Neuropeptide VF neurons promote sleep via the serotonergic raphe
Figures
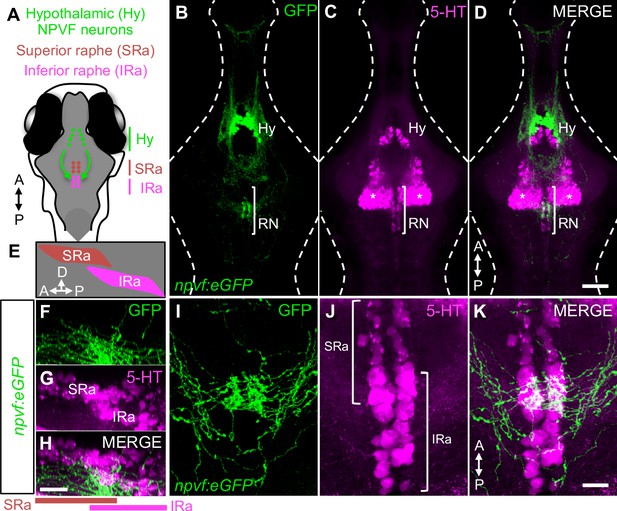
Hypothalamic NPVF neurons project to the serotonergic IRa.
(A,E) Schematic: 6-dpf zebrafish brain showing location of hypothalamic (Hy) NPVF neurons (green), and the serotonergic superior raphe (SRa, red) and inferior raphe (IRa, magenta). A, anterior; P, posterior; D, dorsal. (B–D) Maximum intensity projection of a brain from a 6-dpf Tg(npvf:eGFP) animal (78 μm thick). npvf-expressing neurons in the hypothalamus project to the serotonergic raphe nuclei (RN) in the hindbrain (bracket). 5-HT immunohistochemistry labels the RN (bracket), as well as serotonergic populations in the ventral hypothalamus (asterisks) and pretectum. The bracketed region in (B–D) is shown at higher magnification in (I–K) as a maximum intensity projection (50.5 μm thick), with a sagittal view shown in (F–H). Single optical sections are shown in Figure 1—figure supplement 1. Scale: 50 μm (B–D), 20 μm (F–H), and 10 μm (I–K).
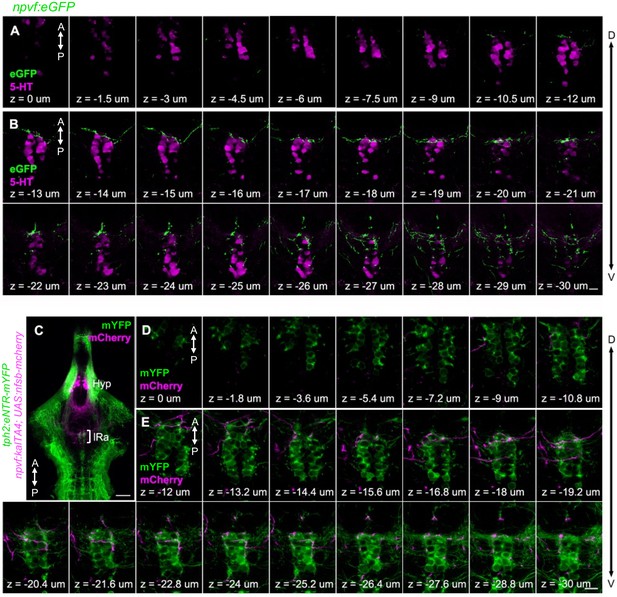
Projections of NPVF neurons to the serotonergic IRa shown in single optical sections.
(A,B) Serial optical sections (0.5 μm thick) in the hindbrain of a 6-dpf Tg(npvf:eGFP) animal labeled with a 5-HT-specific antibody (magenta), which were used to generate the image shown in Figure 1K. Fibers from npvf-expressing neurons (green) do not innervate SRa soma (A) but do innervate IRa soma (B). (C) A 4-μm-thick optical section of a brain from a 6-dpf Tg(npvf:KalTA4); Tg(UAS:nfsb-mCherry); Tg(tph2:eNTR-mYFP) animal. White bracket indicates the IRa and is magnified in panels (D,E), which show 0.6-μm-thick serial optical sections. Fibers from npvf-expressing neurons (magenta) do not innervate SRa neurons (green, D) but do innervate IRa neurons and their fibers (green, E). Hyp, hypothalamus, IRa, inferior raphe; D, dorsal; V, ventral; A, anterior; P, posterior. Every third section (A,D) and every other section (B,E) is shown. Scale: 50 μm (C), 10 μm (B,E).
