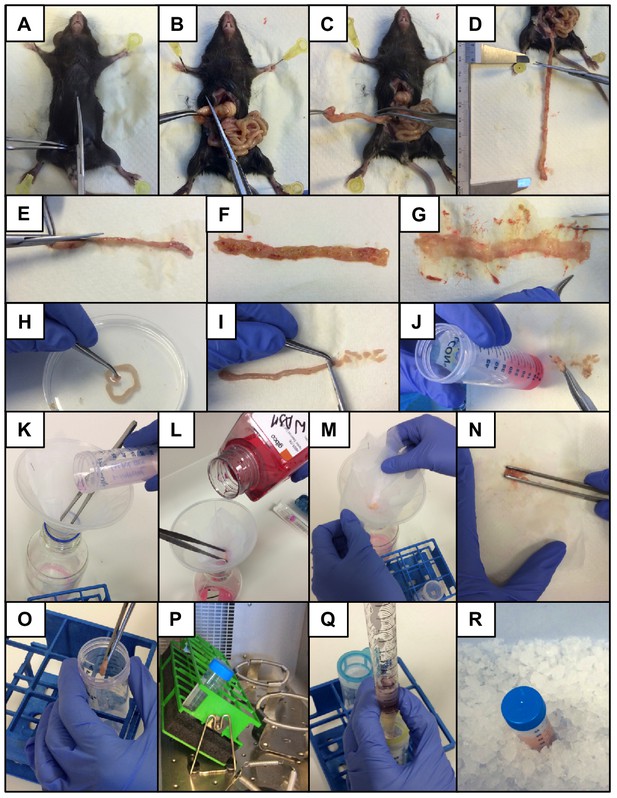
High-dimensional analysis of intestinal immune cells during helminth infection
Figures
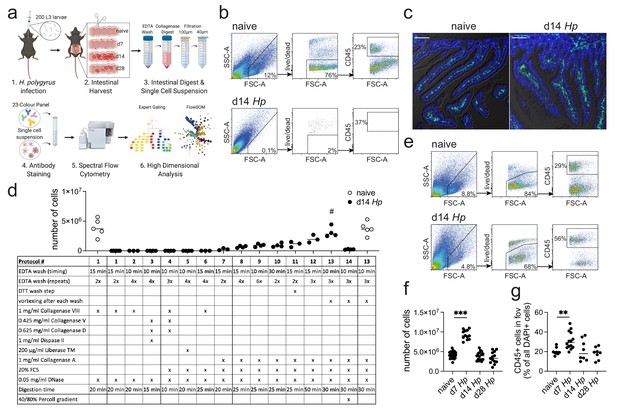
Optimization of a standard intestinal digestion protocol for the heavily infected duodenum.
(a) Schematic of a general intestinal digestion protocol (created with biorender.com). (b) Digest of naïve and day 14 hr. polygyrus (Hp)-infected duodenal segments using the standard digestion protocol. (c) Intestinal cryosections stained with CD45-FITC (green) and DAPI (blue) from naïve and day 14 infected intestines. Scale bar = 100 µm (representative of >10 sections from 3 to 5 mice per group and two independent experiments). (d) Number of live cells isolated from naïve or day 14 infected duodenal segments during the systematic optimization of the standard digestion protocol. Further details can be found in Figure 1—figure supplement 1 and Figure 1—figure supplement 2 (n = 3–5 samples per group, combined data from at least two independent experiments; # depicts the digestion protocol that yielded comparable cell numbers between naïve and infected samples, all other protocols showed a significant difference to the naïve group when compared by ordinary one-way ANOVA followed by Holm-Sidak’s multiple comparisons test). (e) Digest of naïve and day 14 infected duodenal segments using the optimised Hp digestion protocol (#13). (f) Number of live cells isolated from naïve, day 7, day 14 and day 28 infected duodenal segments using the optimized Hp digestion protocol (n > 12 samples per group, combined data from at least three independent experiments; Kruskal-Wallis followed by Dunn’s multiple comparisons test compared to the naïve group; ***p≤0.001). (g) Quantification of CD45+ cells present in the field of view (fov, 635.90µm x 635.90µm) in cryosections from the same timepoints (representative of >10 sections from 3 to 5 mice per group from two independent experiments; Kruskal-Wallis followed by Dunn’s multiple comparisons test compared to the naïve group; **p≤0.01).
