Duodenum Intestine-Chip for preclinical drug assessment in a human relevant model
Figures
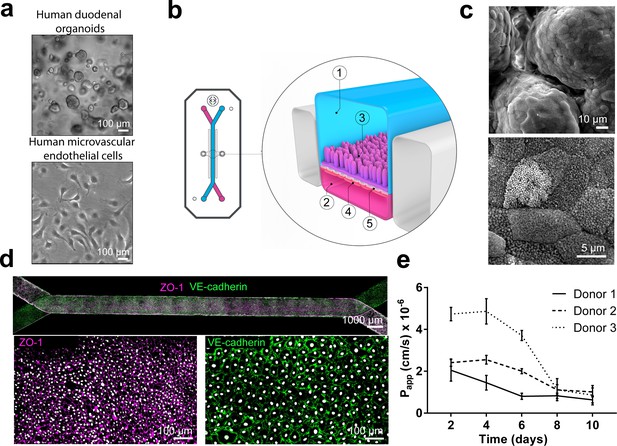
Duodenum Intestine-Chip: a microengineered model of the human duodenum.
(a) Brightfield images of human duodenal organoids (top) and human microvascular endothelial cells (bottom) acquired before their seeding into epithelial and endothelial channels of the chip, respectively. (b) Schematic representation of Duodenum Intestine-Chip, including its top view (left) and vertical section (right) showing: the epithelial (1; blue) and vascular (2; pink) cell culture microchannels populated by intestinal epithelial cells (3) and endothelial cells (4), respectively, and separated by a flexible, porous, ECM-coated PDMS membrane (5). (c) Scanning electron micrograph showing complex intestinal epithelial tissue architecture achieved by duodenal epithelium grown for 8 days on the chip (top) in the presence of constant flow of media (30 µl/hr) and cyclic membrane deformations (10% strain, 0.2 Hz). High magnification of the apical epithelial cell surface with densely packed intestinal microvilli (bottom). See Figure 1-figure supplement demonstrating the effect of mechanical forces on the cytoarchitecture of epithelial cells and the formation of intestinal microvilli (d) Composite tile scan fluorescence image 8 days post-seeding (top) showing a fully confluent monolayer of organoid-derived intestinal epithelial cells (magenta, ZO-1 staining) lining the lumen of Duodenum Intestine-Chip and interfacing with microvascular endothelium (green, VE-cadherin staining) seeded in the adjacent vascular channel. Higher magnification views of epithelial tight junctions (bottom left) stained against ZO-1 (magenta) and endothelial adherence junctions visualized by VE-cadherin (bottom right) staining. Cells nuclei are shown in gray. Scale bars, 1000 µm (top), 100 µm (bottom) (e) Apparent permeability values of Duodenum Intestine-Chips cultured in the presence of flow and stretch (30 µl/hr; 10% strain, 0.2 Hz) for up to 10 days. Papp values were calculated from the diffusion of 3 kDa Dextran from the luminal to the vascular channel. Data represent three independent experiments performed with three different chips/donor, total of three donors; Error bars indicate s.e.m.
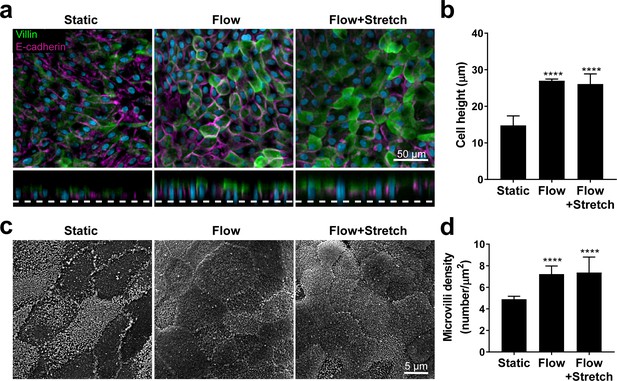
Flow-induced increase in primary intestinal epithelial cells height and microvilli formation.
