Transcriptome dynamics of the Myxococcus xanthus multicellular developmental program
Figures
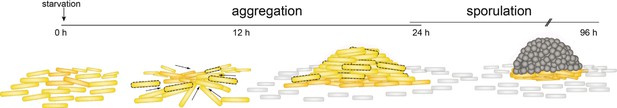
Schematic of the M. xanthus developmental program.
The time line indicates aggregation and sporulation phases. M. xanthus cells (yellow rods) aggregate into mounds (arrows indicate gliding to aggregation centers) and then differentiate into resistant spores (gray circles) to produce mature fruiting bodies. Peripheral rods (gray rods) remain outside of the fruiting bodies as a distinct differentiated state. Cells undergoing lysis are depicted with dashed lines.
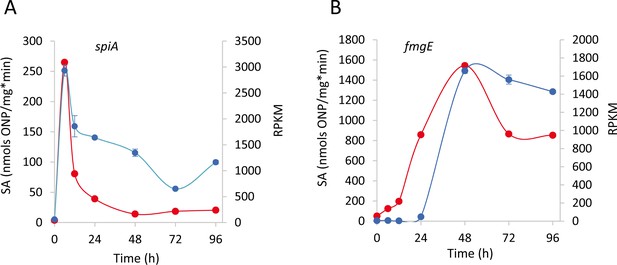
Validation of the RNA-Seq transcription patterns for genes spiA (MXAN_RS20760) (A) and fmgE (MXAN_RS16790) (B).
β-galactosidase specific activity (SA) of the strains harboring lacZ fusions to the respective genes (blue lines) compared to RNA-Seq RPKM values (red lines) at each developmental time point (h). Error bars indicate standard deviations for β-galactosidase specific activity determination.
-
Figure 2—source data 1
Comparison of expression profiles of several developmental genes described in the literature and included in the DGs with the RPKM profiles from this study.
References are in Figure 3—source data 1.
- https://doi.org/10.7554/eLife.50374.007
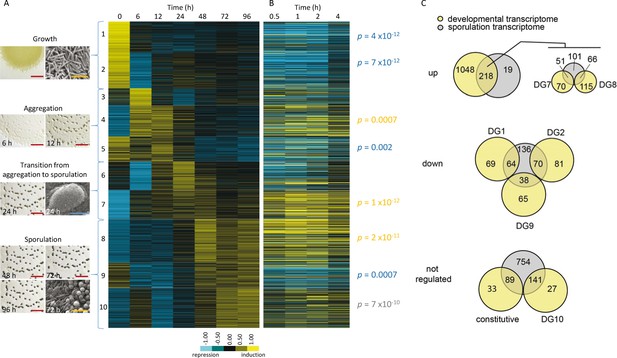
The relative expression profiles of M. xanthus genes observed during the developmental program compared to those previously observed during chemical-induction of sporulation.
(A) Relative expression profiles of significantly regulated genes at the indicated hours after induction of starvation. Genes were clustered into 10 developmental groups based on the time of peak expression and then organized according to the temporal progression of development. Developmental group number and the phase of the developmental program (with photographs of aggregates under the dissecting microscope and cells under the scanning electron microscope) are indicated to the left of the heat map. In the photographs, red, blue and yellow bars represent 2 mm, 100 µm, and 5 µm, respectively. (B) Relative expression levels of the genes in panel A during the indicated hours after chemical-induction of sporulation (Müller et al., 2010). The position of individual genes in panel B is matched to panel A. Relative expression levels for panels A and B are indicated by color code according to the legend. DGs significantly represented in up-, down-, or not-regulated sporulation gene sets are indicated by the probability values in yellow, blue or gray, respectively (C) Comparison of up-, down-, or not-regulated starvation-induced development and chemical-induced sporulation gene tallies. Tally of glycerol-induced sporulation genes up- (top), down- (middle) or not-regulated (bottom) that are significantly enriched in the indicated DGs.
-
Figure 3—source data 1
RPKM values for developmentally controlled genes and distribution of the genes in the ten developmental groups.
Genes with >2 fold change in mRNA levels and >0.7 R2 correlation between the two replicate developmental time courses are listed with their corresponding RPKM mRNA level measurements, following the same order shown in Figure 3A. The old (MXAN_) locus tags, new gene identifiers (MXAN_RS), gene name and predicted functions or pathways in which they have been previously implicated are included.
- https://doi.org/10.7554/eLife.50374.013
-
Figure 3—source data 2
Genes with no reads at time 0 hr.
Since no data existed at 0 hr for these genes (and therefore not included in Figure 3A), we compared correlation scores of the developmental time points and found that 7 out of 13 genes had high correlations and passed the developmental regulation criteria (highlighted in blue). Four of these genes are most likely forming operons with other genes included in DG4 and DG7 based on the genomic environment and relative expression profiles (written in red). The profiles have been represented assuming that relative time at 0 hr = 0 RPKM. Genes excluded because of low correlation values are highlighted in yellow.