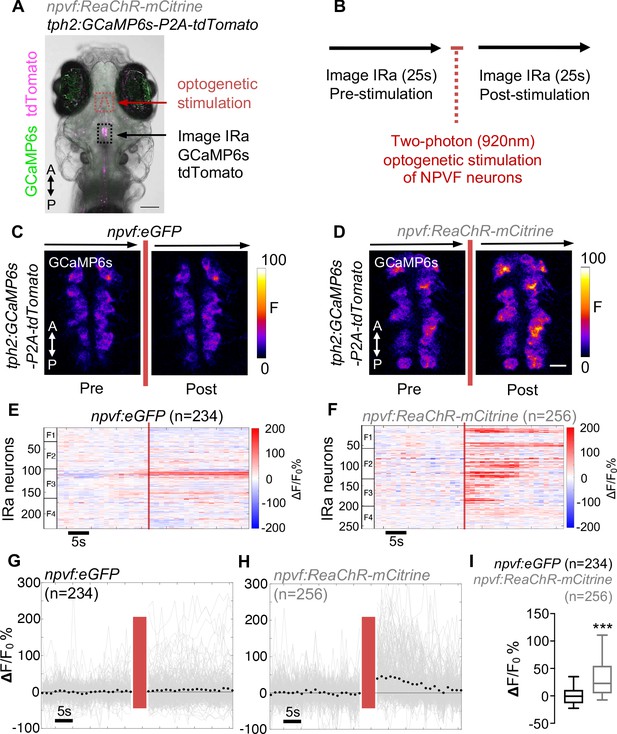
Optogenetic stimulation of NPVF neurons activates serotonergic IRa neurons.
6-dpf Tg(npvf:ReaChR-mCitrine); Tg(tph2:GCaMP6s-P2A-tdTomato) and Tg(npvf:eGFP); Tg(tph2:GCaMP6s-P2A-tdTomato) control animals were analyzed for GCaMP6s/tdTomato fluorescence levels in IRa neurons before and after optogenetic stimulation of NPVF neurons. (A) Example of a 6-dpf Tg(npvf:ReaChR-mCitrine); Tg(tph2:GCaMP6s-P2A-tdTomato) animal showing GCaMP6s and tdTomato fluorescence in IRa neurons with overlayed NPVF neurons (red circles, different z-plane) indicating region of optogenetic stimulation (red box). Black box indicates region where IRa neurons were imaged and analyzed in subsequent panels. Green and magenta signals in the eyes are due to autofluorescence. (B) Normalized GCaMP6s fluorescence in the IRa was first recorded for 25 s (20 frames). NPVF neurons were then optogentically stimulated for 3.7 s (red line), and normalized GCaMP6s fluorescence in the IRa was then immediately imaged again for 25 s. (C,D) Average GCaMP6s fluorescence in IRa neurons for 10 imaging frames before (Pre) and after (Post) optogenetic stimulation of NPVF neurons in Tg(npvf:eGFP); Tg(tph2:GCaMP6s-P2A-tdTomato) (C) and Tg(npvf:ReaChR-mCitrine); Tg(tph2:GCaMP6s-P2A-tdTomato) (D) animals. (E–H) Normalized GCaMP6s fluorescence of individual IRa neurons shown as heat maps (E,F), in which each horizontal line represents an IRa neuron, and as line graphs (G,H) showing individual (gray lines) and mean (dotted line) IRa neuron responses before and after optogenetic stimulation (red lines) in Tg(npvf:eGFP); Tg(tph2:GCaMP6s-P2A-tdTomato) control (E,G) and Tg(npvf:ReaChR-mCitrine); Tg(tph2:GCaMP6s-P2A-tdTomato) (F,H) animals. F1-F4 in (E,F) indicate neurons from four different fish. (I) Box plot of normalized GCaMP6s ΔF/F0 values from the average of the first 10 imaging frames post-stimulation of each neuron for Tg(tph2:GCaMP6s-P2A-tdTomato) animals that also contain either a Tg(npvf:eGFP) (black) or Tg(npvf:ReaChR-mCitrine) (gray) transgene. n = number of neurons quantified from four animals of each genotype. ***p<0.001, Mann-Whitney test. Scale: 100 μm (A), 10 μm (D).
