Jagged and Delta-like ligands control distinct events during airway progenitor cell differentiation
Figures
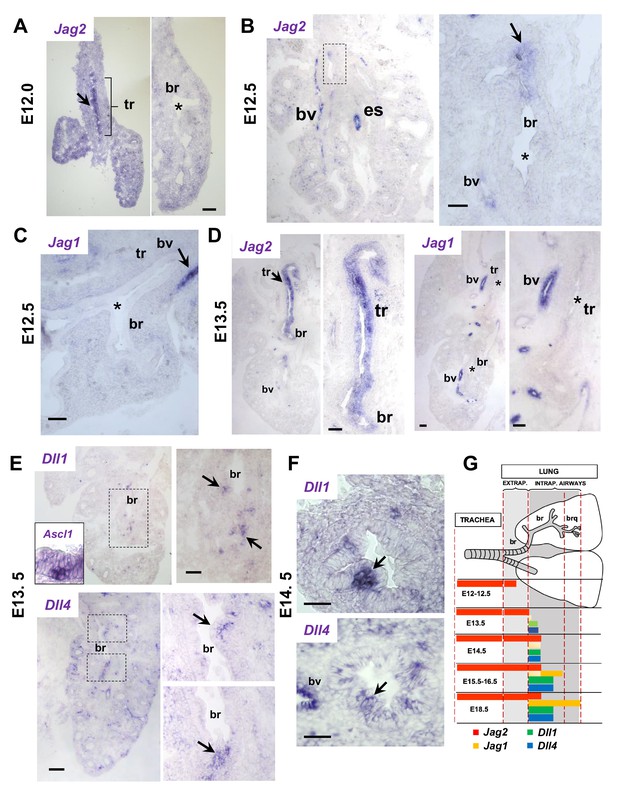
Jag and Dll ligands arise in distinct spatial and temporal patterns in airway epithelial progenitors undergoing cell fate commitment.
In situ hybridization of E12-E14.5 lungs. (A–C) E12-E12.5: Jag2 is the first and only Notch ligand detected in the epithelium, initially restricted to the trachea (tr) and later extending to extrapulmonary airways but not intrapulmonary large airways (bronchi: br)(B, boxed area enlarged in right panel). Jag1 is absent from both trachea and bronchi. Strong Jag ligand signals in blood vessels (bv) and the esophageal epithelium (es). (D) E13.5: Prominent epithelial Jag2 signals still restricted to trachea and extrapulmonary bronchi; abundant Jag1 in vascular but not in the epithelial compartment. (E–F) E13.5- E14.5: Dll1 and Dll4 signals are first detected at E13.5 in epithelial cell clusters of intrapulmonary main bronchi at sites of NEB formation marked by Ascl1 (inset) and signals become highly localized and prominent from E14.5 onward (G) Schematic summarizing the time and sites of Notch ligand expression throughout the developing respiratory tract epithelium. Arrows depict representative signals in airway epithelium; (*) depicts absence or near background signal. Bars in A-F = 40 μm.
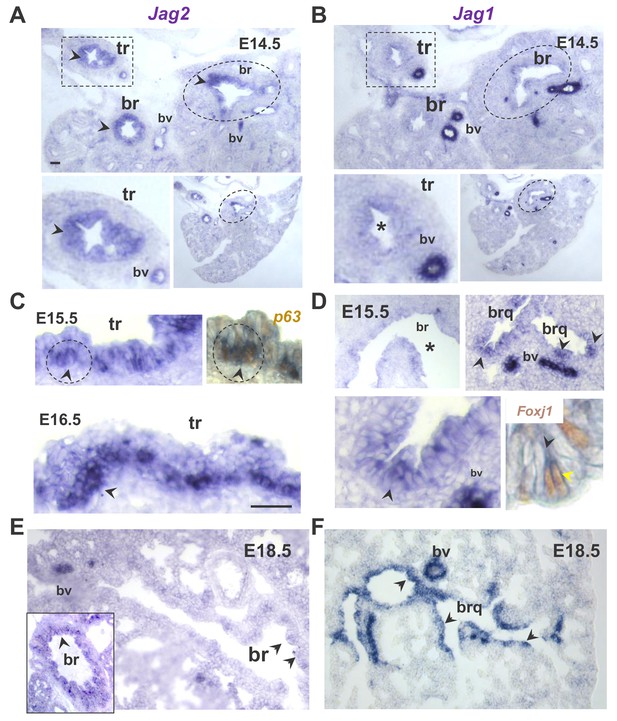
Distinct spatial and temporal patterns of Jag2 and Jag1 in the developing airway epithelium.