- https://doi.org/10.7554/eLife.50374.014
-
Figure 3—source data 3
Previously reported developmental genes and identification in the 10 developmental groups shown in Figure 3A.
Genes have been highlighted by color (see color code) to depict known or implied roles in developmental processes and/or membership of protein families involved in development. References are the same as in Figure 3—source data 1.
- https://doi.org/10.7554/eLife.50374.015
-
Figure 3—source data 4
Genes involved in development that are not included in the DGs shown in Figure 3A and reasons for their exclusion.
- https://doi.org/10.7554/eLife.50374.016
-
Figure 3—source data 5
Genes included for comparison of developmental and sporulation transcriptomes (Figure 3B).
Developmental transcriptome genes (this study) include those assigned to DGs and those which passed requirements for all data points with >0.7 R2 correlation (considered constitutive). Sporulation transcriptome data were from Müller et al. (2010), in which vegetative cells were induced to artificially sporulate by addition of glycerol to 0.5 M. The sporulation gene data set included genes significantly down-, up-, or not-regulated. Up-regulated genes were classified as up1 (peak expression within 2 hr of sporulation induction) or up2 (peak expression at 2–4 hr of sporulation induction). Only genes present in both studies are included in this table.
- https://doi.org/10.7554/eLife.50374.017
-
Figure 3—source data 6
Tally of enrichment of sporulation transcriptome genes (Müller et al., 2010) found in each of the DGs (Figure 3B).
Only genes identified as having reliable expression patterns in both developmental and sporulation transcriptome studies (Figure 3—source data 5) were included. Spore transcriptome genes enriched or depleted in any DG were tested for significance (p<0.01) using a chi square test [as per Müller et al. (2010) generating probability (p) values that this enrichment was due to chance. Sporulation transcriptome up-, down- or not-regulated gene categories significantly over-enriched in any DGs are highlighted in yellow, genes significantly depleted from any DG category are highlighted in blue. Text in red: sporulation up-regulated genes are enriched in DGs 7 and 8.
- https://doi.org/10.7554/eLife.50374.018
-
Figure 3—source data 7
Tally of genes observed in both the developmental (this study) and sporulation [Müller et al., 2010 BMC Genomics 11:264] transcriptomes as supporting data for Figure 3C.
Total number indicates genes with expression profiles considered reliable in both data sets. Developmental DG1 was considered down-regulated genes, while up-regulated genes were considered all genes in DGs 2–10. Developmental constitutive genes were those whose expression level was not more than 2-fold different from vegetative cells at any time point.
- https://doi.org/10.7554/eLife.50374.019
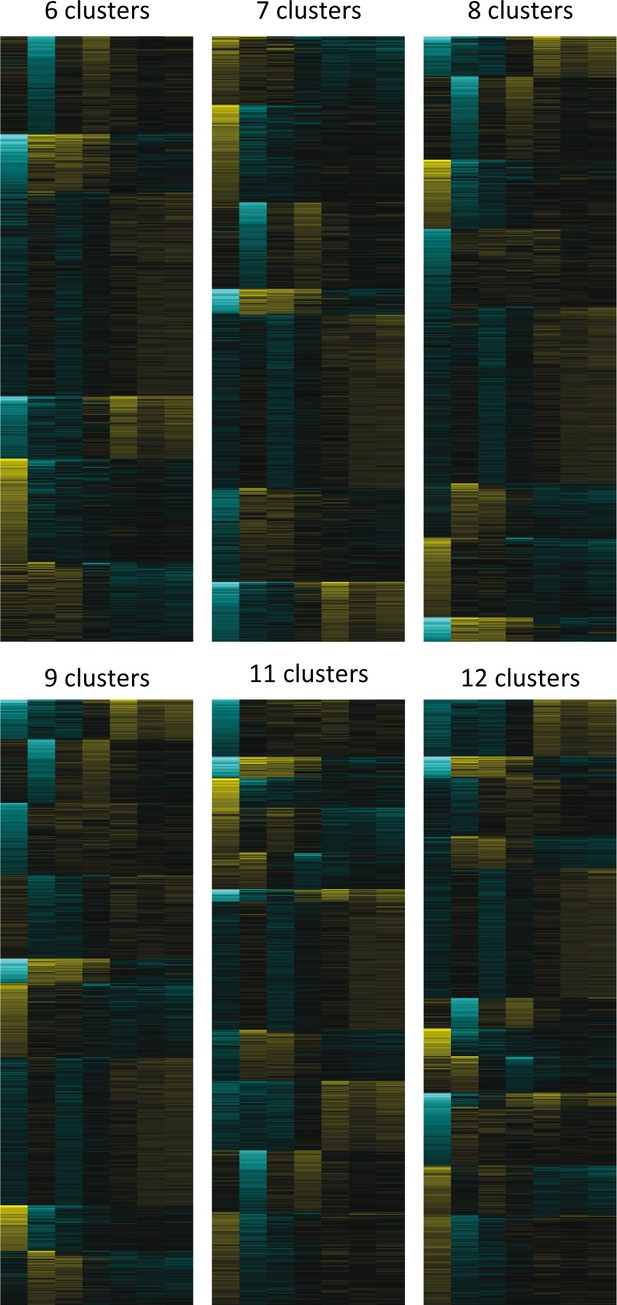

Relative expression profiles of significantly regulated genes at the indicated hours after induction of starvation.
