Structure of human Frizzled5 by fiducial-assisted cryo-EM supports a heterodimeric mechanism of canonical Wnt signaling
Figures
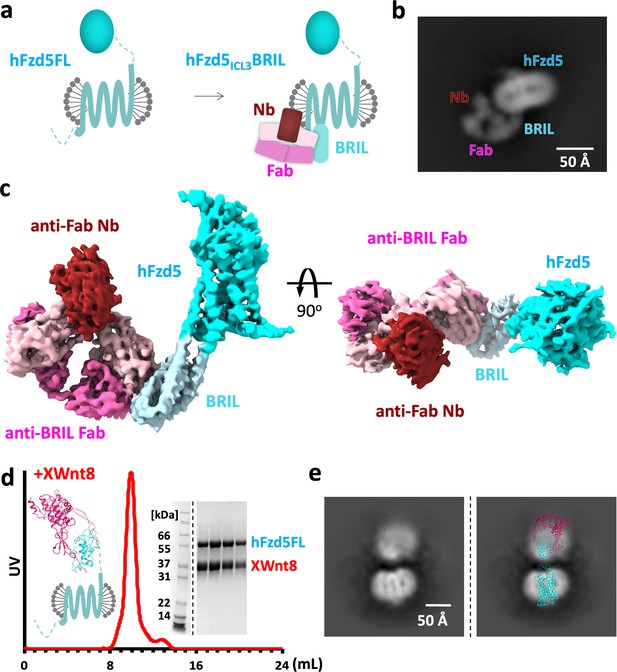
Design scheme and cryo-EM analysis of hFzd5.
(a) Cartoon representation of the strategy for hFzd5 particle decoration by anti-BRIL Fab and anti-Fab Nb. These chaperones double hFzd5 molecular weight and render it asymmetric in detergent micelle, thus working as a fiducial maker for image alignment. (b) A selected 2D class average showing the side view of monomeric hFzd5ICL3BRIL/Fab/Nb. (c) Overall EM volume around the structural model of hFzd5 (cyan), BRIL and the linker between BRIL and Fzd5 (light blue), Fab heavy chain (pink), Fab light chain (light pink) and Nb (wine). (d) The size-exclusion chromatography profile, SDS-PAGE of peak fractions, and (e) the selected 2D class average of XWnt8/hFzd5FL. The full SDS-PAGE and 2D classes are shown in Figure 1—figure supplement 5. Models of XWnt8/mouse Fzd8CRD (mFzd8CRD) (PDB ID: 4F0A) and hFzd5 are overlaid on the blobs to show the relative size of densities.

Multiple sequence alignment of human class F GPCRs.
Sequence alignment of CRD and transmembrane regions of human class F GPCRs. The CRD region and individual transmembrane helix is labelled above the amino acid sequence. The alignments for CRD-hinge domain was generated using MAFFT (https://www.ebi.ac.uk/Tools/msa/mafft), and for TM1-H8 was taken from GPCR data base (www.gpcrdb.org). The alignment result was displayed using MView (www.ebi.ac.uk/Tools/msa/mview).
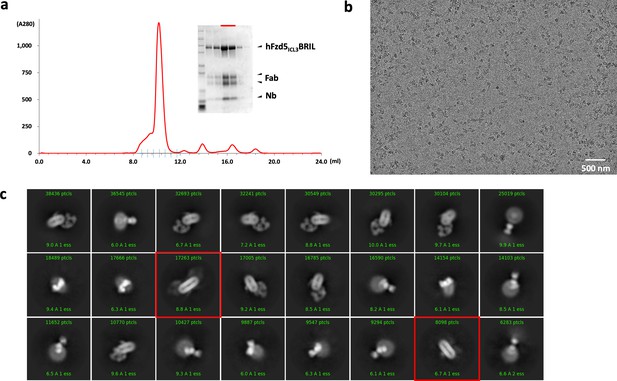
Purification, data collection and 2D classification of hFzd5ICL3BRIL/Fab/Nb.