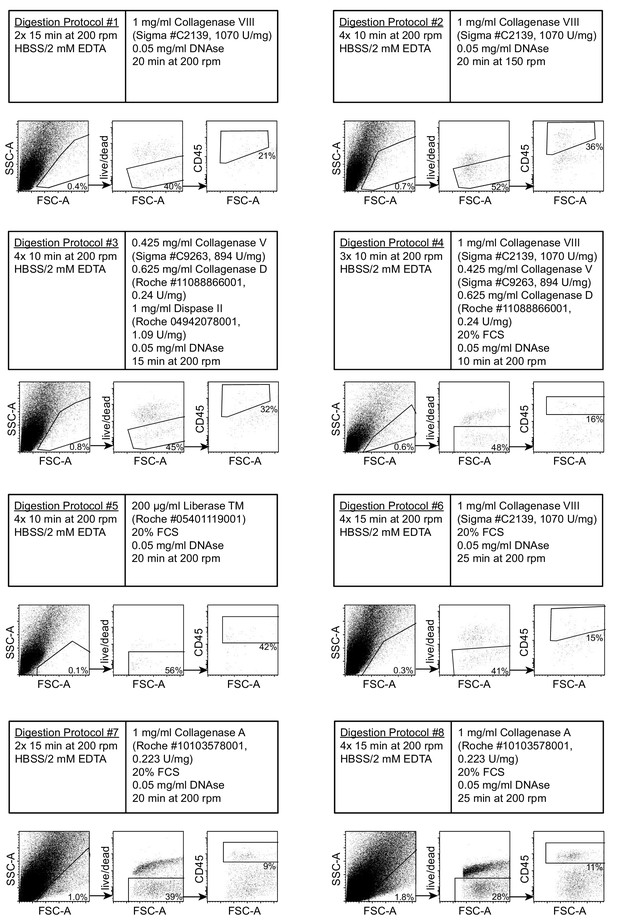
Modification of a standard intestinal digestion protocol to isolate single cells from heavily infected duodenal segments.
Dots plots of acquired events from day 14 hr. polygyrus-infected duodenal segments using digestion protocols #1–8 (representative of 3–5 samples per group from at least two independent experiments). Details for each digestion protocol are annotated. The gating strategy shows cells of interest, viability and CD45 staining.
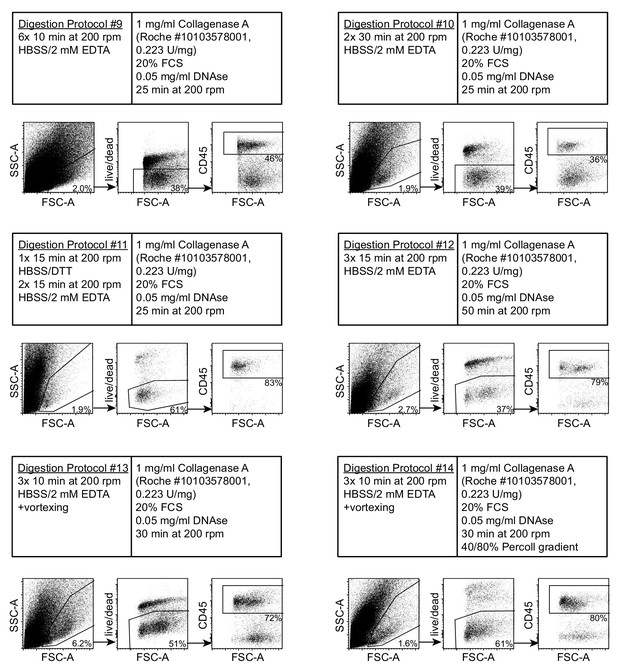
Further optimization of a single cell isolation protocol from heavily infected duodenal segments based on Collagenase A digestion.