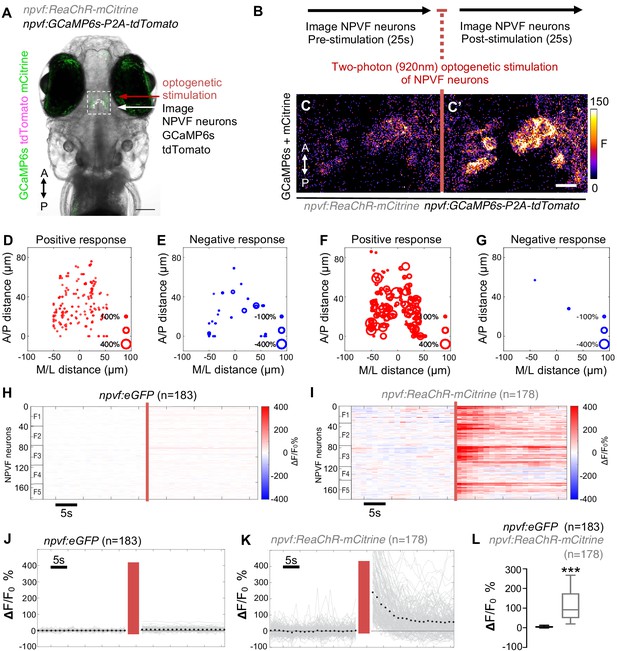
Validation of two-photon-induced optogenetic stimulation of NPVF neurons.
6-dpf Tg(npvf:ReaChR); Tg(npvf:GCaMP6s-P2A-tdTomato) animals and Tg(npvf:eGFP); Tg(npvf:GCaMP6s-P2A-tdTomato) controls were analyzed for GCaMP6s fluorescence normalized to tdTomato fluorescence in NPVF neurons pre- and post-optogenetic stimulation of NPVF neurons. (A) Example of a 6-dpf Tg(npvf:ReaChR); Tg(npvf:GCaMP6s-P2A-tdTomato) animal showing GCaMP6s and tdTomato fluorescence in NPVF neurons. Green and magenta signals in the eyes are due to autofluorescence. White box indicates the region of the hypothalamus that was stimulated and analyzed in subsequent panels. (B) Normalized GCaMP6s fluorescence in NPVF neurons was first recorded for 25 s (20 frames) at low-laser power. NPVF neurons were then optogenetically stimulated at higher laser power, and then normalized GCaMP6s fluorescence in NPVF neurons was immediately imaged again for 25 s (20 frames) at low-laser power. (C,C’) GCaMP6s and mCitrine fluorescence in NPVF neurons recorded one frame before (C) and after (C’) optogenetic stimulation of NPVF neurons. (D–G) Positive or negative normalized GCaMP6s fluorescence responses to optogenetic stimulation of individual NPVF neurons plotted according to their spatial location along the anterior/posterior and medial/lateral axes of the brain. The size of red or blue circles indicates the magnitude of increase or decrease of normalized GCaMP6s fluorescence, respectively. A, anterior; P, posterior; L, lateral; M, medial. ‘0’ along the A/P axis indicates location of the most posterior NPVF neurons. ‘0’ along the M/L axis indicates the midline of NPVF neuron population. (H-K) Normalized GCaMP6s fluorescence of individual NPVF neurons shown as heat maps (H,I), in which each horizontal line represents an NPVF neuron, and as line graphs (J,K) showing individual (gray lines) and mean (dotted line) normalized GCaMP6s ΔF/F0 % values before and after optogenetic stimulation (red lines) in Tg(npvf:eGFP); Tg(tph2:GCaMP6s-P2A-tdTomato) control (H,J) and Tg(npvf:ReaChR-mCitrine); Tg(tph2:GCaMP6s-P2A-tdTomato) (I,K) animals. F1-F5 in (H,I) indicate neurons from five different fish. (L) Box plot of normalized GCaMP6s ΔF/F0 values averaged for the first 10 imaging frames post-stimulation in Tg(npvf:GCaMP6s-P2A-tdTomato) animals that also contain either a Tg(npvf:eGFP) (black) or Tg(npvf:ReaChR-mCitrine) (gray) transgene. n = number of neurons. ***p<0.0005, Mann-Whitney test. Scale: 100 μm (A), 10 μm (C’).