(a) Representative size-exclusion chromatography profile of the hFzd5ICL3BRIL/Fab/Nb complex with the Coomassie stained SDS-PAGE around the peak fractions. SDS samples were prepared in reducing condition. The fractions used for EM are indicated by the red bar. (b) Representative raw micrograph of the hFzd5ICL3BRIL/Fab/Nb complex in detergent. (c) 2D class averages of selected particles. The dimer classes are marked by red boxes.
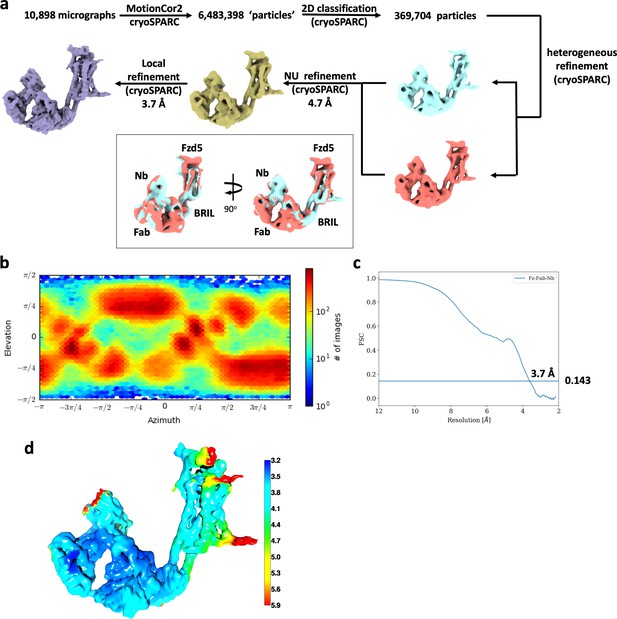
Cryo-EM data analysis of hFzd5ICL3BRIL/Fab/Nb.
(a) Cryo-EM data processing scheme. The resulting maps from the final heterogeneous refinement are superimposed for comparison (cyan and brick). (b) Orientation distribution of particles for the final 3D reconstruction. (c) The gold standard FSC curve for the final map. (d) Local resolution estimation of the final map colored in rainbow.
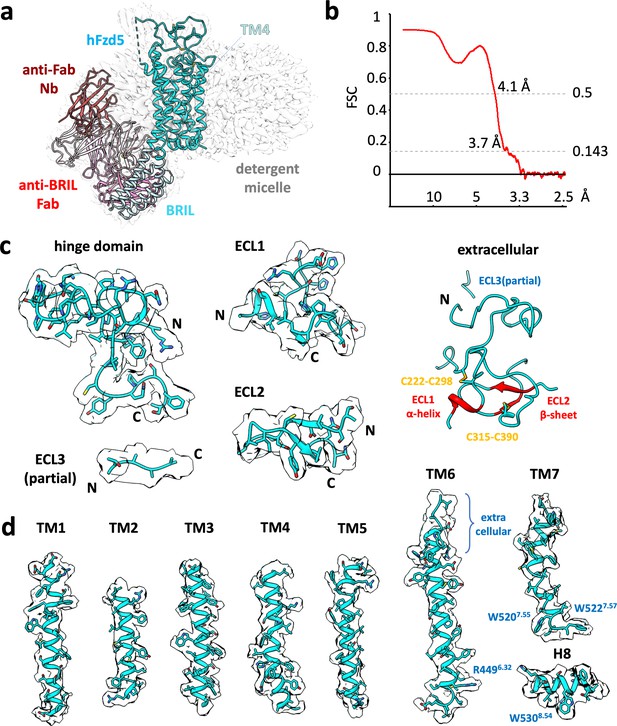
Cryo-EM map of representative built in model.
(a) Cryo-EM map (gray) and structural model of hFzd5ICL3BRIL in detergent complexed with anti-BRIL Fab and anti-Fab Nb. The model is colored as follow: the sequences from Fzd5 (cyan), BRIL and the linker between BRIL and Fzd5 (light blue), Fab heavy chain (pink), Fab light chain (light pink) and Nb (brown). (b) The map-model FCS curve generated by Phenix refinement (masked). Cryo-EM volumes (pale cyan) around the (c) extracellular regions and (d) individual transmembrane helices of hFzd5. Except for the whole extracellular region (c right) which only shows disulfide cysteine resides with red colored secondary structures assigned by DSSP, all sidechains are displayed regardless of their quality or model placement.