(a) Representative confocal images of x-y (top) and x-z (bottom) optical sections of duodenal organoid-derived epithelial cells cultured for 72 hr under static (Static), fluid flow (30 µl/hr; Flow) or flow and stretch (30 µl/hr; 10% strain, 0.2 Hz; Flow+Stretch) conditions and stained for apical marker villin (green) and basolateral protein E-cadherin (magenta). Nuclei were counterstained with DAPI (gray). Scale bar, 50 µm. (b) Quantitative analysis of the average cell height measured from Z-stack images as the distance between apical marker villin (green) and PDMS membrane (dotted line). Data represent the mean ± s.e.m; One-way ANOVA, ****p<0.0001. (c) Scanning electron microscopy surface images of duodenal organoid-derived epithelium cultured for 72 hr under static or flow +/- stretch conditions. Cells seeded in the top channel of Intestine-Chip were maintained with (Flow) or without (Static) medium perfusion (30 µl/hr) in both channels and 10% of mechanical stretch (0.2 Hz) (Flow+Stretch). Images were captured at the center area of the chamber. Scale bar, 5 µm. (d) Quantification of microvilli. Density of microvilli per µm2 was measured from the SEM images (100 µm2, 20 FOV) as described in the Materials and methods. Data represent the mean ± s.e.m; one-way ANOVA, ****p<0.0001.
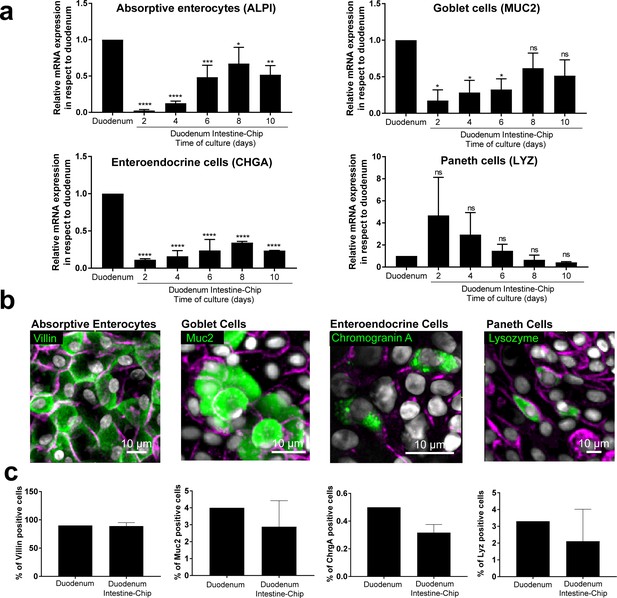
Duodenum Intestine-Chip emulates multi-lineage differentiation of native human intestine.
(a) Comparison of the relative gene expression levels of markers specific for differentiated intestinal cell types, including mucin 2 (MUC2) for goblet cells, alkaline phosphatase (ALPI) for absorptive enterocytes, chromogranin A (CHGA) for enteroendocrine cells, lysozyme (LYZ) for Paneth cells in Duodenum Intestine-Chip and RNA isolated directly from the duodenal tissue (Duodenum). Expression of these genes at different time points (days 2, 4, 6, 8, 10) of Duodenum Intestine-Chip culture is shown. In each graph, values represent average gene expression ± s.e.m (error bars) from three independent experiments, each using different donors of biopsy-derived organoids and at least three different chips per time point. Values are shown relative to duodenal tissue expressed as 1. EPCAM expression is used as normalizing control. One-way ANOVA, ****p<0.0001, ***p<0.001, **p<0.01, *p<0.05, ns p>0.05. (b) Representative confocal fluorescent micrographs demonstrating the presence of all major intestinal cell types (green) in Duodenum Intestine-Chip at day 8 of fluidic culture, including goblet cells stained with anti-mucin-2; enteroendocrine cells visualized with anti-chromogranin A, absorptive enterocytes stained with anti-villin and Paneth cells labeled with anti-lysozyme. Cell-cell borders were stained with anti-E-cadherin and are shown in magenta. Cells nuclei are shown in gray. Scale bar, 10 µm. (c) Quantification of the different intestinal epithelial cell types present in Duodenum Intestine-Chip at day eight and identified by immunostaining, as described in (b). Cell ratios are based on 10 different fields of view (10 FOV) counted in three individual chips (each from a different donor) per staining. DAPI staining was used to evaluate the total cell number. Duodenum values, represent cell ratios observed in the histological sections and are based on the literature (Karam, 1999).
