Breaking antimicrobial resistance by disrupting extracytoplasmic protein folding
Figures
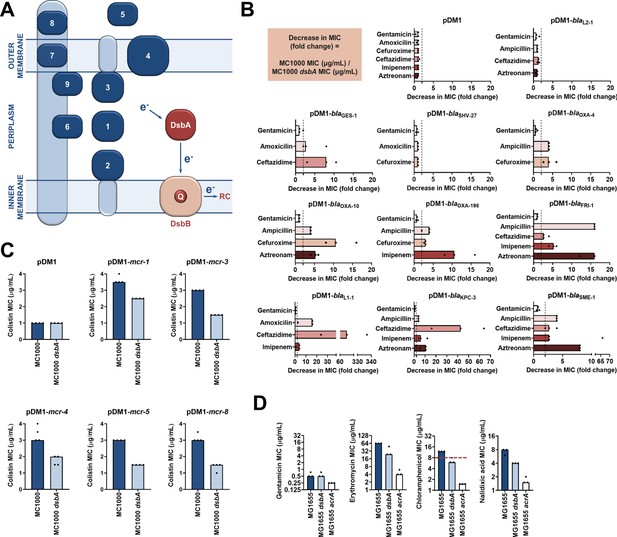
Several antimicrobial resistance mechanisms depend on disulfide bond formation.
(A) DsbA introduces disulfide bonds into extracytoplasmic proteins containing two or more cysteine residues. After each round of oxidative protein folding, DsbA is regenerated by the quinone (Q)-containing protein DsbB, which in turn transfers the reducing equivalents to the respiratory chain (RC) (Kadokura et al., 2003). DsbA substrates (in dark blue) are distributed throughout the extracytoplasmic space of Gram-negative bacteria. Disulfides are introduced to (1) soluble periplasmic proteins (e.g. alkaline phosphatase, β-lactamases; Bardwell et al., 1991), (2) periplasmic domains of inner-membrane proteins (e.g LptA-like enzymes (Piek et al., 2014), (3) periplasmic domains of outer-membrane proteins (e.g. RcsF; Denoncin and Collet, 2013), (4) outer-membrane proteins (e.g. OmpA, LptD; Denoncin and Collet, 2013; Heras et al., 2009), (5) secreted proteins (e.g. toxins or enzymes; Heras et al., 2009), (6–9) protein components of macromolecular assemblies like secretion systems, pili or flagella (Heras et al., 2009) (e.g. (6) GspD, (7) EscC, (8) BfpA, (9) FlgI); all examples are E. coli proteins with the exception of LptA. (B) β-lactam MIC values for E. coli MC1000 expressing diverse disulfide-bond-containing β-lactamases (Ambler classes A, B and D) are substantially reduced in the absence of DsbA (MIC fold changes: > 2, fold change of 2 is indicated by the black dotted lines); no effect is observed for SHV-27, which is further discussed in Figure 1—figure supplement 3. DsbA dependence is conserved within phylogenetic groups (see Figure 1—figure supplement 2). No changes in MIC values are observed for the aminoglycoside antibiotic gentamicin (white bars) confirming that absence of DsbA does not compromise the general ability of this strain to resist antibiotic stress. No changes in MIC values are observed for strains harboring the empty vector control (pDM1) or those expressing the class A β-lactamase L2-1, which contains three cysteines but no disulfide bond (top row). Graphs show MIC fold changes for β-lactamase-expressing E. coli MC1000 and its dsbA mutant from three biological experiments each conducted as a single technical repeat; the MIC values used to generate this panel are presented in Supplementary file 2a. (C) Colistin MIC values for E. coli MC1000 expressing diverse MCR enzymes (Figure 1—figure supplement 1) are reduced in the absence of DsbA. Graphs show MIC values (µg/mL) from four biological experiments, each conducted in technical quadruplicate, to demonstrate the robustness of the observed effects. Gentamicin control data are presented in Figure 1—figure supplement 6. (D) Deletion of dsbA reduces the erythromycin, chloramphenicol and nalidixic acid MIC values for E. coli MG1655, but no effects are detected for the non-substrate antibiotic gentamicin. The essential pump component AcrA serves as a positive control. Graphs show MIC values (µg/mL) from three biological experiments, each conducted as a single technical repeat. Red dotted lines indicate the EUCAST clinical breakpoint for chloramphenicol.
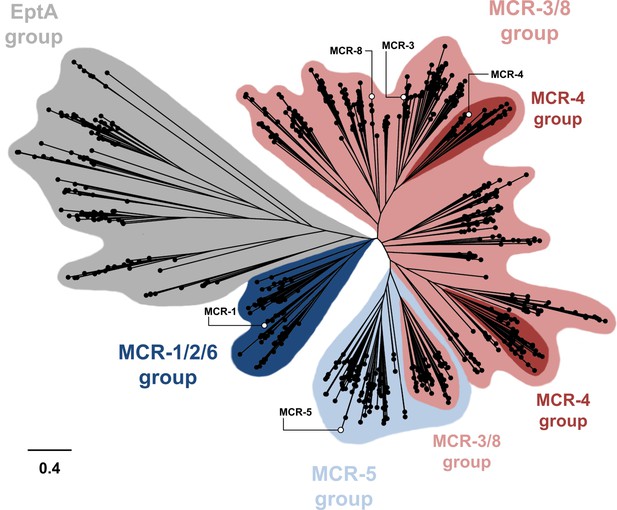
Phylogenetic analysis of MCR- and EptA-like enzymes found in Proteobacteria.
A phylogenetic tree was built based on the alignment of 781 sequences from Proteobacteria. The assignment of each sequence to a specific group was done using Hidden Markov Models built from confirmed sequences of MCR- and EptA-like proteins; EptA-like enzymes are chromosomally encoded phosphoethanolamine transferases that belong to the same extended protein superfamily as MCR enzymes (Zhang et al., 2019). The different MCR groups are broadly indicated in different colors, however it should be noted that there is significant overlap between groups. Open circles mark the enzymes tested in this study which are distributed throughout the MCR phylogeny.
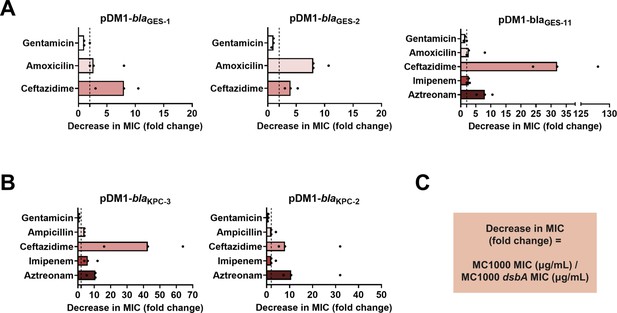
DsbA dependence is conserved within phylogenetic groups of disulfide-bond-containing β-lactamases.
β-lactam MIC values for E. coli MC1000 expressing disulfide-bond-containing β-lactamases belonging to the same phylogenetic family (Supplementary file 1) are substantially reduced in the absence of DsbA for all tested members of each family (MIC fold changes: > 2, fold change of 2 is indicated by the black dotted lines). No changes in MIC values are observed for the aminoglycoside antibiotic gentamicin (white bars) confirming that absence of DsbA does not compromise the general ability of this strain to resist antibiotic stress. (A) GES β-lactamase enzymes GES-1,–2, and –11; the data for GES-1 presented here are also shown as part of Figure 1B. (B) KPC β-lactamase enzymes KPC-3 and –2; the data for KPC-3 presented here are also shown as part of Figure 1B. (C) Graphs in panels (A) and (B) show MIC fold changes for β-lactamase-expressing E. coli MC1000 and its dsbA mutant. MIC assays were performed in three biological experiments each conducted as a single technical repeat; the MIC values used to generate this figure are presented in Supplementary file 2a.
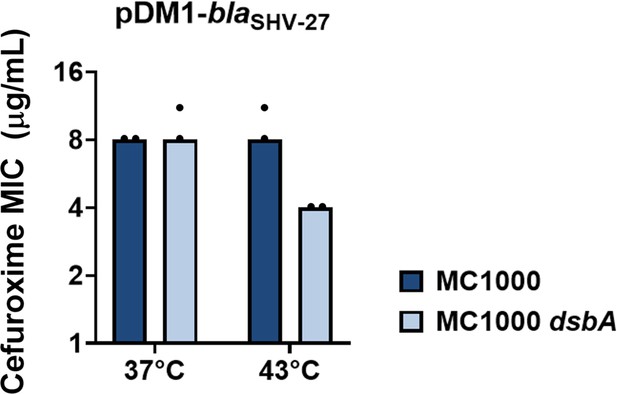
SHV-27 function is dependent on DsbA at temperatures higher than 37°C.
The ESBL SHV-27 differs from the canonical SHV-1 enzyme by a single amino acid substitution (D156G) (Corkill et al., 2001). At 37°C deletion of dsbA does not affect the cefuroxime MIC for E. coli MC1000 harboring pDM1-blaSHV-27. However, at 43°C the cefuroxime MIC for E. coli MC1000 dsbA harboring pDM1-blaSHV-27 is notably reduced. The graph shows MIC values (µg/mL) and is representative of three biological experiments, each conducted as a single technical repeat.
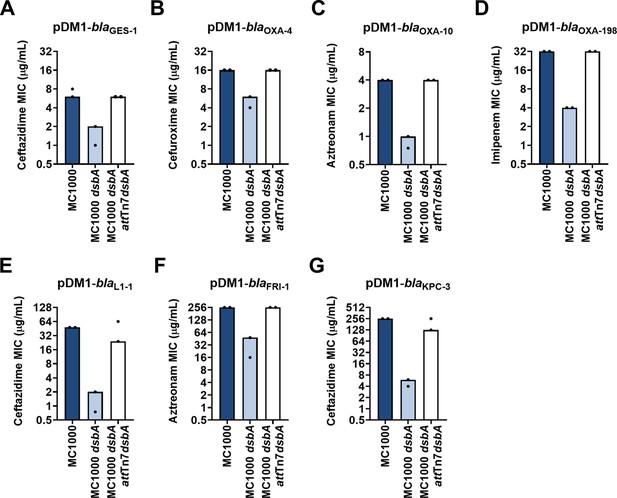
Complementation of dsbA restores the β-lactam MIC values for E. coli MC1000 dsbA expressing β-lactamases.