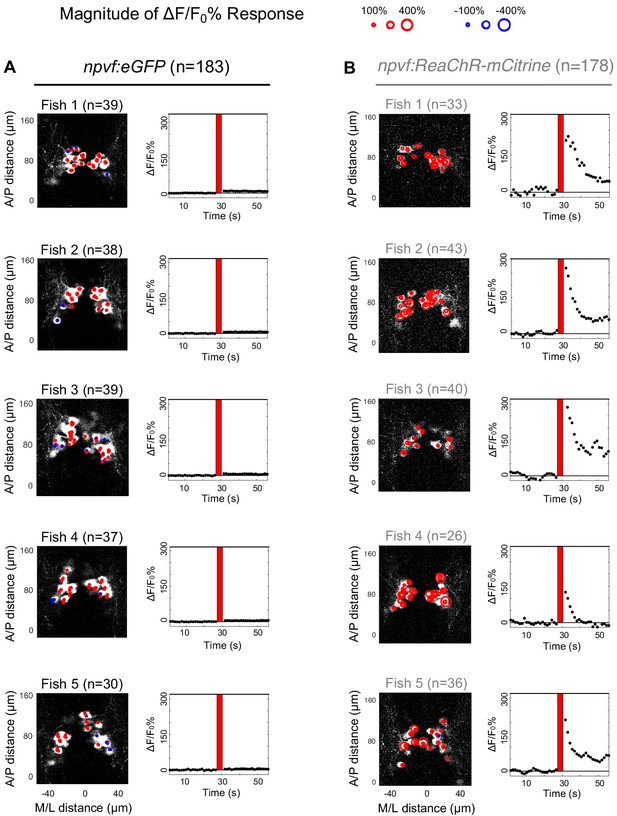
Responses of individual NPVF neurons to optogenetic stimulation of NPVF neurons in individual fish.
6-dpf Tg(npvf:eGFP); Tg(npvf:GCaMP6s-P2A-tdTomato) (A) and Tg(npvf:ReaChR-mCitrine); Tg(npvf:GCaMP6s-P2A-tdTomato) (B) animals were analyzed for normalized GCaMP6s fluorescence levels in NPVF neurons pre- and post-optogenetic stimulation of NPVF neurons. Data from Figure 2—figure supplement 1 are displayed here as separate fish. Maximum intensity projection images (left) show GCaMP6s fluorescence averaged over the first 10 imaging frames post-stimulation and provide spatial information along the anterior/posterior and medial/lateral axes of the brain. Size of red or blue circle indicates magnitude of normalized GCaMP6s fluorescence increase or decrease, respectively, in response to optogenetic stimulation. Dot plots (right) show average normalized GCaMP6s fluorescence of all NPVF neurons for each frame before and after optogenetic stimulation (red line). A, anterior; P, posterior; M, medial; L, lateral. n = number of NPVF neurons imaged.

Optogenetic stimulation of NPVF neurons activates anterior IRa neurons.
(A,B) 6-dpf Tg(npvf:eGFP); Tg(npvf:GCaMP6s-P2A-tdTomato) and (C,D) Tg(npvf:ReaChR-mCitrine); Tg(npvf:GCaMP6s-P2A-tdTomato) animals were analyzed for normalized GCaMP6s fluorescence in IRa neurons before and after optogenetic stimulation of NPVF neurons. Positive (red) or negative (blue) normalized GCaMP6s fluorescence responses of individual IRa neurons is plotted according to their spatial location along the anterior/posterior and medial/lateral axes of the brain. The size of red or blue circles indicates the magnitude of increase or decrease of normalized GCaMP6s fluorescence, respectively. The horizontal dashed line indicates division of the IRa into anterior and posterior halves. A, anterior; P, posterior; L, lateral; M, medial. ‘0’ along the A/P axis indicates location of the most posterior IRa neurons. ‘0’ along the M/L axis indicates the midline of the IRa neuron population. (E–L) Normalized GCaMP6s fluorescence of individual neurons in the anterior or posterior halves of the IRa shown as heat maps (E–H), in which each horizontal line represents an IRa neuron, and as line graphs (I–L) showing individual (gray lines) and mean (dotted line) IRa neuron responses before and after optogenetic stimulation (red lines) in Tg(npvf:eGFP); Tg(tph2:GCaMP6s-P2A-tdTomato) (E,G,I,J) and Tg(npvf:ReaChR-mCitrine); Tg(tph2:GCaMP6s-P2A-tdTomato) (F,H,K,L) animals. F1-F4 in (E–H) indicate neurons from four different fish. (M) Box plot of normalized GCaMP6s ΔF/F0 % values in anterior or posterior IRa neurons averaged for the first 10 imaging frames after stimulation of NPVF neurons in Tg(npvf:eGFP); Tg(tph2:GCaMP6s-P2A-tdTomato) (black) and Tg(npvf:ReaChR-mCitrine); Tg(npvf:GCaMP6s-P2A-tdTomato) (gray) animals. n = number of neurons quantified from four animals of each genotype. ***p<0.001, Two-way ANOVA, Holm-Sidak test.

Responses of individual IRa neurons to optogenetic stimulation of NPVF neurons in individual fish.
