Multimeric single-domain antibody complexes protect against bunyavirus infections
Figures
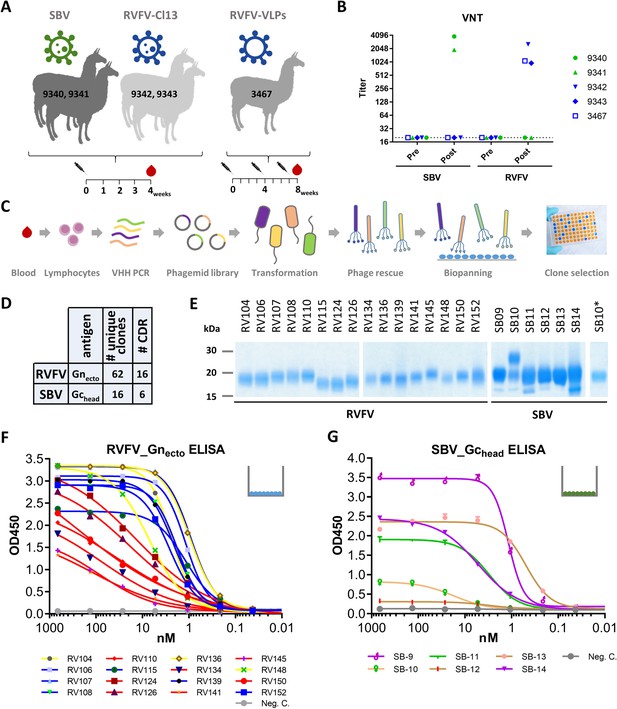
Selection and expression of RVFV and SBV-specific VHHs.
(A) Illustration of the llama immunization strategy to obtain a source for RVFV and SBV-specific VHHs. (B) RVFV and SBV-specific virus neutralizing responses in llama sera determined by virus neutralization test (VNT) before (pre) and post immunization. Neutralizing antibody titers were measured by endpoint titration and calculated as 50% neutralization. (C) Schematic presentation of the VHH library construction and phage display selections. Blood lymphocyte RNA was used as a template for VHH-specific PCRs and purified fragments were cloned into phagemids generating a phagemid library. Following phagemid transduction, the purified phage library was screened for antigen-specific phages. (D) Summarizing table of the VHH clone selection showing the number (#) of unique clones identified and the number of CDR groups per antigen. (E) SDS-PAGE of yeast-expressed RVFV and SBV-specific VHHs. SB10 was shown to be partly N-glycosylated as confirmed by PNGase F based deglycosylation (lane SB10*). (F) RVFV-Gnecto and (G) SBV-Gchead-specific indirect ELISAs. The color coding of the individual VHHs is based on the outcome of the competition ELISA result as presented in Figure 2A and B.

VHH amino acid sequence alignment.
RVFV and SBV-specific VHHs belonging to different CDR groups were aligned, amino acids were numbered and CDR regions were defined according to the IMGT system (Lefranc, 2003). For comparison the older Kabat numbering system is also indicated. The framework regions are indicated in light green and the CDR regions in pink. The CDR3 sequence of RV126 is highly similar to the CDR3 sequence of RV124, however RV126 contains an unusual insertion in FR2 at IMGT position 48. SB-9 is a conventional-like VHH. RV141 and RV145 are predicted to be N-glycosylated at IMGT position 57 whereas SB10 is predicted to be N-glycosylated at IMGT position 84.
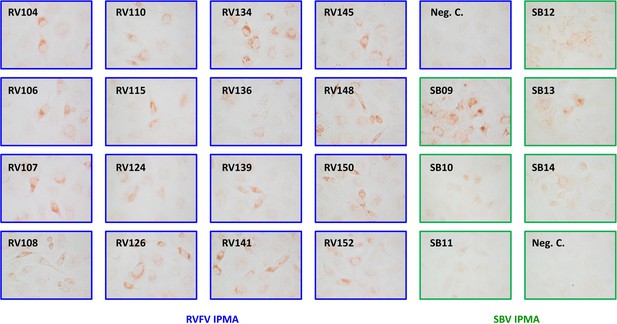
VHH specificity determined by immunoperoxidase monolayer assay (IPMA).
