Magnetic resonance measurements of cellular and sub-cellular membrane structures in live and fixed neural tissue
Figures
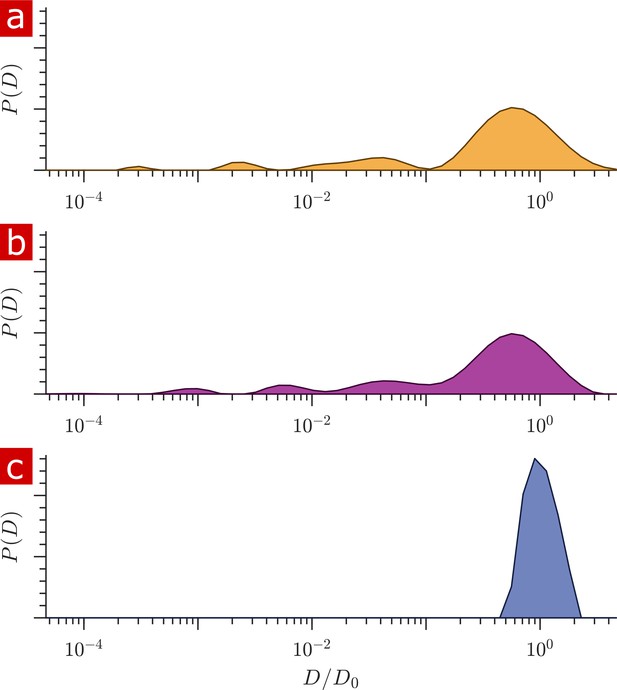
Diffusion coefficient distributions.
(a) Distribution of diffusion coefficients from a fixed spinal cord bathed in aCSF. (b) Distribution from the same spinal cord after removing the aCSF bath. (c) Distribution from only aCSF filling the RF coil.
-
Figure 1—source data 1
1-D Diffusion signal attenuation () and values for the measurements on fixed spinal cord bathed in aCSF (wet), the same spinal cord after removing aCSF (dry), and for aCSF (MATLAB structure array).
1-D Diffusion signal attenuation () and values for repeated measurements on aCSF (MATLAB structure array).
- https://cdn.elifesciences.org/articles/51101/elife-51101-fig1-data1-v2.mat
-
Figure 1—source data 2
1-D Diffusion signal attenuation () and values for repeated measurements on aCSF (MATLAB structure array).
- https://cdn.elifesciences.org/articles/51101/elife-51101-fig1-data2-v2.mat
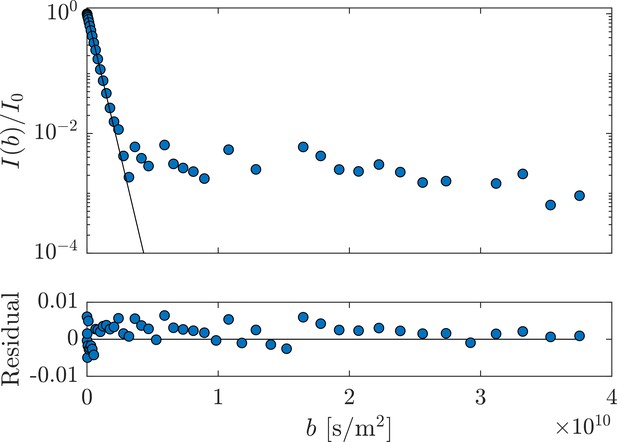
Measurement of aCSF at 25°C.
Single exponential fits of the first 15 of the 50 points, attenuating the signal to , provides D0 = 2.153 ± 0.014 × 10−9 m2/s from three measurements performed on only aCSF filling the coil. Fit residuals are also shown.
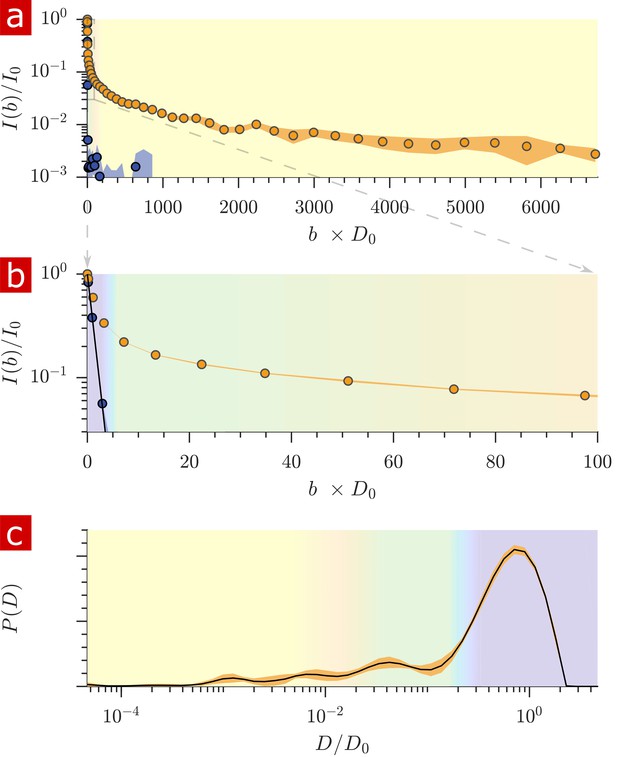
Diffusion measurements performed on a fixed spinal cord.