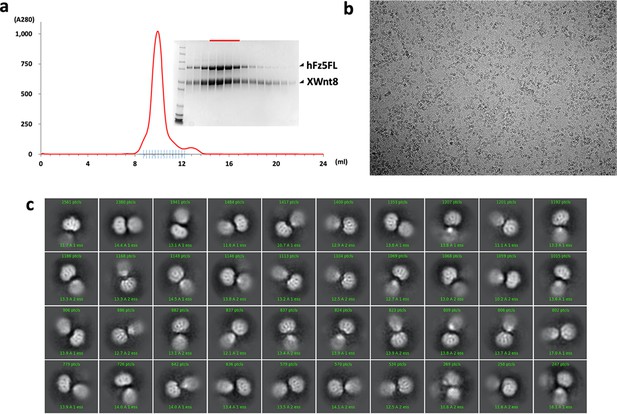
Purification, data collection and 2D class averages of the XWnt8/hFzd5FL complex.
(a) A size-exclusion chromatography profile of the XWnt8/hFzd5FL complex with the Coomassie stained SDS-PAGE. SDS samples were prepared in reducing condition. The peak fractions for the EM analysis are indicated by the red bar. (b) Representative raw micrograph of the XWnt8/hFzd5 complex. (c) 2D class averages of selected particles.
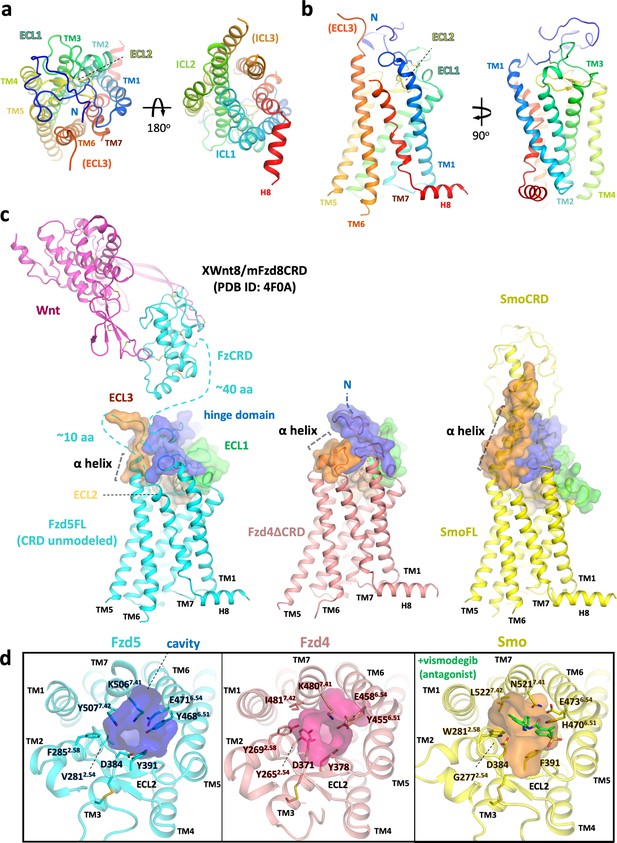
Overall structure of hFzd5 and structural comparison with hFzd4 and hSmo.
(a) Top (left) and bottom (right) views, and (b) side views of hFzd5 colored by rainbow with blue on the N-terminus and red on the C-terminus of the structural model. (c) Comparison of the extracellular regions between hFzd5, hFzd4 (pink, PDB ID: 6BD4) and hSmo (yellow, PDB ID: 5L7D). Transparent surfaces are overlaid on the cartoon models and colored separately by the hinge domain (blue), ECL1 (green), ELC2 (wheat) and ELC3 (orange). XWnt8/mFzd8CRD structure (PDB ID: 4F0A) is displayed and connected to the hFzd5 structure by dashed line, to show the size of the LRP6 binding module relative to 7TM. The α-helices at TM6-ECL3 are indicated with gray dashed lines. (d) Top views of hFzd5, hFzd4, and hSmo around the Smo-7TM ligand binding site with potential gatekeeping residues. The cavity volume calculated using the CavityPlus server (Xu et al., 2018) are displayed as transparent surface representations, with a focus on the constrictions made by bulky resides of Fzd at the upper-core site. The antagonist vismodegib (from PDB ID: 5L7I) is overlaid on the hSmo structure in its inactive and apo state (PDB ID: 5L7D).