Dots plots of acquired events from day 14 hr. polygyrus-infected duodenal segments using digestion protocols #9–14 (representative of 3–5 samples per group from at least two independent experiments). Details for each digestion protocol are annotated. The gating strategy shows cells of interest, viability and CD45 staining.
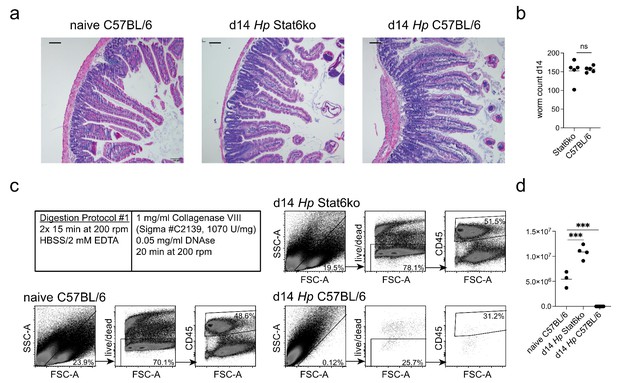
Intestines from H. polygyrus-infected Stat6ko mice can be digested with the standard cell isolation protocol.
(a) H and E stained FFPE (Formalin fixed paraffin embedded) sections from naïve C57BL/6 and day 14 hr. polygyrus-infected C57BL/6 and Stat6ko mice. Scale bar = 100 µm (representative of >10 sections from 3 to 5 mice per group and two independent experiments). (b) Worm counts from day 14 infected C57BL/6 and Stat6ko mice (n = 5 mice per group, representative for two independent experiments; unpaired t-test). (c,d) Dots plots of acquired events and number of live cells isolated from naïve C57BL/6 and day 14 hr. polygyrus-infected C57BL/6 and Stat6ko mice using the standard digestion protocol (representative of 3–5 samples per group from two independent experiments; ordinary one-way ANOVA followed by Holm-Sidak’s multiple comparisons test compared to the naïve group; ***p≤0.001). Details for the standard digestion protocol are annotated. The gating strategy shows cells of interest, viability and CD45 staining.
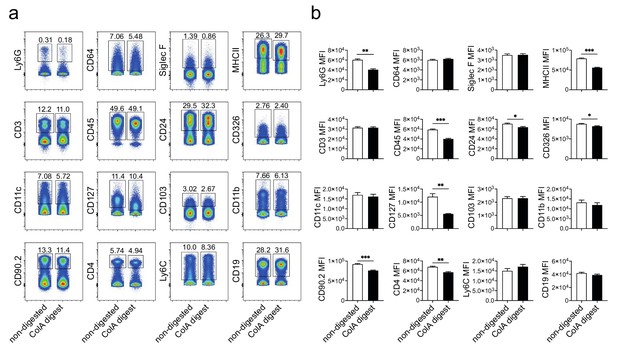
Assessment of epitope integrity of digested and non-digested splenocytes.
(a) Comparison of Collagenase A digested and non-digested splenocytes stained for each surface marker used in our 23-color spectral flow cytometry panel. Gates and percentages of positively stained populations are shown (representative of two independent experiments). (b) Comparison of MFIs for populations shown in a (unpaired t-test; *p≤0.05, **p≤0.01, ***p≤0.001).
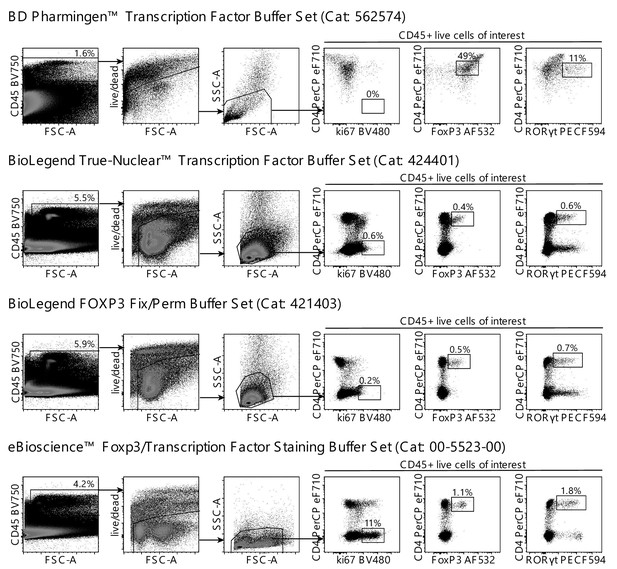
Comparison of different commercial intracellular staining kits on digested lamina propria cells.