6-dpf Tg(npvf:eGFP); Tg(tph2:GCaMP6s-P2A-tdTomato) (A) and Tg(tph2:ReaChR-mCitrine); Tg(npvf:GCaMP6s-P2A-tdTomato) (B) animals were analyzed for normalized GCaMP6s fluorescence levels in IRa neurons pre- and post-optogenetic stimulation of NPVF neurons. Data from Figure 2 are displayed here as separate fish. Maximum intensity projection images (left) show GCaMP6s fluorescence averaged over the first 10 imaging frames post-stimulation and provide spatial information along the anterior/posterior and medial/lateral axes of the brain. Size of red or blue circle indicates magnitude of normalized GCaMP6s fluorescence increase or decrease, respectively, in response to optogenetic stimulation. Dot plots (right) show average normalized GCaMP6s fluorescence of all IRa neurons for each frame before and after optogenetic stimulation (red line). A, anterior; P, posterior; M, medial; L, lateral. n = number of IRa neurons imaged.
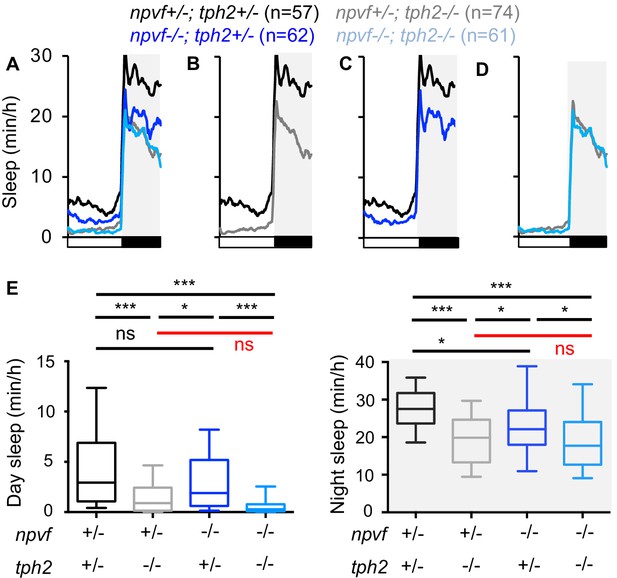
Loss of npvf does not enhance the tph2 mutant sleep phenotype.
(A–D) Sleep for npvf +/-; tph2 +/- (black), npvf -/-; tph2 +/- (dark blue), npvf +/-; tph2 -/- (gray), and npvf -/-; tph2 -/- (light blue) siblings. The graph on the left shows data for all four genotypes. The three other graphs show the same data as separate pair-wise comparisons. (E) Box plots quantify sleep during the day (left) and night (right). White boxes indicate day. Black boxes and gray shading indicate night. Data during the fifth day and night of development from four experiments combined is shown. n = number of animals. ns p>0.05, *p<0.05, ***p<0.005, Two-way ANOVA with Holm-Sidak test for each indicated pair-wise comparison. The comparison in red indicates no significant difference between npvf +/-; tph2 -/- and npvf -/-; tph2 -/- siblings.
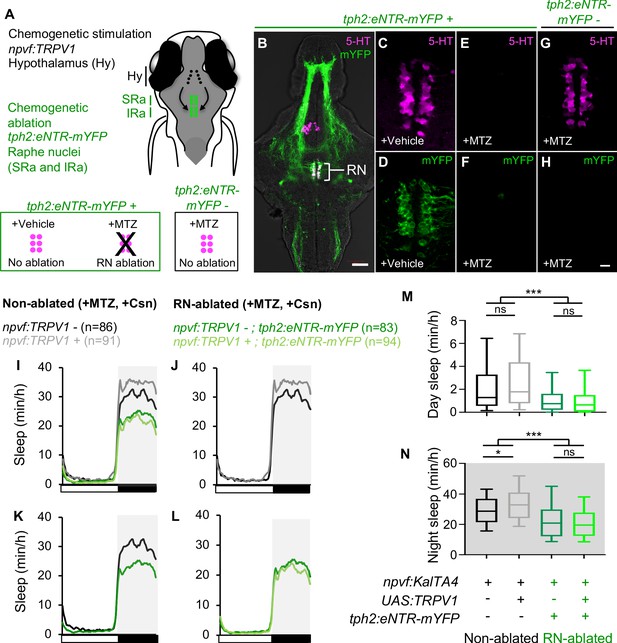
Chemogenetic ablation of the RN abolishes sleep induced by chemogenetic stimulation of NPVF neurons.