Vero E6 cells were infected at MOI 0.1 with either RVFV Clone 13 (blue boxes) or SBV NL-F6 (green boxes) and fixed 1 day post infection following permeabilization and VHH incubation at 10 µg/ml. VHH binding was visualized with a HRP conjugated goat anti-llama IgG conjugate and a AEC (3-amino-9-ethylcarbazole) substrate.
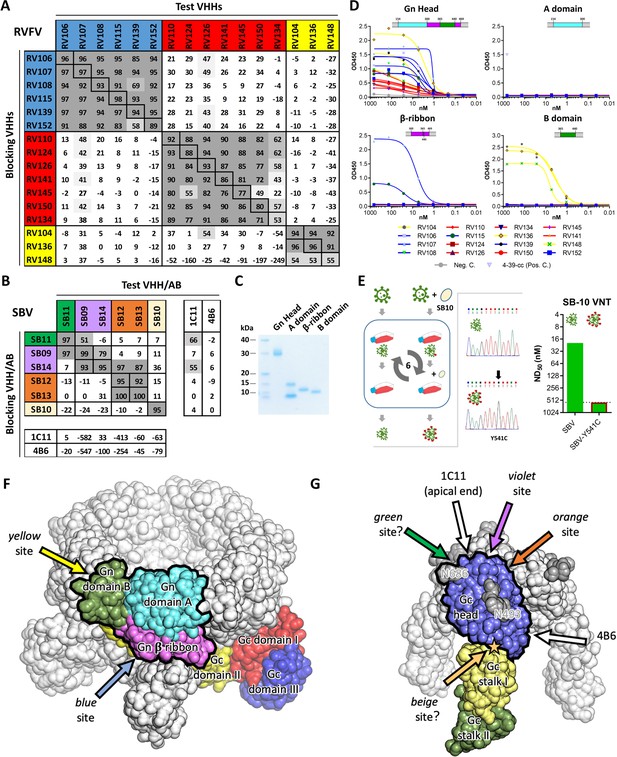
Characterization of VHH binding sites.
(A) RVFV and (B) SBV-specific VHHs and antibodies were tested in competition ELISA using RVFV-Gnecto and SBV-Gchead as antigens. Competition is expressed as percentage of blocking. A percentage above 50% is considered efficient blocking and strongly suggests that the test VHH binds at an overlapping site in the antigen as the blocking VHH. For SBV, competition was additionally assessed for the 1C11 and 4B6 mouse mAbs (Hellert et al., 2019). The RVFV-specific VHHs clustered into three groups each targeting a different antigenic site in RVFV-Gnecto and the SBV VHH panel was shown to target 4 different antigenic sites in SBV-Gchead (of which the green, violet and orange sites are shown to be partly overlapping). Since SB11 and SB14 showed efficient competition (although non-reciprocal) with 1C11, the green, violet and orange antigenic sites are expected to be located at the apical end of the SBV-Gchead domain. (C) SDS-PAGE of the baculovirus produced head, A, β-ribbon and B subdomains of RVFV-Gnecto. The lowest band in the subdomain A lane is expected to be the result of proteolytic cleavage. (D) RVFV-Gnecto subdomain-based VHH ELISAs. Wells were coated with the indicated subdomains and VHH binding was assessed with a HRP-conjugated goat anti-llama IgG. (E) SB10 escape mutant selection. Briefly, wildtype SBV was sequentially passaged in the absence or presence of SB10. Following passage 6, the Gchead coding region was sequenced and SB10 neutralization was quantitatively assessed. (F) Model of a pentameric RVFV spike. The RVFV Gn/Gc heterodimer within a penton (PDB: 6F9F, Halldorsson et al., 2018) is color-coded by domains. The Gn head domain is outlined in black. Arrows indicate approximate positions of the two mapped antigenic sites. (G) Model of a trimeric SBV spike based on the monomeric SBV crystal structure (PDB: 6H3S, Hellert et al., 2019) and the trimeric BUNV cryo-ET map (EMDB: EMD-2352, Bowden et al., 2013). A protomer is color-coded by domains and the Gc head domain is outlined in black. The two N-glycans are labelled with the positions of their respective asparagine residues and the position of the SB10 escape mutation is indicated with a star. Arrows indicate approximate positions of the antigenic sites. Antigenic sites for which evidence is inconclusive are indicated with question marks (green and beige sites).
