Parvalbumin interneurons provide spillover to newborn and mature dentate granule cells
Figures
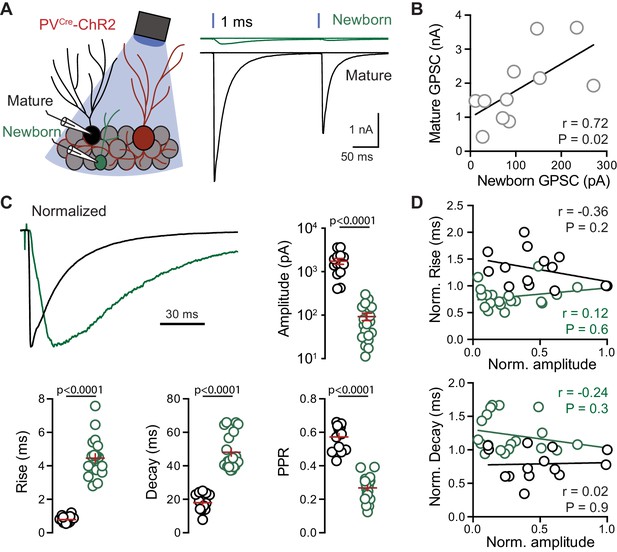
PVs generate distinct GPSCs in mature and newborn GCs.
(A) Left, recording configuration. Right, light-evoked (200 ms ISI) GPSCs in newborn (green) and mature (black) GCs in response to 1 ms light pulseswere blocked by gabazine (10 µM). (B) Correlation of the average peak GPSC amplitude from dual recordings, n = 11. Spearman rank correlation. (C) GPSCs in mature (black) and newborn (green) GCs normalized to the peak amplitude highlights the difference in kinetics. Summary of amplitude, 20–80% rise time, weight decay τ, and PPR includes data from simultaneous (n = 11) and single cell recordings (n = 3 mature and 9 newborn GCs). Unpaired t-tests. Ampl = 1744 ± 267 pA mature, 93 ± 18 pA newborn; rise = 0.79 ± 0.05 ms mature, 4.5 ± 0.3 ms newborn; decay = 18 ± 1.4 ms mature, 48.1 ± 2.4 ms newborn; PPR = 0.57 ± 0.02 mature, 0.27 ± 0.016 newborn. (D) No correlation between GPSC amplitude and rise or decay time across cell pairs. n = 11.
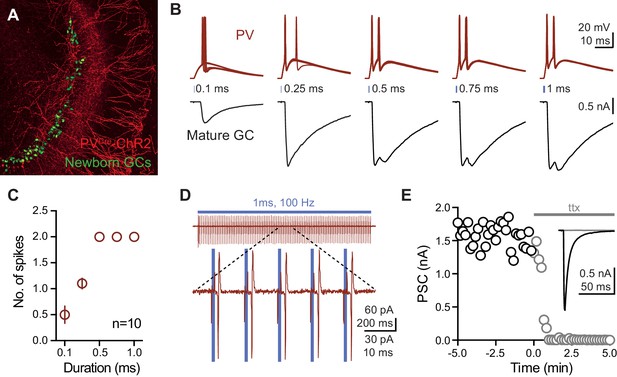
Reliable recruitment of PVs by brief light pulses.
(A) Example confocal image (z-projection) showing ChR2 expression (red) and Pome-EGFP expression (green) in a pseudo-colored image of the dentate gyrus of PvalbCre:ChR2 (H134R)-EYFP (Ai14):Pomc-EGFP mouse. Mature GCs are unlabeled. (B) Simultaneous recordings from ChR2-expressing PVs (red) and mature GCs (black) in response to low frequency light pulses (0.1 Hz; 470 nm) between 0.1 and 1 ms. 10 traces overlaid (top) or averaged (bottom). (C) Summary data showing that PVs typically spike twice in response to light pulses longer than 0.5 ms. n = 10 cells. (D) Top, cell-attached recordings from PVs in response to high-frequency light pulses (100 Hz; 1 ms duration) shows reliable spikes throughout the train. Bottom, expansion of current trace showing 5 light pulses. (E) Low-frequency ChR2-evoked PSCs in mature GCs (black) are blocked with application of TTX (gray). n = 7.
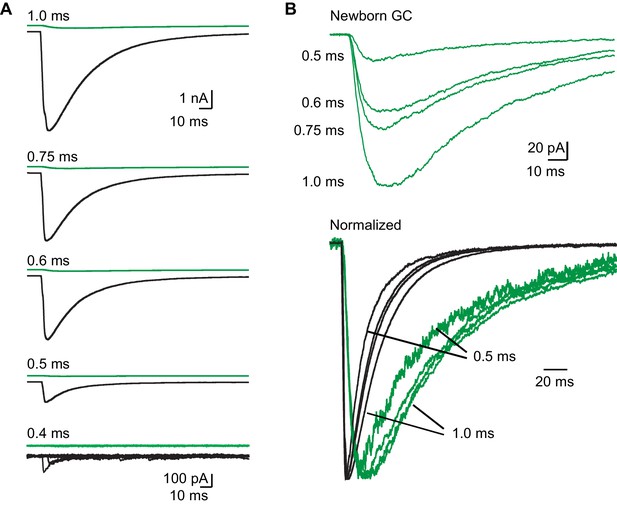
Kinetic differences persist with sub-millisecond light pulses.
