Dermomyotome-derived endothelial cells migrate to the dorsal aorta to support hematopoietic stem cell emergence
Figures

Cell-type-specific endothelial cell markers highlight cellular diversity within the vasculature.
(A) Uniform manifold approximation projection (UMAP) plots of scRNA-seq data of total endothelial lineage cells collected from TgBAC(etv2:Kaede)ci6, Tg(fli1:DsRed)um13; Tg(tp1:GFP)um14, and Tg(drl:H2B-dendra) embryos at 22–24 hpf. Clusters were named according to their gene expression: Erythroid, Lymphoid, General Endothelium (GE), Hemogenic Endothelium (HE), Pre-HSCs, Brain Vascular Endothelial Cells (BVECs-I and BVECs-II), Kidney Vascular Endothelial cells (KVECs), Endocardial Endothelial Cells (EECs), and somite-derived endothelial cells (SDECs). Color-coded marker gene expression levels are shown on corresponding clusters. A pink circle highlights the SDEC cluster. (B) Expression heatmap of 22–24 hpf single-cell transcriptome shows the top predicted differentially expressed marker genes across the different clusters. A red box highlights the SDEC cluster. (C’,C”) A list of somite-annotated genes was curated from the AmiGo annotation database and compared with the SDEC transcriptome. 32 genes were commonly expressed. Interestingly, several of these 32 genes were enriched within the SDEC cluster (C”; white boxed genes, D; enlarged circles).
-
Figure 1—source data 1
Transcriptomes of all endothelial cell clusters, myeloid, and erythroid cells.
The transcriptomes were extracted and read from cells purified collected from TgBAC(etv2:Kaede)ci6, Tg(fli1:DsRed)um13; Tg(tp1:GFP)um14, and Tg(drl:H2B-dendra) embryos at 22–24 hpf.
- https://cdn.elifesciences.org/articles/58300/elife-58300-fig1-data1-v1.xlsx
-
Figure 1—source data 2
Comparison between Genes expressed in EC clusters (e.g. SDECs) and gene annotation of the same anatomical structure (e.g., somite) based on annotation from AmiGO (Consortium, 2019).
Expression of overlapping genes was compared in the same cluster (e.g. SDECs) between 15 ss and 22 hpf and divided into DE genes that are upregulated (Up) on downregulated (Down). Each DE group was then annotated using AmiGO (Consortium, 2019).
- https://cdn.elifesciences.org/articles/58300/elife-58300-fig1-data2-v1.xlsx
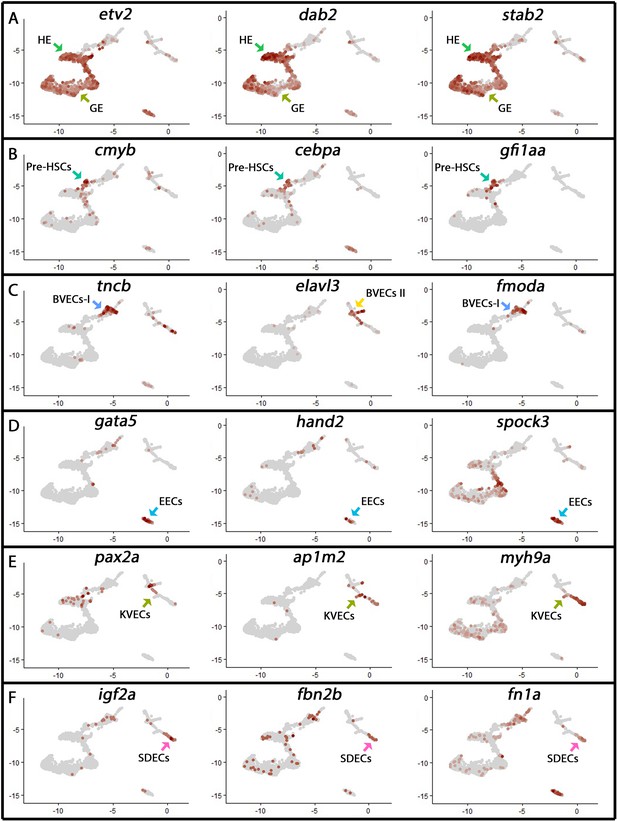
Cluster identity was assigned based on known marker genes.
(A–F) Following unsupervised clustering of single-cell transcriptomes, cluster identity was given based on known marker genes within established tissue lineages. Selected marker genes and the eight distinct endothelial cell clusters are shown (arrows, color-coded by their original cluster color in Figure 1A).