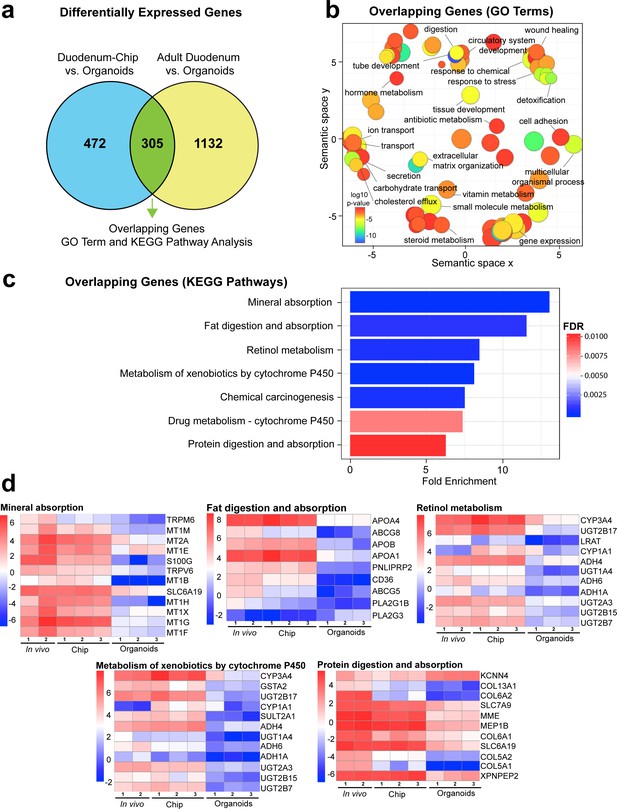
Duodenum Intestine-Chip exhibits higher transcriptomic similarity to adult duodenal tissue than organoid culture.
(a) Differential gene expression analysis was carried out to identify genes that are up- or down-regulated in Duodenum Intestine-Chip compared to organoids (blue circle) (Figure 3—source data 1) and adult duodenum compared to organoids (yellow circle) (Figure 3—source data 2). The gene lists were then compared to determine how many genes overlap between those two comparisons (Figure 3—source data 3), and the results are shown as a Venn diagram. 305 genes were identified as common and responsible for the closer resemblance of Duodenum Intestine-Chip to human adult duodenum than organoids from which chips were derived. Sample sizes were as follows: Duodenum Intestine-Chip, n = 3 (independent donors); Organoids, n = 3 (independent donors); Adult duodenum, n = 2 (independent biological specimens). Intestinal crypts derived from the same three independent donors were used for the establishment of Duodenum Intestine-Chip and organoid cultures. Both chips and organoids were cultured in parallel, in the presence of expansion media for 6 days, followed by 2 days of differentiation media. Experiment was terminated and samples were processed for analyses 8 days post-seeding. (b) The list of overlapping genes was subjected to GO analysis to identify enriched biological processes (GO terms) (Figure 3—source data 4). The results are shown as REVIGO scatterplots in which similar GO terms are grouped in arbitrary two-dimensional space based on semantic similarity. Each circle corresponds to a specific GO term and circle sizes are proportional to the number of genes included in each of the enriched GO terms. Finally, the color of a circle indicates the significance of the specific GO term enrichment. GO terms enriched in the overlapping gene set demonstrate that Duodenum Intestine-Chip is more similar to human duodenum with respect to important biological functions of the intestine, including digestion, transport and metabolism. (c) The results of the KEGG pathway analysis using the 305 differentially expressed genes showed seven significantly enriched (FDR adjusted p-value<0.05) pathways related to absorption, metabolism, digestion and chemical carcinogenesis. The size of the bars indicates the fold-enrichment of the corresponding pathways. (d) Curated heatmaps were generated to examine particular genes that belong to the enriched KEGG pathways and to show the expression levels (loge(FPKM)) of these genes across different samples. Genes belonging to five different pathways, including: ‘mineral