In situ hybridization (ISH) of Jag genes in E14.5-E18.5 wild type mouse lungs. (A–B) Prominent Jag2 signals in tracheal (tr, rectangles) and bronchial (br, circles) epithelium, but nearly background Jag1 signals at E14.5 (lower panels: different magnifications). (C–D) Strong Jag2 expression in the E15.5-E16.5 tracheal epithelium enriched in basal cell progenitors as shown by overlapping Jag2-p63 signals (right panel: double Jag2 ISH- p63 immunostaining); (D) Jag1 epithelial signals are weak in main bronchi but increased more distally in bronchioles (brq) preferentially in multiciliated cells (right lower panel: double Jag1 ISH-Foxj1 immunostaining). (E–F) At 18.5 Jag2 signals are mostly in main bronchi (inset) and nearly absent in intrapulmonary airways. By contrast Jag1 is strongly expressed throughout the airway epithelium. Arrowheads depict representative epithelial signals; bv: blood vessels: are always positive for Jag1 and Jag2. (*) depicts absence of signal or nearly background expression levels. Bars in A and C = 40 μm.
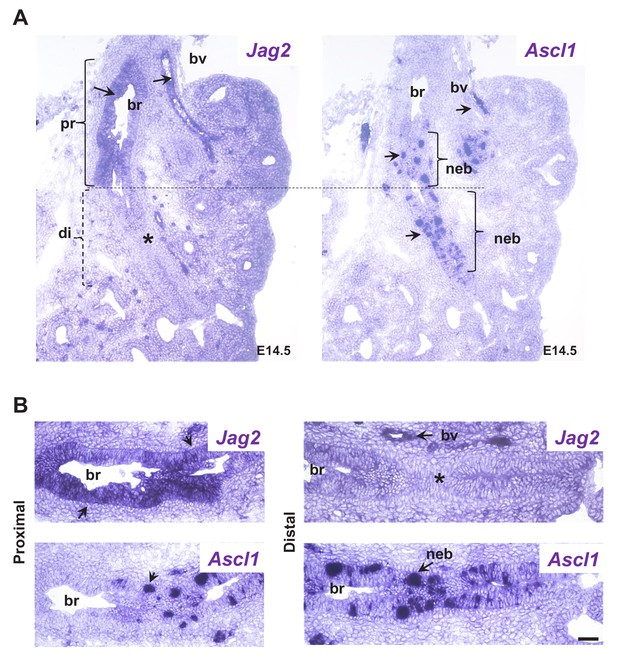
NEBs emerge in non-Jag expressing domains of the developing airway epithelium.
In situ hybridization (Jag2, Ascl1) in serial sections of E14.5 lungs (A) low magnification image showing both Ascl1-expressing cell clusters (NEBs) and Jag2 epithelial signals in proximal (pr) but not in more distal (di) main bronchus (br), where NEBs emerge in Jag2 negative (*) epithelium. Dotted line depicts the Jag2 positive and negative domains (in brackets). (B) Higher magnification of the bracketed areas in (A). Strong signals in blood vessels (bv) and nerves (nv) confirm specificity of Jag2 and Ascl1 staining, respectively (see also Figure 1—figure supplement 1 for Jag1 expression). Scale bar represents 40 μm.
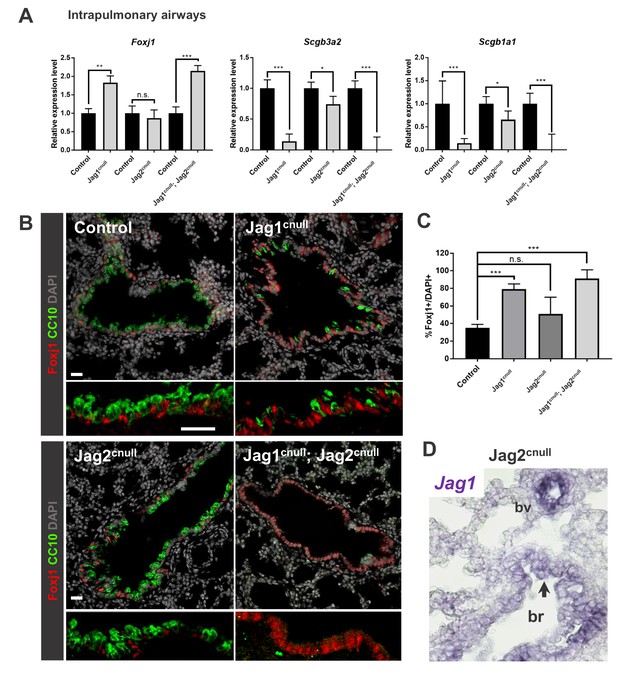
Jag1 and Jag2 collectively contribute to balance multiciliated and secretory cell fate in intrapulmonary airways.