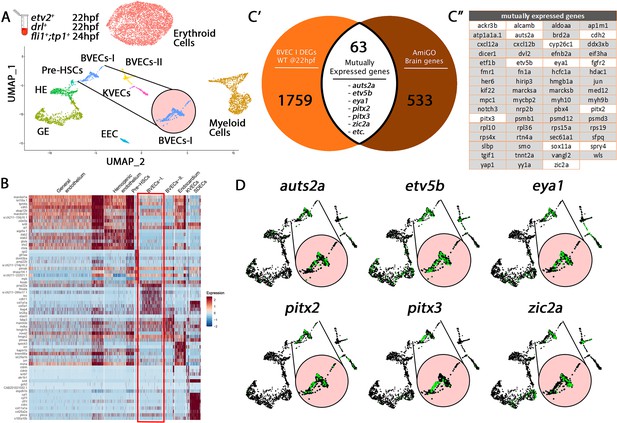
Comparison of BVECs-I cluster genes to brain annotated genes validates cluster origin.
(A) Uniform manifold approximation projection (UMAP) plots of scRNA-seq data of total endothelial lineage cells collected from TgBAC(etv2:Kaede)ci6, Tg(fli1:DsRed)um13; Tg(tp1:GFP)um14, and Tg(drl:H2B-dendra) embryos at 22–24 hpf. Clusters were named according to their gene expression: Erythroid, Lymphoid, General Endothelium (GE), Hemogenic Endothelium (HE), Pre-HSCs, Brain Vascular Endothelial Cells (BVECs-I and BVECs-II), Kidney Vascular Endothelial cells (KVECs), Endocardial Endothelial Cells (EECs), and somite-derived endothelial cells (SDECs). Color-coded marker gene expression levels are shown on corresponding clusters. A pink circle highlights the BVECs-I cluster. (B) Expression heatmap of 22–24 hpf single-cell transcriptome showing the top predicted differentially expressed marker genes across the different clusters. A red box highlights the BVECs-I cluster. (C’,C”) A list of brain-annotated genes was curated from the AmiGo annotation database and compared with the BVECs-I transcriptome. 63 genes were commonly expressed. Interestingly, several of these 63 genes were enriched within the BVECs-I cluster (C”; white boxed genes, D; enlarged circles).
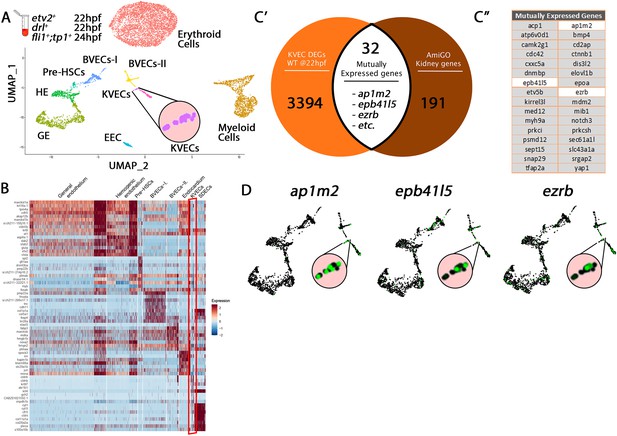
Comparison of KVEC Cluster genes to kidney annotated genes validates cluster origin.
(A) Uniform manifold approximation projection (UMAP) plots of scRNA-seq data of total endothelial lineage cells collected from TgBAC(etv2:Kaede)ci6, Tg(fli1:DsRed)um13; Tg(tp1:GFP)um14, and Tg(drl:H2B-dendra) embryos at 22–24 hpf. Clusters were named according to their gene expression: Erythroid, Lymphoid, General Endothelium (GE), Hemogenic Endothelium (HE), Pre-HSCs, Brain Vascular Endothelial Cells (BVECs-I and BVECs-II), Kidney Vascular Endothelial cells (KVECs), Endocardial Endothelial Cells (EECs), and somite-derived endothelial cells (SDECs). Color-coded marker gene expression levels are shown on corresponding clusters. A pink circle highlights the KVECs cluster. (B) Expression heatmap of 22–24 hpf single-cell transcriptome showing the top predicted differentially expressed marker genes across the different clusters. A red box highlights the KVECs cluster. (C’,C” ) A list of kidney-annotated genes was curated from the AmiGo annotation database and compared with the KVECs transcriptome. 32 genes were commonly expressed. Interestingly, several of these 32 genes were enriched within the KVECs cluster (C”; white boxed genes, D; enlarged circles).