Genes were clustered into a different number of developmental groups to decide which number of groups is the most appropriate.
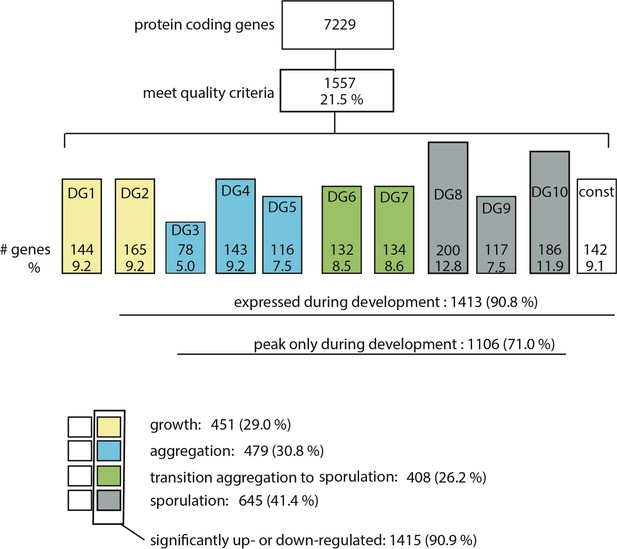
Tally of reliable genes.
Number (#) genes with >50 reads for each time point and replicate correlations > 0.7). Bars are colored coded based on the indicated stages of development assigned in Figure 3. Bar height is proportional to number of genes. DG, developmental group; const, constitutive expression.
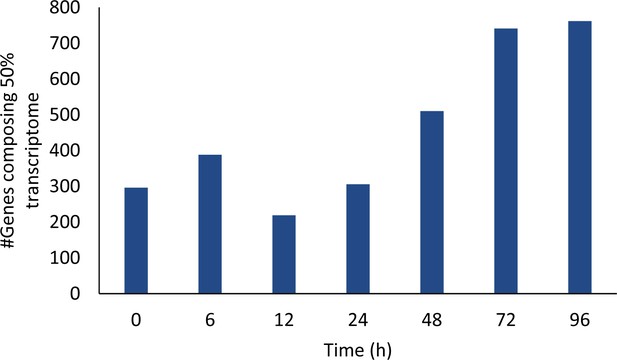
Number of genes composing 50% of the transcriptome throughout the developmental program.
https://doi.org/10.7554/eLife.50374.011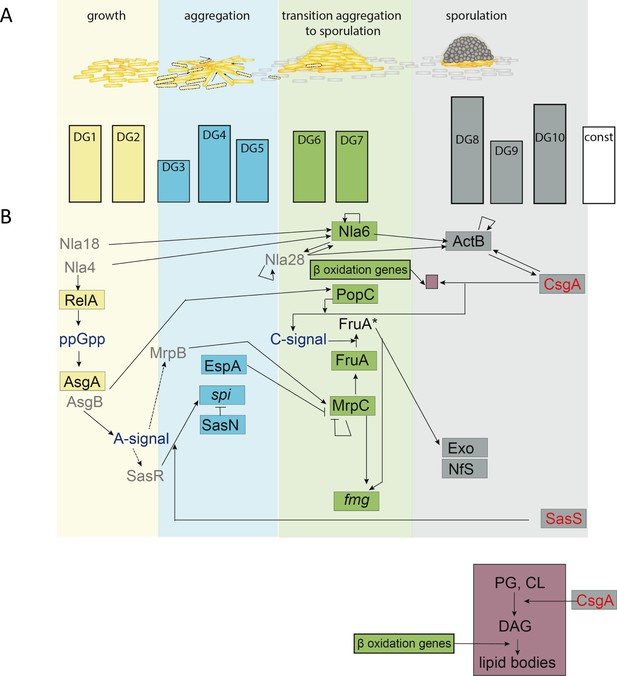
A: Relationship between M. xanthus developmental phases, developmental groups, and a known functional interaction network.