(A) qPCR analysis: markers of multiciliated (Foxj1) and secretory cell fate (Scgb3a2, Scgb1a1 [CC10]) in mutant and respective control lung homogenates (n = 4 Jag1 control, n = 3 Jag1cnull; n = 4 Jag2 control; n = 4 Jag2cnull; n = 4 Jag1/Jag2 control; n = 4 Jag1cnull; Jag2cnull). Decreased expression of secretory markers and increased Foxj1 predominantly in Jag1cnull and in double Jag1cnull; Jag2cnull. Graphs represent mean ± SEM. Student’s t-test: differences statistically significant at *p<0.05, **p<0.005, ***p<0.005, n.s., not significant. (B) Immunofluorescence of lungs from E18.5 control and Jag mutant mice showing changes in expression of Foxj1 and CC10 in intrapulmonary airways consistent with that revealed by qPCR (DAPI in gray). (C) Morphometric analysis: percentage of Foxj1+ cells in control and mutant intrapulmonary airways (normalized by DAPI). Significant increase in the number of Foxj1 labeled cells in Jag1cnull and in Jag1cnull; Jag2cnull, but not in Jag2cnull mutants. (D) In situ hybridization of Jag1 in E18.5 Jag2cnull showing epithelial signals in intrapulmonary airways (arrow, bronchi: br) and in blood vessels (bv). Scale bar in B = 40 μm.
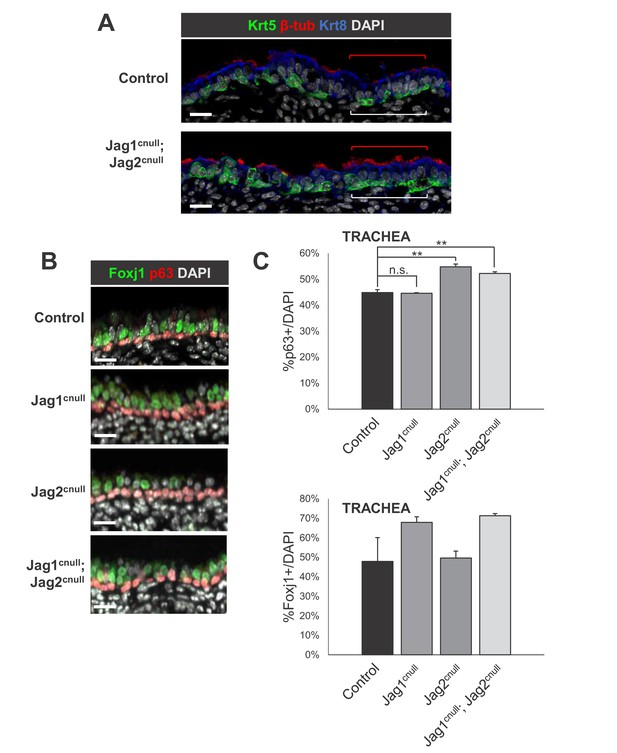
Loss of Jag ligands results in expansion of the embryonic basal cells and multiciliated cells in the E18.5 extrapulmonary airways (trachea).
IF of basal cells (A: Krt5, B: p63) and multiciliated cells (A: beta-tubulin 4; B: Foxj1) in control and double Jag1cnull; Jag2cnull tracheas. (C) Morphometric analysis of basal cells (top: p63+) and multiciliated cells (bottom: Foxj1 +) in single and double Jag mutant tracheas. Sections of whole trachea were counted for p63+ and DAPI+ cells and the percentage of p63+ cells was determined in airway epithelium. Graphs are mean ± SEM; **p<0.05. Scale bars = 20 μm.