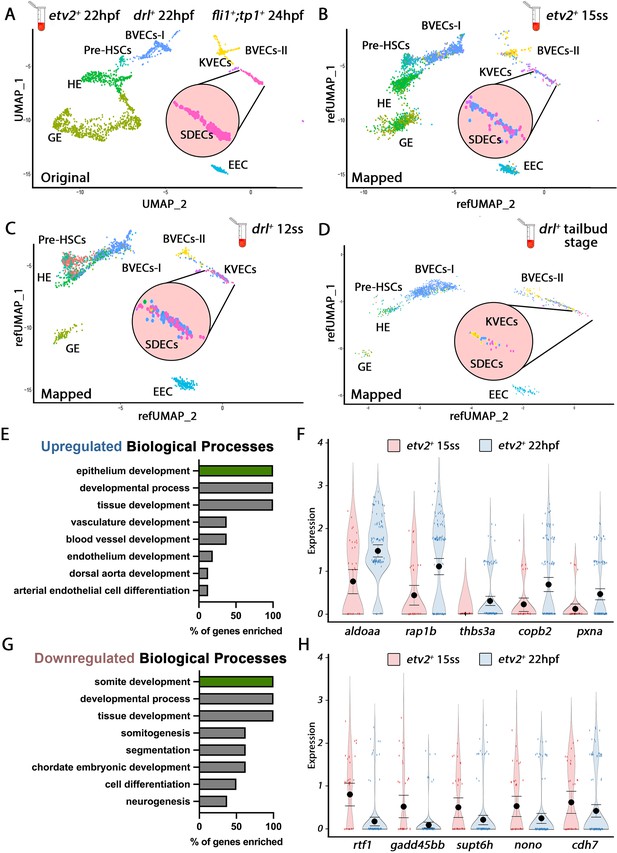
Cellular diversity within the vasculature can be traced back to the tailbud stage.
(A) Uniform manifold approximation projection (UMAP) plots of scRNA-seq data of total endothelial lineage cells collected from TgBAC(etv2:Kaede)ci6, Tg(fli1:DsRed)um13; Tg(tp1:GFP)um14, and Tg(drl:H2B-dendra) embryos at 22–24 hpf. Clusters were named according to their gene expression: General Endothelium (GE), Hemogenic Endothelium (HE), Pre-HSCs, Brain Vascular Endothelial Cells (BVECs-I and BVECs-II), Kidney Vascular Endothelial cells (KVECs), Endocardial Endothelial Cells (EEC), and somite-derived endothelial cells (SDECs). Color-coded marker gene expression levels are shown on corresponding clusters. A pink circle highlights the SDEC cluster. (B–D) Referenced uniform manifold approximation projection (RefUMAP) plots of scRNA-seq data of total endothelial lineage cells collected from etv2:Kaede+ embryos at 15 ss (B) and drl:H2B-dendra+ embryos at 12 ss (C) and tailbud stage (D). By cross-referencing the transcriptomes of EC subsets at each developmental stage to the 22–24 hpf ECs, we identified EC clusters with distinct transcriptomes as early as the tailbud stage. (E–H) Comparison of expression patterns of EC populations from early TgBAC(etv2:Kaede)ci6 15 ss, and later 22 hpf etv2:Kaede+ in the 32 overlapping genes between the SDEC transcriptome data and the AmiGo somite annotated genes. (E,F) Representative genes that were upregulated in the etv2:Kaede+ 22 hpf samples compared to the 15 ss sample (F) and their suggested role in EC differentiation, according to GO biological processes (E). (G,H) Representative genes that were downregulated in the etv2:Kaede+ 22 hpf samples compared to the 15 ss sample (H) and their suggested role in somitogenesis, according to GO biological processes (G). The expression and downregulation of somitic genes within etv2+ ECs between 15 ss and 22 hpf highlight their somitic origin and loss of myogenic cell fate.
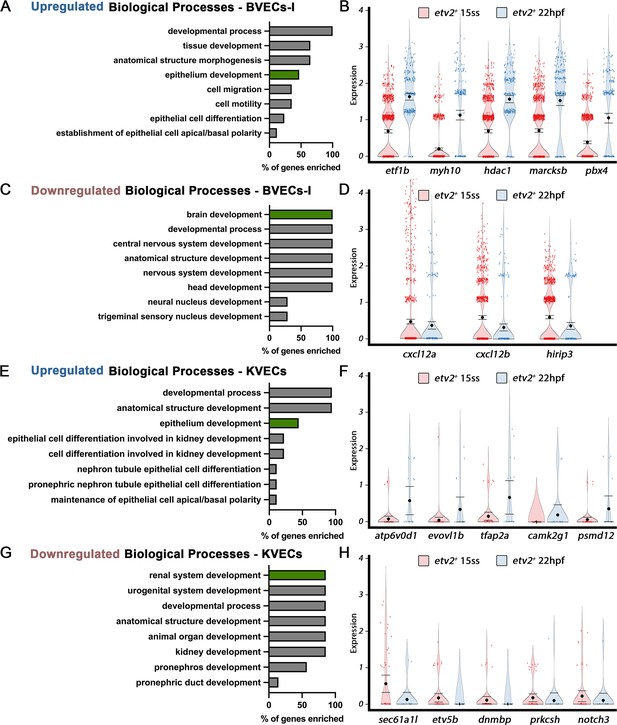
Differentially expressed genes between early and late ECs in BVECs-I or KVECs clusters highlight an early commitment to EC fate.