(A) Schematic of experiment. Animals were first treated with MTZ to ablate serotonergic RN neurons in Tg(tph2:eNTR-mYFP) animals but not in non-transgenic sibling controls. Behavior was then monitored during chemogenetic stimulation of npvf-expressing neurons in Tg(npvf:KalTA4); Tg(UAS:TRPV1-TagRFP-T) animals in parallel with their non-stimulated Tg(npvf:KalTA4) sibling controls. (B) 5-dpf Tg(tph2:eNTR-mYFP) zebrafish brain showing serotonergic RN neurons and some of their projections (green) and labeled with a 5-HT-specific antibody (magenta). The bracketed region is magnified in (C–H). Treatment of Tg(tph2:eNTR-mYFP) animals with MTZ results in the loss of both 5-HT immunoreactivity (E) and mYFP (F) in the RN, but treatment with DMSO vehicle control does not (C,D). MTZ treatment of Tg(tph2:eNTR-mYFP) negative siblings does not result in loss of RN neurons (G). Images are single 4-μm- (B) and 0.6-μm- (C–H) thick optical sections. Scale: 50 μm (B), 10 μm (C–H). (I–L) Sleep of 5-dpf Tg(npvf:KalTA4) (black), Tg(npvf:KalTA4); Tg(UAS:TRPV1-TagRFP-T) (gray), Tg(npvf:KalTA4); Tg(tph2:eNTR-mYFP) (dark green), and Tg(npvf:KalTA4); Tg(UAS:TRPV1-TagRFP-T); Tg(tph2:eNTR-mYFP) (light green) siblings treated with 2 μM Csn. White and black bars under behavioral traces indicate day and night, respectively. (M,N) Box plots quantify sleep during day (M) and night (N). n = number of animals. ns p>0.05, *p<0.05, ***p<0.005, Two-way ANOVA with Holm-Sidak test.
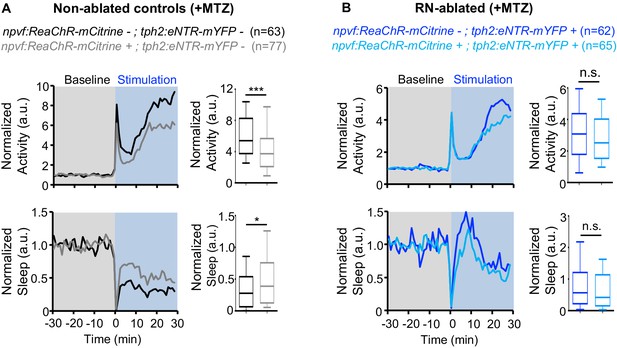
Chemogenetic ablation of the RN abolishes sleep induced by optogenetic stimulation of NPVF neurons.
Normalized locomotor activity (top) and sleep (bottom) of 5-dpf Tg(npvf:ReaChR-mCitrine) (gray and light blue) and non-transgenic sibling control (black and blue) animals before (Baseline) and during blue light exposure (Stimulation) in Tg(tph2:eNTR-mYFP) negative (A) or positive (B) siblings. Because the animals see the blue light, they exhibit a brief startle at light onset that is excluded from analysis, followed by a gradual increase in activity that plateaus after ~15 min. Box plots quantify locomotor activity and sleep for each animal during optogenetic stimulation normalized to the baseline of all animals of the same genotype. n = number of animals. ns p>0.05, *p<0.05, ***p<0.005, Mann-Whitney test.
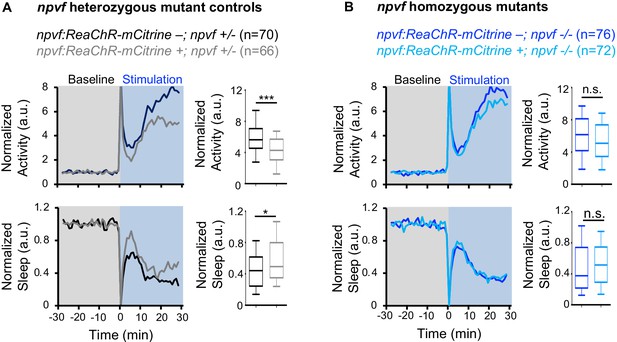
Sleep induced by optogenetic stimulation of NPVF neurons is abolished in npvf mutant animals.
Normalized locomotor activity (top) and sleep (bottom) of 5-dpf Tg(npvf:ReaChR-mCitrine) (gray and light blue) and non-transgenic sibling control (black and blue) animals before (Baseline) and during exposure to blue light (Stimulation) in npvf +/- (A) and npvf -/- (B) animals. Box plots quantify locomotor activity and sleep for each animal during optogenetic stimulation normalized to the baseline of all animals of the same genotype. Stimulation of NPVF neurons decreases locomotor activity and increases sleep compared to non-transgenic sibling controls in npvf +/- animals (A) but not in npvf -/- siblings (B). n = number of animals. ns p>0.05, *p<0.05, ***p<0.005, Mann-Whitney test.