RVFV and SBV-specific neutralization by monovalent VHHs and bacterial superglue-assembled VHH complexes.
(A) RVFV- and (B) SBV-specific monovalent VHHs, VHH combinations and bivalent and bispecific ELP-based VHH complexes were tested in a RVFV or SBV-specific VNT assay. Complexes were formed by mixing the tagged versions of the VHHs with the indicated Catcher-containing ELP scaffolds of different lengths (Figure 4). Mixtures were subsequently pre-incubated with virus before adding them to susceptible cells. The ND50 values refer to total VHH per reaction as neutralization might be the result of cooperative action of VHH complexes and free VHHs. Averages of two biological replicate experiments are presented. Individual VHHs are color-coded based on the antigenic site they recognize as determined by competition ELISA (Figure 2A and B). Generally, most efficient neutralization was observed with multimerization of VHHs that each target a different antigenic site.
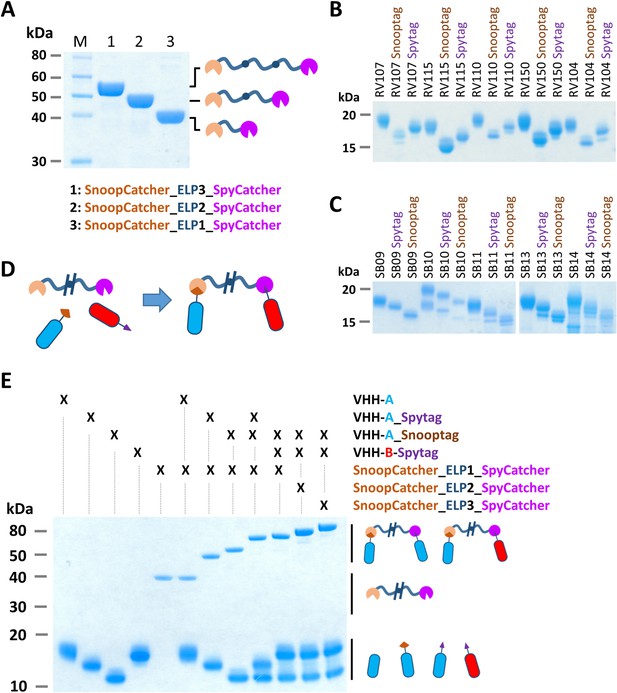
Bacterial superglue-based VHH formatting.
(A) SDS-PAGE of ELP-based scaffolds with an N-terminal SnoopCatcher and a C-terminal SpyCatcher domain. (B) SDS-PAGE of RVFV and (C) SBV-specific VHHs expressed with a C-terminal Snooptag or Spytag. The untagged variants were expressed with the llama antibody long hinge region. (D) Illustration of the bacterial superglue-based site-directed formation of a bispecific or bivalent VHH complex. (E) SDS-PAGE of the bacterial superglue-based formation of bivalent and bispecific VHH complexes using ELPs of various length. VHH RV115 and VHH RV150 were used as an example for VHH-A and VHH-B, respectively.
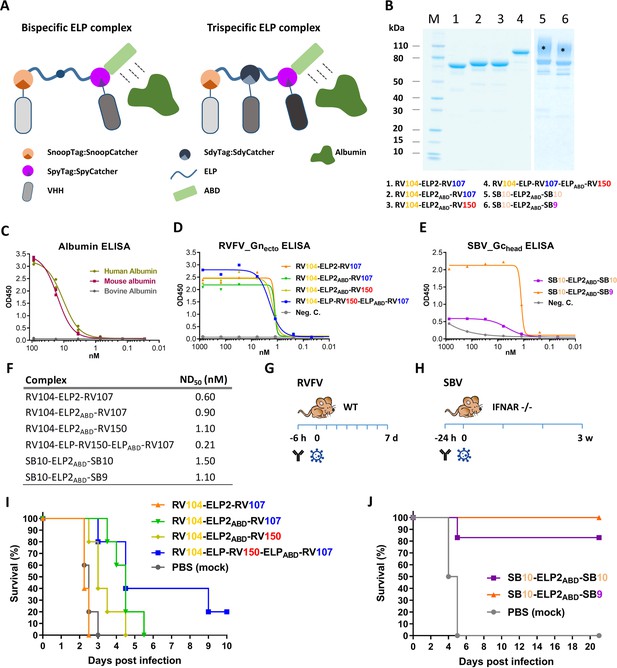
Efficacy of RVFV and SBV-specific ELP-based VHH complexes in mice.