Dots plots of Collagenase A digested lamina propria cells from naïve C57BL/6 mice stained with Zombie NIR, CD45, CD4, ki67, FoxP3 and RORγt using four different commercial intracellular staining kits following the respective manufacturers’ instructions (representative of two independent experiments). The eBioscience FoxP3/Transcription Factor Staining Buffer showed the best separation of our cell populations of interest and was used henceforth.
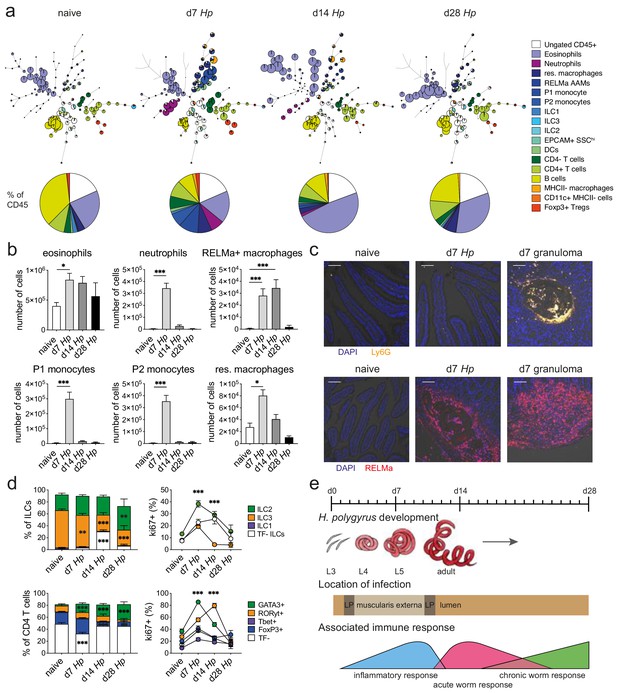
Spectral flow cytometric analysis of isolated intestinal immune cells during the course of H. polygyrus infection.
(a) FlowSOM (top) and manual (bottom) analysis of live CD45+ cells isolated from naïve, day 7, day 14 and day 28 infected duodenal segments stained with 23 surface and intracellular antibodies and gated as described in Figure 2—figure supplement 1 (n = 3–8 samples per group, combined data from two independent experiments). (b) Quantification of different innate immune cell populations during the course of H. polygyrus infection (mean ± s.e.m.; Kruskal-Wallis followed by Dunn’s multiple comparisons test compared to the naïve group; *p≤0.05, ***p≤0.001). (c) Representative images from intestinal cryosections stained with Ly6G-PECF594 (orange) and DAPI (blue) from naïve and day 7 infected duodenal segments (top) or stained with RELMα-APC (red) and DAPI (blue) from naïve, day 7 and day 14 infected duodenal segments (bottom). Scale bar = 100 µm (representative of >10 sections from 3 to 5 mice per group and two independent experiments). (d) Proportions of ILC and CD4 T cell populations and their expression of the proliferation marker ki67 during the course of infection (mean ± s.e.m.; 2-way ANOVA followed by Dunnett’s multiple comparisons test compared to each of the naïve groups (stacked bar graphs) or compared to the combined naïve group (line graphs); **p≤0.01, ***p≤0.001). (e) Schematic of H. polygyrus development, location and associated immune responses during the course of infection.