Re-insertion of dsbA at the attTn7 site of the chromosome restores the β-lactam MIC values for E. coli MC1000 dsbA harboring (A) pDM1-blaGES-1 (ceftazidime MIC), (B) pDM1-blaOXA-4 (cefuroxime MIC), (C) pDM1-blaOXA-10 (aztreonam MIC), (D) pDM1-blaOXA-198 (imipenem MIC), (E) pDM1-blaL1-1 (ceftazidime MIC), (F) pDM1-blaFRI-1 (aztreonam MIC) and (G) pDM1-blaKPC-3 (ceftazidime MIC). Graphs show MIC values (µg/mL) from two biological experiments, each conducted as a single technical repeat.
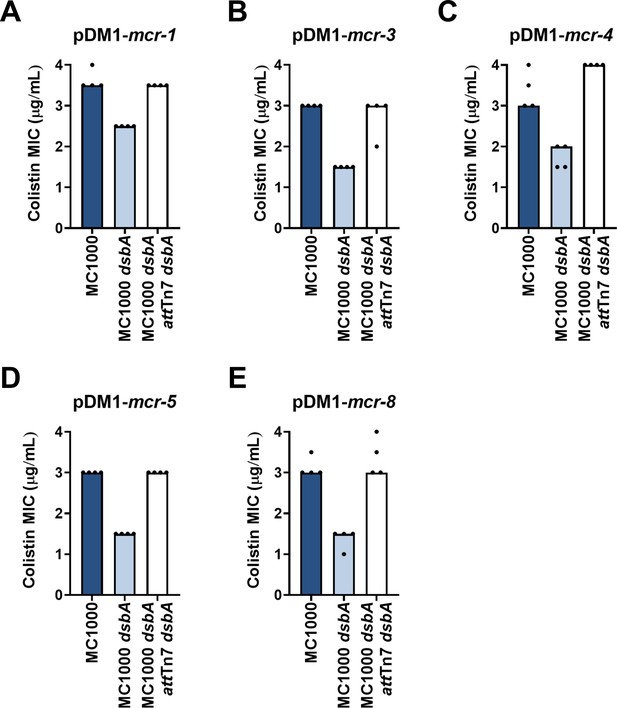
Complementation of dsbA restores the colistin MIC values for E. coli MC1000 dsbA expressing MCR enzymes.
Re-insertion of dsbA at the attTn7 site of the chromosome restores the colistin MIC values for E. coli MC1000 dsbA harboring (A) pDM1-mcr-1 (B) pDM1-mcr-3 (C) pDM1-mcr-4 (D) pDM1-mcr-5 (E) pDM1-mcr-8. Graphs show MIC values (µg/mL) from four biological experiments, each conducted in technical quadruplicate, to demonstrate the robustness of the observed effects.
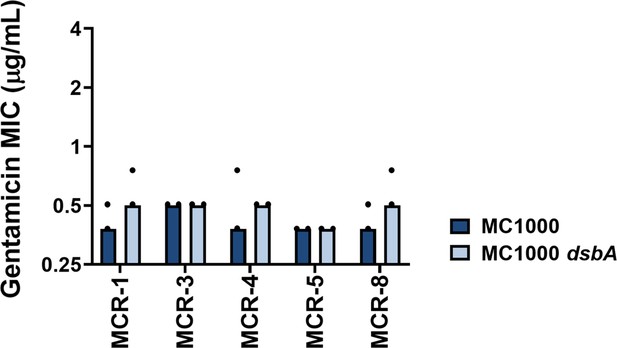
Gentamicin MIC values for E. coli MC1000 strains expressing MCR enzymes.
Deletion of dsbA does not affect the gentamicin MIC values for E. coli MC1000 strains expressing MCR enzymes, confirming that absence of DsbA does not compromise the general ability of this strain to resist antibiotic stress. Graphs show MIC values (µg/mL) from two biological experiments, each conducted as a single technical repeat.
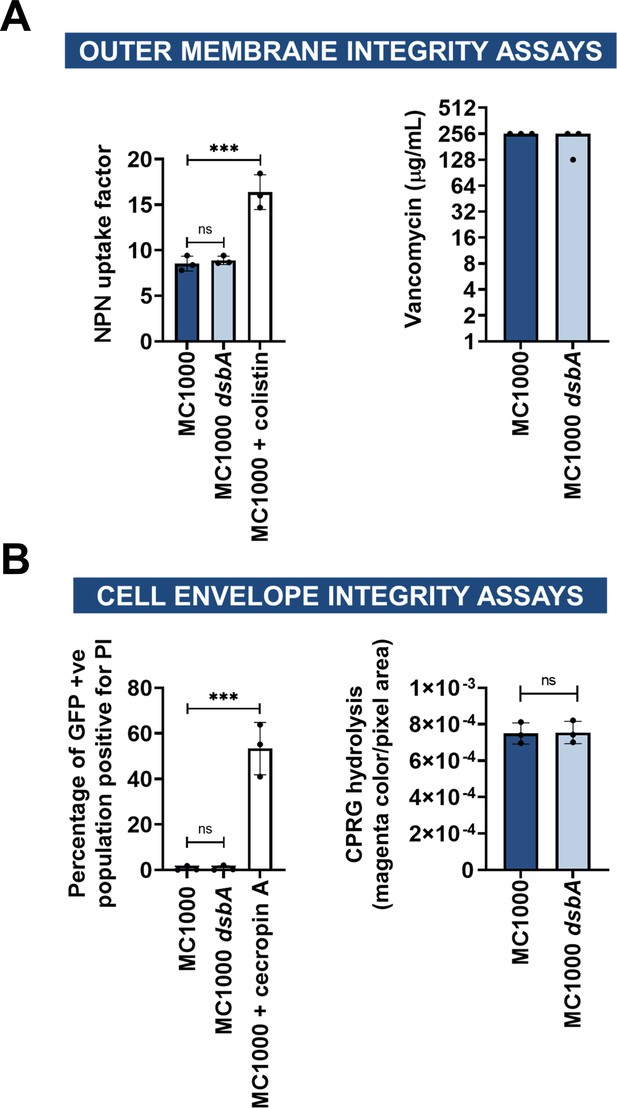
Deletion of dsbA has no effect on membrane permeability in E. coli MC1000.
(A) Outer membrane integrity assays. (left) The bacterial outer membrane acts as a selective permeability barrier to hydrophobic molecules. Deletion of dsbA has no effect on the outer membrane integrity of E. coli MC1000, as the hydrophobic fluorescent dye NPN crosses the outer membrane of E. coli MC1000 and its dsbA mutant to the same extent. Conversely, exposure to the outer-membrane-permeabilizing antibiotic colistin results in a significant increase in NPN uptake. (right) Outer membrane porins of Gram-negative bacteria are too small to allow the passage of large glycopeptides, such as vancomycin, and therefore increase in vancomycin susceptibility in E. coli indicates outer membrane defects. Deletion of dsbA has no effect on the outer membrane integrity of E. coli MC1000, as vancomycin MIC values for both strains do not present major differences. (B) Cell envelope integrity assays. (left) PI is a cationic hydrophilic dye that fluoresces upon intercalation with nucleic acids. Under normal conditions PI freely crosses the outer membrane but is unable to cross the inner membrane. Deletion of dsbA does not result in damage to the bacterial inner membrane, as no difference in basal PI uptake is seen between E. coli MC1000 and its dsbA mutant. Both strains harbor pUltraGFP-GM (Mavridou et al., 2016) for superfolder GFP (sfGFP) expression, and fluorescence was used to distinguish live from dead cells. Addition of the inner-membrane-permeabilizing antimicrobial peptide cecropin A (Silvestro et al., 2000) to E. coli MC1000 induces robust inner-membrane permeabilization in the sfGFP-positive population indicating that the inner membrane becomes compromised. (right) CPRG is excluded from the cytoplasm by the cell envelope, and therefore its hydrolysis by the cytosolic β-galactosidase is prevented. If both the inner and outer membranes are compromised, release of β-galactosidase results in CPRG breakdown and the appearance of red color. The red coloration of E. coli MC1000 dsbA colonies was comparable to those of the parent strain, showing that the cell envelope is not compromised in the mutant strain. E. coli MC1000 does not express the cytosolic β-galactosidase LacZ (Casadaban and Cohen, 1980), so for this assay the MC1000 strains harbor pCB112 (Paradis-Bleau et al., 2014), which expresses LacZ exogenously. For NPN and PI assays, n = 3 (each conducted in technical triplicate), graph shows means ± SD, significance is indicated by *** = p < 0.001, ns = non-significant. For vancomycin and CPRG hydrolysis assays, n = 3 (each conducted as a single technical triplicate). For CPRG hydrolysis assays, graph shows means ± SD, ns = non-significant.
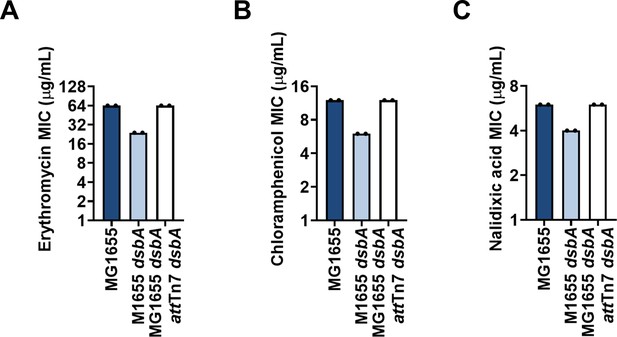
Complementation of dsbA restores efflux-pump substrate MIC values for E. coli MG1655 dsbA.
Re-insertion of dsbA at the attTn7 site of the chromosome restores (A) erythromycin, (B) chloramphenicol and (C) nalidixic acid MIC values for MG1655 dsbA. Graphs show MIC values (µg/mL) from two biological experiments, each conducted as a single technical repeat.