(A) Illustration of ELP-based VHH complexes comprising an albumin binding domain (ABD). (B) SDS-PAGE of purified RVFV and SBV-specific ELP-VHH complexes. Uncoupled VHHs were removed by a Strep-tag based purification following the coupling. The asterisks (*) indicate the glycosylated variants with higher molecular weights. (C) Species-specific albumin ELISA of the ABD containing ELP scaffold. Human, mouse or bovine albumin was coated at a 10 µg/ml concentration and the trispecific ELP scaffold containing an ABD domain (see A) was incubated in a dilution series. Binding was visualized with a Strep-Tactin-HRP conjugate. (D) Indirect RVFV-Gnecto and (E) SBV-Gchead ELISA of purified RVFV and SBV-specific VHH complexes. (F) Neutralizing activity of the purified VHH complexes as determined by RVFV and SBV-specific VNTs. The ND50 values refer to the concentration of complexes. (G) Illustration of the RVFV and (H) SBV mouse models. (I) Survival curve of RVFV infected mice pre-treated with 150 µg of the RVFV-specific VHH complexes. (J) Survival curve of SBV-infected mice pre-treated with either 200 µg of SB10-ELP2ABD-SB10 or 100 µg SB10-ELP2ABD-SB9.
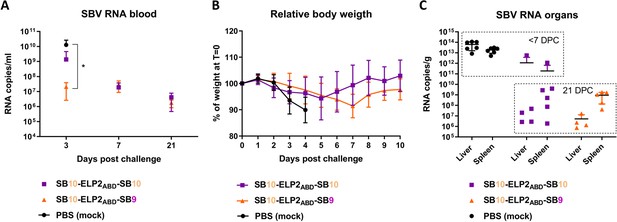
ELP-based SBV-specific VHH complexes reduce viremia.
(A) Systemic SBV viral RNA in blood as determined by RT-qPCR. (B) Relative body weights of mice post challenge. (C) SBV viral RNA in organ suspensions as determined by RT-qPCR. * Statistically significant difference as determined with an unpaired T-test.
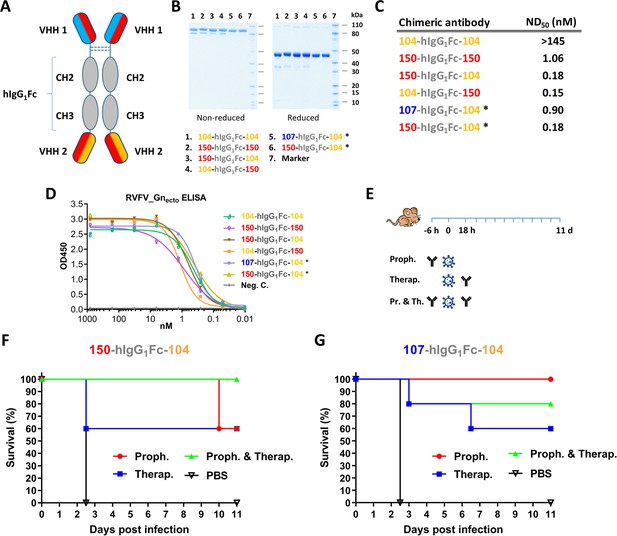
Llama-human chimeric antibodies protect mice from lethal RVFV infection.
(A) Illustration of the VHH and hIgG1Fc-based llama-human chimeric antibodies. (B) SDS-PAGE of the purified chimeric antibodies under non-reducing and reducing conditions. (C) Neutralizing activity of the purified chimeric antibodies expressed as ND50. (D) Indirect RVFV-Gnecto-based ELISA of the chimeric antibodies. (E) Illustration of the mouse experiment setup. (F) Survival curve of RVFV-infected mice treated with 200 µg 150-hIgG1Fc-104. (G) Survival curve of RVFV-infected mice treated with 200 µg 107-hIgG1Fc-104. * These chimeric antibodies were produced at medium scale in CHO cells.