(a) Mean (circles) and SD (shaded bands) of the signal intensity from five diffusion measurements, spaced 54 min apart, performed on a fixed spinal cord (orange) and three measurements performed on aCSF (purple) at 25° C. (b) Signal intensity from the zoomed in area shown in (a). (c) The distribution of diffusion coefficients resulting from inversion of the data. The purple, green, and yellow shading across plots signifies water which is free, less restricted, and more restricted, respectively. Models of signal attenuation (see Appendix 1) are used to define the cutoffs for each of these regimes based on when their signal components would attenuate to exp (−6). Values are inverted to define the color shading on the distribution. This inversion of colors is simply to guide the eye. The color gradient is meant to signify a continuous change between diffusion regimes rather than sharp boundaries.
-
Figure 2—source data 1
1-D Diffusion signal attenuation () and values for measurements repeated every 54 min on a fixed spinal cord (MATLAB structure array).
- https://cdn.elifesciences.org/articles/51101/elife-51101-fig2-data1-v2.mat
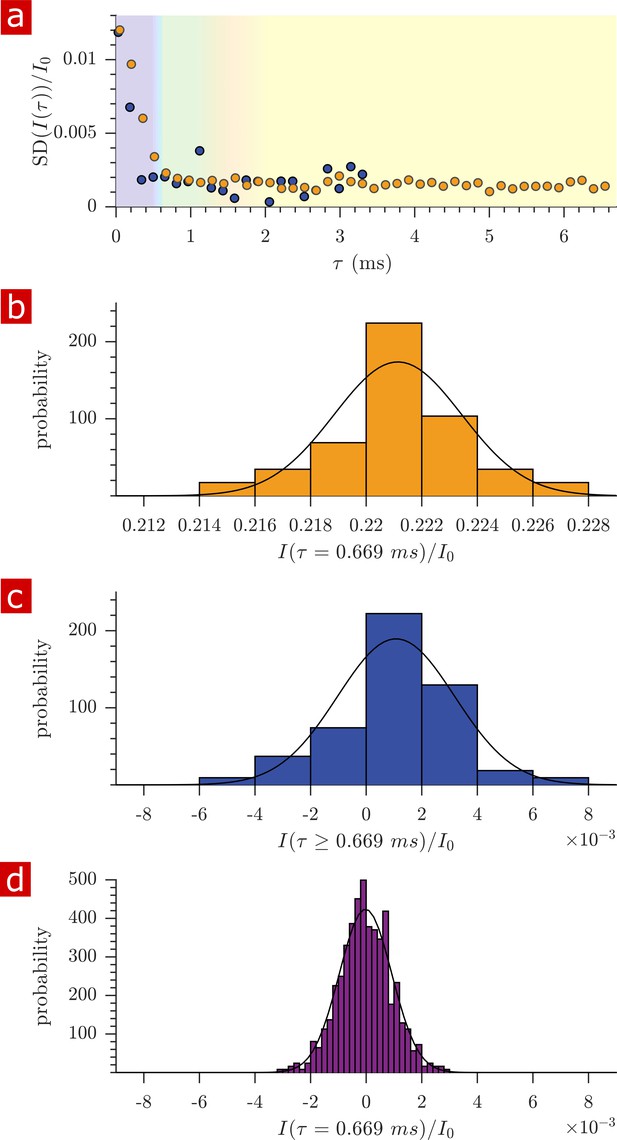
Variability of diffusion data from measurements repeated on the same samples.
(a) Standard deviation (SD) of the signal intensity from 29 diffusion measurements, spaced 54 min apart, performed on a fixed spinal cord (orange) and three measurements performed back-to-back on aCSF (purple) at 25° C. Data is from the same samples as in Figure 2. (b) Histogram of signal intensity from measurements on the fixed spinal cord at ms with mean ± SD 0.2211 ± 0.0023. (c) Histogram from the measurements on aCSF at ms, after signal has decayed away, with mean ± SD 0.0011 ± 0.0021. (d) Histogram of noise from 621 diffusion measurements performed on a dry chamber with nothing filling the solenoid coil, with mean ± SD = −2× 10−5 ± 0.001. In (b), (c) and (d), Gaussian distributions are plotted using the mean and SD as parameters μ and . Note the similar window width in (b), (c), and (d) allow comparison of SD. (See also Appendix 2).