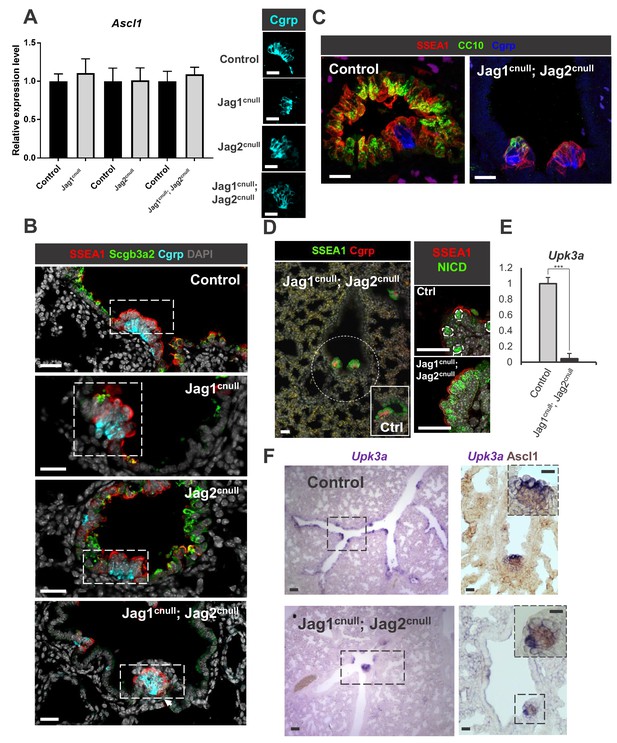
Disruption of Jagged-driven Notch signaling has no detectable impact on PNEC and NEB cell fate specification or maintenance.
(A) qPCR analysis of Ascl1 in E18.5 controls, single and double Jag1cnull; Jag2cnull mutants showing no significant difference in expression. Graph represents mean ± SEM (n = 4 Jag1 control, n = 3 Jag1cnull; n = 4 Jag2 control; n = 4 Jag2cnull; n = 4 Jag1/Jag2 control; n = 4 Jag1cnull; Jag2cnull). Student’s t-test was used to analyze data. Side panels: representative Cgrp immunofluorescence (IF) in controls and mutants (B) IF of secretory (Scgb3a2), NEB-associated SSEA1 (CC) and Cgrp (PNEC/NEB) in control and Jag mutants. (C, D) Preserved NEB microenvironment in E18.5 Jag1cnull; Jag2cnull double mutants: IF showing NEBs (Cgrp) and NEB-associated CCs (SSEA1, N1ICD, CC10 low) in double null mutants similar to controls (Ex. inset in D). (E) qPCR analysis of Upk3a in E18.5 control and double Jag1cnull; Jag2cnull mutants showing nearly abolished Upk3a expression in mutants (n = 3 in each group). Graph: mean ± SEM; ***p<0.0005 (F) ISH of Upk3a (left panels) and double labeled with Ascl1 (immunohistochemistry, right panels): Upk3a expression predominantly in NEB-associated CCs with rare signals scattered in CC elsewhere. In double Jag1cnull; Jag2cnull mutants CCs are abolished but in the NEB microenvironment and Upk3a becomes restricted to those surrounding NEBs. Scale bar in A-D, F = 40 μm. Scale bar in F inset = 20 μm.
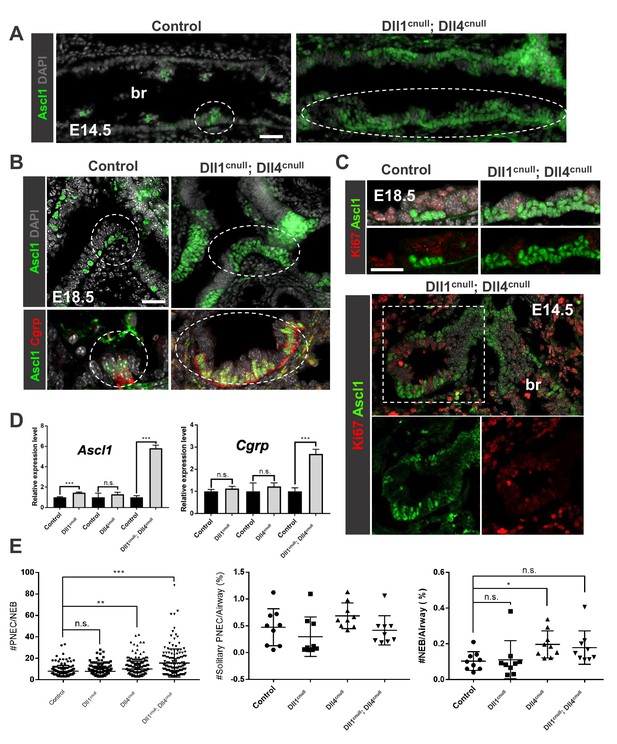
Loss of Dll-driven Notch signaling results in expansion of the NEBs.