(A) Developmental clusters with peak expressions corresponding to the indicated developmental phases. Size of the bars correspond to the number of genes in each category (see Figure 3—source data 1). Yellow bars, peak expression during growth. Blue bars, peak expression during aggregation; green bars, peak expression during transition from aggregation to sporulation; gray bars, peak expression during sporulation phases. (B) Limited functional interaction network of certain well defined genes and their known regulatory networks. Proteins/genes are color coded to the developmental phase from A and aligned with the specific DG in which they were identified. Gray proteins did not meet quality control criteria. Blue text represents small molecule or protein signals. Red proteins were identified in DGs with peak expression profiles that did not match their predicted function. The maroon inset is expanded below the figure. Arrows indicate positive regulation and blunt-ended lines indicate negative regulation.
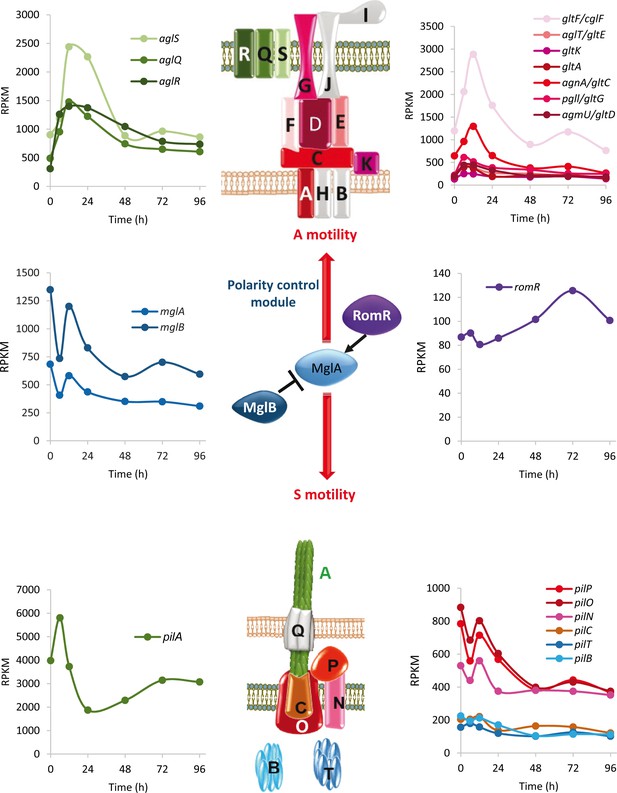
Developmental expression levels of M. xanthus motility proteins.
Schematic representation of the focal adhesion motor complexes necessary for A motility (top), the type IV pili motor complexes necessary for S motility (bottom), and the proteins involved in controlling polarity of both engines (polarity control module; center). The developmental expression levels (RPKM) of significantly regulated motility genes at the indicated times (h) of development are depicted. Gene expression profiles are colored to match the proteins shown in the schematic. Proteins depicted in gray represent genes that were not included in the developmental groups.
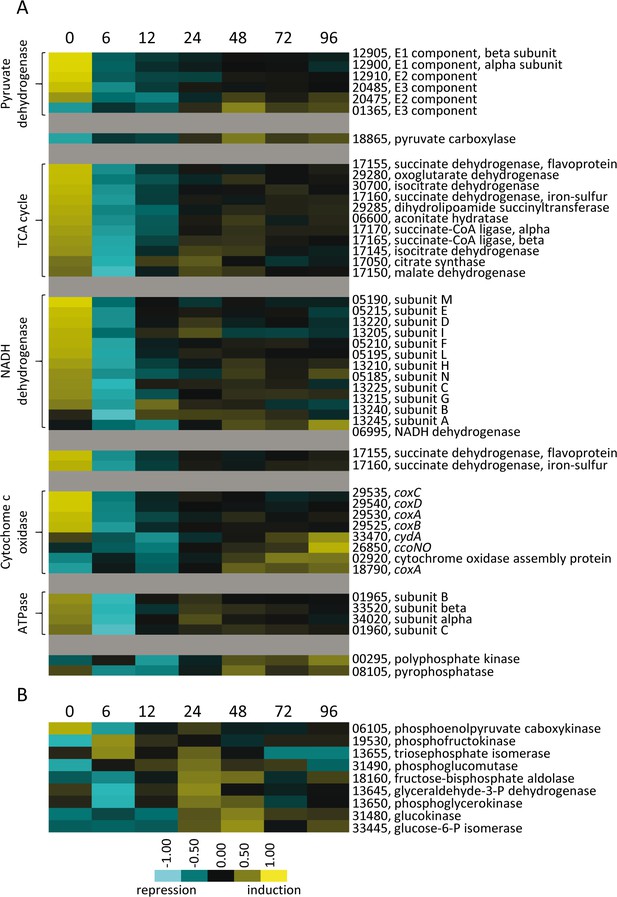
Relative developmental expression profiles of genes involved in energy generation.