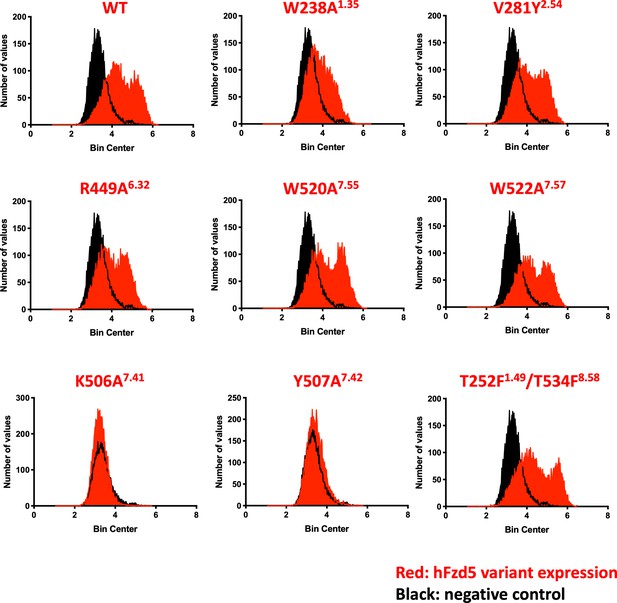
Confirmation of hFzd5 receptor expression in HEK293T transfected cells.
Cell-surface hFzd5 variants were fluorescently labeled, and frequency distribution of FACS data are presented for each variant (red) with a negative control (black).
-
Figure 2—figure supplement 1—source data 1
Raw FACS histogram data for Figure 2—figure supplement 1 showing cell-surface expression of hFzd5 variants.
- https://cdn.elifesciences.org/articles/58464/elife-58464-fig2-figsupp1-data1-v2.xlsx
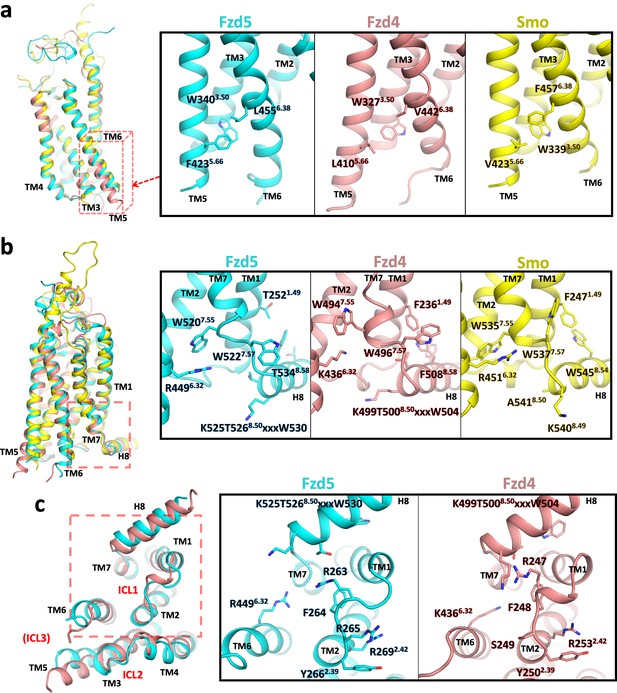
Conformational differences between hFzd5, hFzd4 and hSmo.
(a, b) Structural comparison of hFzd5, hFzd4 and hSmo (a) at the cytoplasmic end of TM5, and (b) the K/R6.32-W7.55 ionic lock with the aromatic network around H8. (c) Bottom views of hFzd5 and hFzd4 showing structural rearrangement of ICL1 and H8 important for Dvl binding. Each molecule is colored by cyan (hFzd5), pink (hFzd4, PDB ID: 6BD4) or yellow (Smo, PDB ID: 5L7D).