(A) Average GPSCs from simultaneous recording of a mature (black) and newborn GC (green) to light pulses of the indicated duration. Each trace is the average of 10 GPSCs, with the exception of responses to 0.4 ms duration where 8 individual GPSCs and failures are overlaid at an expanded scale. GPSCs from 1.0 to 0.5 ms are shown on the same scale. (B) Top, averaged GPSCs from the newborn GCs in (A) overlaid. Bottom, GPSCs from (A) normalized to the peak amplitude to illustrate that kinetic differences between new and mature GCs persist at sub-millisecond light intensities, even as the decay phase accelerates in both cells.
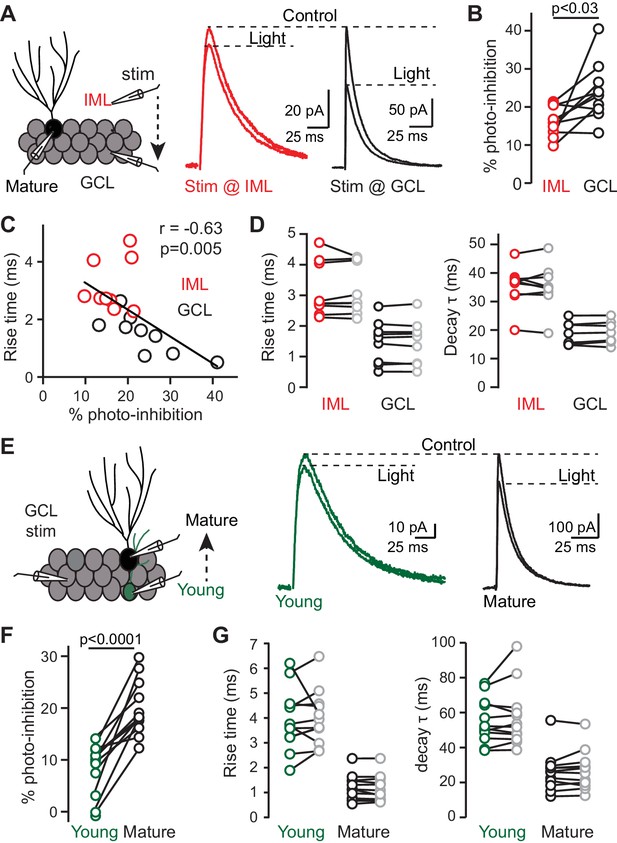
Differential expression of α1 subunits does not account for slow PSCs.
(A) Left, recording from mature GCs with sequential stimulation in the IML (red) and GCL (black). Right, GPSCs showed a smaller α1-GABAAR contribution when stimulating the IML (red) compared to GCL (black). Stimulus artifacts blanked. (B) Photoinhibition reduced IML GPSCs by 16 ± 1% whereas GCL GPSCs were reduced by 24 ± 3%. Paired t-test, n = 9. (C) Slow rise times correlated with less photoinhibition, suggesting a lower contribution of synaptic α1-GABAARs (n = 18). (D) Photoinhibition did not change the rise or decay times. Rise time with IML stimulation: ctrl, 4.7 ± 0.4 ms; blocking α1-GABAAR, 4.8 ± 0.4 ms; with GCL stimulation: ctrl, 2.0 ± 0.3 ms; blocking α1-GABAAR, 2.0 ± 0.3 ms; Decay time with IML stimulation: ctrl, 37.1 ± 2.4 ms; blocking α1-GABAAR, 37.3 ± 2.5 ms; with GCL stimulation: ctrl, 21.9 ± 2.4 ms; blocking α1-GABAAR, 23.4 ± 2.3 ms; n = 9. (E) Left, stimulation in the GCL while sequentially recording from young and mature GCs in G42:α1-LiGABAR mice. Right, slow GPSCs in young GCs (green) showed less photoinhibition compared to mature GCs (black). (F) Photoinhibition reduced GPSCs in young GCs by 8 ± 1% whereas mature GPSCs were reduced by 20 ± 2%; n = 12 each, paired t-test. (G) Photoinhibition did not change the rise or decay time of GPSCs. Rise time: young, ctrl, 4.0 ± 0.4 ms; blocking α1-GABAAR, 4.0 ± 0.3 ms; mature, ctrl, 1.2 ± 0.2 ms; blocking α1-GABAAR, 1.2 ± 0.2 ms; Decay time: young control, 55.2 ± 3.7 ms; blocking α1-GABAAR, 57.4 ± 5.0 ms; mature: control, 26.2 ± 3.2 ms; blocking α1-GABAAR, 27.0 ± 3.2 ms.
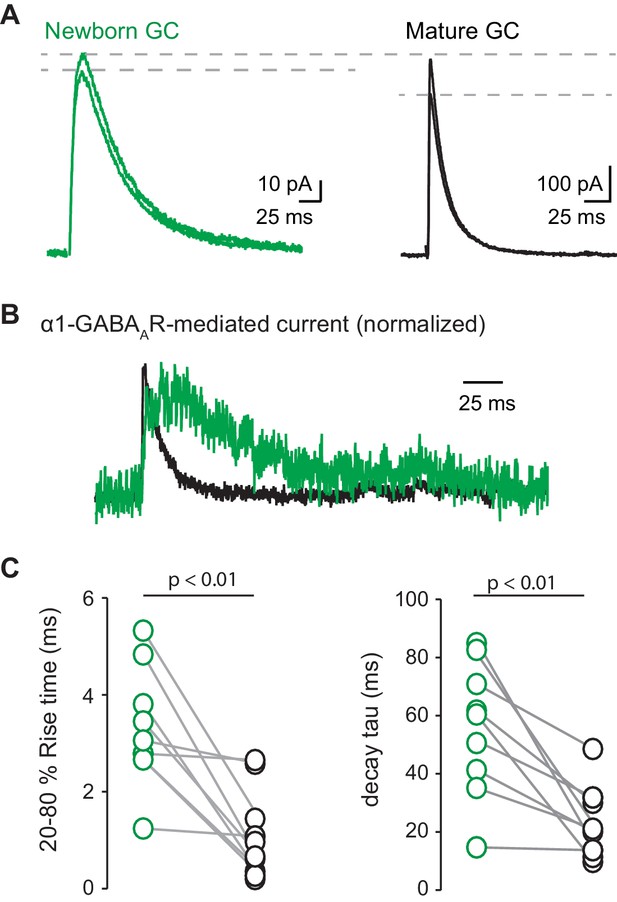
α1-mediated GPSCs in young and mature GCs.