(A) Genes encoding protein homologs for the pyruvate dehydrogenase complex, TCA cycle, and oxidative phosphorylation proteins. (B) Genes necessary for glycolysis/gluconeogenesis. Developmental time points in hours are indicated above each panel. Relative expression levels for panels A and B are indicated by color code according to the legend. For simplicity, the MXAN_RS designation was omitted from the locus tag of each gene.
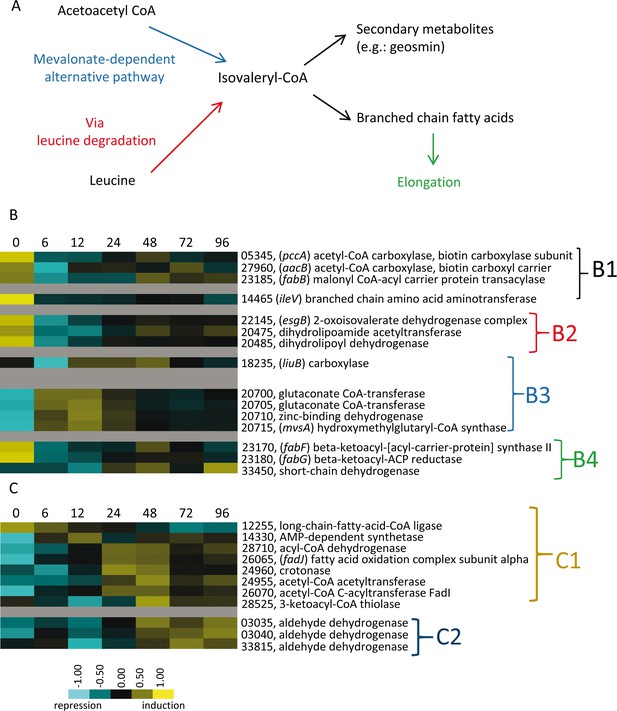
Genes involved in synthesis and degradation of lipids.
(A) Simple representation of the M. xanthus branched fatty acid metabolic pathways depicting leucine degradation and alternative mevalonate-dependent routes. (B) Relative developmental expression profiles of the genes involved in straight-chain and branched-chain fatty acid biosynthesis as designated to the right. (B1) Straight-chain fatty acid primer synthesis; (B2) Branched-chain fatty acid primer synthesis of isovaleryl-CoA via leucine degradation (bkd genes); (B3) Branched-chain fatty acid primer synthesis of isovaleryl-CoA via the alternative pathway (mevalonate); B4: Fatty acid elongation. (C) Lipid degradation via β oxidation (C1) and other pathways (C2). Relative expression levels for panels B and C are indicated by color code according to the legend and developmental time points in hours are indicated above each panel. The MXAN_RS designation was omitted from the locus tag of each gene.
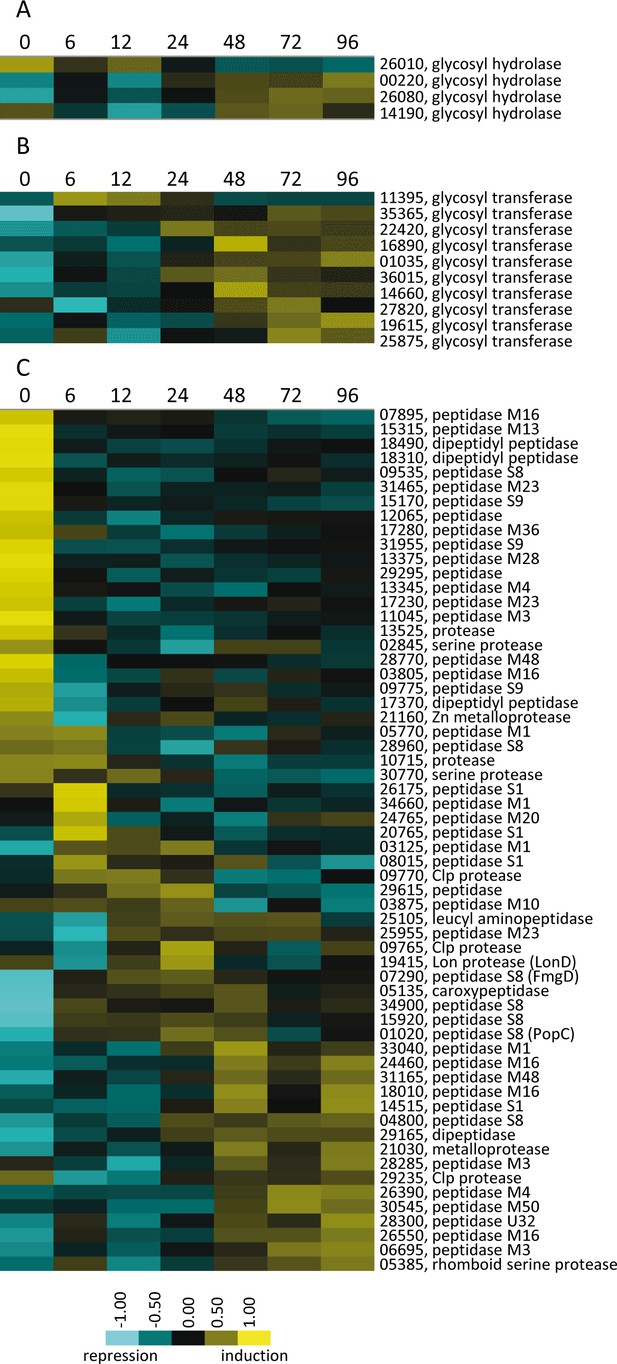
Developmental expression profiles of genes involved in production of polysaccharides and proteins.