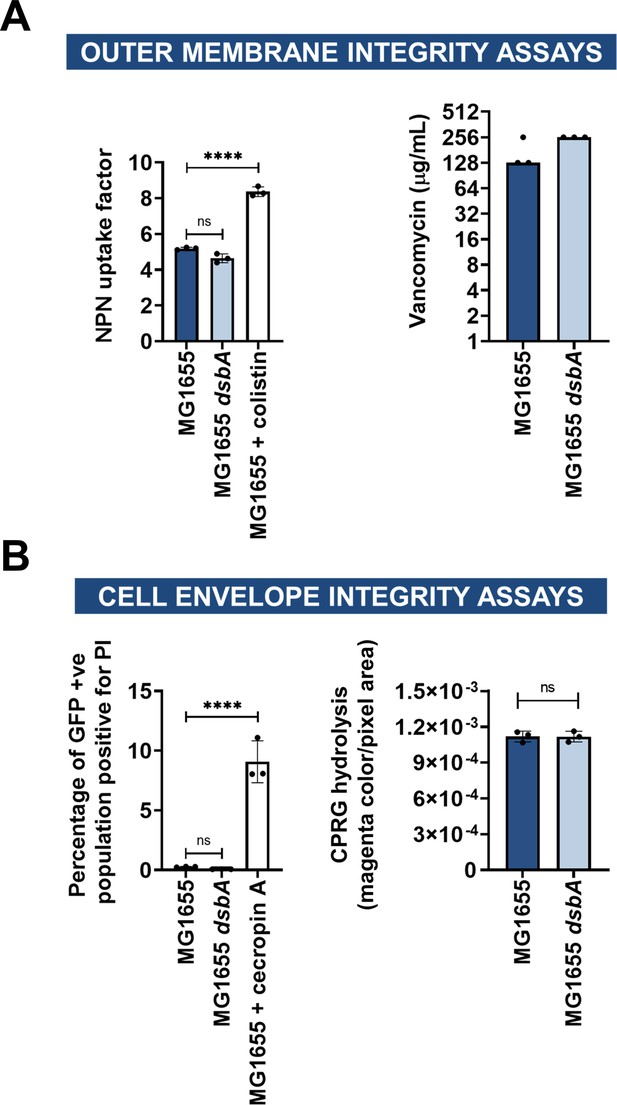
Deletion of dsbA has no effect on membrane permeability in E. coli MG1655.
(A) Outer membrane integrity assays. (left) The bacterial outer membrane acts as a selective permeability barrier to hydrophobic molecules. Deletion of dsbA has no effect on the outer membrane integrity of E. coli MG1655, as the hydrophobic fluorescent dye NPN crosses the outer membrane of E. coli MG1655 and its dsbA mutant to the same extent. Conversely, exposure to the outer-membrane-permeabilizing antibiotic colistin results in a significant increase in NPN uptake. (right) Outer membrane porins of Gram-negative bacteria are too small to allow the passage of large glycopeptides, such as vancomycin, and therefore increased vancomycin susceptibility in E. coli indicates outer membrane defects. Deletion of dsbA has no effect on the outer membrane integrity of E. coli MG1655, as vancomycin MIC values for both strains do not present major differences. (B) Cell envelope integrity assays. (left) PI is a cationic hydrophilic dye that fluoresces upon intercalation with nucleic acids. Under normal conditions, PI freely crosses the outer membrane but is unable to cross the inner membrane. Deletion of dsbA does not result in damage to the bacterial inner membrane, as no difference in basal PI uptake is seen between E. coli MG1655 and its dsbA mutant. Both strains harbor pUltraGFP-GM (Mavridou et al., 2016) for superfolder GFP (sfGFP) expression, and fluorescence was used to distinguish live from dead cells. Addition of the inner-membrane-permeabilizing antimicrobial peptide cecropin A (Silvestro et al., 2000) to E. coli MG1655 induces robust inner-membrane permeabilization in the sfGFP-positive population indicating that the inner membrane becomes compromised. (right) CPRG is excluded from the cytoplasm by the cell envelope, and therefore its hydrolysis by the cytosolic β-galactosidase is prevented. If both the inner and outer membranes are compromised, release of β-galactosidase results in CPRG breakdown and the appearance of red color. The red coloration of E. coli MG1655 dsbA colonies was comparable to those of the parent strain, showing that the cell envelope is not compromised in the mutant strain. For NPN and PI assays, n = 3 (each conducted in technical triplicate), graph shows means ± SD, significance is indicated by *** = p < 0.001, ns = non-significant. For vancomycin and CPRG hydrolysis assays, n = 3 (each conducted as a single technical repeat). For CPRG hydrolysis assays, graph shows means ± SD, ns = non-significant.
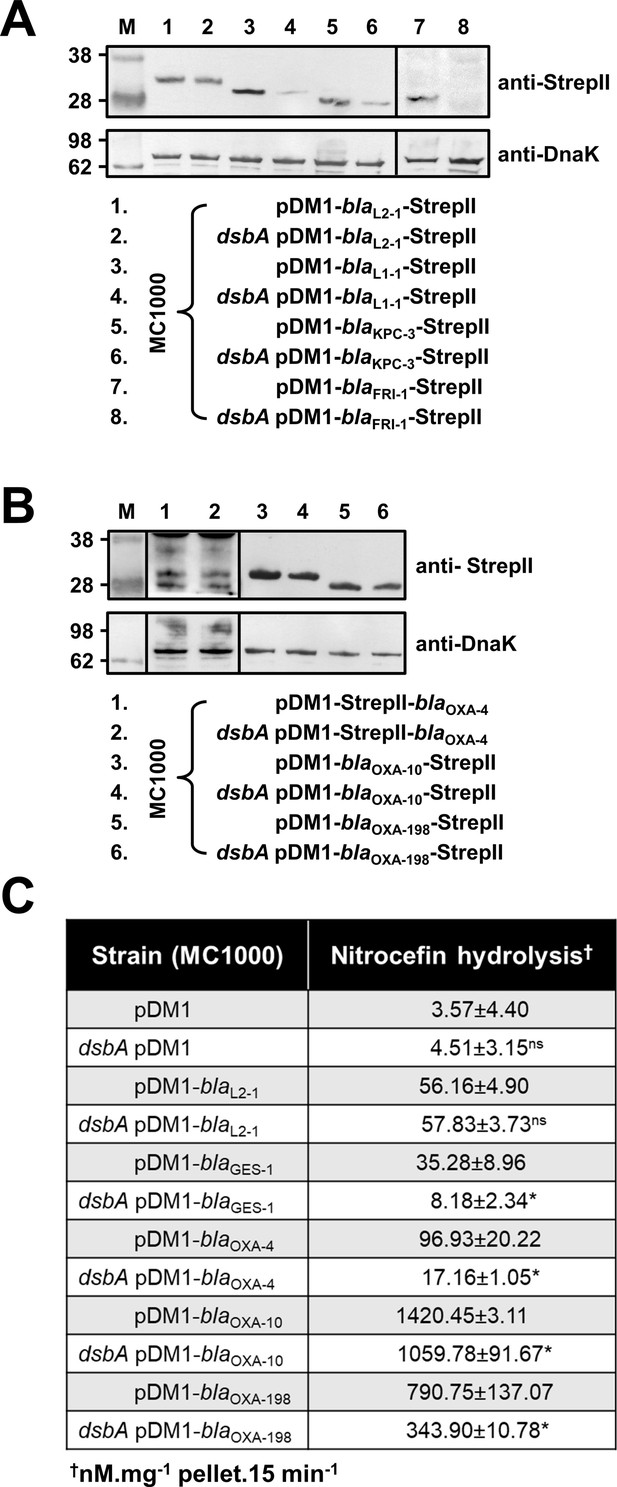
β-lactamase enzymes from most classes become unstable in the absence of DsbA.
(A) Protein levels of disulfide-bond-containing Ambler class A and B β-lactamases are drastically reduced when these enzymes are expressed in E. coli MC1000 dsbA; the amount of the control enzyme L2-1 is unaffected. (B) Protein levels of Class D disulfide-bond-containing β-lactamases are unaffected by the absence of DsbA. OXA-4 is detected as two bands at ~28 kDa. For panels (A) and (B) protein levels of StrepII-tagged β-lactamases were assessed using a Strep-Tactin-AP conjugate or a Strep-Tactin-HRP conjugate. A representative blot from three biological experiments, each conducted as a single technical repeat, is shown; molecular weight markers (M) are on the left, DnaK was used as a loading control and solid black lines indicate where the membrane was cut. (C) The hydrolytic activities of the tested Class D β-lactamases and of the Class A enzyme GES-1, which could not be detected by immunoblotting, are significantly reduced in the absence of DsbA. The hydrolytic activities of strains harboring the empty vector or expressing the control enzyme L2-1 show no dependence on DsbA. n = 3 (each conducted in technical duplicate), table shows means ± SD, significance is indicated by * = p < 0.05, ns = non-significant.
-
Figure 2—source data 1
Original files of the full raw unedited immunoblots used to prepare Figure 2A.
‘Top Panel’ in the file name refers to immunoblots carried out using a Strep-Tactin-AP conjugate, while ‘Bottom Panel’ refers to immunoblots carried out using an anti-DnaK 8E2/2 antibody. ‘Left’ and ‘Right’ in the file names refer to the part of the immunoblot to the left or to the right of the vertical black line shown in the final figure, respectively.
- https://cdn.elifesciences.org/articles/57974/elife-57974-fig2-data1-v2.zip
-
Figure 2—source data 2
Uncropped immunoblots used to prepare Figure 2A.
The figure included in the paper is shown in the center and relevant bands used for each part of the figure are marked with color-coded boxes on the uncropped immunoblots.
- https://cdn.elifesciences.org/articles/57974/elife-57974-fig2-data2-v2.zip
-
Figure 2—source data 3
Original files of the full raw unedited immunoblots used to prepare Figure 2B.
‘Top Panel’ in the file name refers to immunoblots carried out using a Strep-Tactin-AP conjugate or a Strep-Tactin-HRP conjugate, while ‘Bottom Panel’ refers to immunoblots carried out using an anti-DnaK 8E2/2 antibody. ‘Left’, ‘Middle’, and ‘Right’ in the file names refer to the part of the immunoblot to the left, in-between, or to the right of the vertical black lines shown in the final figure, respectively.
- https://cdn.elifesciences.org/articles/57974/elife-57974-fig2-data3-v2.zip
-
Figure 2—source data 4
Uncropped immunoblots used to prepare Figure 2B.
The figure included in the paper is shown in the center and relevant bands used for each part of the figure are marked with color-coded boxes on the uncropped immunoblots.
- https://cdn.elifesciences.org/articles/57974/elife-57974-fig2-data4-v2.zip
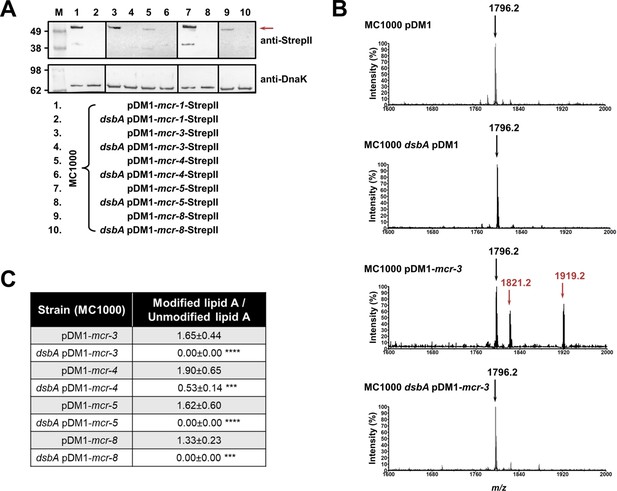
MCR enzymes become unstable in the absence of DsbA.