absorption’, ‘fat digestion and absorption’, ‘retinol metabolism’, ‘metabolism of xenobiotics by cytochrome P450’ and ‘protein digestion and absorption’, are shown. The expression levels of genes associated with ‘chemical carcinogenesis’ (CYP3A4, GSTA2, UGT2B17, CYP1A1, SULT2A1, ADH4, UGT1A4, ADH6, ADH1A, UGT2A3, UGT2B15, UGT2B7) and ‘drug metabolism – cytochrome 450’ CYP3A4, GSTA2, UGT2B17, ADH4, UGT1A4, ADH6, ADH1A, UGT2A3, UGT2B15, UGT2B7) were included in the heatmap representing ‘metabolism of xenobiotics by cytochrome P450’ as they showed to overlap in between three different pathways. Each heatmap has its own color scale, which corresponds to a different range of loge(FPKM) values, as indicated on the color bars located to the left. The provided results further demonstrate that Duodenum Intestine-Chip (Chip) is more similar to adult duodenum (In vivo) than are the organoids (Organoids). Sample sizes were as follows: Duodenum Intestine-Chip, n = 3 (independent donors); Organoids, n = 3 (independent donors); In vivo, adult duodenum, n = 2 (independent biological specimens). Intestinal crypts derived from the same three independent donors were used for the establishment of Duodenum Intestine-Chip and organoid cultures. Both chips and organoids were cultured in parallel as described in (a). See also Figure 3—figure supplement 1, Figure 3—figure supplement 1—source data 1 and Figure 3—figure supplement 1—source data 2 showing the results of DGE analysis followed by functional enrichment performed between Organoids or Duodenum Intestine-Chip and Adult Duodenum.
-
Figure 3—source data 1
Differentially expressed genes in Duodenum Intestine-Chip vs. duodenal organoids, related to Figure 3.
- https://cdn.elifesciences.org/articles/50135/elife-50135-fig3-data1-v1.xlsx
-
Figure 3—source data 2
Differentially expressed genes in adult duodenum vs. duodenal organoids, related to Figure 3.
- https://cdn.elifesciences.org/articles/50135/elife-50135-fig3-data2-v1.xlsx
-
Figure 3—source data 3
Differentially expressed genes common in Duodenum Intestine-Chip and adult duodenum versus duodenal organoids, related to Figure 3.
- https://cdn.elifesciences.org/articles/50135/elife-50135-fig3-data3-v1.xlsx
-
Figure 3—source data 4
Enriched GO terms from a list of differentially expressed genes common in Duodenum Intestine-Chip and adult intestine versus duodenal organoids, related to Figure 3.
- https://cdn.elifesciences.org/articles/50135/elife-50135-fig3-data4-v1.xlsx
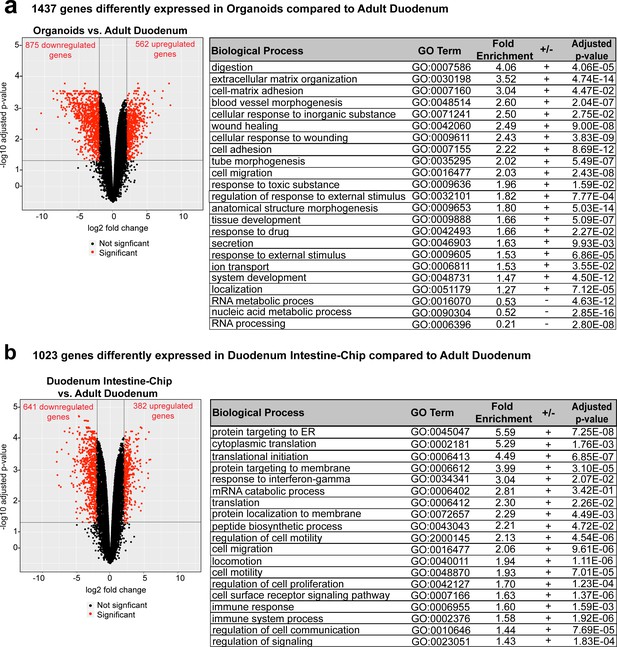
Differentially expressed genes and enriched gene ontology categories in organoids or Duodenum Intestine-Chip with respect to adult duodenum.