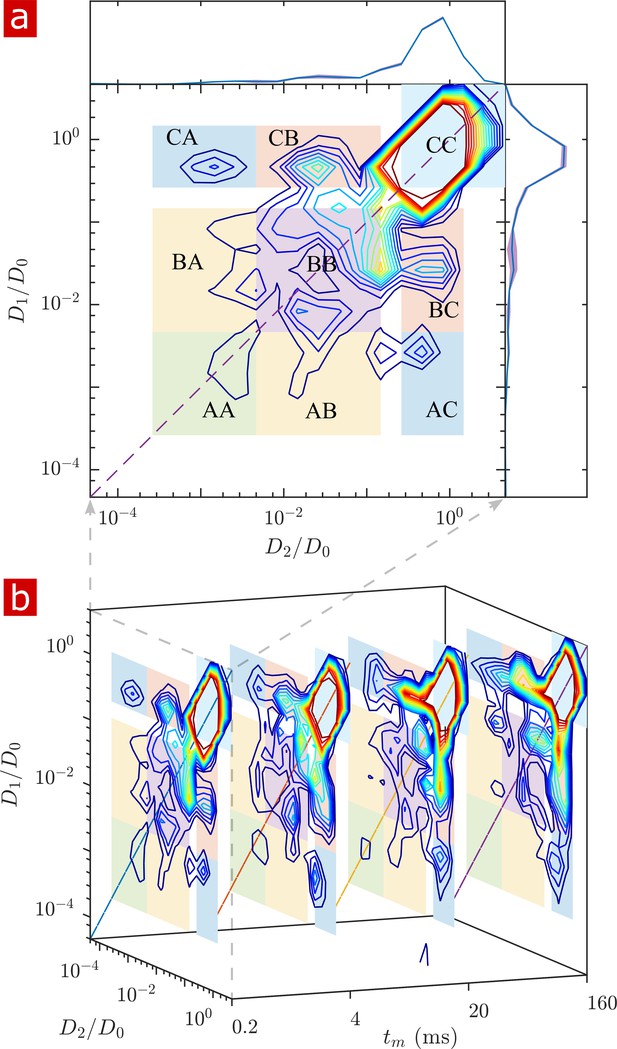
Full 2-D DEXSY diffusion exchange distribution for a fixed spinal cord.
(a) Exchange distribution measured with mixing time . Distributions show exchanging (off-diagonal) and non-exchanging (on-diagonal) components. These components are lumped into regions , , and , shaded and labeled in a 3 × 3 grid for each exchange combination. The range of is set to add detail to components and , but cuts off the top of the most mobile region . The marginal distributions and are presented on the sides, with mean (solid blue lines) and SD (shaded bands around lines) from three measurements. (b) A stacked view of distributions measured with mixing time . With increasing , the probability density builds up in regions of free and restricted water exchange and and decays for non-exchanging restricted water regions and .
-
Figure 3—source data 1
Full 2-D DEXSY datasets from measurements repeated three times at each mixing time on each of the fixed samples (MATLAB structure array).
- https://cdn.elifesciences.org/articles/51101/elife-51101-fig3-data1-v2.mat
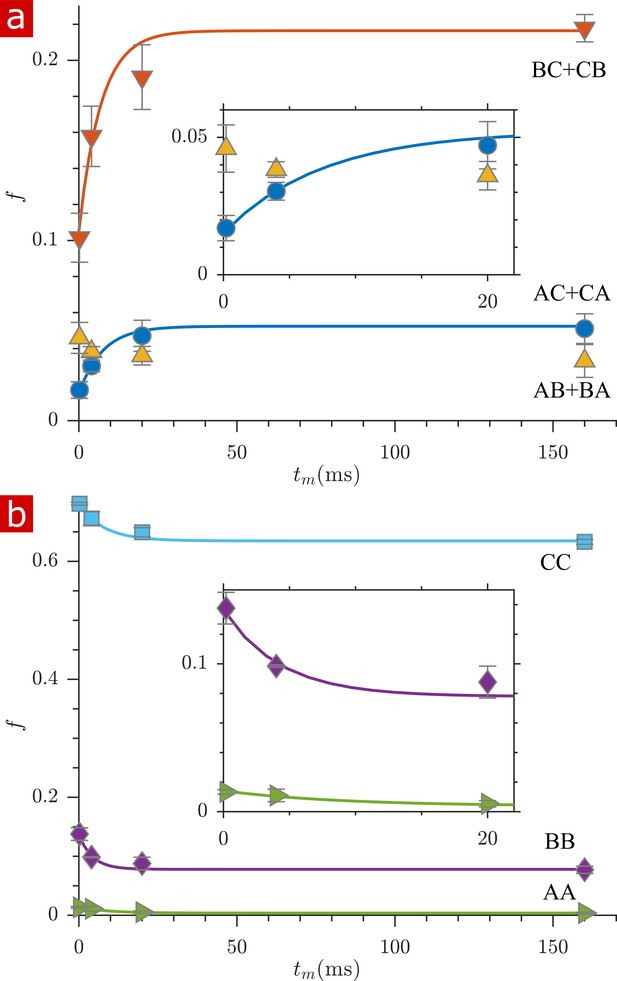
Fractions of exchanging and non-exchanging water.
(a) Fractions of water which exchange between regions , , and , (defined in Figure 3a) as a function of mixing time, . Exchange fractions and are fit with a first-order rate model with estimated apparent exchange rates AXRs (mean ± standard deviation from three measurements) 140 ± 80 and 140 ±140 s−1, respectively. does not follow the expected first-order rate model. (b) Fractions of non-exchanging water. Non-exchange fractions , , and are fit with a first-order rate model with estimated AXRs 110 ± 100, 230 ± 170, and 130 ±100 s−1 respectively. Insets zoom in on the initial rise in exchange. Error bars are the standard deviations from three repeat measurements.