Relative expression profiles of genes predicted to be necessary for polysaccharide hydrolysis (A), polysaccharide synthesis (B), and encoding proteases and peptidases (C). Developmental time points in hours are indicated above each panel and relative expression levels are indicated by color code according to the legend at the bottom. The MXAN_RS designation was omitted from the locus tag of each gene.
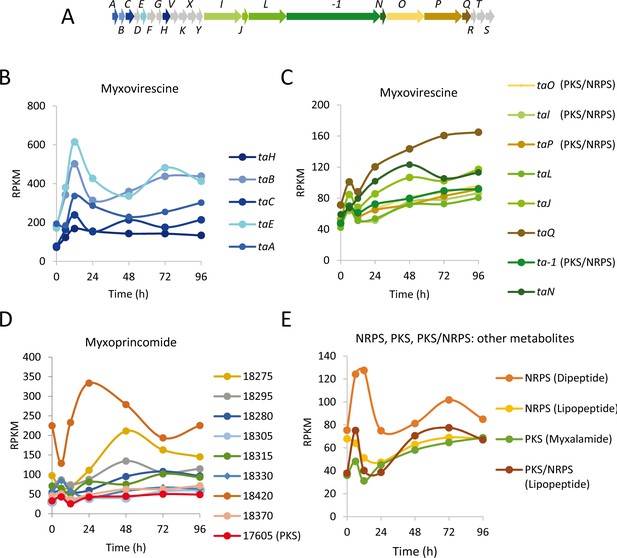
Developmental expression levels of genes involved in myxovirescine (antibiotic TA), myxoprincomide and other secondary metabolite biosynthesis.
(A) Schematic of the myxovirescine gene cluster. Names of genes depicted here are ta followed by the capital letter written with each arrow. (B) and (C) Developmental expression levels (RPKM) of significantly regulated myxovirescine genes plotted against the indicated developmental time points in hours. Gene expression profiles are colored to match the genes depicted in panel A. Genes depicted in gray were not included in the developmental groups. (D) Expression profiles of genes involved in myxoprincomide biosynthesis. (E) Profiles of others NRPS, PKS and PKS/NRPS not included in panels C and D. The MXAN_RS designation was omitted from the locus tag of each gene.
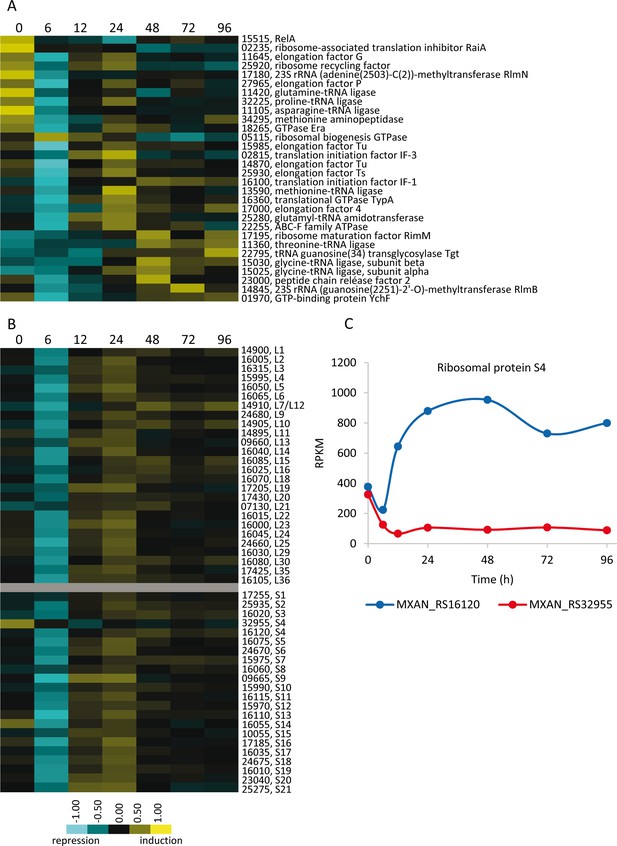
Developmental expression profiles of genes involved in protein production.