(A–B) Ascl1 immunofluorescence (IF) in E14.5 control lungs (A) showing discrete clusters of Ascl1+ NEBs in large intrapulmonary airways (bronchi: br) in contrast to the aberrant NEB expansion in Dll1cnull; Dll4cnull lungs. At E18.5 (B) NEBs at branchpoints are also enlarged in mutants compared to controls and express Cgrp (bottom panel). (C) Double Ki67; Ascl1 IF shows no evidence that NEB expansion results from increased proliferation at E18.5 (top) or E14.5 (bottom panel). (D) qPCR analysis of NE markers at E18.5: significant increase in Ascl1 and Cgrp expression in double null mutants compared to controls; single mutant Dll1cnull but not Dll4cnull showed increased expression of Ascl1 (n = 3 Dll1 control, n = 5 Dll1cnull for Ascl1 and n = 4 Dll1 control, n = 5 Dll1cnull for Cgrp; n = 3 Dll4 control n = 6 Dll4cnull for Ascl1 and Cgrp; n = 3 Dll1/Dll4 control; n = 3 Dll1cnull; Dll4cnull for Ascl1 and Cgrp). Graphs: mean ± SEM. ***p<0.0005; n.s., not significant by Student’s t-test (E) Morphometric analysis of PNECs and NEBs in E14.5 control and Dll mutant lungs. Left panel: NEB size as determined by number of PNECs per NEB. Center panel: solitary PNEC per airway. Right panel: number of NEBs. * P<0.05, ** P<0.005, *** P<0.0005; n.s., not significant by Student’s t-test. Scale bars in A, B = 40 μm.
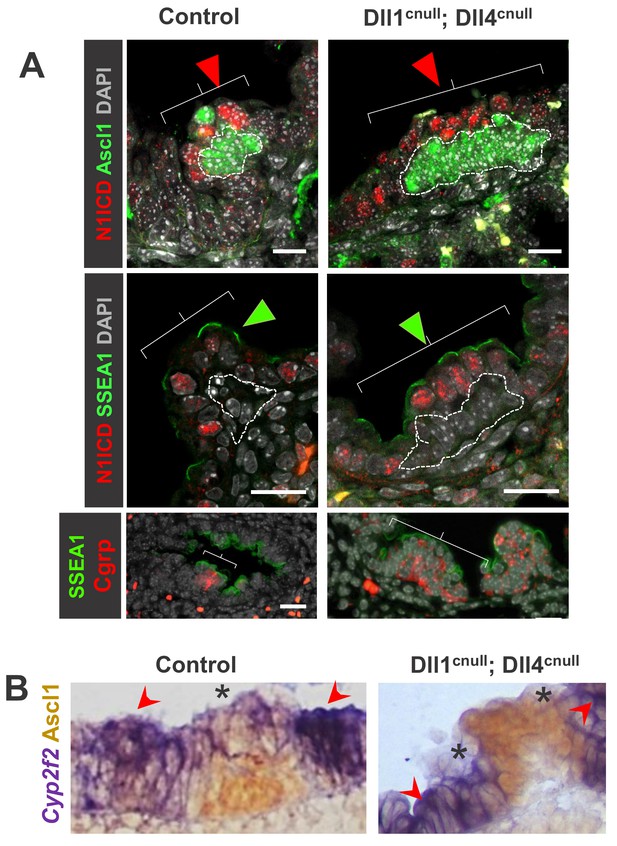
Expansion of the NEB-associated Club cells (CC) in the absence Delta ligands.
(A) Double immunofluorescence of E18.5 control and Dll1cnull; Dll4cnull lungs: Ascl1 and Cgrp expression in control and in the abnormally expanded NEBs (dotted areas) of mutants; robust N1ICD and SSEA1 expression in NEB-associated CCs (arrowheads and brackets). (B) Double ISH (Cyp2f2)/immunohistochemistry (Ascl1) showing strong Cyp2f2 expression (arrowheads) in cells flanking the NE microenvironment but only low signals in the NEB-associated CCs (asterisks) of both control and Dll1cnull; Dll4cnull mutants. Note reciprocal high (arrowhead) and low (*) intensity of signals in areas inside and outside the NEB microenvironment. Scale bars in A = 20 μm.