(A) The amounts of MCR proteins are drastically reduced when they are expressed in E. coli MC1000 dsbA; the red arrow indicates the position of the MCR-specific bands. Protein levels of StrepII-tagged MCR enzymes were assessed using a Strep-Tactin-AP conjugate. A representative blot from three biological experiments, each conducted as a single technical repeat, is shown; molecular weight markers (M) are on the left, DnaK was used as a loading control and solid black lines indicate where the membrane was cut. (B) The ability of MCR enzymes to transfer phoshoethanolamine to the lipid A portion of LPS is either entirely abrogated or significantly reduced in the absence of DsbA. This panel shows representative MALDI-TOF mass spectra of unmodified and MCR-modified lipid A in the presence and absence of DsbA. In E. coli, MC1000 and MC1000 dsbA the major peak for native lipid A is detected at m/z 1796.2 (first and second spectrum, respectively). In the presence of MCR enzymes (E. coli MC1000 expressing MCR-3 is shown as a representative example), two additional peaks are observed, at m/z 1821.2 and 1919.2 (third spectrum). For dsbA mutants expressing MCR enzymes (E. coli MC1000 dsbA expressing MCR-3 is shown), these additional peaks are not present, whilst the native lipid A peak at m/z 1796.2 remains unchanged (fourth spectrum). Mass spectra are representative of the data generated from four biological experiments, each conducted as a technical duplicate. (C) Quantification of the intensities of the lipid A peaks recorded by MALDI-TOF mass spectrometry for all tested MCR-expressing strains. n = 4 (each conducted in technical duplicate), table shows means ± SD, significance is indicated by *** = p < 0.001 or **** = p < 0.0001.
-
Figure 3—source data 1
Original files of the full raw unedited immunoblots used to prepare Figure 3A for which a Strep-Tactin-AP conjugate and an anti-DnaK 8E2/2 antibody were used.
The file names indicate the lanes of the immunoblot included in the paper that each of these files corresponds to.
- https://cdn.elifesciences.org/articles/57974/elife-57974-fig3-data1-v2.zip
-
Figure 3—source data 2
Uncropped immunoblots used to prepare Figure 3A.
The figure included in the paper is shown at the top and relevant bands used for each part of the figure are marked with color-coded boxes on the uncropped immunoblots.
- https://cdn.elifesciences.org/articles/57974/elife-57974-fig3-data2-v2.zip
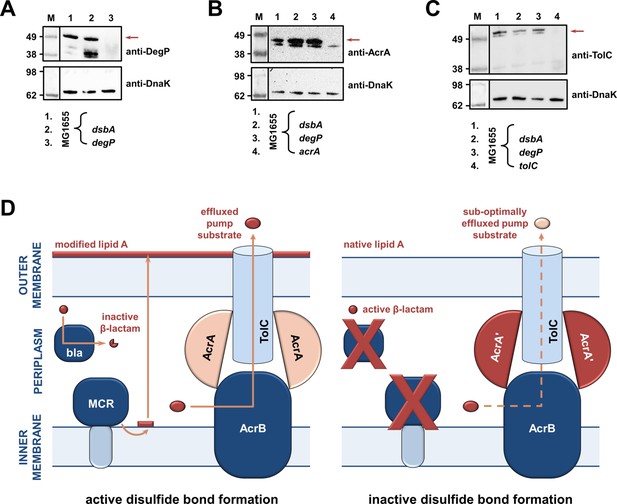
RND efflux pump function is impaired in the absence of DsbA due to accumulation of unfolded AcrA resulting from insufficient DegP activity (A, B, C).
(A) In the absence of DsbA the pool of active DegP is reduced. In E. coli MG1655 (lane 1), DegP is detected as a single band, corresponding to the intact active enzyme. In E. coli MG1655 dsbA (lane 2), an additional lower molecular weight band of equal intensity is present, indicating that DegP is degraded in the absence of its disulfide bond (Hiniker and Bardwell, 2004; Skórko-Glonek et al., 2003). DegP protein levels were assessed using an anti-DegP primary antibody and an HRP-conjugated secondary antibody. E. coli MG1655 degP was used as a negative control for DegP detection (lane 3); the red arrow indicates the position of intact DegP. (B) The RND pump component AcrA accumulates to the same extent in the E. coli MG1655 dsbA and degP strains, indicating that in both strains protein clearance is affected. AcrA protein levels were assessed using an anti-AcrA primary antibody and an HRP-conjugated secondary antibody. E. coli MG1655 acrA was used as a negative control for AcrA detection; the red arrow indicates the position of the AcrA band. (C) TolC, the outer-membrane channel of the AcrAB pump, does not accumulate in a dsbA or a degP mutant. TolC is not a DegP substrate (Werner et al., 2003), hence similar TolC protein levels are detected in E. coli MG1655 (lane 1) and its dsbA (lane 2) and degP (lane 3) mutants. TolC protein levels were assessed using an anti-TolC primary antibody and an HRP-conjugated secondary antibody. E. coli MG1655 tolC was used as a negative control for TolC detection (lane 4); the red arrow indicates the position of the bands originating from TolC. For all panels a representative blot from three biological experiments, each conducted as a single technical repeat, is shown; molecular weight markers (M) are on the left, DnaK was used as a loading control and solid black lines indicate where the membrane was cut. (D) Impairing disulfide bond formation in the cell envelope simultaneously affects distinct AMR determinants. (Left) When DsbA is present, that is, when disulfide bond formation occurs, degradation of β-lactam antibiotics by β-lactamases (marked ‘bla’), modification of lipid A by MCR proteins and active efflux of RND pump substrates lead to resistance. The major E. coli RND efflux pump AcrAB-TolC is depicted in this schematic as a characteristic example. (Right) In the asucess of disulfide bond formation is impaired, most cysteine-containing β-lactamases as well as MCR proteins are unstable and degrade, making bacteria susceptible to β-lactams and colistin, respectively. Absence of DsbA has also a general effect on proteostasis in the cell envelope which results in reduced clearance of nonfunctional AcrA-like proteins (termed ‘AcrA’ and depicted in dark red color) by periplasmic proteases. Insufficient clearance of these damaged AcrA components from the pump complex makes efflux less efficient.
-
Figure 4—source data 1
Original files of the full raw unedited immunoblots used to prepare Figure 4A.
‘Top Panel’ in the file name refers to immunoblots carried out using an anti-HtrA1 (DegP) antibody, while ‘Bottom Panel’ refers to immunoblots carried out using an anti-DnaK 8E2/2 antibody. ‘Left’ and ‘Right’ in the file names refer to the part of the immunoblot to the left or to the right of the vertical black line shown in the final figure, respectively.
- https://cdn.elifesciences.org/articles/57974/elife-57974-fig4-data1-v2.zip
-
Figure 4—source data 2
Uncropped immunoblots used to prepare Figure 4A.
The figure included in the paper is shown in the center and relevant bands used for each part of the figure are marked with color-coded boxes on the uncropped immunoblots.
- https://cdn.elifesciences.org/articles/57974/elife-57974-fig4-data2-v2.zip
-
Figure 4—source data 3
Original files of the full raw unedited immunoblots used to prepare Figure 4B.
‘Top Panel’ in the file name refers to immunoblots carried out using an anti-AcrA antibody, while ‘Bottom Panel’ refers to immunoblots carried out using an anti-DnaK 8E2/2 antibody. ‘Left’ and ‘Right’ in the file names refer to the part of the immunoblot to the left or to the right of the vertical black line shown in the final figure, respectively.
- https://cdn.elifesciences.org/articles/57974/elife-57974-fig4-data3-v2.zip
-
Figure 4—source data 4
Uncropped immunoblots used to prepare Figure 4B.
The figure included in the paper is shown in the center and relevant bands used for each part of the figure are marked with color-coded boxes on the uncropped immunoblots.
- https://cdn.elifesciences.org/articles/57974/elife-57974-fig4-data4-v2.zip
-
Figure 4—source data 5
Original files of the full raw unedited immunoblots used to prepare Figure 4C.
‘Top Panel’ in the file name refers to immunoblots carried out using an anti-TolC antibody, while ‘Bottom Panel’ refers to immunoblots carried out using an anti-DnaK 8E2/2 antibody. ‘Left’ and ‘Right’ in the file names refer to the part of the immunoblot to the left or to the right of the vertical black line shown in the final figure, respectively.
- https://cdn.elifesciences.org/articles/57974/elife-57974-fig4-data5-v2.zip
-
Figure 4—source data 6
Uncropped immunoblots used to prepare Figure 4C.
The figure included in the paper is shown in the center and relevant bands used for each part of the figure are marked with color-coded boxes on the uncropped immunoblots.
- https://cdn.elifesciences.org/articles/57974/elife-57974-fig4-data6-v2.zip
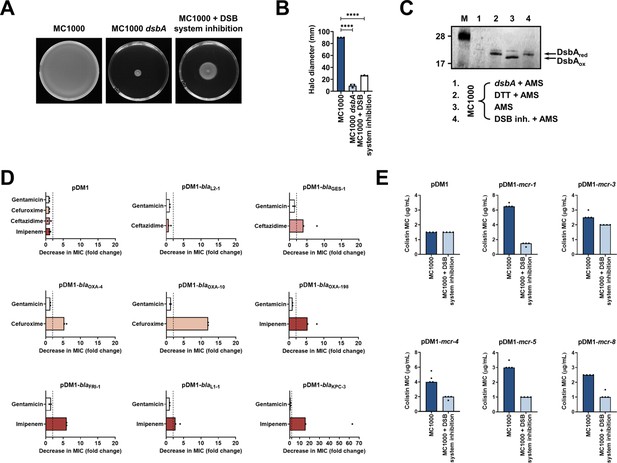
Chemical inhibition of the DSB system impedes DsbA function in E. coli MC1000 and phenocopies the β-lactam and colistin MIC changes that were observed using a dsbA mutant.