(A–D) Comparison of expression patterns of EC populations from early TgBAC(etv2:Kaede)ci6 15 ss and later 22 hpf etv2:Kaede+ ECs in the 63 overlapping genes between the BVECs-I transcriptome data and the AmiGo brain annotated genes. (A,B) Representative genes upregulated in the mixed vasculature 22 hpf samples compared to the 15 ss sample (B) and their suggested role in epithelium development, according to GO biological processes (A). (C,D) Representative genes downregulated in the mixed vasculature 22 hpf samples compared to the 15 ss sample (D) and their suggested role in brain development and neurogenesis, according to GO biological processes (C). (E–H) Comparison of expression patterns of EC populations from early TgBAC(etv2:Kaede)ci6 15 ss and later 22 hpf etv2:Kaede+ ECs in the 32 overlapping genes between the KVECs transcriptome data and the AmiGo kidney annotated genes. (E,F) Representative genes upregulated in the mixed vasculature 22 hpf samples compared to the 15 ss sample (F) and their suggested role in epithelium development, according to GO biological processes (E). (G,H) Representative genes downregulated in the mixed vasculature 22 hpf samples compared to the 15 ss sample (H) and their suggested role in renal system development, according to GO biological processes (G).

Rare SDECs emerge from trunk somites and migrate to the dorsal aorta.
(A–D) Tg(actb2:nls-Eos); Tg(fli1:eGFP)y1 embryos were collected at developmental stages ranging from 4 to 18 ss. (A) Newly developed posterior somite pairs were selected by setting a region of interest and photoconverted by UV light. (A’) A sagittal section through a pair of converted somite showing somite-specific conversion and lack of converted LPM-derived ECs. (B) At 32 hpf, embryos were laterally staged, and images of the dorsal aorta taken. SDECS were quantified in the dorsal aorta by examining individual z-stacks and visualizing colocalization of fli1+; actb2:nlsEosRFP converted cells in Tg(actb2:nls-Eos); Tg(fli1:eGFP)y1 embryos (Ci-Cv). In fli1:eGFP - embryos, we identified SDECs by observing actb2:nlsEosRFP cells on the floor of the DA based on the brightfield channel. (D) We observed that trunk somites (numbers 5–18), located above the yolk tube extension, generated the most SDECs. Each somite pair contributed between 0–6 SDECs to the DA. s, somites; DA, dorsal aorta. In each converted somite pair, , with each point representing the SDEC count from one embryo. The median for each somite pair is indicated as a column, and the standard error of the mean (SEM) is indicated as an error bar.
-
Figure 3—source data 1
A table summarizing all converted somite pairs and SDECs found in Tg(actb2:nls-Eos); Tg(fli1:eGFP)y1 embryos that were included in the final SDECs quantification assay.
Embryo’ somites were converted at developmental stages ranging from 4 to 18 somite stage (Column C) and imaged at 32–36 dpf. The imaging date (Column A), sample number within a cohort (Column B), and the number of observed SDECs (Column D) used for the quantification were documented, and the quantification and presented graph were done in Prism 9 (GraphPad).
- https://cdn.elifesciences.org/articles/58300/elife-58300-fig3-data1-v1.xlsx
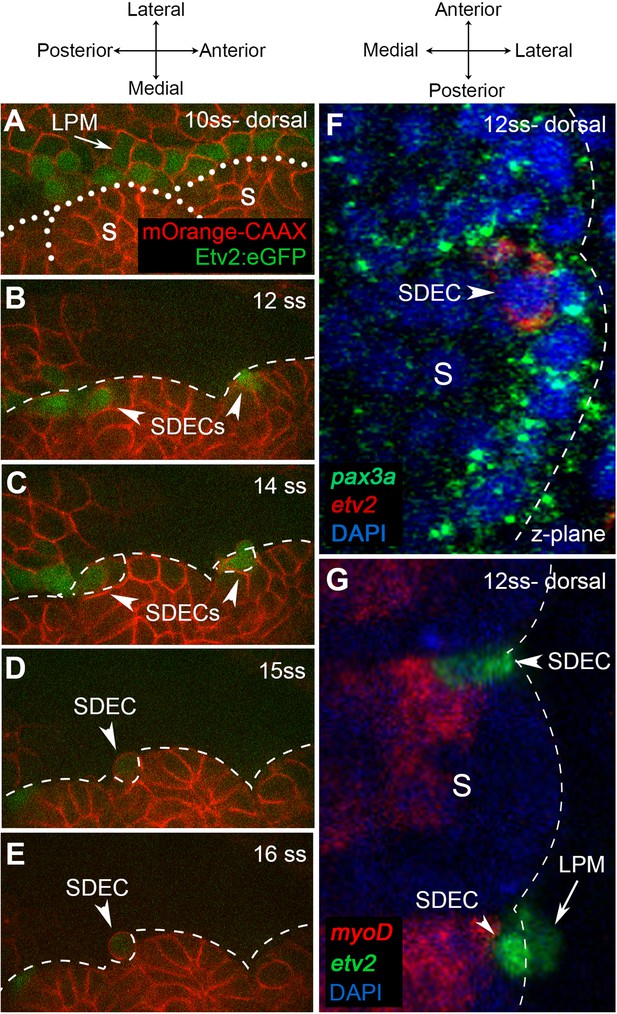
Endothelial cells emerge from the dermomyotome at 12 ss.