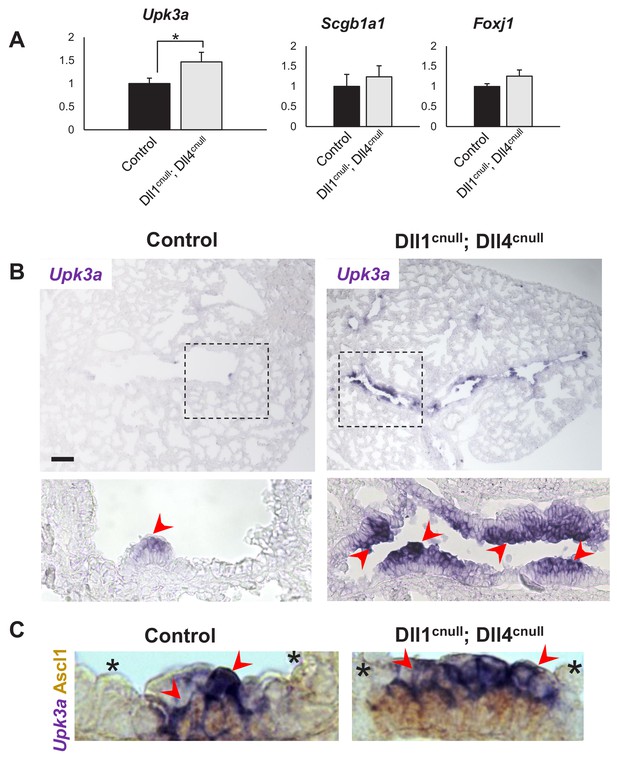
Expansion of the Upk3a expression domain in Dll1cnull; Dll4cnull lungs.
(A) qPCR analysis: significantly increased expression of Upk3a, but not of Scgb1a1 or Foxj1 in mutants relative to controls (n = 3 in both groups). Graphs are mean ± SEM. Student’s t-test *p<0.05. (B) ISH for Upk3a in E18.5 lungs showing marked expansion of the Upk3a expression domain (arrowheads) in intrapulmonary airways of Dll1cnull; Dll4cnull mutants (boxed areas enlarged in the lower panels). (C) Double immunohistochemistry (Ascl1)/ISH (Upk3a) confirms that the Upk3a+ cells (arrowheads) are NEB-associated CCs. Note reciprocal high (arrowhead) and low (*) intensity of signals in areas outside and inside the NEB microenvironment, respectively. Scale bar in B = 40 μm.
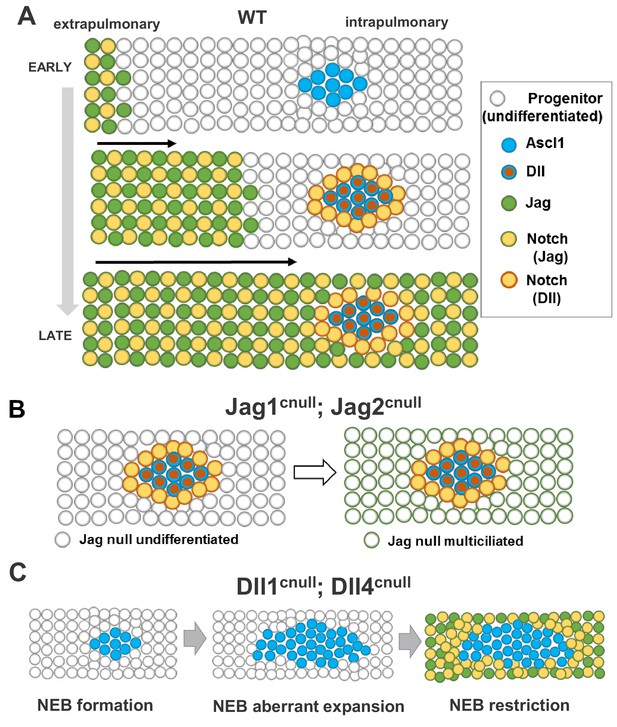
Proposed model for ontogeny and function of Notch ligand families in airway epithelial progenitors.