(A) Chemical inhibition of the DSB system impedes flagellar motility in E. coli MC1000. A functional DSB system is necessary for flagellar motility in E. coli because folding of the P-ring component FlgI requires DsbA-mediated disulfide bond formation (Dailey and Berg, 1993). In the absence of DsbA, or upon addition of a chemical inhibitor of the DSB system, the motility of E. coli MC1000 is significantly impeded. Representative images of motility plates are shown. (B) Quantification of the growth halo diameters in the motility assays shown in panel (A). n = 3 (each conducted as a single technical repeat), graph shows means ± SD, significance is indicated by **** = p < 0.0001. (C) Chemical inhibition of the DSB system impedes DsbA re-oxidation in E. coli MC1000. Addition of the reducing agent DTT to E. coli MC1000 bacterial lysates allows the detection of DsbA in its reduced form (DsbAred) during immunoblotting; this redox state of the protein, when labeled with the cysteine-reactive compound AMS, shows a 1 kDa size difference (lane 2) compared to oxidized DsbA as found in AMS-labeled but not reduced lysates of E. coli MC1000 (lane 3). Addition of a small-molecule inhibitor of DsbB to growing E. coli MC1000 cells also results in accumulation of reduced DsbA (lane 4). E. coli MC1000 dsbA was used as a negative control for DsbA detection (lane 1). A representative blot from two biological experiments, each conducted as a single technical repeat, is shown; DsbA was visualized using an anti-DsbA primary antibody and an AP-conjugated secondary antibody. Molecular weight markers (M) are shown on the left. (D) MIC experiments using representative β-lactam antibiotics show that chemical inhibition of the DSB system reduces the MIC values for E. coli MC1000 expressing disulfide-bond-containing β-lactamases in a similar manner to the deletion of dsbA (compare with Figure 1B). Graphs show MIC fold changes (i.e. MC1000 MIC (µg/mL) / MC1000 + DSB system inhibitor MIC (µg/mL)) for β-lactamase-expressing E. coli MC1000 with and without addition of a DSB system inhibitor to the culture medium from two biological experiments, each conducted as a single technical repeat. Black dotted lines indicate an MIC fold change of 2. The aminoglycoside antibiotic gentamicin serves as a control for all strains; gentamicin MIC values (white bars) are unaffected by chemical inhibition of the DSB system (MIC fold changes: < 2). No changes in MIC values (MIC fold changes: < 2) are observed for strains harboring the empty vector control (pDM1) or expressing the class A β-lactamase L2-1, which contains three cysteines but no disulfide bond (PDB ID: 1O7E) (top row). The MIC values used to generate this panel are presented in Supplementary file 2b. (E) Colistin MIC experiments show that chemical inhibition of the DSB system reduces the MIC values for E. coli MC1000 expressing MCR enzymes in a similar manner to the deletion of dsbA (compare with Figure 1C). Colistin MIC values for strains harboring the empty vector control (pDM1) are unaffected by chemical inhibition of the DSB system. Graphs show MIC values (µg/mL) from four biological experiments, each conducted in technical quadruplicate, to demonstrate the robustness of the observed effects.
-
Figure 5—source data 1
Original file of the full raw unedited immunoblot used to prepare Figure 5C, for which an anti-DsbA antibody was used.
- https://cdn.elifesciences.org/articles/57974/elife-57974-fig5-data1-v2.zip
-
Figure 5—source data 2
Uncropped immunoblot used to prepare Figure 5C.
The figure included in the paper is shown at the bottom and relevant bands used for each part of the figure are marked with a red box on the uncropped immunoblot.
- https://cdn.elifesciences.org/articles/57974/elife-57974-fig5-data2-v2.zip
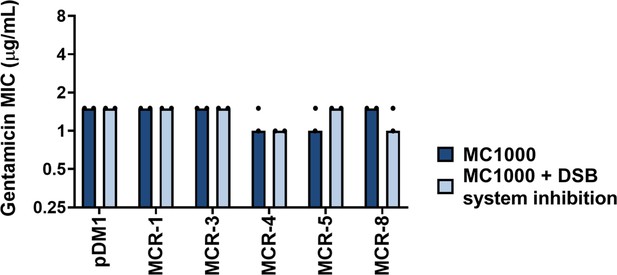
Gentamicin MIC values for E. coli MC1000 strains expressing MCR enzymes.
Chemical inhibition of the DSB system does not affect the gentamicin MIC values for E. coli MC1000 strains expressing MCR enzymes, confirming that inactivation of DsbA does not compromise the general ability of this strain to resist antibiotic stress. Graphs show MIC values (µg/mL) from two biological experiments, each conducted as a single technical repeat.
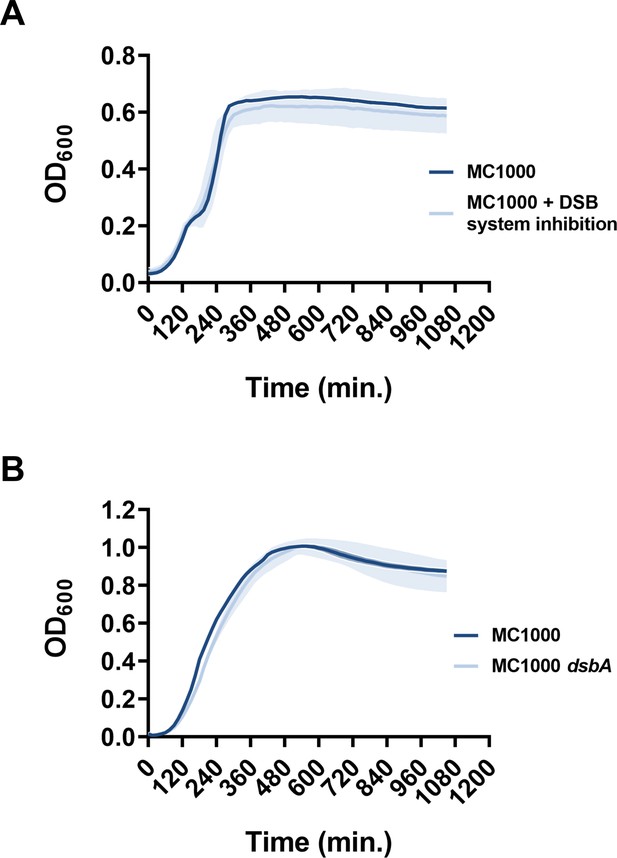
Chemical inhibition of the DSB system or deletion of dsbA does not compromise the growth of E. coli MC1000.
Growth curves of (A) E. coli MC1000 with and without chemical inhibition of the DSB system and (B) E. coli MC1000 and its dsbA mutant show that bacterial growth remains unaffected by the DSB system inhibitor compound used in this study, or by the absence of DsbA. n = 3 (each conducted as a technical triplicate), solid lines indicate mean values, shaded areas indicate SD.
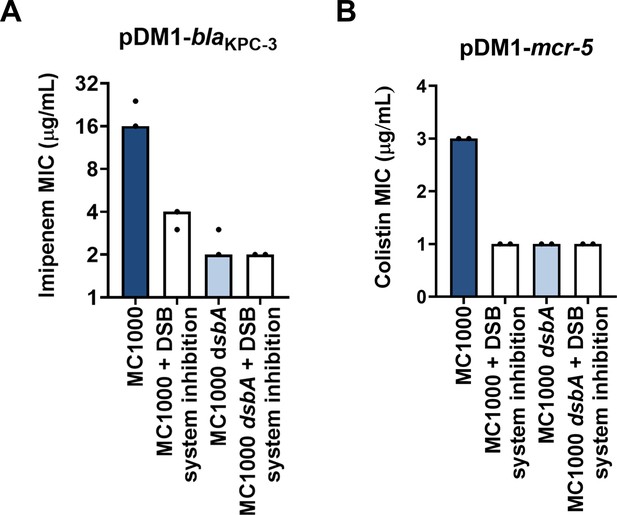
Changes in MIC values observed using the DSB system inhibitor are due solely to inhibition of the DSB system.
(A) E. coli MC1000 harboring pDM1-blaKPC-3 has an imipenem MIC value of 24 μg/mL. Upon chemical inhibition of the DSB system the imipenem MIC for this strain drops to 4 μg/mL, and accordingly the imipenem MIC for E. coli MC1000 dsbA harboring pDM1-blaKPC-3 is 2 μg/mL. The imipenem MIC for E. coli MC1000 dsbA harboring pDM1-blaKPC-3 when exposed to the chemical inhibitor of the DSB system is also 2 μg/mL, indicating that the chemical compound used in this study does not have any off-target effects and only affects the function of the DSB system proteins. (B) Chemical inhibition of the DSB system does not lead to any cumulative effects when tested on an E. coli MC1000 strain expressing MCR-5. The colistin MIC for E. coli MC1000 harboring pDM1-mcr-5 is 3 μg/mL and it drops to 1 μg/mL when the DSB system is chemically inhibited or dsbA is deleted. The same drop in colistin MIC is observed when the E. coli MC1000 dsbA strain harboring pDM1-mcr-5 is exposed to the chemical inhibitor of the DSB system. Data shown in both panels are from two biological experiments, each conducted as a single technical repeat.
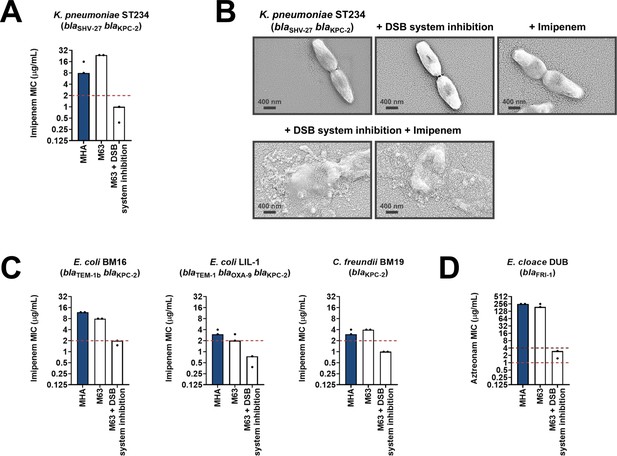
Chemical inhibition of the DSB system sensitizes multidrug-resistant clinical isolates to currently available β-lactam antibiotics.