(a) Volcano plot (left) and functional enrichment analysis (right) of differentially expressed genes between organoids and adult duodenum (Figure 3—figure supplement 1—source data 1). The red dots represent genes that were significantly (adjusted p-value<0.05) up- or down-regulated. The black dots correspond to the non-differentially expressed genes. The vertical lines correspond to 2.0 up/down fold change and the horizontal line indicates the adjusted p-value<0.05. Sample sizes were as follows: Organoids, n = 3, Adult Duodenum, n = 2. All samples are biologically independent (derived from a different donor). Organoids were cultured in the presence of expansion media for 6 days, followed by 2 days of differentiation media. Analysis was performed in samples collected 8 days post-seeding. Functional enrichment analysis demonstrated over (+) and under (-) represented biological processes in the GO categories concerning digestion, extracellular matrix organization, angiogenesis, cell adhesion, tissue development, and cell response to drugs and toxic substances, while nucleic acid metabolic process and RNA processing were underrepresented. GO, Gene Ontology. (b) Differential gene expression and functional enrichment analysis between Duodenum Intestine-Chip and adult human tissue (Figure 3—figure supplement 1—source data 2) demonstrating up- and down-regulated genes (volcano plot, left) and annotated biological processes (table, right) involving but not limited to protein synthesis and targeting as well as cell cycle and cell proliferation. Red dots: significant genes (adj. p-value<0.05). Black dots: non-differentially expressed genes. Sample sizes were as follows: Duodenum Intestine-Chip, n = 3; Adult Duodenum, n = 2. All samples were biologically independent (derived from a different donor). Duodenum Intestine-Chips were grown in the presence of expansion media for 6 days, followed by 2 days of differentiation media. Analysis was performed in samples collected 8 days post-seeding.
-
Figure 3—figure supplement 1—source data 1
Differentially expressed genes in duodenal organoids vs. adult duodenum, related to Figure 3—figure supplement 1.
- https://cdn.elifesciences.org/articles/50135/elife-50135-fig3-figsupp1-data1-v1.xlsx
-
Figure 3—figure supplement 1—source data 2
Differentially expressed genes in Duodenum Intestine-Chip vs. adult duodenum, related to Figure 3—figure supplement 1.
- https://cdn.elifesciences.org/articles/50135/elife-50135-fig3-figsupp1-data2-v1.xlsx
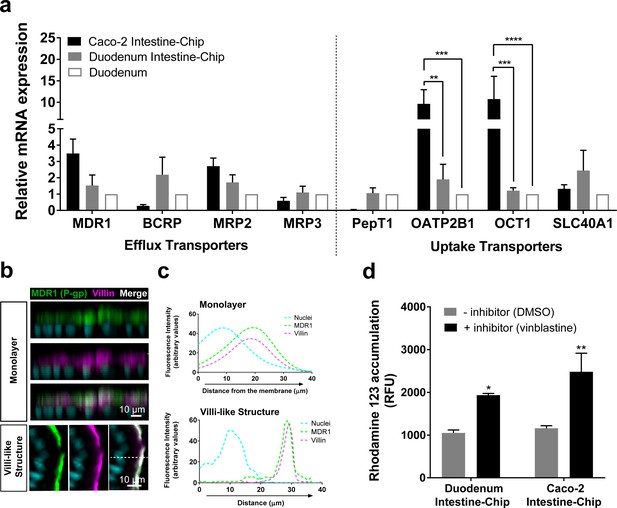
Duodenum Intestine-Chip shows the presence of major intestinal drug transporters and correct localization and function of efflux pump MDR1 (P–gp).