(A) From Figure 2E, slow GPSCs in GAD67-GFP young GCs (green) showed less photoinhibition compared to GPSCs in neighboring mature GCs (black). (B) Subtracted and normalized photosensitive component of GPSCs from (A) showing that the kinetics in young GCs (green) remains slower than that of mature GCs (black). (C) Rise time of α1-mediated GPSCs in young GCs was 3.3 ms ± 0.4 ms vs.1.1 ms ± 0.3 ms in mature GCs. Decay time of the α1-mediated GPSC in young GCs was 55.9 ± 7.6 ms vs. 23.7 ± 3.9 ms in mature GCs. n = 9 pairs of mature and young cells, 3 pairs were excluded due to lack of an α1-mediated component in the young GC. Paired t-tests.
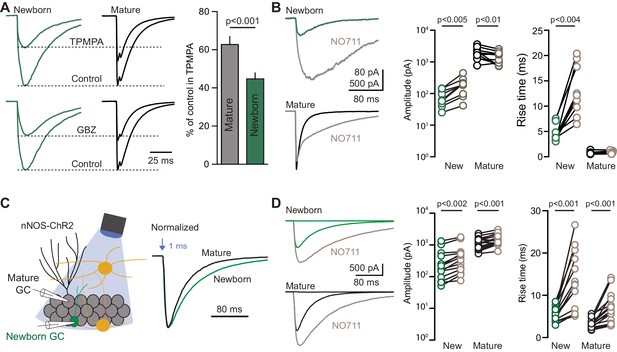
Slow GPSCs in newborn GCs have characteristics of spillover.
(A) Representative PV-evoked GPSCs in simultaneous recordings of newborn (green) and mature GCs (black) in TPMPA (200 µM; top) and GBZ (80 nM; bottom). TPMPA reduced GPSCs in mature GCs to 63 ± 4% of control and in newborn GCs to 45 ± 3% of control, n = 4, p<0.001, paired t-test. (B) In newborn GCs, NO711 (2 µM) increased the GPSC amplitude and prolonged both the rise time and weighted decay τ (from 52 ± 2.6 ms to 136 ± 12.3 ms; n = 9, paired t-tests). In mature GCs, NO711 decreased GPSC amplitude (1980 ± 244 pA to 1476 ± 187 pA) and prolonged the decay τ (18 ± 1 ms to 69 ± 5 ms) but did not alter the rise time of GPSCs (n = 11; paired t-tests). (C) Recording configuration to identify GPSCs evoked by nNOS-expressing interneurons in Pomc-EGFP expressing newborn (green) or mature (black) GCs. Representative GPSCs evoked by 1 ms light pulse normalized to peak amplitude highlight slow rise and decay phases. (D) In contrast to GPSCs evoked by PVs (B). NO711 (gray) increased the amplitude and rise time of nNOS-evoked GPSCs in both newborn (n = 14) and mature GCs (n = 18, paired t-tests). Recordings from newborn and mature GCs were interleaved in separate slices.
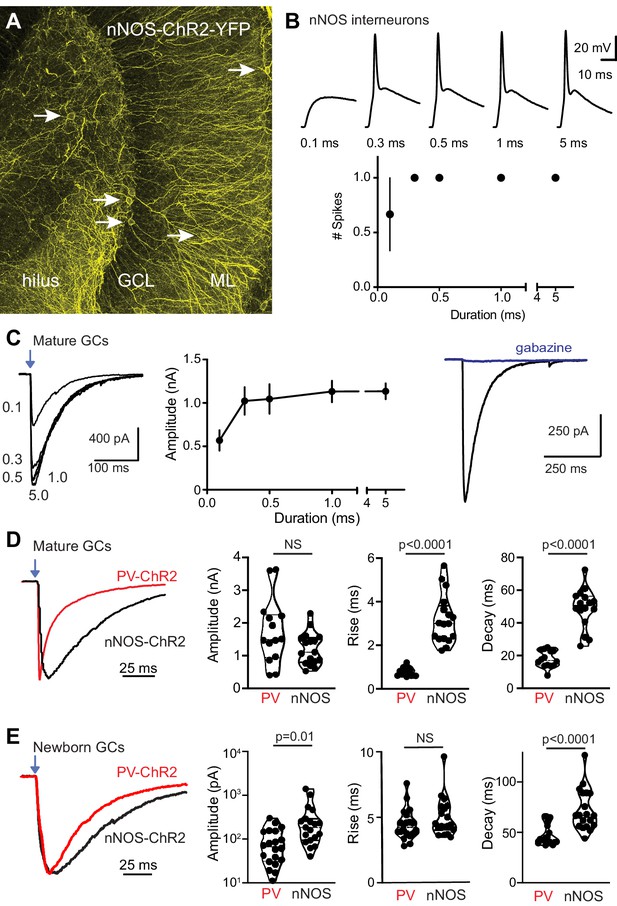
Reliable recruitment of nNOS interneurons by brief light pulses.