(A) Relative expression levels of genes involved in translation or ribosome assembly. (B) Relative expression levels of genes encoding ribosomal proteins. Relative expression levels for panels A and B are indicated by color code according to the legend at the bottom, and developmental time points in hours are indicated above each panel. The MXAN_RS designation was omitted from the locus tag of each gene. (C) Developmental expression levels (RPKM) of the paralogous genes encoding protein S4 plotted against developmental time points in hours.
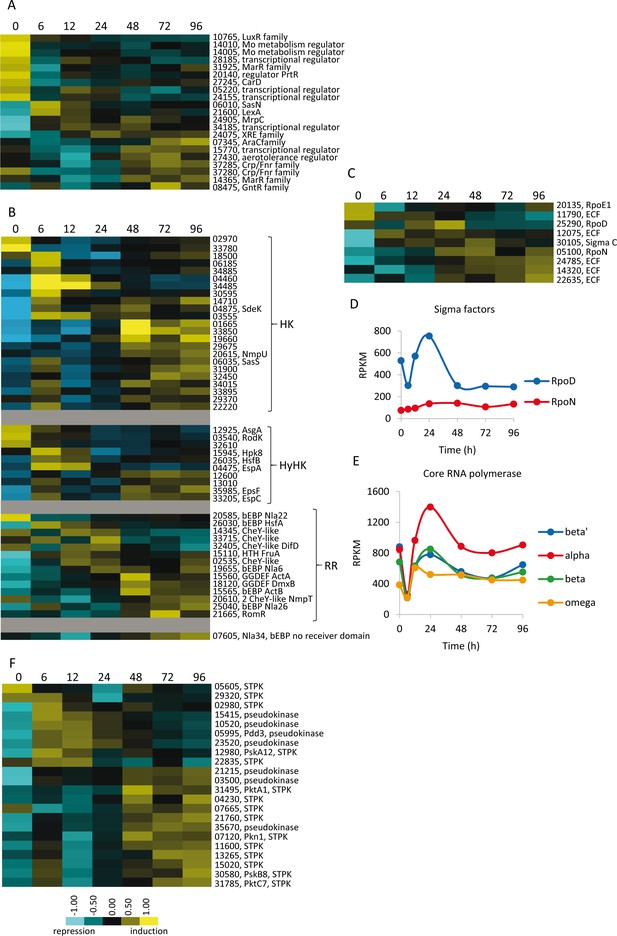
Developmental expression profiles of genes involved in transcriptional regulation and signal transduction.
Relative expression levels of genes encoding one-component regulators (A), two-component signal transduction proteins (B), sigma factors (C), and serine/threonine protein kinases (F) are depicted. Relative expression levels for panels A, B, C, and F are indicated by color code according to the legend at the bottom, and developmental time points in hours are indicated above each panel. The MXAN_RS designation was omitted from the locus tag of each gene. Expression levels (RPKM) of genes encoding the major sigma factors (D) and the subunits of the RNA polymerase (E) plotted against developmental time points in hours.
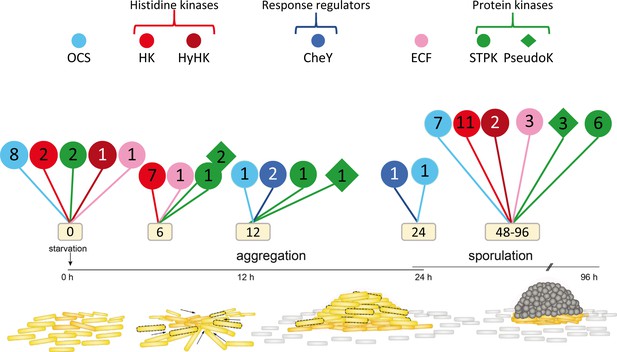
New signaling proteins that are developmentally regulated in M. xanthus identified in this study.
Code used to distinguish among types of regulators is indicated in the upper part, where OCS indicates one-component systems; HK, histidine kinase: hyHK, hybrid histidine kinase; CheY, CheY-likeresponse regulator; RR, response regulator; ECF, ECF sigma factor; STPK, active Ser/Thr protein kinase; PseudoK, pseudokinase. Numbers inside each symbol indicate the number of each type of regulator that have not been previously identified as being developmentally regulated. Information about proteins depicted here is shown in Figure 11—source data 1.
-
Figure 11—source data 1
Regulatory elements included in the developmental groups.
Time point of maximum expression is highlighted in green when genes where previously identified to play a role in development; in yellow if these genes have been previously identified but their role in development have not been well established; and in blue if they have been first identified in this study. References for genes previously identified and shown with names here are found in Figure 3—source data 1.
- https://doi.org/10.7554/eLife.50374.028
Tables
Statistical analysis of the M. xanthus DK1622 transcriptome raw data.