In all panels cells expressing Jag (1, 2) or Dll (1, 4) ligands are represented collectively; Notch activated by Jag or Dll are depicted distinctly (box, right). (A) Distinct spatial and temporal onset of expression of Jag and Dll in wild type differentiating airways. Top row: Jag expression (green dots) and Jag-Notch activation (yellow dots outlined in green) initiate in trachea/extrapulmonary airways but no Jag or Dll is present in intrapulmonary airways where NE/NEBs exist (Ascl1, blue circles). Middle row: Jag expression continues to advance into intrapulmonary airways in a proximal-to-distal fashion to activate Notch signaling and establish the balance of secretory vs multiciliated cell fates. In intrapulmonary airways Dll ligands are induced in NEBs (red dots outlined in blue) and Dll-Notch signaling is activated in adjacent cells to form NEB-associated club cells (CC, yellow dots outlined in red), limiting NEB expansion. Bottom row: As differentiation proceeds, Jag-Notch activation advances more distally and surrounds the NEB and associated CCs. (B) In Jag1cnull; Jag2cnull mutants the size and composition of the NEB microenvironment in intrapulmonary airways are unaffected since Jag ligands are not required for NE specification or Dll induction. Later, the NEB microenvironment is surrounded by Jag null multiciliated cells and remains unperturbed (right). (C) Dll1cnull; Dll4cnull double mutants: NE/NEB formation is initiated in intrapulmonary airways (left, compare to A). However the inability of Ascl1 cells to induce Dll ligands prevents induction of Notch-Dll signaling in surrounding undifferentiated progenitors, leading to aberrant expansion of NEBs (middle). Later, as Jag expression advances in intrapulmonary airways, Jag-Notch is activated in the progenitor cells surrounding the NEB, generating CCs and restricting its expansion (right, see Discussion).
Tables
| Reagent type (species) or resource | Designation | Source or reference | Identifiers | Additional information |
|---|---|---|---|---|
| Genetic reagent (M. musculus) | Dll1flox | PMID: 16495313 | MGI: 3044907 | Dr. Julian Lewis |
| Genetic reagent (M. musculus) | Dll4flox | PMID: 18824585 | MGI: 3828266 | Dr. Freddy Radtke (École polytechnique fédérale de Lausanne) |
| Genetic reagent (M. musculus) | Jag1flox | PMID: 16495313 | MGI: 3623344 | Dr. Julian Lewis |
| Genetic reagent (M. musculus) | Jag2flox | PMID: 20533406 | MGI: 4829504 | Dr. Thomas Gridley (Tufts University) |
| Genetic reagent (M. musculus) | Shhcre | Jackson Laboratory | Stock# 005622, RRID: IMSR_JAX:005622 | PMID: 15315763 |
| Antibody | anti-beta IV tubulin (mouse monoclonal) | Abcam | Cat#ab11315, RRID:AB_297919 | IF (1:100) |
| Antibody | anti-Ascl1 (mouse monoclonal) | Thermo Fisher Scientific | Cat# 14-5794-82, RRID: AB_2572887 | IHC/IF (1:100), IF requires tyramide amplification |
| Antibody | anti-CC10 (goat polyclonal) | Santa Cruz Biotechnology | Cat# sc-9772, RRID: AB_2238819 | IF (1:150) |
| Antibody | anti-Cgrp (rabbit polyclonal) | Sigma-Aldrich | Cat# C8198, RRID: AB_259091 | IF (1:2500) |
| Antibody | anti-Foxj1 (mouse monoclonal) | Thermo Fisher Scientific | Cat# 14-9965-82, RRID: AB_1548836 | IHC/IF (1:50) |
| Antibody | anti-Ki67 (rabbit monoclonal) | Cell Signaling Technology | Cat # 9129, RRID: AB_2687446 | IF (1:100) |
| Antibody | anti-Krt5 (rabbit polyclonal) | Biolegend | Cat# 905501, RRID: AB_2565050 | IF (1:500) |
