-associated mutations.
(A, D, G) ELISA assays were used to detect IL-18 and (B, E, H) IL-1β in monocytes from HC (n = 10), patients with CF (n = 10), SAID (n = 4) and NCFB (n = 4) and (C, F) IL-18 in HBECs (n = 3 independent experiments). Cell stimulation was as follows: (A–C) Amiloride (100 mM or 10 mM, 1 hr), (D–F) EIPA (10 mM, 1 hr), (G–I) S18 derived peptide (25 mM, 4 hr) were used as a pre-treatment before a stimulation with LPS (10 ng/mL, 4 hr) and Nigericin (1µM) for the final 30 min. A 2-way ANOVA with Tukey’s multiple comparison test was performed (p values * = 0.05, ** = 0.01, *** = 0.001 and **** = 0.0001) (*) indicate significance when comparing HC with CF-associated mutants. (•) indicate significance between treatments within the same cell line.
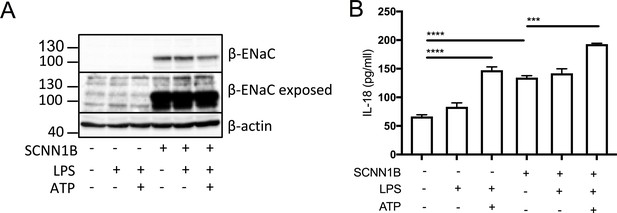
SCNN1B over-expression in BEAS-2B cells increases pro-inflammatory cytokine secretion.
BEAS-2B cells were transiently transfected with 10µg SCNN1B cDNA (+) or a pcDNA3.1 vector only control (-) for 48 hr then stimulated with LPS (10 ng/mL, 4 hr) and ATP (5 mM) for the final 30 min (n = 3 independent experiments). Cells were lysed and immunoblotted for β-ENaC and β-actin a). ELISA assays were used to detect IL-18 in the supernatant b) fraction. A 2-way ANOVA with Tukey’s multiple comparison test was performed (p values * =≤ 0.05, ** =≤ 0.01, *** =≤ 0.001 and **** =≤ 0.0001).
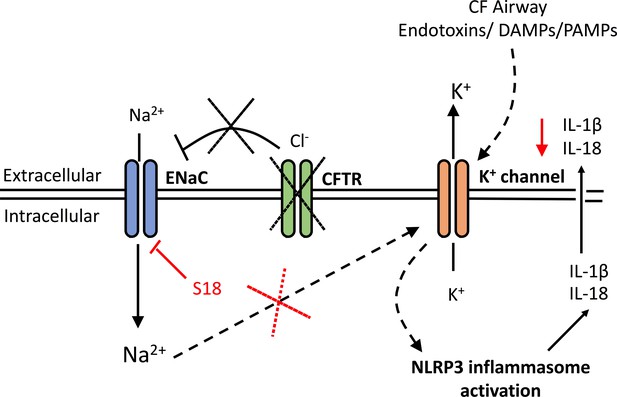
A schematic diagram of the proposed excessive NLRP3 inflammasome activation observed in individuals and cells with CF-associated mutations.
Without functional CFTR, inhibition of ENaC currents is diminished leading to increased intracellular Na+ levels. Dysregulation of ENaC-dependent Na2+ influx leads to increased K+ efflux (via unknown mechanism) and NLRP3 inflammasome activation, with subsequent release of IL-1β and IL-18. In the CF airway, K+ efflux is exacerbated upon K+ channel stimulation by endotoxins, DAMPs or PAMPs, leading to aberrant NLRP3 inflammasome activation and excessive IL-1β and IL-18 secretion. Blocking ENaC currents with S18 peptide restores Na+ and K+ levels which reduces NLRP3-mediated production of IL-1β and IL-18.
Tables
| Reagent type (species) or resource | Designation | Source or reference | Identifiers | Additional information |
|---|---|---|---|---|
| Antibody | Rabbit polyclonal anti-SCNN1B | Avia Systems Biology, San Diego | Cat# ARP72375_P050; RRID: AB_2811256 | WB 1:500 |
| Antibody | Goat polyclonal anti-Rabbit IgG (H+L) Poly-HRP Secondary Antibody | ThermoFisher Scientific | Cat# 32260; RRID: AB_1965959 | WB 1:4000 |
| Antibody | Rabbit polyclonal anti-actin-β | GeneTex | Cat# GTX109639, RRID: AB_1949572 | WB 1:20:000 |
| Antibody | Mouse monoclonal Phycoerythrin anti-ASC (TMS-1) | Biolegend | Cat# 653903, RRID: AB_2564507 | 5 µL/ ml |
| Cell line (Homo-sapiens) | BEAS-2B cell line | ATCC | ATCC CRL-9609 | |
| Cell line (Homo-sapiens) | IB3-1 | ATCC | ATCC CRL-2777 | |
| Cell line (Homo-sapiens) | CuFi-1 cell line | ATCC | ATCC CRL-4013 | |
| Cell line (Homo-sapiens) | CuFi-4 cell line | ATCC | ATCC CRL-4015 | |
| Commercial assay or kit | MycoAlertTM | Lonza | Cat# LT07-118 | |
| Biological samples (Homo-sapiens) | Human Blood Samples | St James's University Hospital | Health Research Authority REC reference 17/YH/0084 | |
| Chemical compound, drug | Lymphoprep | Axis Shield | Cat# 1114544 | |
| Chemical compound, drug | Pan Monocyte Isolation Kit, human | Miltenyi Biotec | Cat# 130-096-537 | |
| Chemical compound, drug | Lipopolysacchride Ultrapure EK | InvivoGen | Cat# tlrl-eklps | 10ng/ml |
| Chemical compound, drug | MCC950 | Cayman Chemical | Cat# CAY17510-1 | 15 nM, 1 hr |
| Chemical compound, drug | YVAD | InvivoGen | Cat# inh-yvad | 2 μg/ mL, 1 hr |