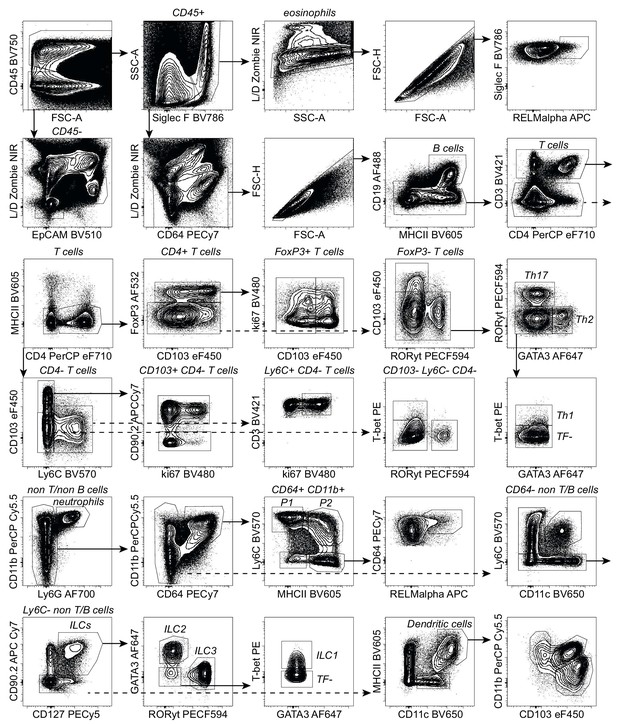
Gating strategy for duodenal lamina propria cells.
Contour plots of Collagenase A digested lamina propria cells from day 7 hr. polygyrus-infected C57BL/6 mice stained with our 23-color spectral flow cytometry panel. The gating strategy was used to identify innate and adaptive immune cells that have been associated with helminth infection and used for manual analysis of all stages of infection.
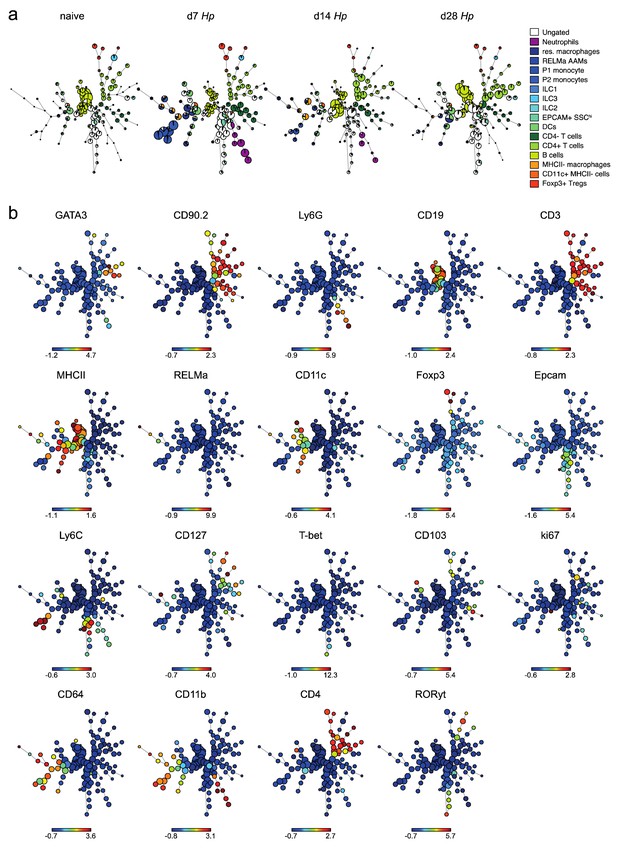
FlowSOM analysis of duodenal lamina propria cells.
(a) FlowSOM analysis of live CD45+ cells isolated from naïve, day 7, day 14 and day 28 hr. polygyrus-infected duodenal segments stained with 23 surface and intracellular antibodies and gated as described in Figure 2—figure supplement 6 (n = 3–8 samples per group, combined data from two independent experiments). (b) Range of expression of each antibody for the combined FlowSOM analysis shown in a). Due to the uptake of Zombie NIR by eosinophils, only live CD45+ Siglec F- cells are shown.