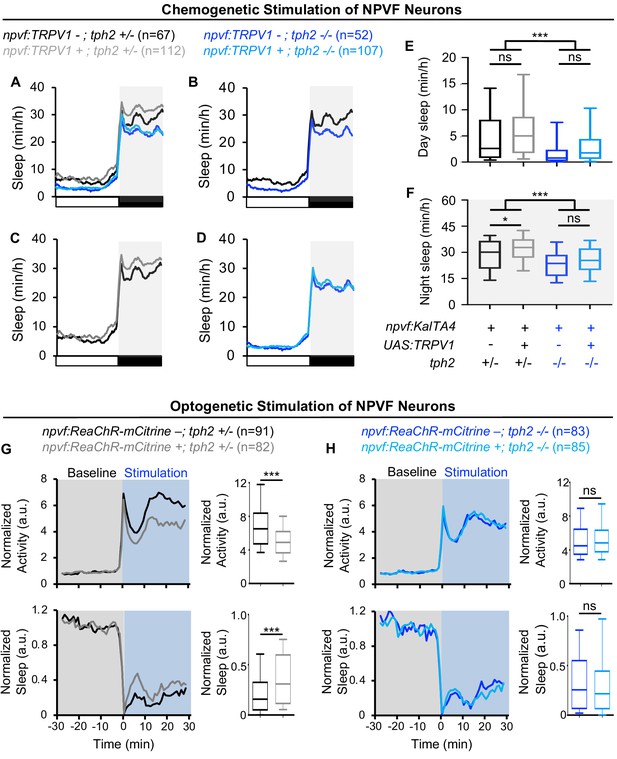
Sleep induced by chemogenetic or optogenetic stimulation of NPVF neurons is abolished in tph2 mutant animals.
(A) Sleep of 5-dpf Tg(npvf:KalTA4); tph2+/- (black), Tg(npvf:KalTA4); tph2-/- (dark blue), Tg(npvf:KalTA4); Tg(UAS:TRPV1-TagRFP-T); tph2+/- (gray), and Tg(npvf:KalTA4); Tg(UAS:TRPV1-TagRFP-T); tph2-/- (light blue) siblings treated with 2 μM Csn. White and black bars under behavioral traces indicate day and night, respectively. (E,F) Box plots quantify sleep during the day (E) and night (F). Chemogenetic stimulation of NPVF neurons increases sleep at night compared to non-transgenic sibling controls in tph2+/- animals (C,F) but not in tph2-/- siblings (D,F). n = number of animals. ns p>0.05, *p<0.05, ***p<0.005, Two-way ANOVA with Holm-Sidak test. (G,H) Normalized locomotor activity (top) and sleep (bottom) of Tg(npvf:ReaChR-mCitrine) (gray and light blue) and non-transgenic sibling control (black and dark blue) animals before (Baseline) and during exposure to blue light (Stimulation) in (G) tph2+/- or (H) tph2-/- animals. Box plots quantify locomotor activity and sleep for each animal during optogenetic stimulation normalized to the baseline of all animals with the same genotype. Optogenetic stimulation of NPVF neurons decreases locomotor activity and increases sleep compared to non-transgenic sibling controls in tph2+/- animals (G) but not in tph2-/- siblings (H). n = number of animals. ns p>0.05, ***p<0.005, Mann-Whitney test.