Data for each of the replicas at 0, 6, 12, 24, 48, 72 and 96 hr of development are shown.
| Sample name | #Gb | #mapped reads | #rRNA-reads | #clean reads (Non-rRNA) | %rRNA rate | Coverage (x) | R2 correlation |
|---|---|---|---|---|---|---|---|
| WT_0_1 | 5.70 | 61906718 | 60713758 | 1192960 | 98.07 | 13.05 | 0.99 |
| WT_0_2 | 5.62 | 61117544 | 59826984 | 1290560 | 97.89 | 14.12 | |
| WT_6_1 | 5.02 | 54468281 | 53756387 | 711894 | 98.69 | 7.79 | 1.00 |
| WT_6_2 | 4.80 | 52436258 | 51759879 | 676379 | 98.71 | 7.40 | |
| WT_12_1 | 5.04 | 54054132 | 53271826 | 782306 | 98.55 | 8.56 | 0.99 |
| WT_12_2 | 5.37 | 57646891 | 56798096 | 848795 | 98.53 | 9.29 | |
| WT_24_1 | 3.09 | 33003343 | 32498394 | 504949 | 98.47 | 5.52 | 0.80 |
| WT_24_2 | 5.38 | 53962216 | 53018166 | 944050 | 98.25 | 10.33 | |
| WT_48_1 | 6.32 | 62796702 | 61500435 | 1296267 | 97.94 | 14.18 | 0.99 |
| WT_48_2 | 5.14 | 34693098 | 34020090 | 673008 | 98.06 | 7.36 | |
| WT_72_1 | 6.43 | 63625946 | 62530758 | 1095188 | 98.28 | 11.98 | 0.99 |
| WT_72_2 | 5.20 | 51866662 | 50779793 | 1086869 | 97.90 | 11.89 | |
| WT_96_1 | 6.33 | 63255801 | 62039033 | 1216768 | 98.08 | 13.31 | 0.99 |
| WT_96_2 | 6.11 | 61187496 | 60080963 | 1106533 | 98.19 | 12.11 |
-
Table 1—source data 1
Number of reads for each ORF of Myxococcus xanthus at 0, 6, 12, 24, 48, 72 and 96 hr of development.
- https://doi.org/10.7554/eLife.50374.004
-
Table 1—source data 2
RPKM values of the developmental time course and correlation scores.
RPKM values reported here were calculated from the total number of non-tRNA/rRNA containing reads. The old (MXAN_) locus tags, new gene identifiers (MXAN_RS), gene name and predicted functions or pathways in which they have been previously implicated are included. The number of missing data points and fold change were used as criteria for the developmental gene analysis. DG or reason that genes were not included in the DGs is indicated.
- https://doi.org/10.7554/eLife.50374.005
| Reagent type (species) or resource | Designation | Source or reference | Identifiers | Additional information |
|---|---|---|---|---|
| Strain, strain background (Myxococcus xanthus) | DK1622 | Kaiser, 1979; Goldman et al., 2006 | Genome: NC_008095.1 | Wild-type strain used to obtain RNA |
| Strain, strain background (Myxococcus xanthus) | DK4322 | Kroos et al., 1986 | MXAN_RS20760; MXAN_4276 | spiA::Tn5-lacZ; Tn5 lac (Kmr) Ω4521 |
| Strain, strain background (Myxococcus xanthus) | DK4294 | Kroos et al., 1986 | MXAN_RS16790; MXAN_ 3464 | fmgE::Tn5-lacZ; Tn5 lac (Kmr) Ω4406 |
| Chemical compound, drug | DNase I | Sigma-Aldrich | Cat No./ID: AMPD1 | |
| Chemical compound, drug | SuperScript III Reverse Transcriptase | Invitrogen | Cat No./ID: 18080044 | |
| Chemical compound, drug | Escherichia coli DNA polymerase | New England Biolabs | D1806 | |
| Chemical compound, drug | E. coli RNAse H | Invitrogen | Cat No./ID: 18021071 | |
| Chemical compound, drug | Proteinase K | Ambion | Cat No./ID: 25530–015 | |
| Chemical compound, drug | E. coli DNA ligase | New England Biolabs | Cat No./ID: M0205L | |
| Chemical compound, drug | lysozyme | Roche | Cat No./ID: 10837059001 | |
| Commercial assay or kit | RNeasy Midi Kit | Qiagen | Cat No./ID: 75142 | |
| Commercial assay or kit | RNA Protect Bacteria Reagent | Qiagen | Cat No./ID: 76506 | |
| Software, algorithm | BWA software | Li and Durbin, 2009 | ||
| Software, algorithm | SAMtools | Li et al., 2009 | ||
| Software, algorithm | Artemis v.16.0.0 | Rutherford et al., 2000 | ||
| Software, algorithm | Cluster 3 Software Package | de Hoon et al., 2004 |
Additional files
-
Transparent reporting form
- https://doi.org/10.7554/eLife.50374.029