(A) Addition of a small-molecule inhibitor of DsbB results in sensitization of a K. pneumoniae clinical isolate to imipenem. (B) Chemical inhibition of the DSB system in the presence of imipenem (final concentration of 6 μg/mL) results in drastic changes in cell morphology for the K. pneumoniae clinical isolate used in panel (A), while bacteria remain unaffected by single treatments (DSB inhibitor or imipenem). Images show representative scanning electron micrographs of untreated cells (top row, left), cells treated with the DSB inhibitor (top row, middle), cells treated with imipenem (top row, right), and cells treated with both the DSB inhibitor and imipenem (bottom row). Scale bars are at 400 nm. (C) Addition of a small-molecule inhibitor of DsbB results in sensitization of E. coli and C. freundii clinical isolates to imipenem. (D) Chemical inhibition of the DSB system of an E. cloacae clinical isolate harboring blaFRI-1 results in reduction of the aztreonam MIC value by over 180 µg/mL, resulting in intermediate resistance as defined by EUCAST. For panels (A), (C) and (D) graphs show MIC values (μg/ml) from two biological experiments, each conducted as a single technical repeat. MIC values determined using Mueller-Hinton agar (MHA) in accordance with the EUCAST guidelines (dark blue bars) are comparable to the values obtained using defined media (M63 agar, white bars); use of growth media lacking small-molecule oxidants is required for the DSB system inhibitor to be effective. Red dotted lines indicate the EUCAST clinical breakpoint for each antibiotic, and purple dotted lines indicate the EUCAST threshold for intermediate resistance.
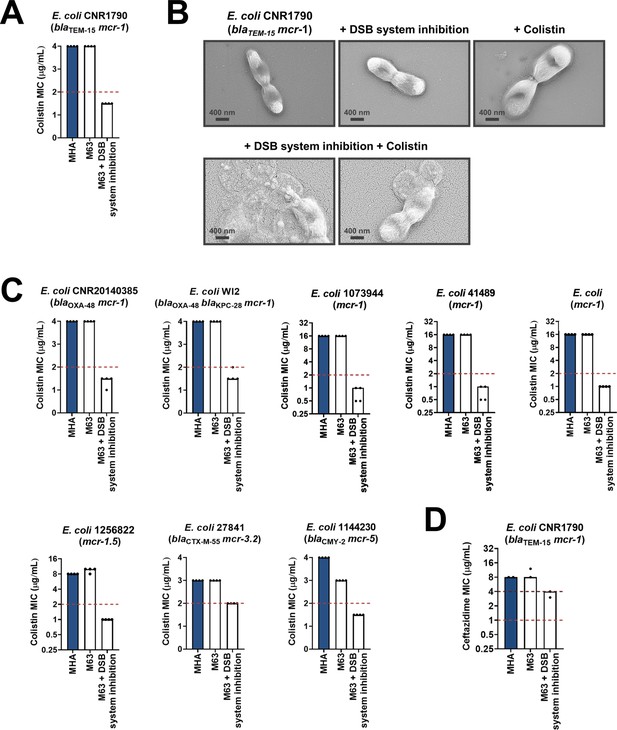
Chemical inhibition of the DSB system sensitizes multidrug-resistant clinical isolates to colistin.
(A) Addition of a small-molecule inhibitor of DsbB to a colistin-resistant clinical E. coli isolate expressing MCR-1 results in sensitization to colistin. (B) Chemical inhibition of the DSB system in the presence of colistin (final concentration of 2 μg/mL) results in drastic changes in cell morphology for the E. coli clinical isolate used in panel (A), while bacteria remain unaffected by single treatments (DSB inhibitor or colistin). Images show representative scanning electron micrographs of untreated cells (top row, left), cells treated with the DSB inhibitor (top row, middle), cells treated with colistin (top row, right), and cells treated with both the DSB inhibitor and colistin (bottom row). Scale bars are at 400 nm. (C) Chemical inhibition of the DSB system results in sensitization of four additional colistin-resistant E. coli strains expressing MCR enzymes. For panels (A) and (C), graphs show MIC values (µg/mL) from four biological experiments, each conducted in technical quadruplicate, to demonstrate the robustness of the observed effects. (D) Use of the DSB system inhibitor on the same clinical E. coli isolate tested in panel (A), results in intermediate resistance for ceftazidime as defined by EUCAST. The graph shows MIC values (μg/ml) from two biological experiments, each conducted as a single technical repeat. For panels (A), (C), (D), MIC values determined using Mueller-Hinton agar (MHA) in accordance with the EUCAST guidelines (dark blue bars) are comparable to the values obtained using defined media (M63 agar, white bars); use of growth media lacking small-molecule oxidants is required for the DSB system inhibitor to be effective. For all panels, red dotted lines indicate the EUCAST clinical breakpoint for each antibiotic, and purple dotted lines indicate the EUCAST threshold for intermediate resistance.
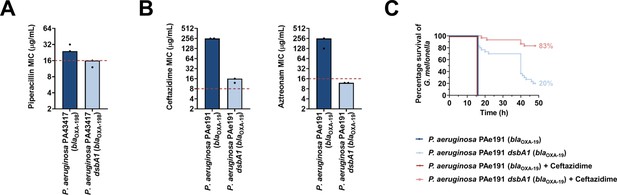
Absence of the principal DsbA analogue (DsbA1) from P. aeruginosa clinical isolates expressing OXA enzymes sensitizes them to existing β-lactam antibiotics and dramatically increases the survival of infected G. mellonella larvae that undergo antibiotic treatment.
(A) Absence of DsbA1 sensitizes the P. aeruginosa PA43417 clinical isolate expressing OXA-198 to the first-line antibiotic piperacillin. (B) Absence of DsbA1 sensitizes the P. aeruginosa PAe191 clinical isolate expressing OXA-19 to aztreonam and results in reduction of the ceftazidime MIC value by over 220 µg/mL. For panels (A) and (B) the graphs show MIC values (μg/ml) from two biological experiments, each conducted as a single technical repeat; red dotted lines indicate the EUCAST clinical breakpoint for each antibiotic. (C) 100% of the G. mellonella larvae infected with P. aeruginosa PAe191 (blue curve) or infected with P. aeruginosa PAe191 and treated with 7.5 µg/mL ceftazidime (red curve) die 18 hr post infection, and only 20% of the larvae infected with P. aeruginosa PAe191 dsbA1 (light blue curve) survive 50 hr post infection. Treatment of larvae infected with P. aeruginosa PAe191 dsbA1 with 7.5 µg/mL ceftazidime (pink curve) results in 83% survival, 50 hr post infection. The graph shows Kaplan-Meier survival curves of infected G. mellonella larvae after different treatment applications; horizontal lines represent the percentage of larvae surviving after application of each treatment at the indicated time point (a total of 30 larvae were used for each curve). Statistical analysis of this data was performed using a Mantel-Cox test; n = 30; p =< 0.0001 (significance) (P. aeruginosa versus P. aeruginosa dsbA1), p > 0.9999 (non-significance) (P. aeruginosa vs P. aeruginosa treated with ceftazidime), p =< 0.0001 (significance) (P. aeruginosa treated with ceftazidime versus P. aeruginosa dsbA1), p =< 0.0001 (significance) (P. aeruginosa dsbA1 versus P. aeruginosa dsbA1 treated with ceftazidime).
Tables
Overview of the β-lactamase enzymes investigated in this study.
Enzymes GES-1, –2 and –11 as well as KPC-2 and –3 belong to the same phylogenetic cluster (GES-42 and KPC-44, respectively, see Supplementary file 1). All other tested enzymes belong to distinct phylogenetic clusters (Supplementary file 1). The ‘Cysteine positions’ column states the positions of cysteine residues after position 30 and hence, does not include amino acids that would be part of the periplasmic signal sequence. All β-lactamase enzymes except L2-1 (shaded in grey; PDB ID: 1O7E) have one disulfide bond. The ‘Mobile’ column refers to the genetic location of the β-lactamase gene; ‘yes’ indicates that the gene of interest is located on a plasmid, while ‘no’ refers to chromosomally encoded enzymes. All tested enzymes have a broad hydrolytic spectrum and are either Extended Spectrum β-Lactamases (ESBLs) or carbapenemases. The ‘Inhibition’ column refers to classical inhibitor susceptibility that is, susceptibility to inhibition by clavulanic acid, tazobactam, or sulbactam.