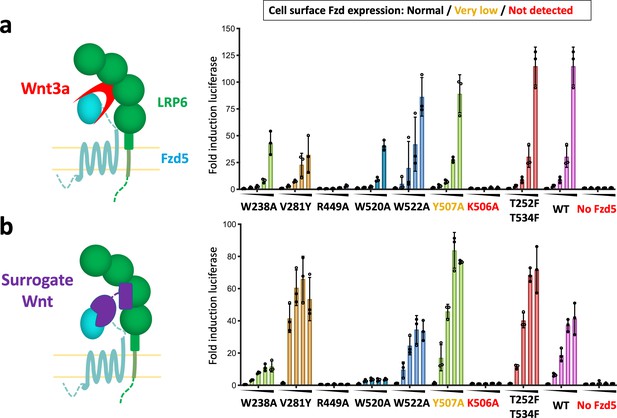
Structure-guided hFzd5 mutant signaling assays.
Schematic presentation of the ternary complex formation, and hFzd5-mediated signaling upon stimulation by (a) Wnt3a and (b) surrogate Wnt agonist. Non-lipidated, water-soluble surrogate Wnt would form the complex in a different geometry from natural Wnt ligands. Each series was stimulated (a) by Wnt3a at the concentrations of 0, 1.6, 3.1, 6.3 or 12.5 nM, or (b) by surrogate Wnt at the concentrations of 0, 0.25, 1, 4 or 16 nM, displayed from left (no agonist) to right (the highest concentration) and indicated by black triangles. The bars and error bars represent means and standard deviations, respectively, of data points from three technical replicates shown as circles.
-
Figure 4—source data 1
Raw data points for the signaling experiments shown as circles in Figure 4.
- https://cdn.elifesciences.org/articles/58464/elife-58464-fig4-data1-v2.xlsx
Tables
Cryo-EM data collection and refinement statistics
| hFzd5ICL3BRIL/anti-BRIL Fab/anti-Fab Nb (EMD-21927) (PDB ID: 6WW2) | |
|---|---|
| Data collection and processing | |
| Magnification | 81,000x |
| Voltage (kV) | 300 |
| Electron exposure (e/Å2) | 50 |
| Defocus range (μm) | −0.8 to −2.0 |
| Pixel size (Å) | 1.078 |
| Symmetry imposed | C1 |
| Initial particle images (no.) | 6,483,398 |
| Final particle images (no.) | 369,704 |
| Map resolution (Å) FSC threshold | 3.70 0.143 |
| Map resolution range (Å) | 3.2 to 5.9 |
| Refinement | |
| Initial model used (PDB code) | 5L7D, 6BD4, 6ANI, 6CBV |
| Model resolution (Å) FSC threshold | 3.7/4.1 0.143/0.5 |
| Map sharpening B factor (Å2) | −124.4 |
| Model composition Non-hydrogen atoms Protein residues | 7,594 982 |
| B factors (Å2) Protein | 89.59 |
| R.m.s. deviations Bond lengths (Å) Bond angles (°) | 0.010 1.083 |
| Validation MolProbity score Clashscore Poor rotamers (%) | 1.87 5.73 0 |
| Ramachandran plot Favored (%) Allowed (%) Disallowed (%) | 89.59 10.31 0.10 |
| Reagent type (species) or resource | Designation | Source or reference | Identifiers | Additional information |
|---|---|---|---|---|
| Cell line (Spodoptera frugiperda) | Sf9 | ATCC | CTL-1711 (RRID:CVCL_0549) | Insect cells used for baculovirus production |
| Cell line (Homo sapiens) | HEK293S GnTI- | Gift from Prof. H Gobind Khorana (PMID:12370423) | Mammalian cells used for baculovirus expression of hFzd5FL and hFzd5ICL3BRIL | |
| Cell line (Homo sapiens) | HEK293T with Fzd1/2/4/5/7/8 KO | Gift from Prof. Michael Boutros (PMID:28733458) | For signaling assay | |
| Cell line (Mus musculus) | Mouse monoclonal M1 hybridoma | Gift from Prof. Brian K Kobilka (PMID:17962520) | To purify anti-FLAG M1 (mouse IgG2a) antibody to prepare FLAG affinity column (15 mg/ml resin). This was used for purification of hFzd5FL and hFzd5ICL3BRIL. | |
| Strain, strain background (Escherichia coli) | BL21(DE3) | New England Biolabs | C2527I | E. coli strain for expression of the nanobody |