(a) Comparison of the relative average gene expression levels of drug efflux (MDR1, BCRP, MRP2, MRP3) and uptake (PEPT1, OATP2B1, OCT1, SLC40A1) transporters in Caco-2 Intestine-Chip, Duodenum Intestine-Chip, both assessed on day 8 of culture, and RNA isolated directly from the duodenal tissue (Duodenum). The results show that Duodenum Intestine-Chip expresses drug transport proteins at the levels close to human duodenal tissue. Note, the expression of OATP2B1 and OCT1 in Caco-2 Intestine-Chip were significantly higher than in human duodenum while the difference between Duodenum Intestine-Chip and adult duodenum is not significant. Each value represents average gene expression ± s.e.m (error bars) from three independent experiments, each involving Duodenum Intestine-Chip established from a tissue of three different donors (three chips/donor), RNA tissue from three independent biological specimens, and Caco-2 Intestine-Chip (three chips). Values are shown relative to the duodenal tissue expressed as 1, two-way ANOVA, ****p<0.0001, ***p<0.001, **p<0.01. EPCAM expression was used as normalizing control. (b) Representative confocal immunofluorescence micrographs showing apical localization of the efflux transporter MDR1 (green) and the cell surface marker villin (magenta) in a vertical cross section of monolayer (top) formed in Duodenum Intestine-Chip at day 4 and later formed villi-like structure (bottom) at day 8. Cell nuclei are visualized in cyan. Scale bar, 10 µm. (c) Line plots corresponding to confocal images in (b) showing the distribution of fluorescent intensities for three different channels: MDR1 (green), villin (magenta) and nuclei (cyan) along the basal–apical axis of enterocytes forming a monolayer or villi-like structures in Duodenum Intestine-Chip. The fluorescent intensity was analyzed in 3D reconstructed confocal images of Duodenum Intestine-Chip and plotted as average across 20 different z-stacks. Distribution of MDR1 and villin shows significant overlap. See also Figure 4—figure supplement 1 showing luminal localization of additional efflux (BCRP) and uptake (PEPT1) transporters in Duodenum Intestine-Chip. (d) Activity of efflux pump proteins in Caco-2 and Duodenum Intestine-Chip. The intracellular accumulation of the fluorescent substrate of MDR1 - Rhodamine 123 is significantly increased in response to the MDR1 inhibitor vinblastine (black bars) in comparison to vehicle (DMSO) control (gray bars) in Caco2 and Duodenum Intestine-Chip. Data were represented as mean ± s.e.m (error bars) of at least three independent experiments involving chips generated from organoids of three individual donors or Caco-2 cells, all assessed 8 days post-seeding. Two-way ANOVA, **p<0.01, *p<0.05.
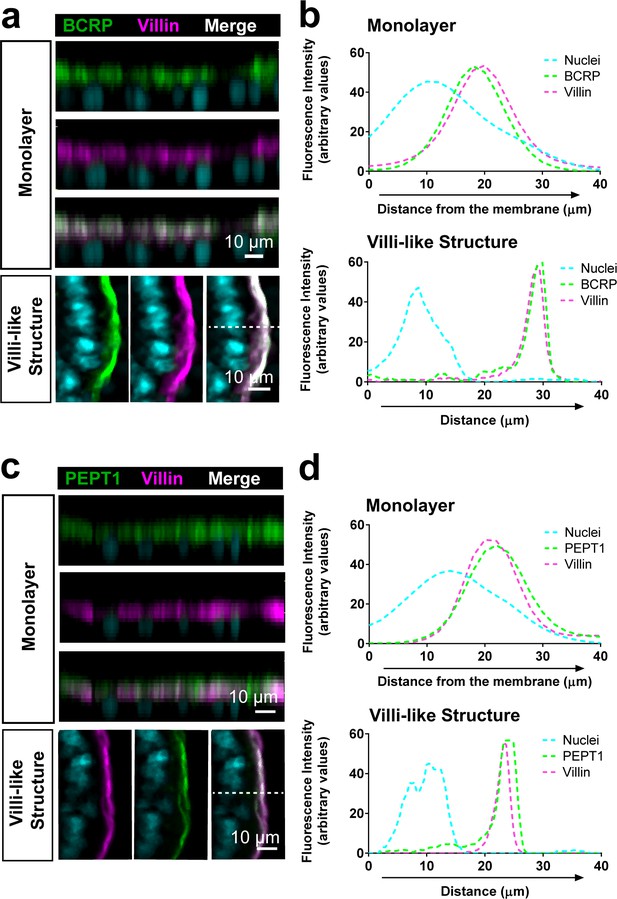
Luminal localization of efflux (BCRP) and uptake (PEPT1) transporters in Duodenum Intestine-Chip.