(A) Confocal image showing ChR2-expressing processes in the hilus and ML of nNOS-CreER:ChR2 mice. Arrows point to soma. (B) nNOS interneurons show single spikes in response to light pulses (0.1 Hz, 470 nm) between 0.1 and 5 ms duration (n = 3–6 cells). (C) Mature GCs exhibit slow GPSCs in response to light pulses (n = 7) and block by gabazine (10 µM, n = 14). Recordings in CGP55845 (10 µM) to block GABAB receptors (Gonzalez et al., 2018). (D) Comparison of GPSCs in mature GCs evoked by PVs (from Figure 1) and nNOS interneurons. Left, GPSCs normalized to peak highlight kinetic differences. Amplitude = 1744 ± 267 pA versus 1222 ± 120 pA, p=0.09. Rise time = 0.79 ± 0.05 versus 3.2 ± 0.3 ms. Decay time = 18 ± 1.4 ms versus 52.7 ± 4.7 ms. Unpaired t-tests, n = 14 PVs and 18 nNOS. (E) Comparison of GPSCs in newborn GCs evoked by PVs (from Figure 1) and nNOS interneurons. Left, GPSCs normalized to peak amplitude. Amplitude = 93 ± 18 pA versus 300 ± 80 pA. Rise time = 4.5 ± 0.3 ms versus 5.0 ± 0.3 ms. Decay time = 48 ± 2 ms versus 72 ± 5 ms. Unpaired t-tests, n = 20 PVs and 19 nNOS.
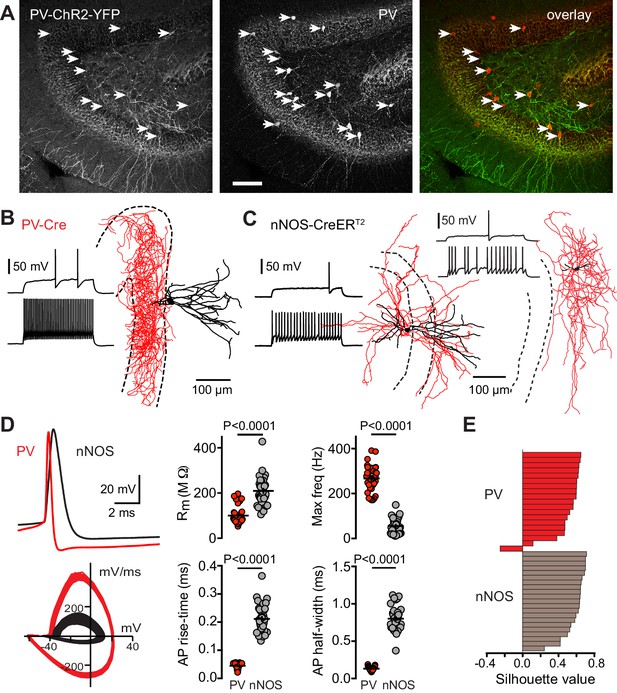
PV-Cre targets fast-spiking basket cells.
(A) Left, confocal z-projection (16 images at 2 µm steps, 20X, 0.5 NA) showing PV-ChR2-YFP expression and immunolabeling of PV (middle). Arrows indicate labeled soma. Right, overlay showing high co-localization between ChR2-YFP and PV, with arrows indicating co-localized soma. Scale bar, 100 µm. (B) Biocytin reconstruction of a PV showing typical basket cell morphology with axon (red) targeting the granule cell layer (dotted lines). Inset shows spiking pattern at threshold and 2x threshold current injections. (C) Partial reconstructions of two nNOS interneurons, with spiking patterns as in (B). (D) Top, PVs showed rapid action potential kinetics (red) compared to nNOS interneurons (black). Bottom, overlaid phase plots of voltage responses at 2x threshold. Right, summary of action potential properties of PVs (n = 38) and nNOS interneurons (n = 39). Max frequency was measured in response to a step current injection of 700 pA. (E) Silhouette plot from a 2-step cluster analysis of 15 parameters that resulted in two clusters comprised of nNOS and PVs (see Figure 4—figure supplement 3). Within each cluster, cells are ranked on the vertical axis in decreasing order of silhouette value (n = 40 cells).
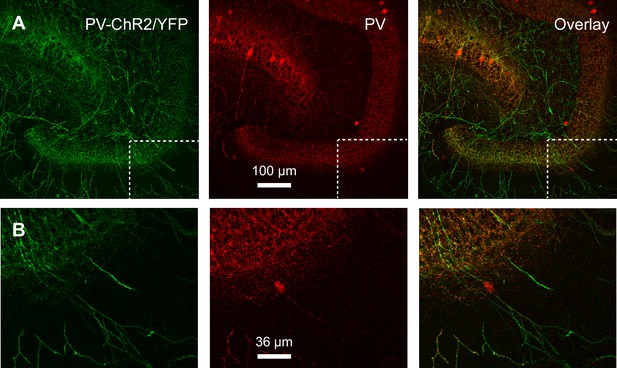
PV-ChR2 co-labeling with PV immunoreactivity.
(A) Confocal z-projection showing expression of PV-ChR2-YFP (left, green), PV immunoreactivity (middle, red) and overlay (right). Scale bar, 100 µm. Note the high density of labeled processes. (B) Region boxed from (A) at higher magnification shows ChR2-YFP (left, green) excluded from the soma of a PV+ interneuron (middle, red), while the overlay (right) suggests this PV+ cell expresses ChR2-YFP in processes. Scale bar, 36 µm.