Tables
| Reagent type (species) or resource | Designation | Source or reference | Identifiers | Additional information |
|---|---|---|---|---|
| Strain, strain background (virus, Ovis aries) | RVFV-35/74 | (Kortekaas et al., 2011) | RVFV-35/74 | Recombinant virus |
| Strain, strain background (virus, Homo sapiens) | RVFV-Clone 13 | (Muller et al., 1995) | RVFV-Clone 13 | Natural isolate lacking 69% of the NSs gene |
| Strain, strain background (virus, Bos taurus) | SBV NL-F6 | (Van Der Poel et al., 2014; Hulst et al., 2013) | SBV NL-F6 | Natural isolate |
| Strain, strain background (virus, Ovis aries) | SBV BH619 | (Wernike et al., 2015b) | SBV BH619 | Natural isolate |
| Strain, strain background (E. coli) | TG1 cells | Immunosource | 60502–2 | Electrocompetent cells |
| Strain, strain background (E. coli) | BL21 (DE3) | New England Biolabs | C2527H | Competent cells |
| Strain, strain background (Mus musculus) | BALB/cAnNCrl mice | Charles River Laboratories | BALB/cAnNCrl | |
| Strain, strain background (Mus musculus) | IFNAR-/-mice C57BL/6 | FLI | B6.129S2-Ifnar1tm1Agt/Mmjax | |
| Strain, strain background (yeast) | S. cerevisiae | (Harmsen and De Haard, 2007a) | S. cerevisiae | |
| Cell line Chlorocebus aethiops | Vero E6 | ATCC | CRL-1586 | |
| Cell line Spodoptera frugiperda | Sf9-ET cells | ATCC | CRL-3357 | |
| Cell line Trichoplusia ni | High Five cells | Thermo Fisher Scientific | B855-02 | |
| Recombinant DNA reagent | pRL144 | (Harmsen et al., 2005) | pRL144 | Phage display vector |
| Peptide, recombinant protein | RVFV-Gnecto | (de Boer et al., 2010) | ||
| Peptide, recombinant protein | SBV-Gchead | (Wernike et al., 2017) | ||
| Recombinant DNA reagent | Coding regions RVFV-Gn-ecto | This paper | Genscript | Table 1 |
| Recombinant DNA reagent | Coding region SBV-Gc-head | This paper | Genscript | Table 1 |
| Recombinant DNA reagent | pRL188 | (Harmsen et al., 2007b) | AJ811567 | Yeast expression vector |
| Recombinant DNA reagent | pQE-80L | Qiagen | Expression plasmid | |
| Recombinant DNA reagent | pBAC3 | Merck | 70088 | Baculo transfer plasmid |
| Commercial assay, kit | ELISA Streptactin coated microplates | IBA Lifesciences | 2-1501-001 | |
| Commercial assay, kit | FlashBac ULTRA system | Oxford Expression Technologies | 100300 | |
| Commercial assay, kit | Lightning-Link HRP Conjugation Kit | Innova Biosciences | AB102890 | |
| Other | Gravity Flow Strep-tactin Sepharose column | IBA | 2-1202-001 | |
| Other | Amicon Ultra centrifugal filters | Merck Millipore | UFC900324 | |
| Other | RVFV VLPs | (de Boer et al., 2010) | Virus-like particles | |
| Other | Ni-NTA resin | Qiagen | 30210 | |
| Other | Protein A agarose Fast Flow 50% | Sigma | P3476 | |
| Other | CHO transient expression system | (Daramola et al., 2014) | Expression system | |
| Other | Human albumin | Sigma | A9511 | |
| Other | Mouse albumin | Sigma | A3139 | |
| Other | Bovine albumin | Sigma | A7906 | |
| Other | Bis-Tris NuPAGE Novex Gels | Life Technologies | 4–12% NP0322 12% NP0342 | |
| Other | TMB One Component HRP Microwell Substrate | SurModics | TMBW-1000–01 | |
| Other | HRP-conjugated Strep-Tactin | IBA | 2-1502-001 | 1:5000 |
| Commercial assay, kit | QIAamp Viral RNA kit | Qiagen | 52904 | |
| Commercial assay, kit | RNA Clean and Concentrator −5 kit | Zymo | R1013 | |
| Commercial assay, kit | Phusion Flash High-Fidelity PCR Master Mix | Thermo Fisher Scientific | F548 | |
| Commercial assay, kit | MagAttract Virus mini M48 kit | Qiagen | 955336 | |
| Commercial assay, kit | DNA Clean and Concentrator-5 kit | Zymo | D4014 | |
| Commercial assay, kit | Superscript III First-Strand Synthesis System | Invitrogen | 18080051 | |
| Sequence-based reagent | JR565 | This paper | PCR primers | GACAATTGATGACACATATAGCTT |
| Sequence-based reagent | JR829 | This paper | PCR primers | ACAGAGCCTCTGAGAAATGTCTG |
| Sequence-based reagent | JR830 | This paper | PCR primers | GATTTGCATACCAGTATTGGTG |
| Antibody | Polyclonal HRP-conjugated goat anti-llama IgG-H+L | Bethyl | A160-100P | IPMA (1:1000), ELISA (1:2000) |
Amino acid sequences of used domains and tags.