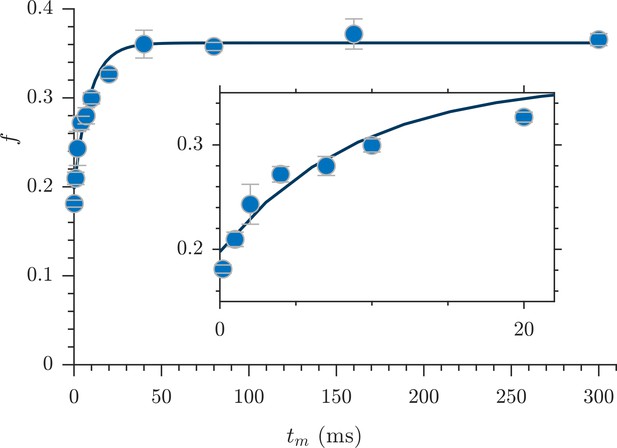
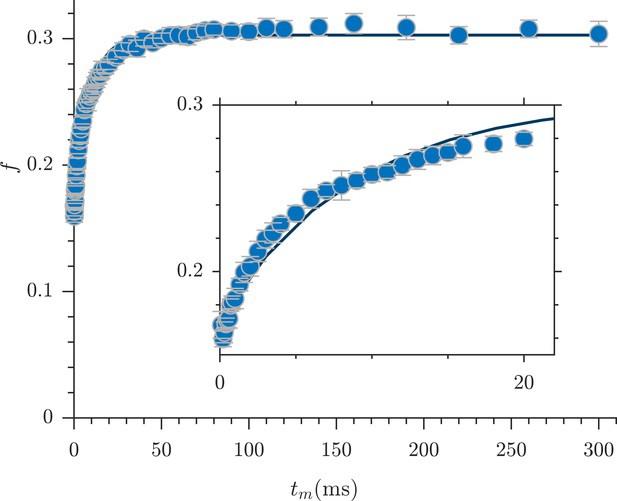
Rapid measurement of exchange fractions.
A fit of the first-order rate model estimated an apparent exchange rate, AXR = 110 ± 30 s−1 (mean ± SD from three measurements performed on one specimen at 25°C). Inset shows a zoomed-in region of the initial rise in exchange.
-
Figure 4—source data 1
Rapid exchange data for all fixed samples, including raw data from the four-point method () and the exchange fractions () (MATLAB structure array).
- https://cdn.elifesciences.org/articles/51101/elife-51101-fig4-data1-v2.mat
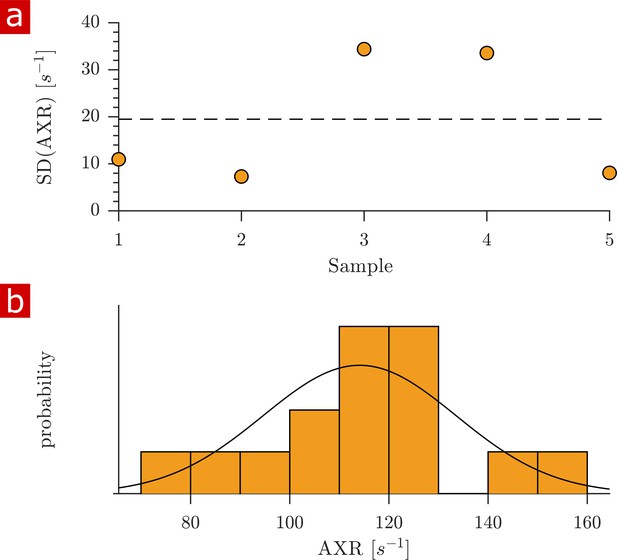
Variability of apparent exchange rates (AXRs).
(a) Standard deviations of AXRs from three rapid exchange measurements shown for five fixed spinal cord samples. The dashed line shows the SD from all (3 × 5) measurements. (b) Histogram of all measured AXRs with mean = 114 and SD = 19 s−1. A Gaussian distribution is plotted using the mean and SD as parameters μ and . An Anderson–Darling test indicates that the distribution is not dissimilar from a Gaussian distribution ().
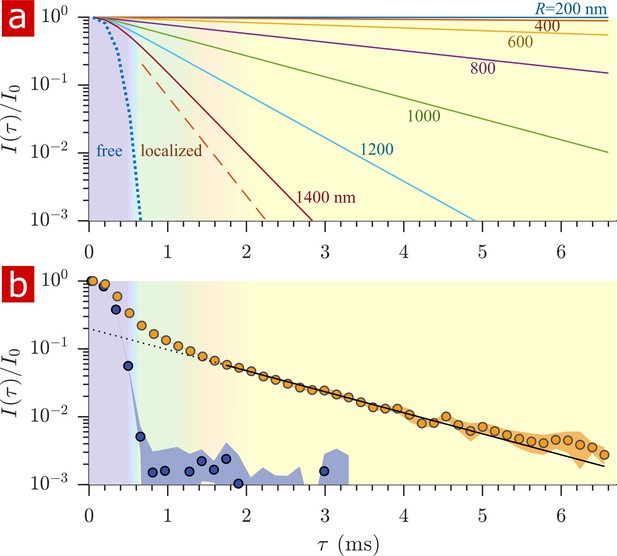
Sensitivity to membrane structure sizes from the diffusion signal attenuation.
(a) Signal intensity is simulated for water restricted in spherical compartments of varying radius between R = 200 − 1400 nm (solid lines) (Neuman, 1974), for water localized near surfaces in larger restrictions (red dashed line) (de Swiet and Sen, 1994; Hurlimann et al., 1995), as well as water diffusing freely (purple dotted line) (Woessner, 1961). Signal is plotted as a function of the variable (rather than ). (b) Signal is re-plotted from Figure 2. Signal at ms is fit with the model for water restricted in spherical compartments in the limit of long (solid black line) (Neuman, 1974) incorporating the AXR = 110 s−1(Carlton et al., 2000), estimating a radius R = 900 nm. The dotted black line extrapolates back to . (See Appendix 1 for model equations.) Color shading is similar to Figure 2.