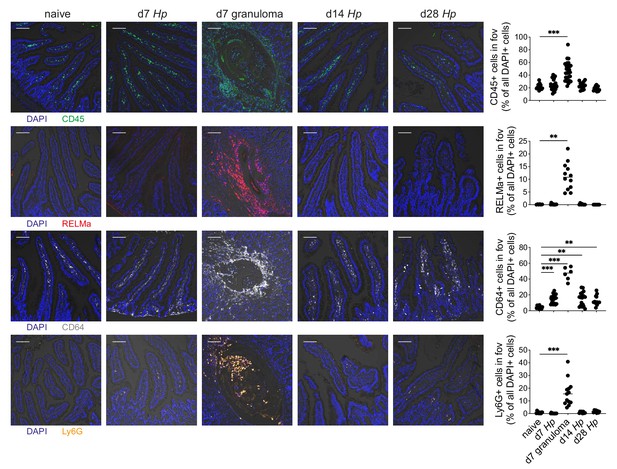
Intestinal cryosections highlight infiltration of innate immune cells within larval granulomas.
Representative images from intestinal cryosections stained with CD45-FITC (green), RELMα-APC (red), CD64-PE (white), Ly6G-PECF594 (orange) and DAPI (blue) from naïve, day 7, day 14 and day 28 hr. polygyrus-infected duodenal segments and their quantification per field of view (fov, 635.90µmx635.90µm). Scale bar = 100 µm (representative of >10 sections from 3 to 5 mice per group from two independent experiments; Kruskal-Wallis followed by Dunn’s multiple comparisons test compared to the naïve group; **p≤0.01, ***p≤0.001).
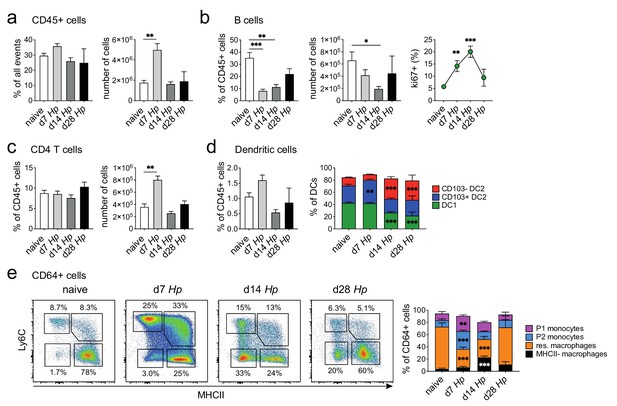
Analysis of isolated intestinal immune cells during the course of H. polygyrus infection.
Analysis of live CD45+ cells (a), B cells (b), CD4 T cells (c), Dendritic cells (d) and CD64+ cells (e) isolated from naïve, day 7, day 14 and day 28 infected H. polygyrus duodenal segments as gated in Figure 2—figure supplement 6 (n = 3–8 samples per group, combined data from two independent experiments; mean ± s.e.m.; Kruskal-Wallis followed by Dunn’s multiple comparisons test compared to the naïve group (bar graphs), 2-way ANOVA followed by Dunnett’s multiple comparisons test compared to each of the naïve groups (stacked bar graphs) or compared to the combined naïve group (line graphs); *p≤0.05, **p≤0.01, ***p≤0.001).
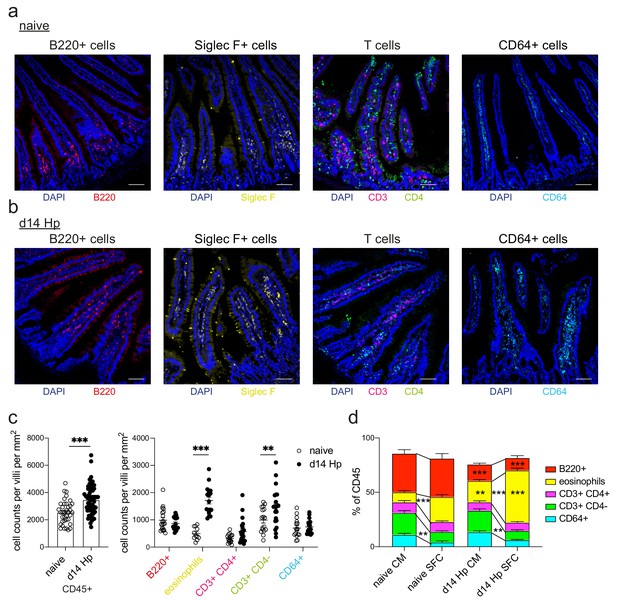
Quantification of intestinal immune cells detected by confocal microscopy or spectral flow cytometry.