| Enzyme | Amblerclass | Cysteine positions | Mobile | Spectrum | Inhibition |
|---|---|---|---|---|---|
| L2-1 | A | C82 C136 C233 | no | ESBL | yes |
| GES-1 | A | C63 C233 | yes | ESBL | yes |
| GES-2 | A | C63 C233 | yes | ESBL | yes |
| GES-11 | A | C63 C233 | yes | Carbapenemase | yes |
| SHV-27 | A | C73 C119 | no | ESBL | yes |
| OXA-4 | D | C43 C63 | yes | ESBL | yes |
| OXA-10 | D | C44 C51 | yes | ESBL | no (Aubert et al., 2001) |
| OXA-198 | D | C116 C119 | yes | Carbapenemase | no (El Garch et al., 2011) |
| FRI-1 | A | C68 C238 | yes | Carbapenemase | no (Dortet et al., 2015) |
| L1-1 | B3 | C239 C267 | no | Carbapenemase | no (Palzkill, 2013) |
| KPC-2 | A | C68 C237 | yes | Carbapenemase | no (Papp-Wallace et al., 2010) |
| KPC-3 | A | C68 C237 | yes | Carbapenemase | no (Papp-Wallace et al., 2010) |
| SME-1 | A | C72 C242 | no | Carbapenemase | yes |
| Reagent type (species) or resource | Designation | Source or reference | Identifiers | Additional information |
|---|---|---|---|---|
| Genetic reagent(Escherichia coli) | DH5α | Hanahan and Glover, 1985 | F– endA1 glnV44 thi-1 recA1 relA1 gyrA96 deoR nupG purB20 φ80dlacZ∆M15 ∆(lacZYA-argF)U169 hsdR17(rK–mK+) λ– | - |
| Genetic reagent(Escherichia coli) | CC118λpir | Herrero et al., 1990 | araD Δ(ara, leu) ΔlacZ74 phoA20 galK thi-1 rspE rpoB argE recA1 λpir | - |
| Genetic reagent(Escherichia coli) | HB101 | Boyer and Roulland-Dussoix, 1969 | supE44 hsdS20 recA13 ara-14 proA2 lacY1 galK2 rpsL20 xyl-5 mtl-1 | - |
| Genetic reagent(Escherichia coli) | MC1000 | Casadaban and Cohen, 1980 | araD139 ∆(ara, leu)7697 ∆lacX74 galU galK strA | - |
| Genetic reagent(Escherichia coli) | MC1000 dsbA | Kadokura et al., 2004 | dsbA::aphA, KanR | - |
| Genetic reagent(Escherichia coli) | MC1000 dsbA attTn7::Ptac-dsbA | This study | dsbA::aphA attTn7::dsbA, KanR | Can be obtained from the Mavridou lab |
| Genetic reagent(Escherichia coli) | MG1655 | Blattner et al., 1997 | K-12 F– λ– ilvG– rfb-50 rph-1 | - |
| Genetic reagent(Escherichia coli) | MG1655 dsbA | This study | dsbA::aphA, KanR | Can be obtained from the Mavridou lab |
| Genetic reagent(Escherichia coli) | MG1655 dsbA attTn7::Ptac-dsbA | This study | dsbA::aphA attTn7::dsbA, KanR | Can be obtained from the Mavridou lab |
| Genetic reagent(Escherichia coli) | MG1655 acrA | This study | acrA | Can be obtained from the Mavridou lab |
| Genetic reagent(Escherichia coli) | MG1655 tolC | This study | tolC | Can be obtained from the Mavridou lab |
| Genetic reagent(Escherichia coli) | MG1655 degP | This study | degP::strAB, StrR | Can be obtained from the Mavridou lab |
| Strain, strain background (Escherichia coli) | BM16 | Dortet et al., 2014 | blaTEM-1bblaKPC-2 | Human clinical strain |
| Strain, strain background (Escherichia coli) | LIL-1 | Dortet et al., 2014 | blaTEM-1blaOXA-9 blaKPC-2 | Human clinical strain |
| Strain, strain background (Escherichia coli) | CNR1790 | Dortet et al., 2018 | blaTEM-15 mcr-1 | Human clinical strain |
| Strain, strain background (Escherichia coli) | CNR20140385 | Dortet et al., 2018 | blaOXA-48mcr-1 | Human clinical strain |
| Strain, strain background (Escherichia coli) | WI2 (ST1288) | Beyrouthy et al., 2017 | blaOXA-48blaKPC-28 mcr-1 | Human clinical strain |
| Strain, strain background (Escherichia coli) | 1073944 (ST117) | Wise et al., 2018 | mcr-1 | Human clinical strain |
| Strain, strain background (Escherichia coli) | 41,489 | Dortet et al., 2018 | mcr-1 | Human clinical strain |
| Strain, strain background (Escherichia coli) | - | Dortet et al., 2018 | mcr-1 | Human clinical strain |
| Strain, strain background (Escherichia coli) | 1256822 (ST48) | Wise et al., 2018 | mcr-1.5 | Human clinical strain |
| Strain, strain background (Escherichia coli) | 27,841 (ST744) | Haenni et al., 2018 | blaCTX-M-55mcr-3.2 | Environmental strain from livestock |
| Strain, strain background (Escherichia coli) | 1144230 (ST641) | Wise et al., 2018 | blaCMY-2mcr-5 | Human clinical strain |
| Strain, strain background (Klebsiella pneumoniae) | ST234 | Nordmann et al., 2012 | blaSHV-27blaKPC-2 | Human clinical strain |
| Strain, strain background (Citrobacter freundii) | BM19 | Dortet et al., 2014 | blaKPC-2 | Human clinical strain |
| Strain, strain background (Enterobacter cloacae) | DUB | Dortet et al., 2015 | blaFRI-1 | Human clinical strain |
| Strain, strain background (Pseudomonas aeruginosa) | PA43417 | El Garch et al., 2011 | blaOXA-198 | Human clinical strain |
| Genetic reagent (Pseudomonas aeruginosa) | PA43417 | This study | dsbA1 blaOXA-198 | Can be obtained from the Mavridou lab |
| Strain, strain background (Pseudomonas aeruginosa) | PAe191 | Mugnier et al., 1998 | blaOXA-19 | Human clinical strain |
| Genetic reagent (Pseudomonas aeruginosa) | PAe191 | This study | dsbA1 blaOXA-19 | Can be obtained from the Mavridou lab |
| Recombinant DNA reagent | pDM1 (plasmid) | Lab stock | GenBank MN128719 | pDM1 vector, p15A ori, Ptac promoter, MCS, TetR |
| Recombinant DNA reagent | pDM1-blaL2-1 (plasmid) | This study | - | blaL2-1 cloned into pDM1, TetR; can be obtained from the Mavridou lab |
| Recombinant DNA reagent | pDM1-blaGES-1 (plasmid) | This study | - | blaGES-1 cloned into pDM1, TetR; can be obtained from the Mavridou lab |
| Recombinant DNA reagent | pDM1-blaGES-2 (plasmid) | This study | - | blaGES-2 cloned into pDM1, TetR; can be obtained from the Mavridou lab |
| Recombinant DNA reagent | pDM1-blaGES-11 (plasmid) | This study | - | blaGES-11 cloned into pDM1, TetR; can be obtained from the Mavridou lab |
| Recombinant DNA reagent | pDM1-blaSHV-27 (plasmid) | This study | - | blaSHV-27 cloned into pDM1, TetR; can be obtained from the Mavridou lab |
| Recombinant DNA reagent | pDM1-blaOXA-4 (plasmid) | This study | - | blaOXA-4 cloned into pDM1, TetR; can be obtained from the Mavridou lab |
| Recombinant DNA reagent | pDM1-blaOXA-10 (plasmid) | This study | - | blaOXA-10 cloned into pDM1, TetR; can be obtained from the Mavridou lab |
| Recombinant DNA reagent | pDM1-blaOXA-198 (plasmid) | This study | - | blaOXA-198 cloned into pDM1, TetR; can be obtained from the Mavridou lab |
| Recombinant DNA reagent | pDM1-blaFRI-1 (plasmid) | This study | - | blaFRI-1 cloned into pDM1, TetR; can be obtained from the Mavridou lab |
| Recombinant DNA reagent | pDM1-blaL1-1 (plasmid) | This study | - | blaL1-1 cloned into pDM1, TetR; can be obtained from the Mavridou lab |
| Recombinant DNA reagent | pDM1-blaKPC-2 (plasmid) | This study | - | blaKPC-2 cloned into pDM1, TetR; can be obtained from the Mavridou lab |
| Recombinant DNA reagent | pDM1-blaKPC-3 (plasmid) | This study | - | blaKPC-3 cloned into pDM1, TetR; can be obtained from the Mavridou lab |
| Recombinant DNA reagent | pDM1-blaSME-1 (plasmid) | This study | - | blaSME-1 cloned into pDM1, TetR; can be obtained from the Mavridou lab |
| Recombinant DNA reagent | pDM1-mcr-1 (plasmid) | This study | - | mcr-1 cloned into pDM1, TetR; can be obtained from the Mavridou lab |
| Recombinant DNA reagent | pDM1-mcr-3 (plasmid) | This study | - | mcr-3 cloned into pDM1, TetR; can be obtained from the Mavridou lab |
| Recombinant DNA reagent | pDM1-mcr-4 (plasmid) | This study | - | mcr-4 cloned into pDM1, TetR; can be obtained from the Mavridou lab |
| Recombinant DNA reagent | pDM1-mcr-5 (plasmid) | This study | - | mcr-5 cloned into pDM1, TetR; can be obtained from the Mavridou lab |
| Recombinant DNA reagent | pDM1-mcr-8 (plasmid) | This study | - | mcr-8 cloned into pDM1, TetR; can be obtained from the Mavridou lab |
| Recombinant DNA reagent | pDM1-blaL2-1-StrepII (plasmid) | This study | - | blaL2-1 encoding L2-1 with a C-terminal StrepII tag cloned into pDM1, TetR; can be obtained from the Mavridou lab |
| Recombinant DNA reagent | pDM1-blaGES-1-StrepII (plasmid) | This study | - | blaGES-1 encoding GES-1 with a C-terminal StrepII tag cloned into pDM1, TetR; can be obtained from the Mavridou lab |
| Recombinant DNA reagent | pDM1-StrepII-blaOXA-4 (plasmid) | This study | - | blaOXA-4 encoding OXA-4 with an N-terminal StrepII tag cloned into pDM1, TetR; can be obtained from the Mavridou lab |
| Recombinant DNA reagent | pDM1-blaOXA-10-StrepII (plasmid) | This study | - | blaOXA-10 encoding OXA-10 with a C-terminal StrepII tag cloned into pDM1, TetR; can be obtained from the Mavridou lab |
| Recombinant DNA reagent | pDM1-blaOXA-198-StrepII (plasmid) | This study | - | blaOXA-198 encoding OXA-198 with a C-terminal StrepII tag cloned into pDM1, TetR; can be obtained from the Mavridou lab |
| Recombinant DNA reagent | pDM1-blaFRI-1-StrepII (plasmid) | This study | - | blaFRI-1 encoding FRI-1 with a C-terminal StrepII tag cloned into pDM1, TetR; can be obtained from the Mavridou lab |
| Recombinant DNA reagent | pDM1-blaL1-1-StrepII (plasmid) | This study | - | blaL1-1 encoding L1-1 with a C-terminal StrepII tag cloned into pDM1, TetR; can be obtained from the Mavridou lab |
| Recombinant DNA reagent | pDM1-blaKPC-3-StrepII (plasmid) | This study | - | blaKPC-3 encoding KPC-3 with a C-terminal StrepII tag cloned into pDM1, TetR; can be obtained from the Mavridou lab |
| Recombinant DNA reagent | pDM1-mcr-1-StrepII (plasmid) | This study | - | blaMCR-1 encoding MCR-1 with a C-terminal StrepII tag cloned into pDM1, TetR; can be obtained from the Mavridou lab |
| Recombinant DNA reagent | pDM1-mcr-3-StrepII (plasmid) | This study | - | blaMCR-3 encoding MCR-3 with a C-terminal StrepII tag cloned into pDM1, TetR; can be obtained from the Mavridou lab |