Representative cross-sectional confocal images of Duodenum Intestine-Chip (left) showing apical localization of efflux BCRP (a; green) and uptake PEPT1 drug transporters (c; green) that co-localize with luminal cell surface marker villin (magenta) at the time of confluent monolayer formation (day 4 of culture) as well as within successively formed villi-like structures (at day 8). Cell nuclei are visualized in cyan. Scale bar, 10 µm (b and d) Line plots corresponding to confocal images in (a andb) representing distribution of the fluorescence intensities across epithelial cell z-axis revealed close overlap of green (transporters; BRCP and PEPT1) and magenta (apical cell marker; villin) signals confirming co-distribution of these proteins on the luminal cell surface. The fluorescent intensity for each channel was analyzed in 3D reconstructed confocal images of Duodenum Intestine-Chip and plotted as average across 26 different z-stacks (for BCRP) and 21 different z-stacks (for PEPT1).
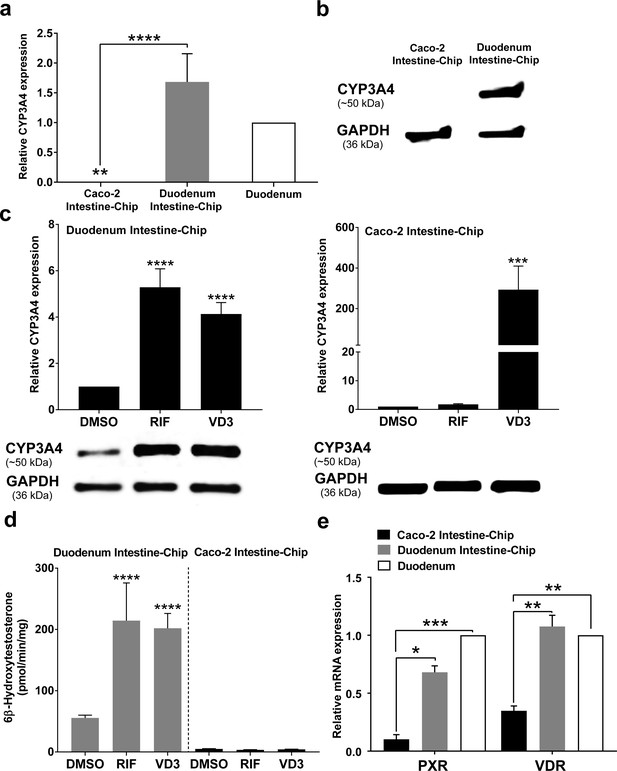
CYP3A4 expression levels and induction in Duodenum Intestine-Chip and Caco-2 Intestine-Chip.
(a) Average gene expression levels of CYP3A4 ± s.e.m (error bars) in Caco-2 Intestine-Chip, Duodenum Intestine-Chip assessed at day 8 of their culture and human duodenum (three independent biological specimens). All values are shown relative to the adult duodenal tissue expressed as 1, one-way ANOVA, ****p<0.0001, **p<0.01. EPCAM expression was used as normalizing control. (b) Protein analysis of CYP3A4 in Caco-2 and Duodenum Intestine-Chip assessed at day eight using western blot. (c) The CYP3A4 induction in Duodenum Intestine-Chip and Caco-2 Intestine-Chip treated at day 6 with solvent (DMSO), 20 μM rifampicin (RIF) or 100 nM 1,25-dihidroxyvitamin D3 (VD3) for 48 hr. The gene expression levels (top) of CYP3A4 were examined by real-time PCR analysis at day 8. On the y-axis, the gene expression levels in the DMSO-treated chips were taken as 1.0. All data are represented as means ± s.e.m. Two-way ANOVA, ****p<0.0001 (compared with DMSO-treated cells). The corresponding CYP3A4 protein expression levels (bottom) were measured at day 8 by western blotting analysis. (d) CYP3A4 enzyme activity was determined by monitoring the formation of 6β-hydroxytestosterone in the medium of Duodenum Intestine-Chip and Caco-2 Intestine-Chip, as measured by LC-MS. For induction studies, 20 µM RIF or 100 nM VD3 was added 48 hr before measurement. Data are expressed as mean ± s.e.m of three independent experiments each involving Duodenum Intestine-Chip established from organoid-derived cells of a different donor and Caco-2 Intestine-Chip. At least three different chips were used per condition. Two-way ANOVA, ****p<0.0001 (e) Gene expression analysis of the receptors PXR and VDR in Duodenum Intestine-Chip and Caco-2 Intestine-Chip examined at day 8 by real-time PCR analysis and compared to their expression in adult duodenal tissue (Duodenum). On the y-axis, the gene expression levels in adult tissue were taken as 1.0. All data are represented as means ± s.e.m. Two-way ANOVA, ***p<0.001, **p<0.01, *p<0.05.