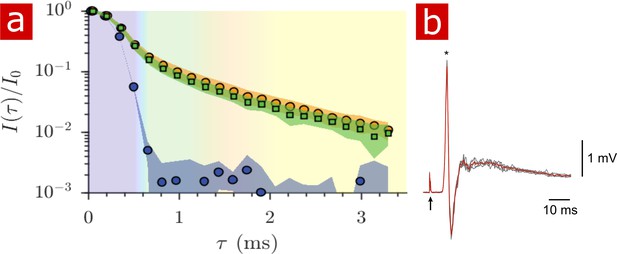
Diffusion in fixed vs. live.
(a) Signal intensity from diffusion measurements performed at 25° C on live samples (n = 9) (green squares), fixed samples (n = 6) (orange circles) and aCSF (purple circles) plotted as a function of the variable . (b) Mono- and poly- synaptic reflexes were recorded from the L6 ventral root of live samples (n = 4) after NMR measurements. Stimulation was done on the homonymous dorsal root. The grey lines are five successive stimuli (30 s interval) and the superimposed red line is the average signal. The arrow indicates the stimulus artifact and the star the monosynaptic reflex.
-
Figure 6—source data 1
1-D diffusion data for all fixed and live samples (MATLAB structure array).
- https://cdn.elifesciences.org/articles/51101/elife-51101-fig6-data1-v2.mat
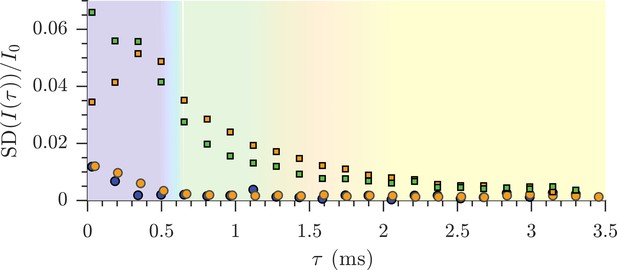
Sample-to-sample variability of diffusion data.
Standard deviations of the diffusion signal attenuation are presented for live (n = 9) (green squares) and fixed (n = 6) (orange circles) sample groups. The standard deviations from repeated measurements are also re-plotted from Figure 2—figure supplement 1a. The sample-to-sample variability is higher that the variability of repeated measurements due to structural and size differences between samples and the sensitivity of the measurement to these differences.
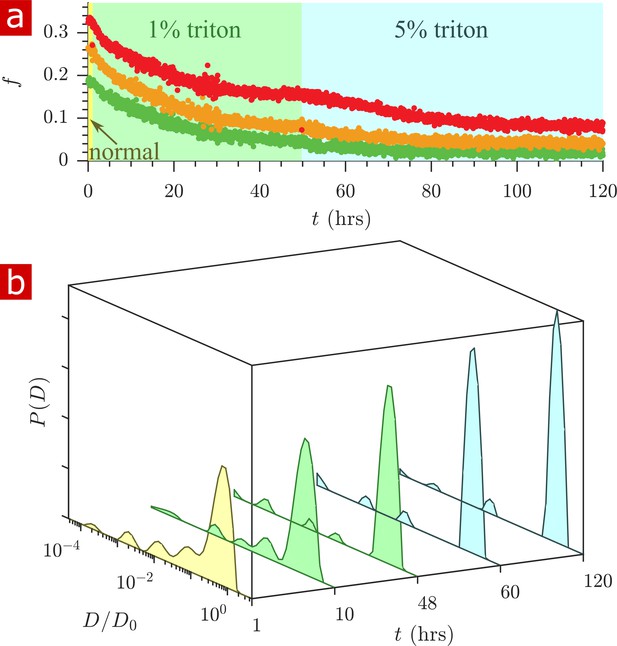
Timecourse study of Triton X delipidation.
(a) Exchange fractions from rapid exchange measurements with (green dots), 4 (orange dots), and 20 ms (red dots) measured throughout the timecourse, as the sample was washed to aCSF with 1% Triton X, and then 5% Triton X. (b) Representative diffusion coefficient distributions from 1-D diffusion measurements performed at different times before and after addition of Triton.
-
Figure 7—source data 1
1-D diffusion data for real-time delipidation of a fixed spinal cord (MATLAB structure array).
- https://cdn.elifesciences.org/articles/51101/elife-51101-fig7-data1-v2.mat
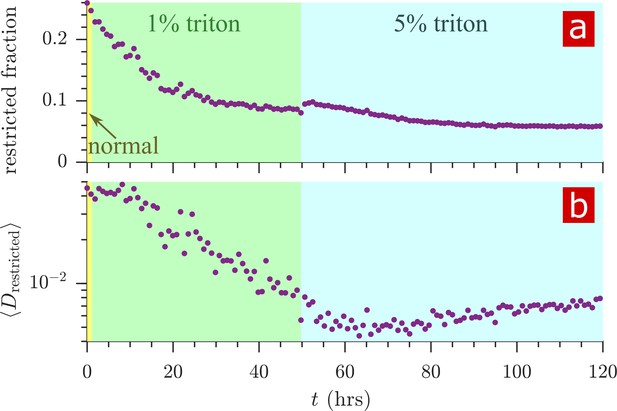
Timecourse of restriction during Triton X delipidation.