Representative images from intestinal cryosections stained with CD45-FITC, B220-PECF594 (red), Siglec F- PECF594 (yellow), CD3-PE (magenta), CD4-APC (green), CD64-PE (cyan) and DAPI (blue) from naïve (a) or day 14 hr. polygyrus infected (b) duodenal segments. Scale bar = 100 µm (representative of >10 sections from 3 to 5 mice per group from two independent experiments). (c) Quantification of CD45+, B220+, eosinophils (counted as non-epithelial Siglec F+ cells), CD3+ CD4+, CD3+ CD4- and CD64+ cells per villi section per mm2 (>15 villi from >10 sections from 3 to 5 mice per group from two independent experiments were quantified; unpaired t-test or 2-way ANOVA followed by Sidak’s multiple comparisons test compared to each of the naïve groups; **p≤0.01, ***p≤0.001). d, Comparison of immune cell frequencies detected by confocal microscopy (CM, as shown in c) or spectral flow cytometry (SFC, as shown in Figure 1a) from naïve or day 14 infected duodenal segments (mean ± s.e.m.; 2-way ANOVA followed by Sidak’s multiple comparisons test comparing detected cell frequencies between technologies or between naïve and infected segments; **p≤0.01, ***p≤0.001).

Gating strategy for mesenteric lymph node cells.
Contour plots of digested duodenal draining mesenteric lymph node cells from day 7 hr. polygyrus-infected C57BL/6 mice stained with our 23-color spectral flow cytometry panel. The gating strategy was used to identify innate and adaptive immune cells that have been associated with helminth infection and used for manual analysis of all stages of infection.
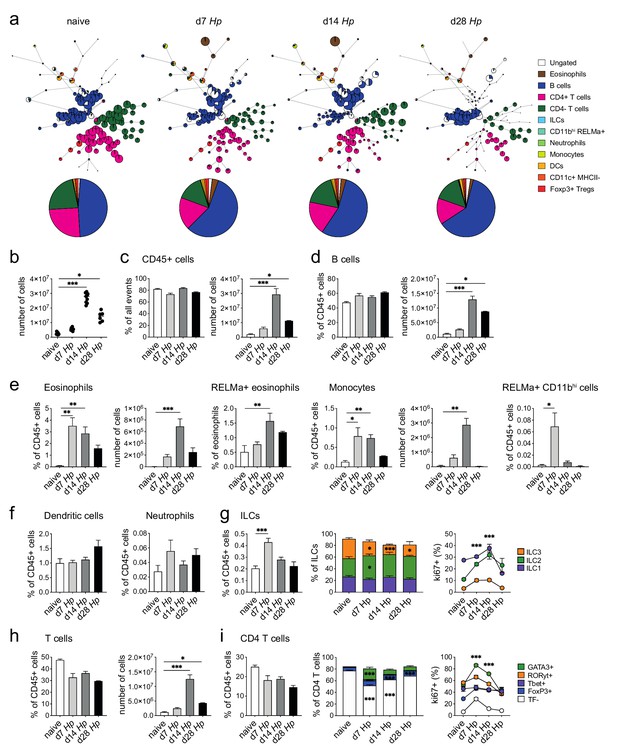
Spectral flow cytometric analysis of mesenteric lymph node cells during the course of H. polygyrus infection.