(A–E) Time-lapse imaging from a dorsal view of Tg(etv2.1:EGFP)zf372 embryos injected with mOrange:CAAX mRNA and imaged between 10 ss and 15 ss. (A) The expression of Etv2:GFP+ cells is visible along the LPM region (arrow) at 10 ss. At this stage, no Etv2:GFP+ cells are visible in the somites. (B) Starting at 12 ss, the first Etv2:GFP+ SDECs are detected in the lateral lip of the dermomyotome (arrowheads). Simultaneously, the LPM Etv2:GFP+ cells start migrating to the midline. (C) Soon after emergence, SDECs change shape and become rounder (arrowheads). (D–E) Etv2:GFP+ SDECs bud off from the somite as individual cells (arrowhead). (F) Dorsal view of a 12 ss embryo that was submitted to double fluorescent in situ hybridization for muscle progenitor maker pax3a (green) and endothelial marker etv2 (red). pax3a expression reveals the dermomyotome compartment that contains muscle progenitor cells. An etv2+ SDEC (red and arrowhead) is found in the dermomyotome, co-expressing pax3a (green), showing colocalization of an endothelial and muscle progenitor cell marker. We observed 1–2 etv2-positive cells per somite in each of the embryos examined (n=6). (G) Somitic etv2+ SDECs (green) do not co-express the muscle differentiation marker myoD (red), suggesting that etv2 expression is restricted to the muscle progenitor region of the somite. Dashed white lines delimitate somite from the LPM (arrow). We observed 1–2 etv2-positive cells per somite in each of the embryos examined (n=6). s, somites; LPM, lateral plate mesoderm; SDECs, somite-derived endothelial cells.
Time-lapse imaging of Tg(etv2.1:eGFP)zf372; Tg(phldb1:mCherry) embryo between 12 and 16 ss.
Lateral view of a transgenic embryo. LPM cells migrate from the left side under the somites. SDECs (Green, arrows) arise from the first and fifth somites (Red) and exit the somites to follow their LPM-derived counterparts to the midline. S, somites; LPM, lateral plate mesoderm; SDECs, somite-derived endothelial cells.
Time-lapse imaging of Tg(etv2.1:eGFP)zf372; Tg(phldb1:mCherry) embryo between 12 and 16 ss.
Lateral view of a transgenic embryo. SDECs (Green, arrows) arise from the somites (S, Magenta). By this time, most of the LPM cells (arrowheads) have ingressed underneath the somites. S, somites; SDECs, somite-derived endothelial cells; LPM, lateral plate mesoderm.

notch is required for the maintenance of a bipotent skeletal muscle progenitor population in the somite.
(A–F) Dorsal view of 12 ss control (A–C) and meox1 morphant embryos (D–F). Embryos were submitted to double fluorescent in situ hybridization for meox1 (green) and etv2 (red). In control and morphant embryos, meox1; etv2 double-positive cells are detected within the somite compartment (arrowheads). (C,F) Overlay of meox1 (green), etv2 (red), and DAPI (blue). (D–F) Knockdown of meox1 results in ectopic formation of double-positive cells within the somite (arrowheads). We observed 3–4 etv2 positive cells per somite in the meox1 morphants compared to 1–2 etv2-positive cells per somite in the siblings (n=3). (G–H) Time-lapse imaging of a 22 hpf Tg(etv2.1:eGFP)zf372; Tg(phldb1:mCherry) embryo, injected with meox1 morpholino and mOrange2:CAAX mRNA to delineate cell boundaries. Knockdown of meox1 results in an extension of the period that the dermomyotome can generate Etv2:GFP+ cells (arrowheads). (I–K) Cross section of 12 ss Tg(etv2.1:eGFP)zf372 embryo. In absence of meox1 (J), ectopic Etv2:GFP+ cells are visible in epithelialized layer of the somites, compared to controls (I). In embryos coinjected with mib and meox1 morpholinos, the number of Etv2:GFP+ cells within the somite compartment (dotted line) is substantially increased (arrowheads) (K), suggesting that Notch signaling is dispensable for SDEC specification. (L–N) Lateral view of 12 ss embryos analyzed by FISH for meox1 (green), etv2 (red), and DAPI (blue). In notch3+/- heterozygote controls (L) and notch3-/- mutant embryos (M), etv2+ SDECs are detected in the somites. (N) notch3-/- mutant embryos co-injected with meox1 morpholino results in ectopic formation of etv2; meox1 double positive cells (arrowheads). We observed 2–4 etv2-positive cells per somite in the notch3 mutants and >6 etv2-positive cells in the notch3 mutants; Mib morphants (n=3). (O) qRT-PCR in 24 hpf notch3-/- mutant embryos and sibling controls. Genetic ablation of notch3 results in decreased expression of muscle progenitor markers pax3a and pax7b; increased expression of muscle differentiation genes, myod and myog, and endothelial markers, etv2, and fli1. Asterisks denote a statistically significant difference (p<0.05, unpaired, two-tailed Student’s t-test; n=3.) (P,Q) notch3-/- mutant embryos show premature expression of MyoHII in 48 hpf embryos (Q) compared to sibling controls (P). (R) Summary cartoon for the role of Notch signaling in the maintenance of bipotent-muscle progenitors (bipotent muscle progenitors in purple and green; muscle cells in green; SDECs in purple). s, somites; LPM, lateral plate mesoderm; SDECs, somite-derived endothelial cells.