Tables
| Reagent type (species) or resource | Designation | Source or reference | Identifiers | Additional information |
|---|---|---|---|---|
| Antibody | Rabbit polyclonal anti-5-HT | MilliporeSigma | Cat# S5545; RRID:AB_477522 | (1:1000) |
| Antibody | Chicken polyclonal anti-GFP | Aves Laboratory | Cat# GFP-1020, RRID:AB_10000240 | (1:1000) |
| Antibody | Rabbit polyclonal anti-DsRed | Takara Bio | Cat# 632496, RRID:AB_10013483 | (1:1000) |
| Antibody | Rabbit polyclonal anti-tagRFP | Evrogen | Cat# AB233, RRID:AB_2571743 | (1:200) |
| Antibody | Goat polyclonal anti-Chicken IgY (H+L) Secondary Antibody, Alexa Fluor 488 | ThermoFisher Sci. | Cat# A-11039; RRID:AB_2534096 | (1:500) |
| Antibody | Goat polyclonal anti-Rabbit IgG (H+L) Cross-Adsorbed Secondary Antibody, Alexa Fluor 568 | ThermoFisher Sci. | Cat# A-11011; RRID:AB_143157 | (1:500) |
| Antibody | Goat polyclonal anti-Rat IgG (H+L) Cross-Adsorbed Secondary Antibody, Alexa Fluor 488 | ThermoFisher Sci. | Cat# A-11006, RRID:AB_2534074 | (1:500) |
| Chemical compound, drug | Metronidazole | MP Biomedicals | Cat# 0215571080 | |
| Strain, strain background (Danio rerio) | npvf ct845 mutant | Lee et al., 2017 | RRID:ZDB-ALT-170927-1 | |
| Strain, strain background (Danio rerio) | tph2 ct817 mutant | Chen et al., 2013a | RRID:ZDB-ALT-131122-14 | |
| Strain, strain background (Danio rerio) | Tg(npvf:eGFP) ct847Tg | Lee et al., 2017 | RRID:ZDB-ALT-170927-3 | |
| Strain, strain background (Danio rerio) | Tg(npvf:GCaMP6s-P2A-tdTomato) ct872Tg | Lee et al., 2019 | ZFIN: ZDB-ALT-190725–5 | |
| Strain, strain background (Danio rerio) | Tg(npvf:ReaChR-mCitrine) ct849Tg | Lee et al., 2017 | RRID:ZDB-ALT-170927-5 | |
| Strain, strain background (Danio rerio) | Tg(npvf:kalta4) ct848Tg | Lee et al., 2017 | RRID:ZDB-ALT-170927-4 | |
| Strain, strain background (Danio rerio) | Zebrafish: Tg(tph2:eNTR-mYFP) ct866Tg | Oikonomou et al., 2019 | RRID:ZDB-ALT-190508-3 | |
| Strain, strain background (Danio rerio) | Tg(tph2:GCaMP6s-P2A-NLS:tdTomato) ct874 | This study; Figure 2 and Figure 2—figure supplements 3 and 4. | ZFIN: ZDB-ALT-200512–2 | GCaMP6s-P2A-NLS:tdTomato expressed under the tph2 promoter; – Prober Lab |
| Strain, strain background (Danio rerio) | Tg(UAS:nfsb-mCherry) rw0144Tg | Agetsuma et al., 2010 | RRID:ZDB-ALT-110215-7 | |
| Strain, strain background (Danio rerio) | Tg(UAS:TRPV1-tagRFP-T) ct851Tg | Lee et al., 2017 | RRID:ZDB-ALT-170927-7 | |
| Sequence-based reagent | Primer: tph2 mutant genotyping primer 1: AGAACTTACAAAACTCTATCCAACTC | Oikonomou et al., 2019 | ||
| Sequence-based reagent | Primer: tph2 mutant genotyping primer 2: AGAGAGGACAACATCTGGGG | Oikonomou et al., 2019 | ||
| Sequence-based reagent | Primer: tph2 mutant genotyping primer 3: TAATCATGCAGTCCGTTAATACTC | Oikonomou et al., 2019 | ||
| Sequence-based reagent | Primer: npvf mutant genotyping primer 1: CAGTGGTGGTGCGAGTTCT | Lee et al., 2017 | ||
| Sequence-based reagent | Primer: npvf mutant genotyping primer 2: GCTGAGGGAGGTTGATGGTA | Lee et al., 2017 | ||
| Sequence-based reagent | Primer: Tg(npvf:ReaChR-mCitrine) genotyping primer 1: CACGAGAGAATGCTGTTCCA | Lee et al., 2017 | ||
| Sequence-based reagent | Primer: Tg(npvf:ReaChR-mCitrine) genotyping primer 2: CCATGGTGCGTTTGCTATAA | Lee et al., 2017 | ||
| Sequence-based reagent | Primer: Tg(UAS:TRPV1-tagRFP-T) genotyping primer 1: CAGCCTCACTTTGAGCTCCT: | Lee et al., 2017 | ||
| Sequence-based reagent | Primer: Tg(UAS:TRPV1-tagRFP-T) genotyping primer 2: TCCTCATAAGGGCAGTCCAG | Lee et al., 2017 | ||
| Software, algorithm | MATLAB R2017b | Mathworks | RRID:SCR_001622 | |
| Software, algorithm | Prism6 | GraphPad | RRID:SCR_002798 | |
| Software, algorithm | Image J/Fiji | Schneider et al., 2012 | RRID:SCR_002285 | |
| Other | 96-well plate | GE Healthcare Life Sciences | Cat#: 7701–1651 | |
| Other | MicroAmp Optical Adhesive Film | Thermo Fisher Scientific | Cat#: 4311971 |