(a) FlowSOM (top) and manual (bottom) analysis of live CD45+ duodenum draining mesenteric lymph node cells from naïve, day 7, day 14 and day 28 hr. polygyrus-infected C57BL/6 mice stained with 23 surface and intracellular antibodies and gated as described in Figure 2—figure supplement 6 (n = 2–9 samples per group, combined data from two independent experiments). Number and percentage of live cells (b) CD45+ cells (c) B cells (d), innate immune cells (e,f) ILCs (g) T cells (h) and CD4 T cells (i) are shown (combined data from two independent experiments; mean ± s.e.m.; Kruskal-Wallis followed by Dunn’s multiple comparisons test compared to the naïve group (bar graphs), 2-way ANOVA followed by Dunnett’s multiple comparisons test compared to each of the naïve groups (stacked bar graphs) or compared to the combined naïve group (line graphs); *p≤0.05, **p≤0.01, ***p≤0.001).
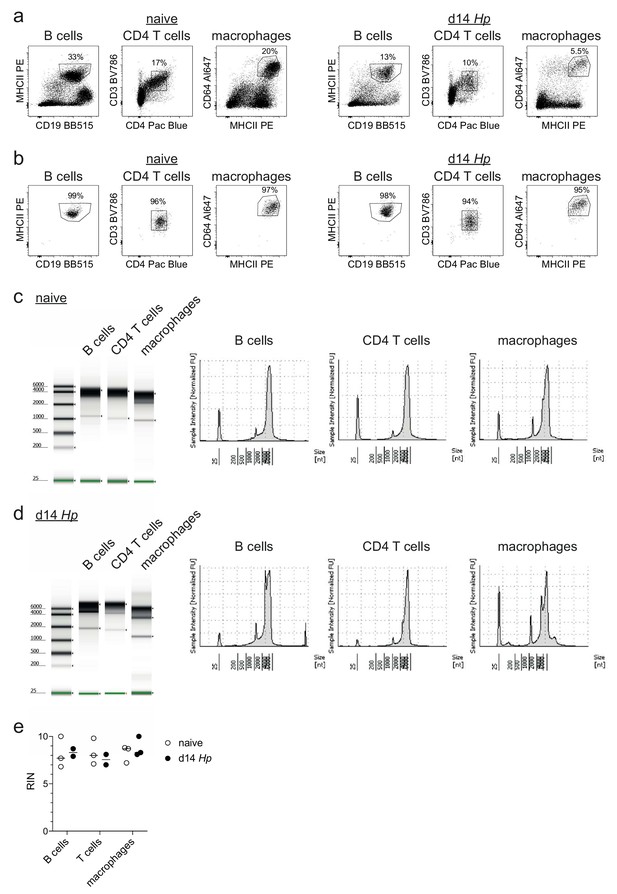
Assessment of RNA quality of sorted intestinal immune cells from naïve and H. polygyrus-infected mice.
(a,b) Frequency of B cells (identified as CD45+ CD19+ CD3- CD64- MHCII+ cells), CD4 T cells (identified as CD45+ CD19- CD3+ CD4+ CD64- MHCII- cells) and macrophages (identified as CD45+ CD19- CD3- CD64+ MHCII+ cells) from naïve or day 14 hr. polygyrus-infected C57BL/6 mice in CD45 enriched single cell suspensions (a) and after cell sorting for each individual population (b) (representative of 3–5 samples from three independent experiments). (c,d,e) RNA quality report from the Agilent TapeStation showing the gel image, electropherogram and reported RIN numbers from RNA extracted from 5,000 cells per population (representative of 2–3 samples from three independent experiments).
Additional files
-
Supplementary file 1
23-color spectral flow cytometry panel for the analysis of intestinal immune cells during helminth infection.
Antibodies used for the high-dimensional analysis of intestinal immune cells are listed by channel, emission, marker, fluorochrome, clone, company, catalog ID, optimal staining dilution and working concentration.
- https://cdn.elifesciences.org/articles/51678/elife-51678-supp1-v1.xlsx
-
Transparent reporting form
- https://cdn.elifesciences.org/articles/51678/elife-51678-transrepform-v1.pdf