| Protein | Amino acid sequence |
|---|---|
| SBV Gchead | INCKNIQSTQLTIEHLSKCMAFYQNKTSSPVVINEIISDASVDEQELIKSLNLNCNVIDRFISESSVIETQVYYEYIKSQLCPLQVHDIFTINSASNIQWKALARSFTLGVCNTNPHKHICRCLESMQMCTSTKTDHAREMSIYYDGHPDRFEHDMKIILNIMRYIVPGLGRVLLDQIKQTKDYQALRHIQGKLSPKSQSNLQLKGFLEFVDFILGANVTIEKTPQTLTTLSLI |
| Twin Strep-tag | GSAWSHPQFEKGGGSGGGSGGSAWSHPQFEK |
| Yeast signal peptide | MMLLQAFLFLLAGFAAKISA |
| SpyTag | AHIVMVDAYKPTK |
| SpyCatcher | DIPTTENLYFQGAMVDTLSGLSSEQGQSGDMTIEEDSATHIKFSKRDEDGKELAGATMELRDSSGKTIS TWISDGQVKDFYLYPGKYTFVETAAPDGYEVATAITFTVNEQGQVTVNGKATKGDAHIDGPQGIWGQLEWKK |
| SnoopTag | KLGDIEFIKVNK |
| SnoopCatcher | SSGLVPRGSHMKPLRGAVFSLQKQHPDYPDIYGAIDQNGTYQNVRTGEDGKLTFKNLSDGKYR LFENSEPAGYKPVQNKPIVAFQIVNGEVRDVTSIVPQDIPATYEFTNGKHYITNEPIPPKGPQGIWGQLDGHGVG |
| ELP (1-2-3) | NL(GVPGVGVPGVGVPGEGVPGVGVPGVGVPGVGVPGVGVPGEGVPGVGVPGVGVPGVGVPGVGVPGEGVPGVGVPGVGVPG)1-3GLL |
| HisTag | MKGSSHHHHHH |
| SdyCatcher | SSGLVPRGSHMASMTGGQQMGRGSSGLSGETGQSGNTTIEEDSTTHVKFSKRDANGKELAGAMIELRNLSGQTIQSWISDGTVKVFYLMPGTYQFVETAAPEGYELAAPITFTIDEKGQIWVDS |
| SdyTag | DPIVMIDNDKPIT |
| RVFV-Gn-A | EDPHLRNRPGKGHNYIDGMTQEDATCKPVTYAGACSSFDVLLEKGKFPLFQSYAHHRTLLEAVHDTIIAKADPPSCDLQSAHGNPCMKEKLVMKTHCPNDYQSAHYLNNDGKMASVKCPPKYELTEDCNFCRQMTGASLKKGSYPLQ |
| RVFV-Gn-ß | QDLFCQSSEDDGSKLKTKMKGVCEVGVQALKKCDGQLSTAHEVVPFAVFKNSKKVYLDKLDLKTEENGSGSGVVQIQVSGVWKKPLCVGYERVVVKRELSA |
| RVFV-Gn-B | NLLPDSFVCFEHKGQYKGTMDSGQTKRELKSFDISQCPKIGGHGSKKCTGDAAFCSAYECTAQYANAYCSHANGSG |