| Antibody | anti-Krt8 (chicken polyclonal) | Abcam | Cat# ab107115, RRID: AB_10976462 | IF (1:500) |
| Antibody | anti-N1ICD (rabbit monoclonal) | Cell Signaling Technology | Cat# 4147, RRID: AB_2153348 | IF (1:100), requires tyramide amplification |
| Antibody | anti-p63 (rabbit polyclonal) | Santa Cruz Biotechnology | Cat# sc8343, RRID: AB_653763 | IHC/IF (1:400) |
| Antibody | anti-Scgb3a2 (goat polyclonal) | R and D Systems | Cat# AF3465, RRID: AB_2183550 | IF (1:100) |
| Antibody | anti-SSEA1 (mouse monoclonal) | EMD Millipore | Cat# MAB4301, RRID: AB_177627 | IF (1:300) |
| Antibody | anti-rabbit, Alexa Fluor 488 (donkey polyclonal) | Thermo Fisher Scientific | Cat# A21206, RRID: AB_2535792 | IF (1:300) |
| Antibody | anti-rabbit, Alexa Fluor 568 (donkey polyclonal) | Thermo Fisher Scientific | Cat# A10042, RRID: AB_2534017 | IF (1:300) |
| Antibody | anti-rabbit, Alexa Fluor 647 (donkey polyclonal) | Thermo Fisher Scientific | Cat# A31573, RRID: AB_2536183 | IF (1:300) |
| Antibody | anti-goat, Alexa Fluor 488 (donkey polyclonal) | Thermo Fisher Scientific | Cat# A11055, RRID: AB_2534102 | IF (1:300) |
| Antibody | anti-goat, Alexa Fluor 568 (donkey polyclonal) | Thermo Fisher Scientific | Cat# A11057, RRID: AB_2534104 | IF (1:300) |
| Antibody | anti-goat, Alexa Fluor 647 (donkey polyclonal) | Thermo Fisher Scientific | Cat# A21447, RRID: AB_2535864 | IF (1:300) |
| Antibody | anti-chicken, Alexa Fluor 488 (donkey polyclonal) | Jackson ImmunoResearch | Cat# 703-545-155, RRID: AB_2340375 | IF (1:300) |
| Chemical compound, drug | BM-Purple | Roche | Cat# 11442074001 | |
| Commercial assay, kit | anti-Mouse IgG (Peroxidase) polymer detection kit, made in horse | Vector Laboratories | Cat# MP-7402, RRID: AB_2336528 | |
| Commercial assay, kit | anti-Rabbit IgG (Peroxidase) polymer detection kit, made in horse | Vector Laboratories | Cat# MP-7401, RRID: AB_2336529 | |
| Commercial assay, kit | TSA Plus Cyanine 3 | Akoya Biosciences | Cat# NEL753001KT | Previously PerkinElmer |
| Commercial assay, kit | TSA Plus Cyanine 5 | Akoya Biosciences | Cat# NEL745001K | Previously PerkinElmer |
| Commercial assay, kit | ImmPACT DAB peroxidase (HRP) substrate | Vector Laboratories | Cat# SK-4105, RRID: AB_2336520 | |
| Commercial assay, kit | RNeasy mini kit | Qiagen | Cat# 74104 | |
| Commercial assay, kit | MAXIscript T7 | Thermo Fisher Scientific | Cat# AM1314M | |
| Commercial assay, kit | SuperScript IV First-Strand Synthesis System | Thermo Fisher Scientific | Cat# 18091050 |
Primers used to generate ISH probes.
T7 or T3 primers were added to the reverse or forward primers of each gene, respectively.
| Gene | Forward (5’ → 3’) | Reverse (5’ → 3’) |
|---|---|---|
| Dll1 | AATTAACCCTCACTAAAGGGAGACTGCTGAGAGAGGAAGGGAG | TAATACGACTCACTATAGGGAGAAGACCCGAAGTGCCTTTGTA |
| Dll4 | AATTAACCCTCACTAAAGGGAGACTACTCAGACACCCAGCTCC | TAATACGACTCACTATAGGGAGAATCCTTTGCAAGCCTCCTCT |
| Jag1 | AATTAACCCTCACTAAAGGGAGACGCCATAGGTAGAGTTTGAGG | TAATACGACTCACTATAGGGAGATAGTTGCTGTGGTTCTGAGC |
| Jag2 | AATTAACCCTCACTAAAGGGAGATGGCACCCAGAACCCTTG | TAATACGACTCACTATAGGGAGAATACTCCGTTGTTTTCCGCC |
| Ascl1 | AATTAACCCTCACTAAAGGGTCGTCCTCTCCGGAACTGAT | TAATACGACTCACTATAGGGAGAAGAAGCAAAGACCGTGGGAG |
| Upk3a | AATTAACCCTCACTAAAGGGGTGGCTGGACTGTGAACCTC | TAATACGACTCACTATAGGGTTGCCCACCCTGACTAGGTA |
| Cyp2f2 | AATTAACCCTCACTAAAGGGGGAACTTTGGAGGCATGAAA | TAATACGACTCACTATAGGGAACTCCTGAGGCGTCTTGAA |