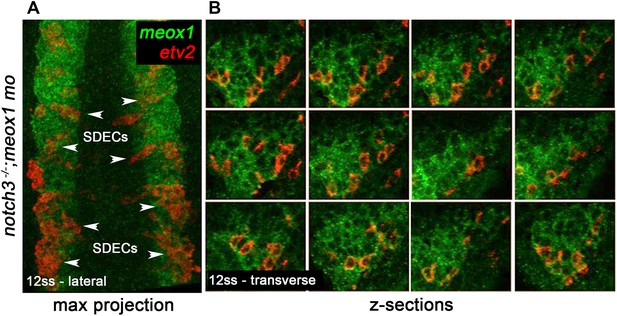
Bipotent muscle progenitor cells contain endothelial potential that can reach the dermomyotome compartment.
(A) Max projection of 12 hpf notch3-/- mutant embryos injected with meox1 morpholino shows broad endothelial potential within the somite compartment by the ectopic formation of double positive meox1 (green) and etv2 (red) SDECs (arrowheads). (B) Z-sections of a representative dermomyotome compartment show the extent of double-positive cells. SDECs, somite-derived endothelial cells.
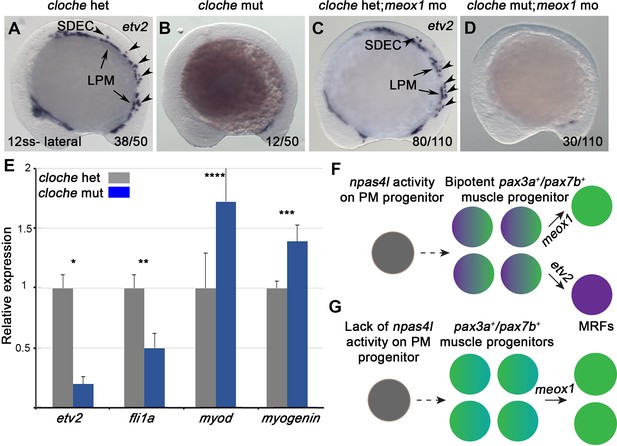
npas4l is required for the specification of SDECs.
(A–D) WISH for etv2 in 12 ss npas4l-/- (cloche) mutant and control embryos. (B) cloche mutant embryos show an absence of etv2 expression along the A-P axis of the embryo, compared to sibling control (A). (D) Similarly, cloche mutant embryos injected with meox1 morpholino show loss of etv2 expression, compared to sibling control (C). (E) qRT-PCR of cloche mutant embryos shows expected loss of endothelial genes (fli1 and etv2) and concomitant increase of muscle differentiation genes (myod and myog), compared to sibling control. All genes analyzed between cloche mutant and cloche het embryos showed a statistically significant difference (p<0.001, unpaired, two-tailed Student’s t-test; n=3.) (F, G) Summary cartoon for the effect of npas4l on endothelial cell competence in PM progenitors (early mesoderm progenitor in grey; bipotent muscle progenitor in purple and green; muscle cells in green; endothelial cells in purple). LPM, lateral plate mesoderm; SDECs, somite-derived endothelial cells.
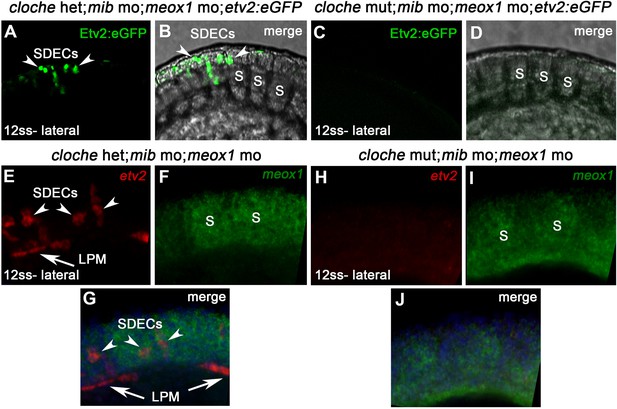
npas4l is required for the specification of SDECs.