The fraction (a) and mean diffusion coefficient (b) of signal arising from restricted water (with ). A bump in the restricted fraction seen upon addition of the 5% Triton is due to the Triton X itself which forms 5 nm micelles (Paradies, 1980).
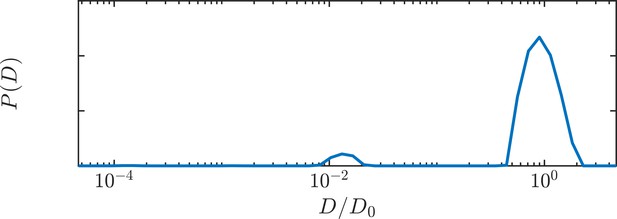
Diffusion coefficient distributions of 5% Triton X in aCSF.
An additional peak at is due to formation of 5 nm micelles (Paradies, 1980).
Timelapse video of diffusion coefficient distributions during delipidation.
Each frame is spaced 54 min apart and the whole video represents 120 hr of delipidation. See Figure 7 for times as which the sample was washed to aCSF with 1% Triton X and 5% Triton X. The final distribution at 120 hr looks very similar to the distribution for 5% TritonX in aCSF.
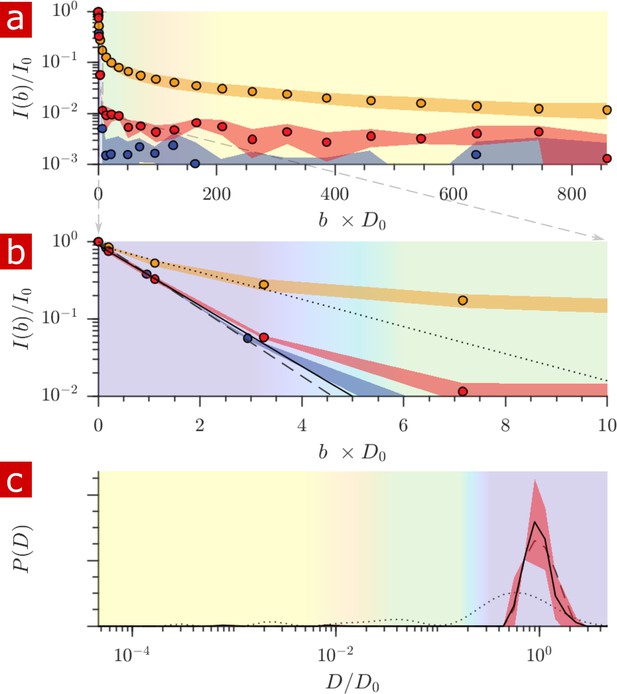
Diffusion measurement after delipidation.
(a) Diffusion signal intensity from measurements on spinal cords performed at 25° C after delipidation (n = 2) with 10% Triton X and after washing the Triton away. The mean (circles) and SD (shaded bands) of the attenuation are plotted for the delipidated samples (red) alongside pure aCSF (purple) and fixed undelipidated spinal cords (n = 6) (orange). (b) The initial attenuation of signal. Monoexponential fits of the attenuation from points 2–4 yielded 2.15 ± 0.02, 1.94 ± 0.02 and 0.87 ± 0.14 ×10−9 m2/s for the aCSF, delipidated, and undelipidated spinal cords and are shown as the dashed, solid, and dotted lines, respectively. (c) Diffusion coefficient distribution of the delipidated spinal cords, for which the mean (solid line) and SD (shaded band around line) are not significantly different from the pure aCSF (dashed line). The distribution from a fixed, undelipidated spinal cord (dotted line) is also shown for comparison. The purple, green, and yellow shading across the plots signifies water which is free, localized, and restricted.
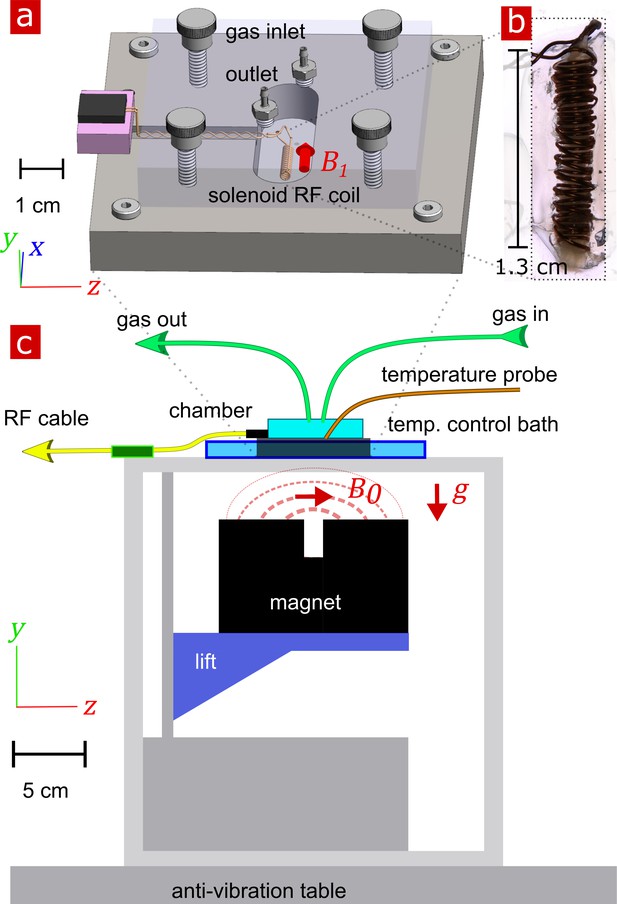
Experimental setup.