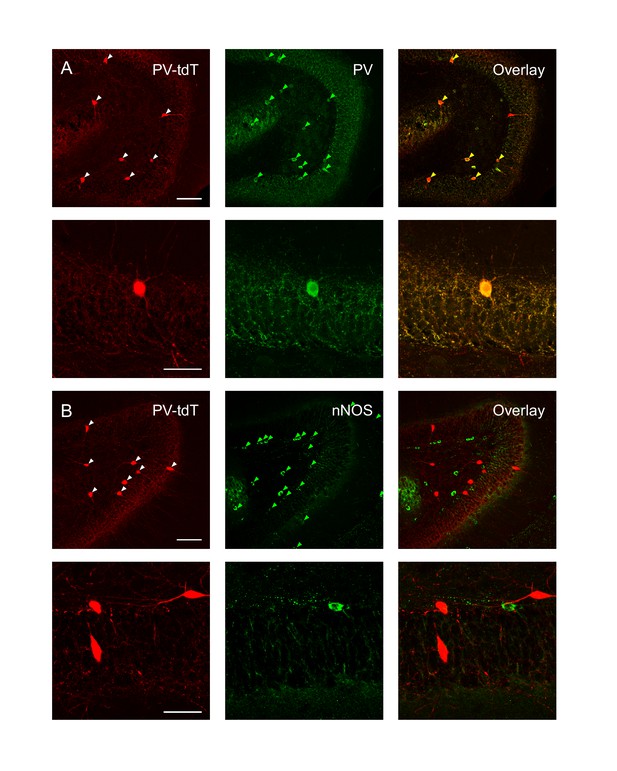
PV-tdT with PV and nNOS immunoreactivity.
(A) Low (upper, 20x) and high (lower, 60x) magnification confocal z-projections showing expression of PV-tdT (left, red) and PV immunoreactivity (middle, green) and overlap (right). Arrowheads indicate cells that were labeled (left, middle) or co-labeled (right). Upper scale bar is 100 µm, lower scale bar is 50 µm. (B) Low (upper) and high (lower) magnification z-projection images showing expression of PV-tdT (left, red) and nNOS immunoreactivity (middle, red) and overlap (right). Same scale as in A.
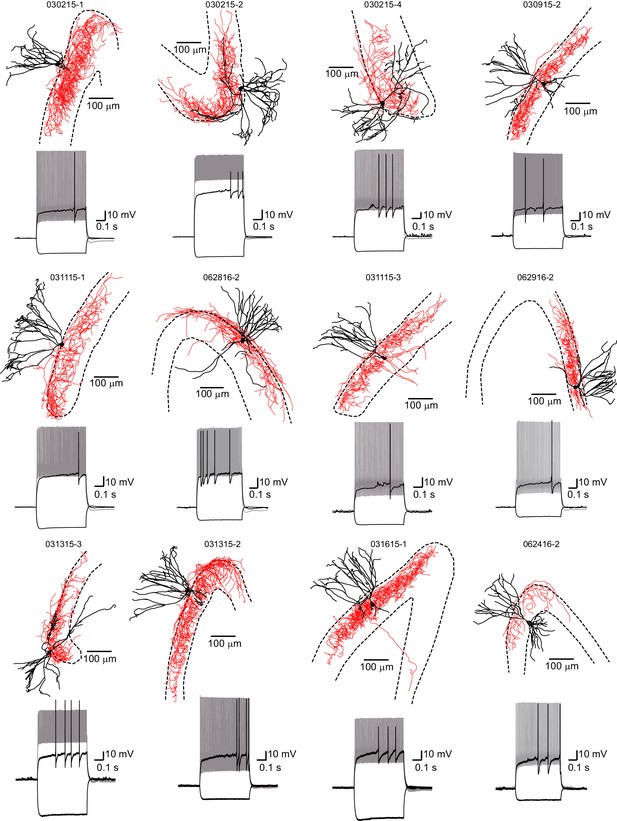
PVs have basket cell morphology.
Reconstructions of recovered PVs from PV-tdT and PV-ChR2 mice showing axonal targeting of the granule cell layer (GCL). The edges of the GCL across all z-levels are indicated by dotted lines. Soma and dendrites in black, axon in red. Insets below show spiking pattern at threshold and 2x threshold current injection.
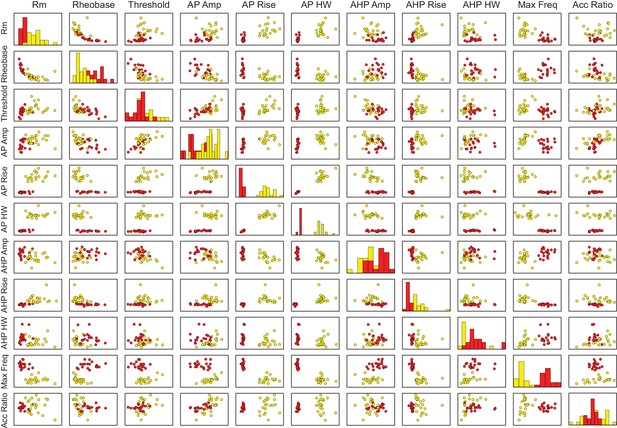
Comparison of cluster analysis parameters.
The two-step cluster analysis of dentate PV and nNOS interneurons was based on molecular and electrophysiological properties. Eleven electrophysiological parameters plotted as pairwise correlations comparing PV and nNos populations, including input resistance (measured from a 10 mV hyperpolarization); rheobase (minimum current injection to elicit an action potential); action potential threshold (voltage at a rate of 20 V/s); action potential amplitude, 20–80% rise time, half-width (measured from threshold); afterhyperpolarization amplitude, 20–80% rise time, half-width (measured from action potential threshold); maximum firing frequency (highest frequency at increasing current injections); accommodation ratio (measured at twice threshold). Four categorical variables (not shown) included molecular marker, cell body location, presence of fast notch in AHP, and firing pattern at twice threshold.