| Chemical compound, drug | OxPAPC | InvivoGen | Cat# tlrl-oxp1 | 30 μg/ mL, 1 hr |
| Chemical compound, drug | Amiloride (hydrochloride) | Cayman Chemical | Cat# 26295 | 10 μM, 100 μM, 1 hr |
| Chemical compound, drug | SPLUNC1-derived peptide, S18 | Gift from Spyryx Biosciences, Inc | 25 μM, 4 hr | |
| Chemical compound, drug | 5-(N-ethyl-N-isopropyl)-Amiloride (EIPA) | Cayman Chemical | Cat# 1154-25-2 | 10 μM, 1 hr |
| Chemical compound, drug | Ouabain | Torcis Bioscience | Cat# 630-60-4 | 100 nM, 24 hr |
| Chemical compound, drug | ATP | InvivoGen | Cat# tlrl-atpl | 5 mM, 30 min |
| Chemical compound, drug | poly(dA:dT) dsDNA | InvivoGen | Cat# tlrl-patn | 1 μg/ mL, 1 hr |
| Chemical compound, drug | TcdB | Cayman Chemical | Cat# CAY19665-50 | 10 ng/mL, 1 hr |
| Chemical compound, drug | Flagellin | InvivoGen | Cat# tlrl-pbsfla | 10 ng/mL, 1 hr |
| Commercial assay or kit | Pierce BCA Protein Assay Kit | ThermoFisher Scientific | Cat# 23225 | |
| Chemical compound, drug | PhosSTOP | Merck | Cat# 4906845001 | |
| Chemical compound, drug | Pierce Protease Inhibitor Mini Tablets | ThermoFisher Scientific | Cat# A32955 | |
| Chemical compound, drug | Immobilon Western Chemiluminescent HRP Substrate | Merck | Cat# WBKLS0500 | |
| Commercial assay or kit | IL-1 beta Human Matched Antibody Pair | ThermoFisher Scientific | Cat# CHC1213 | Assay sensitivity < 31.2 pg/mL |
| Commercial assay or kit | IL-18 Human Matched Antibody Pair | ThermoFisher Scientific | Cat# BMS267/2MST | Assay sensitivity 78 pg/mL |
| Commercial assay or kit | IL-6 Human Matched Antibody Pair | ThermoFisher Scientific | Cat# CHC1263 | Assay sensitivity 15.6 pg/mL |
| Commercial assay or kit | TNF alpha Human Matched Antibody Pair | ThermoFisher Scientific | Cat# CHC1753 | Assay sensitivity < 15.6 pg/mL |
| Commercial assay or kit | IL1RA Human Matched Antibody Pair | ThermoFisher Scientific | Cat# CHC1183 | Assay sensitivity < 31.2 pg/mL |
| Chemical compound, drug | (TMB) substrate solution | Sigma | Cat# T0440 | |
| Commercial assay or kit | Caspase-1 Colorimetrix Assay | R and D Systems | Cat# BF15100 | |
| Commercial assay or kit | High-Capacity cDNA Reverse Transcription Kit | ThermoFisher Scientific | Cat# 4368814 | |
| Recombinant DNA reagent | SCNN1B cDNA plasmid | Addgene | Cat# 83429 | |
| Recombinant DNA reagent | pcDNA3.1 cDNA plasmid | Gift from N.M Hooper, Manchester | ||
| Chemical compound, drug | sodium-sensitive molecule SBFI | ThermoFisher Scientific | Cat# S-1263 | 10 mM, 100 min |
| Chemical compound, drug | potassium-sensitive molecule PBFI | ThermoFisher Scientific | Cat# P-1266 | 10 mM, 100 min |
| Chemical compound, drug | Pluronic F-127 | Sigma | Cat# P2443 | |
| Software, algorithm | GraphPad Prism7 | Graphpad software |
Additional files
-
Supplementary file 1
Donor demographics.
Patients with Cystic Fibrosis (CF), systemic autoinflammatory diseases (SAID), non-CF bronchiectasis (NCFB) and healthy controls (HC). All CF donors were F508del/F508del homozygous (n = 30) with no sign of infection. Three of the patients with NCFB had primary ciliary dyskinesia (PCD) and one patient with NCFB had an unknown genotype. All patients with a SAID had characterised mutations in a known disease-causing gene (Tumor Necrosis Factor Receptor Associated Periodic Syndrome (TRAPS) n = 2, Muckle-Wells n = 2, A20 haploinsufficiency n = 1, Pyrin-Associated Autoinflammation with Neutrophilic Dermatosis (PAAND) n = 1, Familial Mediterranean Fever (FMF) n = 2, Hyper IgD Syndrome (HIDS) n = 2 and Schnitzler syndrome n = 1). BMI: Body Mass Index; FEV: Forced expiratory volume; CRP: C-reactive protein.
- https://doi.org/10.7554/eLife.49248.016
-
Transparent reporting form
- https://doi.org/10.7554/eLife.49248.017