(a) 3-D technical drawing of the test chamber. (b) Image of the solenoid RF coil containing a fixed, delipidated specimen. (c) Technical drawing of the experimental setup. The magnet is drawn in the ‘service’ position to show the field lines extending from one magnetic pole to the other. To perform measurements, the magnet would be raised such that the was correctly positioned relative to the sample. Vectors , and point in the , , and directions respectively.
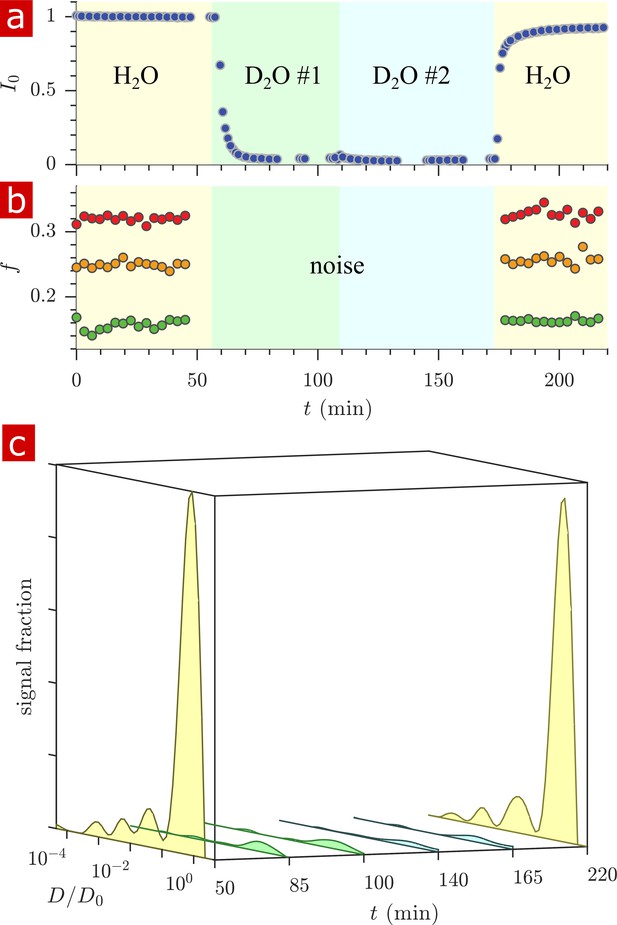
Timecourse study of D2O wash.
The sample was washed from aCSF to aCSF made with deuterated water in two steps and back to aCSF as shown by the yellow, green, blue, and yellow pastel color shadings for H2O, D2O #1, D2O #2, and H2O. (a) The proton signal intensity from rapid exchange measurement data normalized to remove effects at different mixing times. (b) Exchanging fractions from rapid measurements with (blue), 4 (red), and 20 ms (orange). (c) 1-D diffusion measurements were performed at points throughout the timecourse (seen as breaks in the data in (a)) and distributions are presented as signal fractions ().
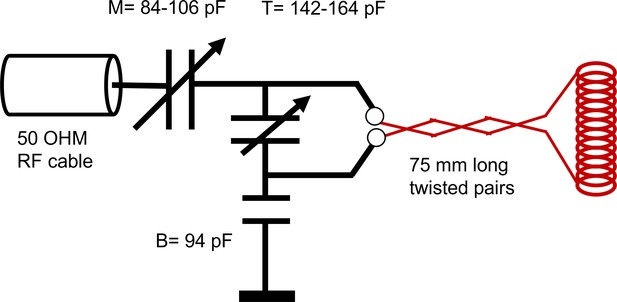
RF circuit design.
Drawing of the circuit for the RF showing the capacitance of the tune (T), match (M), and balance (B). Note a single wrapping of the solenoid was drawn rather than the actual double-wrap for visual simplicity.
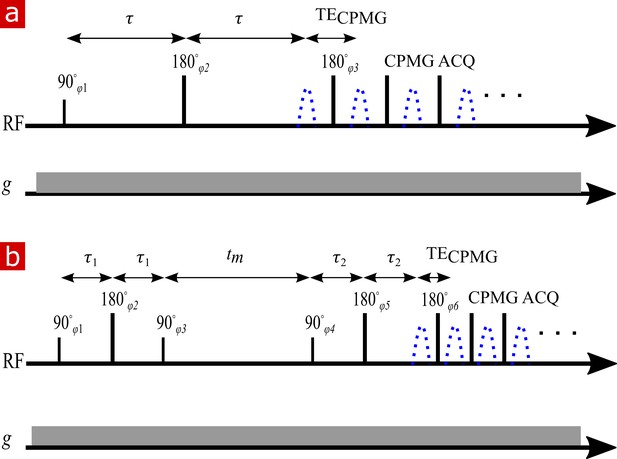
NMR diffusion and exchange pulse sequences.
(a) Spin echo pulse sequence for measuring diffusion with a static gradient. is varied to control . (b) DEXSY pulse sequence for measuring exchange with a static gradient. The two SE encoding blocks with and , varied independently to control and , are separated by . Signal is acquired in a CPMG train for both (a) and (b).