Characteristics of GABA release are consistent with PVs.
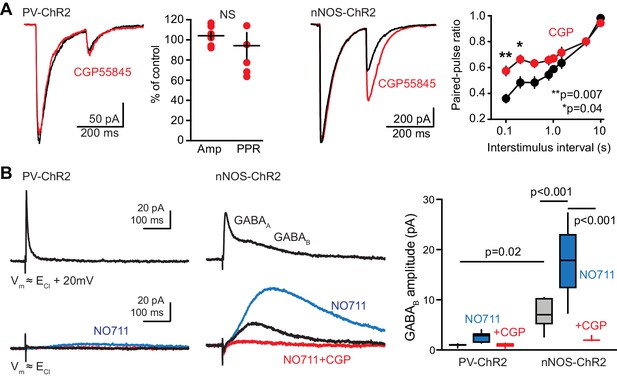
(A) Left, the GABAB receptor antagonist CGP55845 (2 µM, red) had no effect on the amplitude or paired-pulse ratio (PPR) of PV-evoked GPSCs in newborn GCs. Paired t-tests, n = 6, p=0.5 for amplitude and p=0.2 for PPR. In contrast, CGP55845 increased the PPR of GPSCs in slices from nNOS-ChR2 mice at inter-stimulus intervals of 100 ms (**p=0.007) and 200 ms (*p=0.04). Paired t-tests with multiple comparison corrections, n = 7. (B) Mature GCs were held at 20 mV above experimentally-identified ECl to first confirm that brief light pulses (0.1 ms) generated GABA release assayed by GABAA PSCs (top traces). PV-mediated GABAB GPSCs were undetectable in control conditions. In slices from nNOS-ChR2 mice, GABAB IPSCs were apparent at both ECl and ECl+ 20 mV, enhanced by NO711 and blocked by CGP55845 (lower traces). Right, summary data from both mouse lines. ANOVA with Tukey multiple comparisons, n = 4–7 cells.
PV-ChR2 photo-responses in non-PV interneurons.
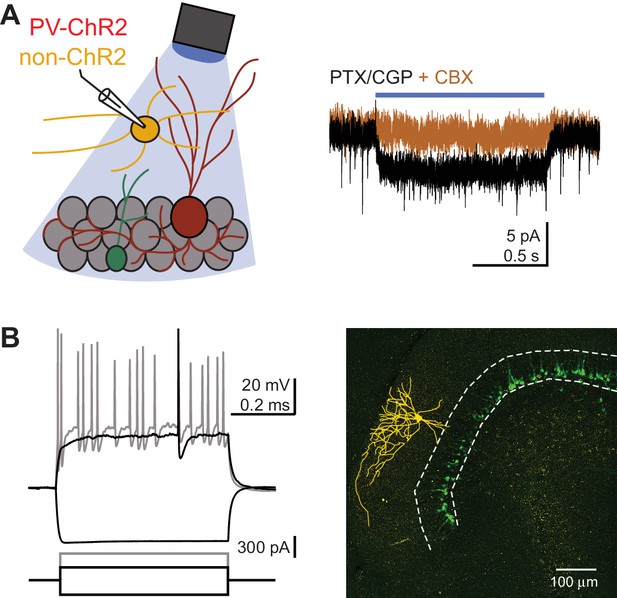
(A) Left, cartoon illustrating recording from a non-ChR2-expressing interneuron in the molecular layer in slices from PV-ChR2:Pomc-EGFP mice. Right, in GABAA and GABAB receptor antagonists, photostimulation (1 s) generates a small steady-state current that is blocked by carbenoxolone (CBX, 100 µM). Small light-evoked CBX-sensitive currents were detected in 4/8 unlabeled molecular layer slow-spiking interneurons. (B) Spike pattern in response to current injections (left) and biocytin reconstruction (right, yellow) from the recording shown in (A). Endogenous ChR2-YFP is bleached, leaving only Pomc-EGFP expression visible.
Spillover GPSCs are generated cooperatively by multiple PVs.
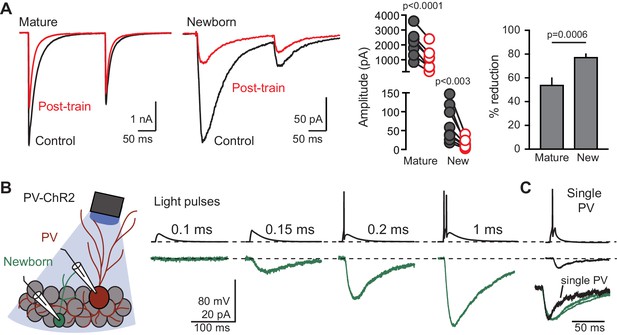
(A) Left, examples GPSCs in mature (black) and newborn GCs (green) in response to a conditioning train that depletes GABA release. Middle and right, the conditioning train reduces GPSCs (paired t-tests) to a greater extent in newborn GCs (78 ± 2%, n = 8) compared to mature GCs (54 ± 5%, unpaired t-test, n = 7). (B) Cartoon and examples of simultaneous recordings from a PV (red) and newborn GC (green) in response to increasing durations of photostimulation at low light intensity. The threshold for this PV was 0.2 ms light duration. Single (top) and average of 3–6 traces (bottom) are shown. (C) In the same pair of cells, direct current injection to the PV (10 ms, 700 pA) generated a small GPSC in the newborn GC, showing that single PVs can generate spillover to newborn GCs (found in 2/5 experiments). Inset, normalized GPSCs in response to single PV stimulation and optogenetic (multiple) PV stimulation shows slower decay phase, as expected for GABA pooling in response to multiple active PVs.