(A–D) Tg(etv2.1:eGFP)zf372; cloche mutant and heterozygous embryos were injected with meox1 and mib morpholinos. (C,D) Cloche mutant embryos showed loss of Etv2:eGFP expression in the LPM and somites at 12 ss compared with control embryos (A,B; arrowheads). (E–J) cloche mutant and heterozygous embryos were injected with both meox1 and mib morpholinos. FISH for meox1 (green) and etv2 (red) shows loss of all etv2 and meox1 double-positive cells in cloche mutants (H–J) compared with control embryos (E-G; arrowheads) at 12 ss. S, somites; LPM, lateral plate mesoderm; SDECs, somite-derived endothelial cells.
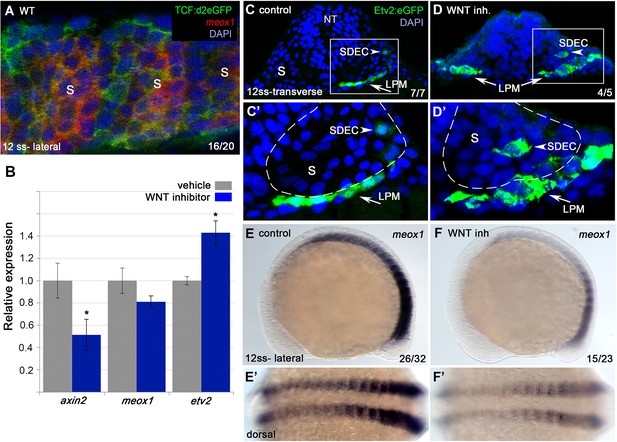
Wnt signaling is required for the regionalization of SDECs.
(A) FISH for meox1 (red) and antibody staining for a destabilized Wnt/TCF reporter line (green) show co-expression of GFP and meox1 within the somite. (B) Inhibition of Wnt signaling using the chemical inhibitor IWP2 from 2 ss to 15 ss results in decreased expression of axin2 and meox1 with a concomitant increase of the expression of etv2 by qRT-PCR. We observed a reduction in meox1 expression, although not statistically significant. All genes analyzed between Wnt inhibitor and control embryos, except meox1, showed a statistically significant difference (p<0.001, unpaired, two-tailed Student’s t-test; n=3.) (C–D’) Cross section of Tg(etv2.1:EGFP)zf372 embryos treated with IWP2 from 2 ss to15 ss. wnt inhibition results in ectopic formation of Etv2:GFP+ cells within the somite (arrowheads). (C’,D’) enlargement of somite compartment (dashed lines). Notice LPM cells migrating under the sclerotome (arrows). (E–F’) IWP2 control and treated embryos. wnt inhibition results in decreased expression of meox1 by WISH, compared to control embryos. (E,F) Lateral view. (E’,F’) Dorsal view. s, somites; LPM, lateral plate mesoderm; SDECs, somite-derived endothelial cells.
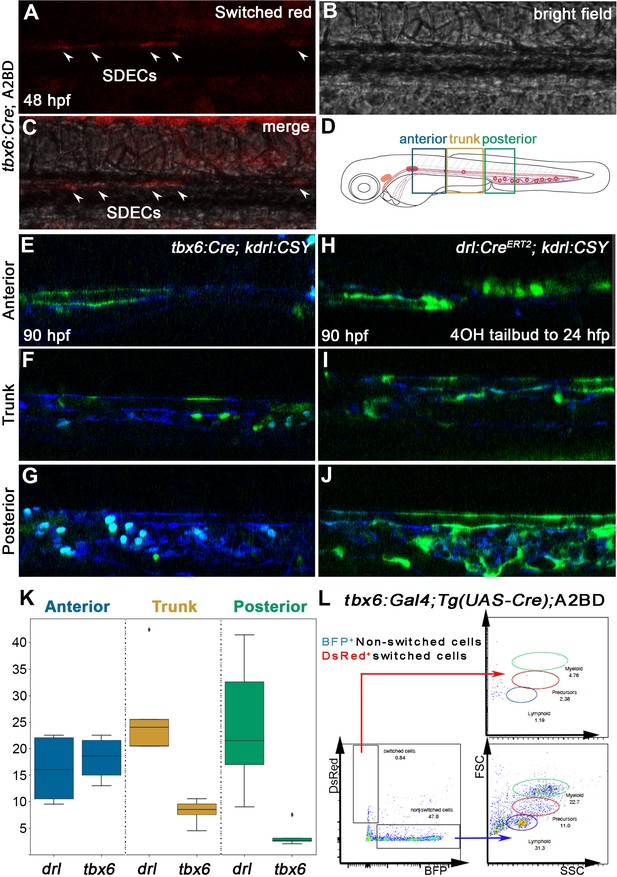
SDECs contribute to the dorsal aorta but do not generate HSPCs.