| Recombinant DNA reagent | pDM1-mcr-4-StrepII (plasmid) | This study | - | blaMCR-4 encoding MCR-4 with a C-terminal StrepII tag cloned into pDM1, TetR; can be obtained from the Mavridou lab |
| Recombinant DNA reagent | pDM1-mcr-5-StrepII (plasmid) | This study | - | blaMCR-5 encoding MCR-5 with a C-terminal StrepII tag cloned into pDM1, TetR; can be obtained from the Mavridou lab |
| Recombinant DNA reagent | pDM1-mcr-8-StrepII (plasmid) | This study | - | blaMCR-8 encoding MCR-8 with a C-terminal StrepII tag cloned into pDM1, TetR; can be obtained from the Mavridou lab |
| Recombinant DNA reagent | pGRG25 (plasmid) | McKenzie and Craig, 2006 | - | Encodes a Tn7 transposon and tnsABCD under the control of ParaB, thermosensitive pSC101 ori, AmpR |
| Recombinant DNA reagent | pGRG25-Ptac::dsbA (plasmid) | This study | - | Ptac::dsbA fragment cloned within the Tn7 of pGRG25; when inserted into the chromosome and the plasmid cured, the strain expresses DsbA upon IPTG induction, AmpR; can be obtained from the Mavridou lab |
| Recombinant DNA reagent | pSLTS (plasmid) | Kim et al., 2014 | - | Thermosensitive pSC101ori, ParaB for λ-Red, PtetR for I-SceI, AmpR |
| Recombinant DNA reagent | pUltraGFP-GM (plasmid) | Mavridou et al., 2016 | - | Constitutive sfGFP expression from a strong Biofab promoter, p15A ori, (template for the accC cassette), GentR |
| Recombinant DNA reagent | pKD4 (plasmid) | Datsenko and Wanner, 2000 | - | Conditional oriRγ ori, (template for the aphA cassette), AmpR |
| Recombinant DNA reagent | pCB112 (plasmid) | Paradis-Bleau et al., 2014 | - | Inducible lacZ expression under the control of the Plac promoter, pBR322 ori, CamR |
| Recombinant DNA reagent | pKNG101 (plasmid) | Kaniga et al., 1991 | - | Gene replacement suicide vector, oriR6K, oriTRK2, sacB, (template for the strAB cassette), StrR |
| Recombinant DNA reagent | pKNG101-dsbA1 (plasmid) | This study | - | PCR fragment containing the regions upstream and downstream P. aeruginosa dsbA1 cloned in pKNG101; when inserted into the chromosome the strain is a merodiploid for dsbA1 mutant, StrR; can be obtained from the Mavridou lab |
| Recombinant DNA reagent | pRK600 (plasmid) | Kessler et al., 1992 | - | Helper plasmid, ColE1 ori, mobRK2, traRK2, CamR |
| Recombinant DNA reagent | pMA-T mcr-3 (plasmid) | This study | - | GeneArt cloning vector containing mcr-3, ColE1 ori, (template for mcr-3), AmpR; can be obtained from the Mavridou lab |
| Recombinant DNA reagent | pMK-T mcr-8 (plasmid) | This study | - | GeneArt cloning vector containing mcr-8, ColE1 ori, (template for mcr-8), KanR; can be obtained from the Mavridou lab |
| Chemical compound, drug | Ampicillin | Melford | A40040-10.0 | - |
| Chemical compound, drug | Piperacillin | Melford | P55100-1.0 | - |
| Chemical compound, drug | Cefuroxime | Melford | C56300-1.0 | - |
| Chemical compound, drug | Ceftazidime | Melford | C59200-5.0 | - |
| Chemical compound, drug | Imipenem | Cambridge Bioscience | CAY16039-100 mg | - |
| Chemical compound, drug | Aztreonam | Cambridge Bioscience | CAY19784-100 mg | - |
| Chemical compound, drug | Kanamycin | Gibco | 11815032 | - |
| Chemical compound, drug | Gentamicin | VWR | A1492.0025 | - |
| Chemical compound, drug | Streptomycin | ACROS Organics | AC612240500 | - |
| Chemical compound, drug | Tetracycline | Duchefa Biochemie | T0150.0025 | - |
| Chemical compound, drug | Colistin sulphate | Sigma | C4461-1G | - |
| Chemical compound, drug | Tazobactam | Sigma | T2820-10MG | - |
| Chemical compound, drug | Isopropyl β-D-1-thiogalactopyranoside (IPTG) | Melford | I56000-25.0 | - |
| Chemical compound, drug | KOD Hotstart DNA Polymerase | Sigma | 71086–3 | - |
| Chemical compound, drug | Nitrocefin | Abcam | ab145625-25mg | - |
| Chemical compound, drug | 1-N-phenylnaphthylamine (NPN) | Acros Organics | 147160250 | - |
| Chemical compound, drug | 4-acetamido-4ˊ-maleimidyl-stilbene-2,2ˊ-disulfonic acid (AMS) | ThermoFisher Scientific | A485 | - |
| Chemical compound, drug | 4,5-dichloro-2-(2-chlorobenzyl)pyridazin-3-one | Enamine | EN300-173996 | - |
| Commercial assay or kit | BugBuster Mastermix | Sigma | 71456–3 | - |
| Commercial assay or kit | Novex ECL HRP chemiluminescent substrate reagent kit | ThermoFisher Scientific | WP20005 | - |
| Commercial assay or kit | SigmaFast BCIP/NBT tablets | Sigma | B5655-25TAB | - |
| Commercial assay or kit | Immobilon Crescendo chemiluminescent reagent | Sigma | WBLUR0100 | - |
| Commercial assay or kit | ETEST - Amoxicillin | Biomerieux | 412,242 | - |
| Commercial assay or kit | ETEST - Cefuroxime | Biomerieux | 412,304 | - |
| Commercial assay or kit | ETEST - Ceftazidime | Biomerieux | 412,292 | - |
| Commercial assay or kit | ETEST - Imipenem | Biomerieux | 412,373 | - |
| Commercial assay or kit | ETEST - Aztreonam | Biomerieux | 412,258 | - |
| Commercial assay or kit | ETEST - Gentamicin | Biomerieux | 412,367 | - |
| Commercial assay or kit | ETEST - Erythromycin | Biomerieux | 412,333 | - |
| Commercial assay or kit | ETEST - Chloramphenicol | Biomerieux | 412,308 | - |
| Commercial assay or kit | ETEST - Nalidixic acid | Biomerieux | 516,540 | - |
| Commercial assay or kit | ETEST - Ciprofloxacin | Biomerieux | 412,310 | - |
| Commercial assay or kit | ETEST - Nitrofurantoin | Biomerieux | 530,440 | - |
| Commercial assay or kit | ETEST - Trimethoprim | Biomerieux | 412,482 | - |
| Antibody | Strep-Tactin-HRP conjugate (mouse monoclonal) | Iba Lifesciences | NC9523094 | (1:3,000) in 3 w/v % BSA/TBS-T |
| Antibody | Strep-Tactin-AP conjugate (mouse monoclonal) | Iba Lifesciences | NC0485490 | (1:3,000) in 3 w/v % BSA/TBS-T |
| Antibody | anti-DsbA(rabbit polyclonal) | Beckwith lab | - | (1:1,000) in 5 w/v % skimmed milk/TBS-T |
| Antibody | anti-AcrA(rabbit polyclonal) | Koronakis lab | - | (1:10,000) in 5 w/v % skimmed milk/TBS-T |
| Antibody | anti-TolC(rabbit polyclonal) | Koronakis lab | - | (1:5,000) in 5 w/v % skimmed milk/TBS-T |
| Antibody | anti-HtrA1 (DegP)(rabbit polyclonal) | Abcam | ab231195 | (1:1,000) in 5 w/v % skimmed milk/TBS-T |
| Antibody | anti-DnaK 8E2/2(mouse monoclonal) | Enzo Life Sciences | ADI-SPA-880-D | (1:10,000) in 5% w/v skimmed milk/TBS-T |
| Antibody | anti-rabbit IgG-AP conjugate (goat polyclonal) | Sigma | A3687-.25ML | (1:6,000) in 5% w/v skimmed milk/TBS-T |
| Antibody | anti-rabbit IgG-HRP conjugate (goat polyclonal) | Sigma | A0545-1ML | (1:6,000) in 5% w/v skimmed milk/TBS-T |
| Antibody | anti-mouse IgG-AP conjugate (goat polyclonal) | Sigma | A3688-.25ML | (1:6,000) in 5% w/v skimmed milk/TBS-T |
| Antibody | anti-mouse IgG-HRP conjugate (goat polyclonal) | Sigma | A4416-.5ML | (1:6,000) in 5% w/v skimmed milk/TBS-T |
| Software, algorithm | FlowJo | Tree Star | - | version 10.0.6 |
| Software, algorithm | Adobe Photoshop CS4 | Adobe | - | extended version 11.0 |
| Software, algorithm | Prism | GraphPad | - | version 8.0.2 |
| Software, algorithm | blastp | Altschul et al., 1990 | - | version 2.2.28+ |
| Software, algorithm | USEARCH | Edgar, 2010 | - | version 7.0 |
| Software, algorithm | MUSCLE | Edgar, 2004 | - | - |
| Software, algorithm | FastTree | Price et al., 2010 | - | version 2.1.7 |
| Software, algorithm | HMMER | Finn et al., 2015 | - | version 3.1b2 |
Additional files
-
Supplementary file 1
Analysis of the cysteine content and phylogeny of all identified β-lactamases.
6,649 unique β-lactamase protein sequences were clustered with a 90% identity threshold and the centroid of each cluster was used as a phylogenetic cluster identified for each sequence (“Phylogenetic cluster” column). All sequences were searched for the presence of cysteine residues (“Total number of cysteines” and “Positions of all cysteines” columns). Proteins with two or more cysteines after the first 30 amino acids of their primary sequence (cells shaded in grey in the “Number of cysteines after position 30” column) are potential substrates of the DSB system for organisms where oxidative protein folding is carried out by DsbA and provided that translocation of the β-lactamase outside the cytoplasm is performed by the Sec system. The first 30 amino acids of each sequence were excluded to avoid considering cysteines that are part of the signal sequence mediating the translocation of these enzymes outside the cytoplasm. Cells shaded in grey in the “Reported in pathogens” column mark β-lactamases that are found in pathogens or organisms capable of causing opportunistic infections. The Ambler class of each enzyme is indicated in the “Ambler class column” and each class (A, B1, B2, B3, C and D) is highlighted with a different color.
- https://cdn.elifesciences.org/articles/57974/elife-57974-supp1-v2.xlsx
-
Supplementary file 2
MIC data used to generate Figure 1B, Figure 1—figure supplement 2, and Figure 5B.
Cells that are shaded in grey represent strain-antibiotic combinations that were not tested. The aminoglycoside antibiotic gentamicin serves as a control for all strains. For the “Supplementary File 2a” tab, values are representative of three biological experiments each conducted as a single technical repeat, and for the “Supplementary File 2b” tab, values are representative of two biological experiments each conducted as a single technical repeat.
- https://cdn.elifesciences.org/articles/57974/elife-57974-supp2-v2.xlsx
-
Supplementary file 3
Supplementary tables 1-6 and relevant citations.
- https://cdn.elifesciences.org/articles/57974/elife-57974-supp3-v2.docx
-
Transparent reporting form
- https://cdn.elifesciences.org/articles/57974/elife-57974-transrepform1-v2.docx