Conditioning stimulation reduces GABA release.
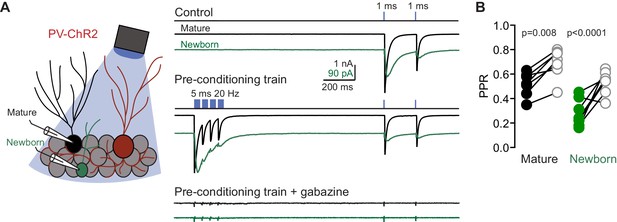
(A) Cartoon depiction of recording configuration from mature (black traces) and newborn GCs (green traces) in response to interleaved pre-conditioning train stimulation (blue). (B) The reduction in GPSC amplitude resulting from train stimulation was associated with an increase in the PPR, suggestive of reduced release probability. n = 7 mature and 8 newborn GCs. ANOVA with Sidek post-hoc tests. GPSCs evoked by the pre-conditioning train were blocked by gabazine (10 µM).
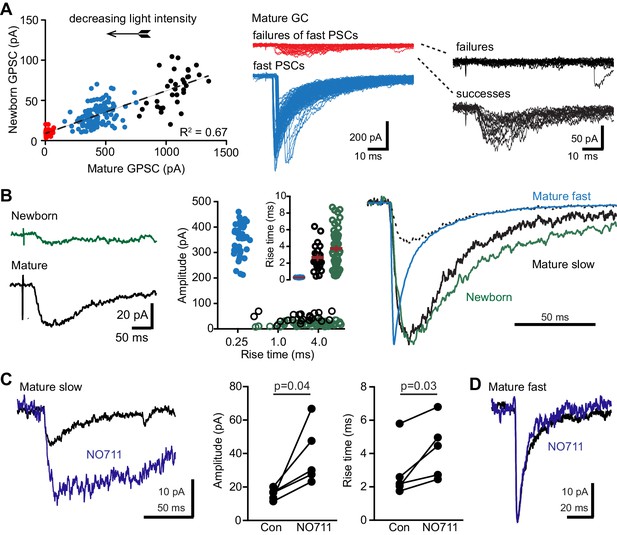
PVs evoke spillover in mature GCs.
(A) Reducing the light intensity (arrow; 0.1 ms duration) decreased the amplitude of GPSCs in mature and newborn GCs. At intermediate intensities, GPSCs in mature GCs had fast rise times (blue symbols and traces; 0.2 ± 0.005 ms) and variable latencies, illustrating variability in threshold for activating axons. Below threshold for fast GPSCs, small slow events were evident (red symbols and traces) that were sorted into failures and successes (right, black traces). (B) Left, average of the subthreshold successes in the mature GC (black) and the average of the correlated traces in the newborn GC (green, successes and failures combined). Middle, comparison of amplitudes and rise times of fast GPSCs in mature (blue), slow GPSCs in mature (black) and slow GPSCs in newborn GCs (green). Right, normalized slow GPSCs in mature and newborn GCs have similar rise and decay phases. Dotted line shows slow GPSC in mature GC scaled to the decay of the fast GPSC. (C) NO711 increased the amplitude and rise time of slow GPSCs in mature GCs, consistent with spillover. Paired t-tests, n = 5. In contrast, in mature GCs that did not exhibit slow GPSCs near threshold light stimulation, NO711 did not affect fast GPSCs (D).
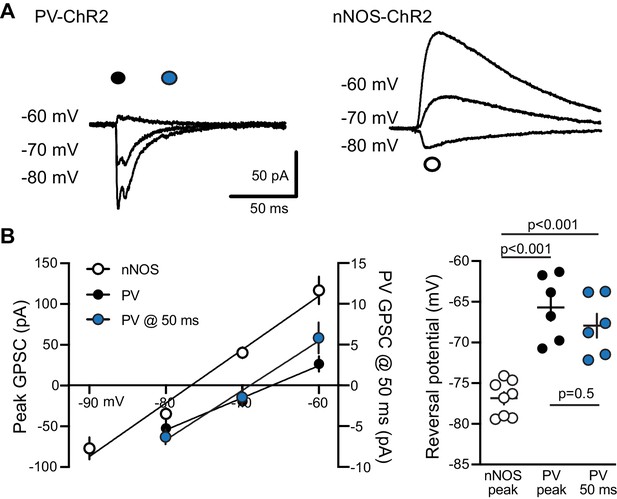
Reversal potentials of fast and slow GPSCs.
(A) Mature GC GPSCs evoked by PV-ChR2 (left) and nNOS-ChR2 (right) had different reversal potentials consistent with somatic and dendritic localization of synapses, respectively. Recordings used KGlu-based intracellular solution (see Materials and methods) and 1 ms light pulses in CGP55845 (10 µM). (B) Left, averaged I/V plots for GPSCs. Right, the reversal potential measured at the peak of PV-evoked GPSCs (solid, −65.7 ± 1.6 mV, n = 6) and at 50 ms after the stimulus (blue, -67.9 ± 1.5 mV) were similar to the calculated choride reversal potential (−77 mV) when the membrane potential was corrected for the junction potential (~12 mV). In contrast, the reversal potential of the peak nNOS-evoked GPSC was more hyperpolarized (−76.9 ± 0.8 mV, n = 8) as expected for GPSCs evoked by dendritic-projecting interneurons (Groisman et al., 2020). ANOVA with Tukey’s multiple comparisons.