(A–C) Lineage tracing of SDECs using tbx6:Gal4; Tg(UAS-Cre); A2BD shows dsRed+ cells in the vasculature region at 48 hpf (arrowheads). (E–J) Using a vasculature-specific switch line TgBACkdrl:LOXP-AmCyan-LOXP-ZsYellow (referred to as kdrl:CSY), we observe the contribution of SDECs or LPM-derived endothelial cells to the vasculature. (E–G) For SDEC labeling, a PM-specific driver tbx6:Gal4; Tg(UAS-Cre) was used. PM-derived YFP+ SDECs are observed in the vasculature of imaged embryos. (H–J) For LPM-specific EC labeling, a Tg(drl:CreERT2) was used and treated with 10 µm tamoxifen starting at 8 hpf. YFP+ ECs are observed in all regions of the vasculature. (K) Quantification of YFP+ SDECs and ECs from tbx6 or drl switched embryos, respectively. Quantifications were based on independent experiments per transgenic background with n=23 for tbx6 switched embryos and n=9 for drl switch embryos. (L) Analysis of the adult kidney marrow of tbx6:Gal4; Tg(UAS-Cre); A2BD animals shows no contribution to hematopoietic cells from switched DsRed+ SDECs through flow cytometry analysis, whereas the FSC/SSC distribution of the unswitched BFP+ ECs corresponds to all blood lineages (quantifications based from independent experiments with a total of n=21 samples). SDECs, somite-derived endothelial cells.
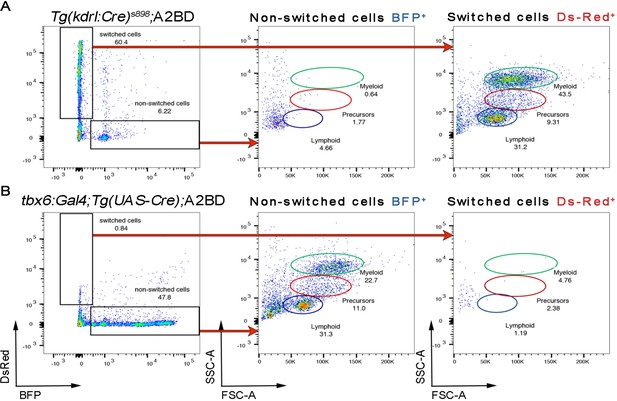
Paraxial mesoderm does not generate HSPCs.
(A) kdrl:Cre; A2BD and (B) tbx6:Gal4; Tg(UAS-Cre); A2BD adult kidney marrow was analyzed by flow cytometry. Top row illustrates that the hematopoietic lineages of kdrl:Cre; A2BD originate from switched DsRed+ ECs (A), whereas in the tbx6:Gal4; Tg(UAS-Cre); A2BD, BFP+ non-switched ECs are the source of the hematopoietic system. DsRED+ SDECs do not give rise to hematopoietic lineages (B).
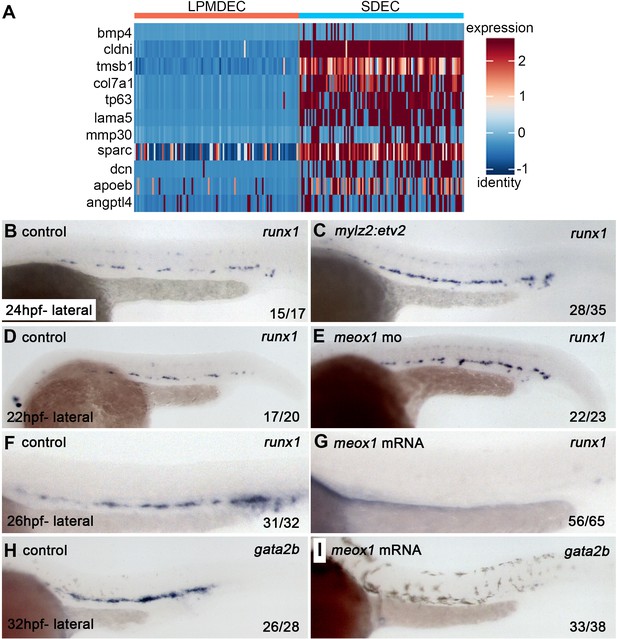
SDECs act as a vascular niche for hemogenic endothelium.
(A) Heatmap of genes differentially regulated between LPM-derived endothelial cells (LPMDEC sample is composed of pre-HSCs and HE clusters) and SDECs. (B,C) Zebrafish embryos were injected with an expression vector containing a somite-specific promoter, mylz2, driving an etv2 transgene to ectopically induced SDECs. By WISH, an increase in runx1 expression was observed at 24 hpf compared to embryos injected with an empty mylz2 vector. (D,E) Similarly, meox1 morphants exhibit an increased expression of runx1 by WISH compared to uninjected control embryos. (F–I) Conversely, overexpression of meox1 by mRNA injection strongly reduces the hemogenic markers runx1 and gata2b. SDECs, somite-derived endothelial cells.