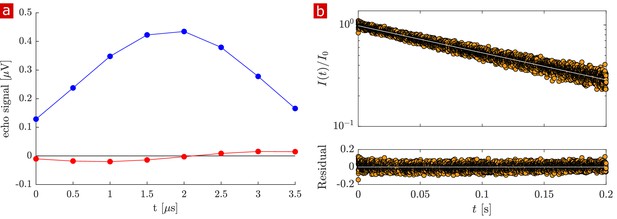
measurement.
Representative Carr-Purcell-Meiboom-Gill (CPMG) (10 s repetition time (TR), 8000 echoes, TE = 25 μs) on fixed spinal cord. (a) The echo shape summed over all echoes (real signal (blue) phased maximum and imaginary signal phased to zero (red)). (b) Real signal decay (orange circles) and exponential fit with ms and (white line) and residuals of the fit.
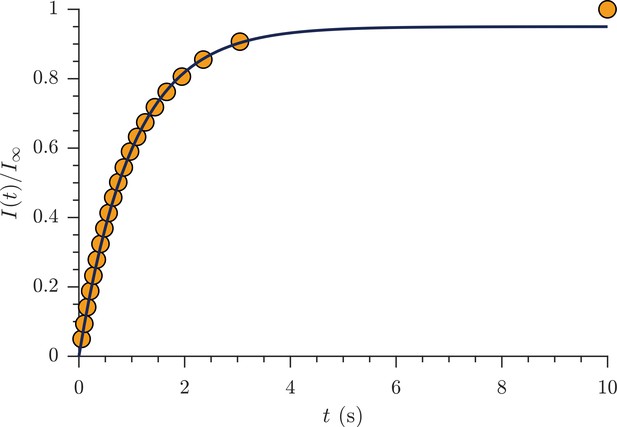
measurement.
Representative saturation recovery experiment on fixed spinal cord (1 s TR, 21 recovery time points logarithmically spaced from 50 ms to 10 s) showing signal intensity normalized by signal at 10 s recovery time (orange circles) and exponential fit with estimated T1 = 990 ms (solid black line).
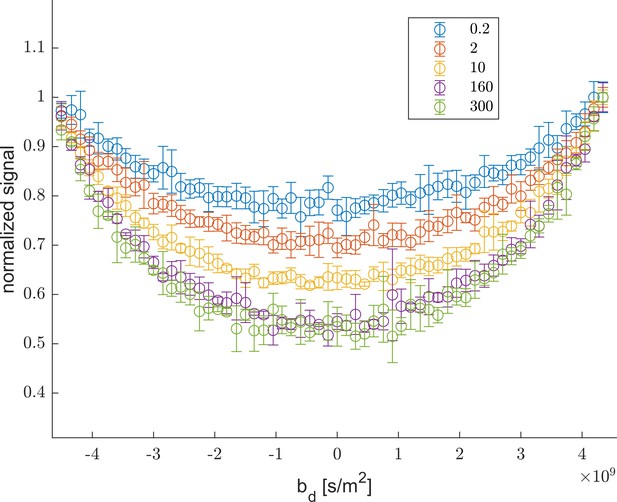
Rapid exchange test #1.
The curvature along slices of as a function of highlighting the increase in the depth of the curvature with increasing , shown in figure legend. The increased depth is due to increased exchange (Song et al., 2016; Cai et al., 2018).
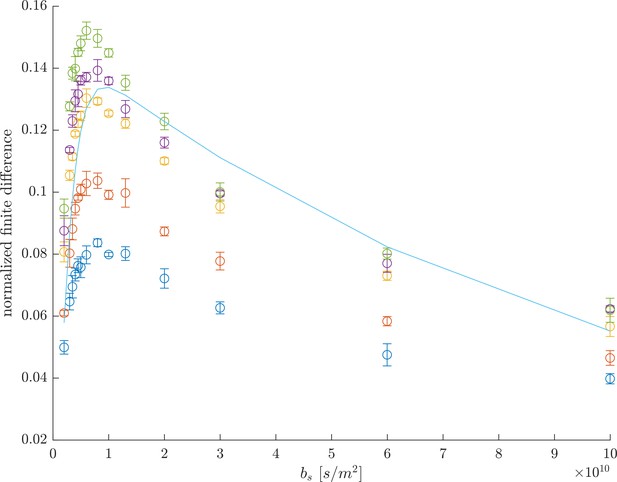
Rapid exchange test #2.
Difference between and , normalized by , as a function of , showing the optimal for measuring the largest curvature response as occurring near . The line is a prediction of the finite difference for a two-site system, from Equation 8 in Cai et al. (2018) using , , and .
Tables
Static gradient spin echo DEXSY Phase Cycles.
| φ1 | φ2 | φ3 | φ4 | φ5 | φ6 | φrec |
|---|---|---|---|---|---|---|
| 0 | +π/2 | 0 | 0 | +π/2 | π/2 | π |
| π | -π/2 | 0 | 0 | +π/2 | π/2 | 0 |
| 0 | +π/2 | π | 0 | +π/2 | π/2 | 0 |
| π | -π/2 | π | 0 | +π/2 | π/2 | π |
| 0 | +π/2 | 0 | π | -π/2 | π/2 | 0 |
| π | -π/2 | 0 | π | -π/2 | π/2 | π |
| 0 | +π/2 | π | π | -π/2 | π/2 | π |
| π | -π/2 | π | π | -π/2 | π/2 | 0 |