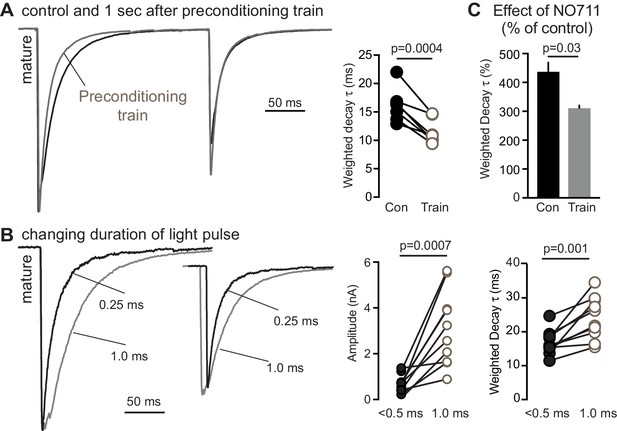
Reducing the number of active synapses limits spillover in mature GCs.
(A) The weighted decay t in mature GCs decreased by 29%, from 16.2 ± 1.1 ms to 11.5 ± 0.8 ms (paired t-test, n = 7) following the condition train, consistent with reduced pooling between active release sites. (B) Similarly, reducing the number of recruited fibers by altering the duration of the light pulse reduced the decay t of GPSCs in mature GCs (from 24.2 ± 2.0 to 17 ± 1.1 ms) as well as the amplitude (from 3105 ± 519 to 643 ± 131 pA). Inset, GPSCs normalized to last peak. Paired t-tests, n = 10. (C) NO711 had a smaller effect on the weighted decay t following the preconditioning train (402 ± 44% compared to 304 ± 10%; n = 4, paired t-test).
Tables
| Reagent type (species) or resource | Designation | Source or reference | Identifiers | Additional information |
|---|---|---|---|---|
| Genetic reagent (M. musculus) | Pomc-EGFP | Jackson Laboratory | Stock #: 009593 RRID:MGI:3776090 | |
| Genetic reagent (M. musculus) | PVCre | Jackson Laboratory | Stock #: 008069 RRID:MGI:3590684 | |
| Genetic reagent (M. musculus) | Ai14 | Jackson Laboratory | Stock #: 007914 RRID:MGI:3809523 | |
| Genetic reagent (M. musculus) | Ai32 | Jackson Laboratory | Stock #: 024109 RRID:MGI:104735 | |
| Genetic reagent (M. musculus) | nNOS-CreER | Jackson Laboratory | Stock #: 014541 RRID:MGI:97360 | |
| Genetic reagent (M. musculus) | α1-LiGABAR | Jackson Laboratory | Stock #: 028965 RRID:MGI:95613 | |
| Genetic reagent (M. musculus) | G42 | Jackson Laboratory | Stock #: 007677 RRID:MGI:3721279 | |
| Antibody | anti-GFP, Alexa Fluor 488 (rabbit polyclonal) | Invitrogen | Cat#: A-21311; RRID:AB_221477 | 1:1000 |
| Antibody | anti-nNOS (rabbit polyclonal) | Chemicon | Cat#: AB5380 | 1:2000 |
| Antibody | anti-PV (rabbit polyclonal) | Abcam | Cat#: ab11427 | 1:1000 |
| Antibody | goat anti-rabbit BIOT | SouthernBiotech | Cat#: 4050–08; RRID:AB_2732896 | 1:800 |
| Antibody | streptavidin, Alexa Fluor 647 | Invitrogen | Cat#: S32357 | 1:200 |
| Chemical compound, drug | carbenoxolone (CBX) | Sigma-Aldrich | Cat#: C4790; CAS: 7421-40-1 | 100 µM |
| Chemical compound, drug | NBQX | Abcam | Cat#: ab120045; CAS: 118876-58-7 | 10 µM |
| Chemical compound, drug | CPP | Abcam | Cat#: ab120159; CAS: 126453-07-4 | 5 μM |
| Chemical compound, drug | SR95531 (gabazine) | Abcam | Cat#: ab120042; CAS: 104104-50-9 | 80 nM, 3–10 µM |
| Chemical compound, drug | NO711 | Sigma-Aldrich | Cat#:N142; CAS: 145645-62-1 | 2–5 μM |
| Chemical compound, drug | TTX | Abcam | Cat#: ab120054; CAS: 4368-28-9 | 1 μM |
| Chemical compound, drug | QX-314 | Abcam | Cat#: ab120118; CAS: 5369-03-9 | 50 μM |
| Chemical compound, drug | TPMPA | Sigma-Aldrich | Cat#: T200; CAS: 182485-36-5 | 200 μM |
| Chemical compound, drug | CGP55845 | Sigma-Aldrich | Cat#: SML0594; CAS: 149184-22-5 | 2–10 μM |
| Chemical compound, drug | PAG-1C | Synthesized in- house; Lin et al., 2018 | 25–50 µM | |
| Chemical compound, drug | tris(2carboxyethyl) phosphine (TCEP) | Sigma-Aldrich | Cat# 646547 | 2.5–5.0 mM |
| Software, algorithm | Axograph X, version | AxoGraph Scientific | axograph.com | |
| Software, algorithm | pClamp 10 | Molecular Devices | moleculardevices.com | |
| Software, algorithm | Prism, version 7/8 | GraphPad | graphpad.com | |
| Software, algorithm | Neurolucida | MBF Bioscience | mbfbioscience.com |