Tables
| Reagent type (species) or resource | Designation | Source or reference | Identifiers | Additional information |
|---|---|---|---|---|
| Antibody | anti-BCRP (Mouse monoclonal) | Millipore | Cat# MAB4155 RRID:AB_95060 | IF(1:100) |
| Antibody | anti-Chromogranin A (Goat polyclonal) | Santa Cruz Biotechnology | Cat# sc-1488 RRID:AB_2276319 | IF(1:100) |
| Antibody | anti-E-Cadherin (Mouse monoclonal) | Abcam | Cat#: ab1416 RRID:AB_300946 | IF(1:100) |
| Antibody | anti-Lysozyme (Rabbit polyclonal) | Agilent | Cat#: A0099 RRID:AB_2341230 | IF(1:500) |
| Antibody | anti-MDR-1 (P-gp) (Mouse monoclonal) | Thermo Fisher Scientific | Cat# MA5-13854 RRID:AB_10979045 | IF(1:100) |
| Antibody | anti-Mucin-2 (Mouse monoclonal) | Santa Cruz Biotechnology | Cat# sc-7314 RRID:AB_627970 | IF(1:400) |
| Antibody | anti-PEPT1 (Mouse monoclonal) | Santa Cruz Biotechnology | Cat# sc-373742 RRID:AB_10918256 | IF(1:100) |
| Antibody | anti-VE-Cadherin (Rabbit polyclonal) | Abcam | Cat#: ab33168 RRID:AB_870662 | IF(1:400) |
| Antibody | anti-Villin (Rabbit monoclonal) | Abcam | Cat#: ab130751 AB_11159755 | IF(1:100) |
| Antibody | anti-ZO-1) (Mouse monoclonal) | Thermo Fisher Scientific | Cat# 33–9100 RRID:AB_2533147 | IF(1:100) |
| Chemical compound, drug | 3 kDa Dextran, Cascade Blue | Thermo Fisher Scientific | Cat# D7132 | 0.1 mg/ml |
| Chemical compound, drug | 1α,25-Dihydroxyvitamin D3 | Sigma | Cat# D1530 | 100 nM |
| Chemical compound, drug | Rifampicin | Sigma | Cat# R3501 | 20 µM |
| Chemical compound, drug | Testosterone | Sigma | Cat# T1500 | 200 µM |
| Commercial assay or kit | MDR1 Efflux Assay Kit | Millipore | Cat# ECM910 | |
| Software, algorithm | Prism | GraphPad | ||
| Software, algorithm | Fiji | RRID:SCR_002285 |
Additional files
-
Supplementary file 1
RNAseq datasets downloaded from public databases.
- https://cdn.elifesciences.org/articles/50135/elife-50135-supp1-v1.xlsx
-
Supplementary file 2
List of human TaqMan gene expression assays used for qRT-PCR.
- https://cdn.elifesciences.org/articles/50135/elife-50135-supp2-v1.xlsx
-
Transparent reporting form
- https://cdn.elifesciences.org/articles/50135/elife-50135-transrepform-v1.docx