| Strain, strain background (Escherichia coli) | BL21 (gold) | Agilent Technologies | 230130 | E. coli strain for expression of the Fabs |
| Transfected construct (Homo sapiens) | pRK5 hFzd5 wild-type | Yu et al., 2012 (PMID:23095888) | For signaling assay | |
| Transfected construct (Homo sapiens) | pRK5 hFzd5 mutants | this study | For signaling assay | |
| Recombinant DNA reagent | 7xTCF-Ffluc | Addgene | 24308 | For signaling assay |
| Recombinant DNA reagent | BestBac Linearized Baculovirus DNA 2.0 | Expression Systems | 91–002 | For baculovirus production |
| Peptide, recombinant protein | Wnt3a | R and D Systems | 5036-WN | For signaling assay |
| Peptide, recombinant protein | Surrogate Wnt | Miao et al. Cell Stem Cell 2020 | For signaling assay | |
| Peptide, recombinant protein | DRPB_Fz8 | Dang et al., 2019 (PMID:31086346) | For signaling assay | |
| Commercial assay or kit | Luciferase assay kit | Promega | E152A | For signaling assay |
| Chemical compound, drug | n-Dodecyl-β-D-Maltoside (DDM) | Anatrace | D310S | Membrane protein purification |
| Chemical compound, drug | Lauryl Maltose Neopentyl Glycol (LMNG) | Anatrace | NG310 | Membrane protein purification |
| Chemical compound, drug | Glyco-Diosgenin (GDN) | Anatrace | GDN101 | Membrane protein purification |
| Chemical compound, drug | Cholesterol Hemisuccinate tris Salt (CHS) | Anatrace | CH210 | Membrane protein purification |
| Chemical compound, drug | Digitonin | EMD Millipore | 300410 | Cryo-EM specimen freezing additive |
| Software, algorithm | DigitalMicrograph | Gatan | Microscope alignment and cryo-EM data collection | |
| Software, algorithm | serialEM | https://bio3d.colorado.edu/SerialEM/ (PMID:16182563) | version 3.6 | Cryo-EM data collection |
| Software, algorithm | MotionCor2 | https://emcore.ucsf.edu/ucsf-motioncor2/ (PMID:28250466) | Motion correction of cryo-EM movies | |
| Software, algorithm | cryoSPARC | https://cryosparc.com/ (PMID:28165473) | version 2.12.14 | Cryo-EM data processing |
| Software, algorithm | Coot | https://www2.mrc-lmb.cam.ac.uk/personal/pemsley/coot/ (PMID:20383002) | Structure model building | |
| Software, algorithm | Isolde | https://isolde.cimr.cam.ac.uk/ (PMID:29872003) | Structure model building/refinement | |
| Software, algorithm | Phenix suite | https://www.phenix-online.org/ (PMID:20124702) | Structure refinement | |
| Software, algorithm | UCSF Chimera | https://www.cgl.ucsf.edu/chimera/ (PMID:15264254) | Initial homology model docking | |
| Software, algorithm | UCSF ChimeraX | https://www.rbvi.ucsf.edu/chimerax/ (PMID:28710774) | Structural visualization/figure preparation/Isolde execution | |
| Software, algorithm | PyMol | Schrödinger | Structural visualization/figure preparation | |
| Software, algorithm | GraphPad Prism 7 | GraphPad | analysis of signaling data | |
| Other | CNBr-Activated Sepharose 4 Fast Flow | GE Healthcare | 17098101 | For preparation of anti-FLAG sepharose |
| Other | Lipofectamine 2000 | Invitrogen | 11668030 | Transfection reagent for signaling assay |





